Fisica Computazionale applicata alle Macromolecole Modelli probabilistici per

Fisica Computazionale applicata alle Macromolecole Modelli probabilistici per Sequenze Biologiche Pier Luigi Martelli Università di Bologna gigi@biocomp. unibo. it 051 2094005 338 3991609

PROLOGUE: Pitfalls of standard alignments
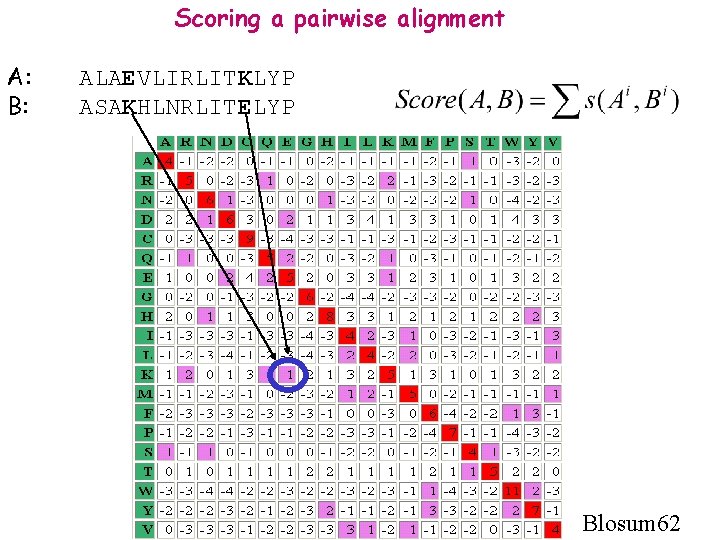
Scoring a pairwise alignment A: B: ALAEVLIRLITKLYP ASAKHLNRLITELYP Blosum 62
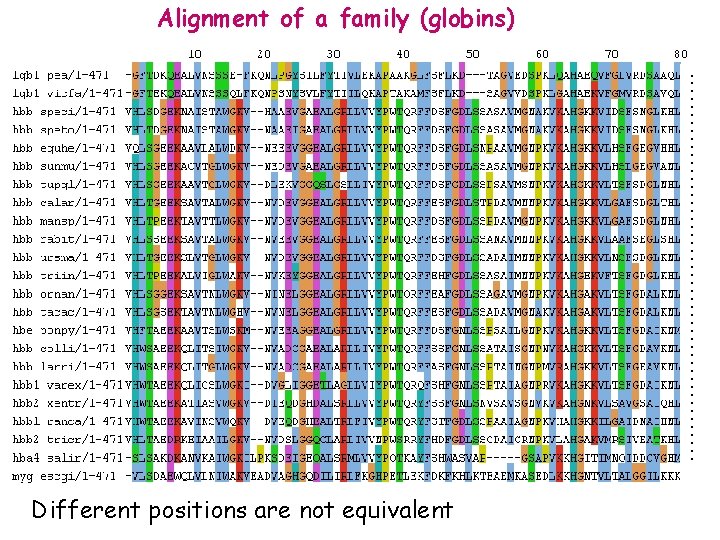
……………………. Alignment of a family (globins) Different positions are not equivalent
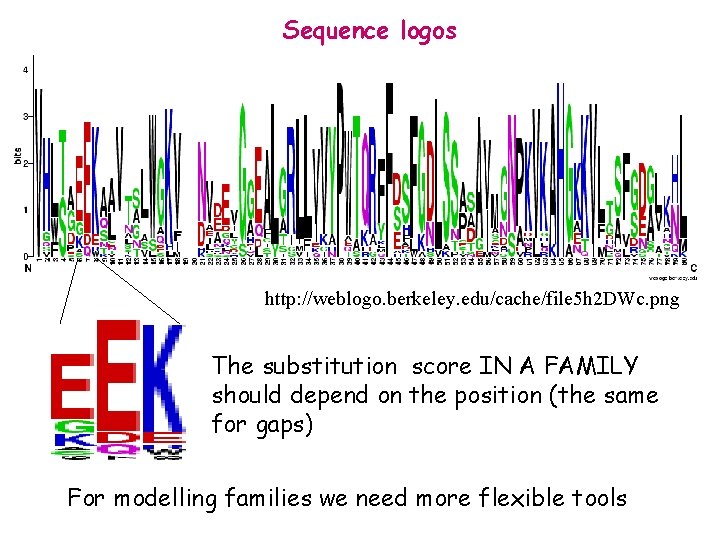
Sequence logos http: //weblogo. berkeley. edu/cache/file 5 h 2 DWc. png The substitution score IN A FAMILY should depend on the position (the same for gaps) For modelling families we need more flexible tools

Probabilistic Models for Biological Sequences • What are they?

Probabilistic models for sequences Generative definition: • Objects producing different outcomes (sequences) with different probabilities Probability • The probability distribution over the sequences space determines the model specificity Sequence space M Generates si with probability P(si | M) e. g. : M is the representation of the family of globins
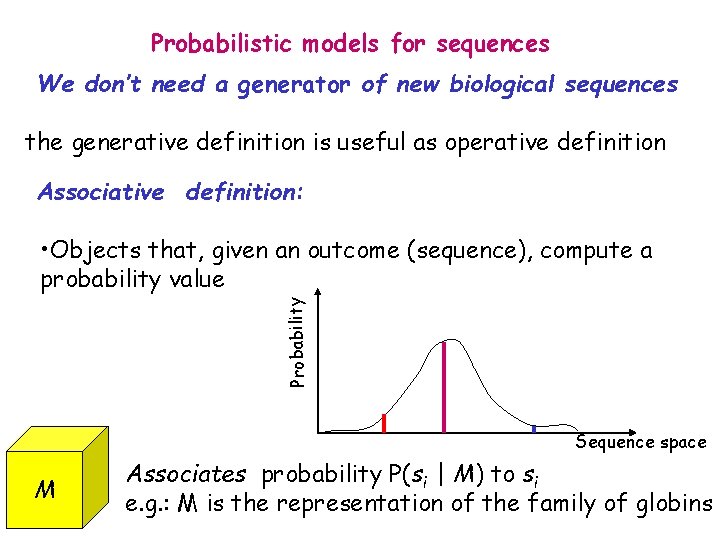
Probabilistic models for sequences We don’t need a generator of new biological sequences the generative definition is useful as operative definition Associative definition: Probability • Objects that, given an outcome (sequence), compute a probability value Sequence space M Associates probability P(si | M) to si e. g. : M is the representation of the family of globins
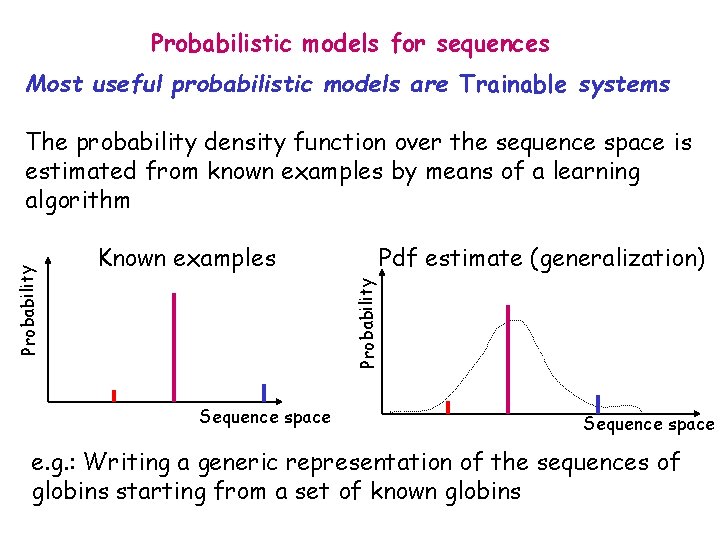
Probabilistic models for sequences Most useful probabilistic models are Trainable systems Known examples Pdf estimate (generalization) Probability The probability density function over the sequence space is estimated from known examples by means of a learning algorithm Sequence space e. g. : Writing a generic representation of the sequences of globins starting from a set of known globins

Probabilistic Models for Biological Sequences • What are they? • Why to use them?

Modelling a protein family Probabilistic model Given a protein class (e. g. Globins), a probabilistic model trained on this family can compute a probability value for each new sequence Seq 1 Seq 2 Seq 3 Seq 4 Seq 5 Seq 6 0. 98 0. 21 0. 12 0. 89 0. 47 0. 78 This value measures the similarity between the new sequence and the family described by the model

Probabilistic Models for Biological Sequences • What are they? • Why to use them? • Which probabilities do they compute?

P( s | M ) or P( M | s ) ? A model M associates to a sequence s the probability P( s | M ) This probability answers the question: Which is the probability for a model describing the Globins to generate the sequence s ? The question we want to answer is: Given a sequence s, is it a Globin? We need to compute P( M | s ) !!
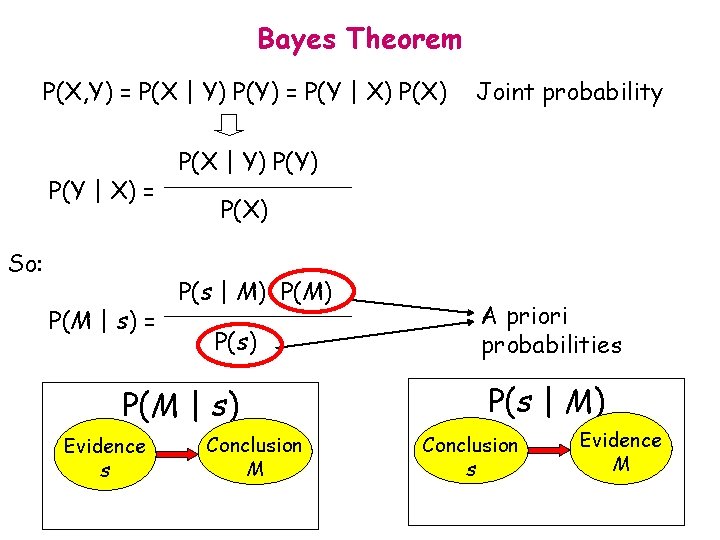
Bayes Theorem P(X, Y) = P(X | Y) P(Y) = P(Y | X) = So: P(M | s) = P(X | Y) P(X) P(s | M) P(s) P(M | s) Evidence s Joint probability Conclusion M A priori probabilities P(s | M) Conclusion s Evidence M

Bayes’ rule: Example • A rare disease affects 1 out of 100, 000 people. • A test shows positive – with probability 0. 99 when applied to an ill person, and – with probability 0. 01 when applied to a healthy person. • What is the probability that you have the disease given that you test positive?

Bayes’ rule: Example P(+|ill) = 0. 99 P(+|healthy) = 0. 01 Happy End: More likely the test is incorrect!! P(ill) = 10 -5

Is the pope an alien? Since the probability P(Pope|Human) =1/(6, 000, 000) do this imply that the Pope is not a human being? Beck-Bornholdt HP, Dubben HH, Nature 381, 730 (1996)

P(Pope|Human) is not the same as P(Human|Pope) but P(Alien) ~ 0 So P(Human|Pope) ~ 1. 0 The pope is (probably) not an alien S Eddy and D Mc. Kay’s answer

The A priori probabilities P(M | s) = P(s | M) P(s) A priori probabilities P(M) is the probability of the model (i. e. of the class described by the model) BEFORE we know the sequence: can be estimated as the abundance of the class P(s) is the probability of the sequence in the sequence space. Cannot be reliably estimated!!

Comparison between models We can overcome the problem comparing the probability of generating s from different models P(M 1 | s) P(M 2 | s) = = P(s | M 1) P(s) P(s | M 2) P(M 2) = P(s | M 1) P(s | M 2) P(M 2) Ratio between the abundance of the classes
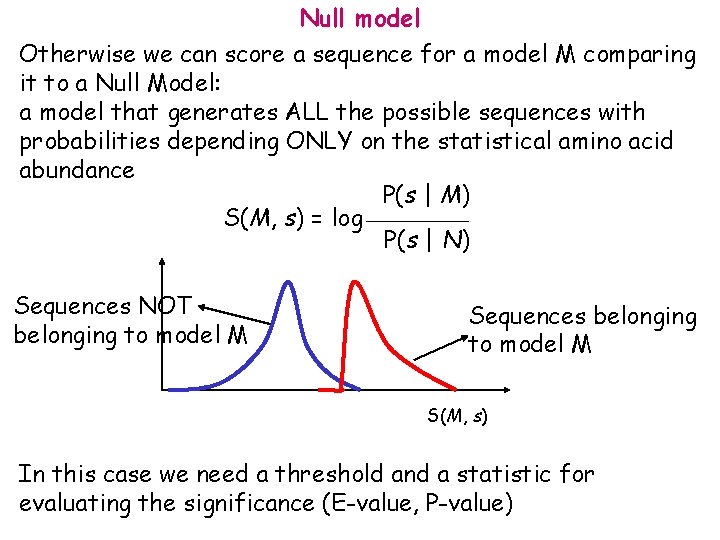
Null model Otherwise we can score a sequence for a model M comparing it to a Null Model: a model that generates ALL the possible sequences with probabilities depending ONLY on the statistical amino acid abundance P(s | M) S(M, s) = log P(s | N) Sequences NOT belonging to model M Sequences belonging to model M S(M, s) In this case we need a threshold and a statistic for evaluating the significance (E-value, P-value)

The simplest probabilistic models: Markov Models • Definition
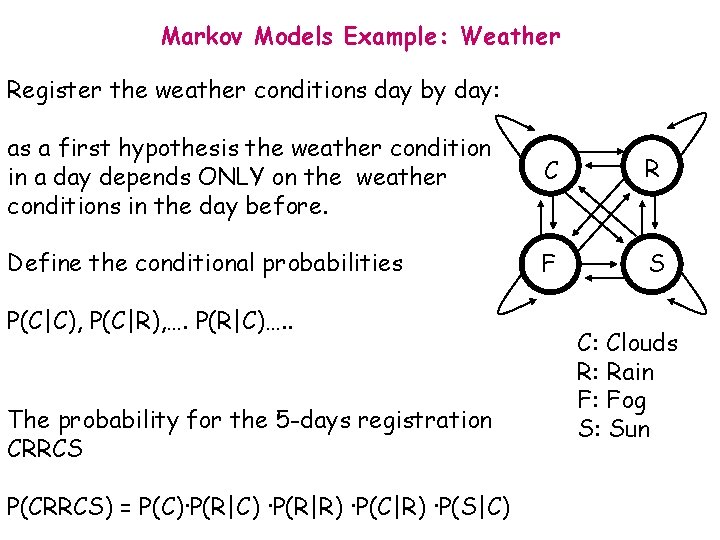
Markov Models Example: Weather Register the weather conditions day by day: as a first hypothesis the weather condition in a day depends ONLY on the weather conditions in the day before. C R Define the conditional probabilities F S P(C|C), P(C|R), …. P(R|C)…. . The probability for the 5 -days registration CRRCS P(CRRCS) = P(C)·P(R|C) ·P(R|R) ·P(C|R) ·P(S|C) C: Clouds R: Rain F: Fog S: Sun
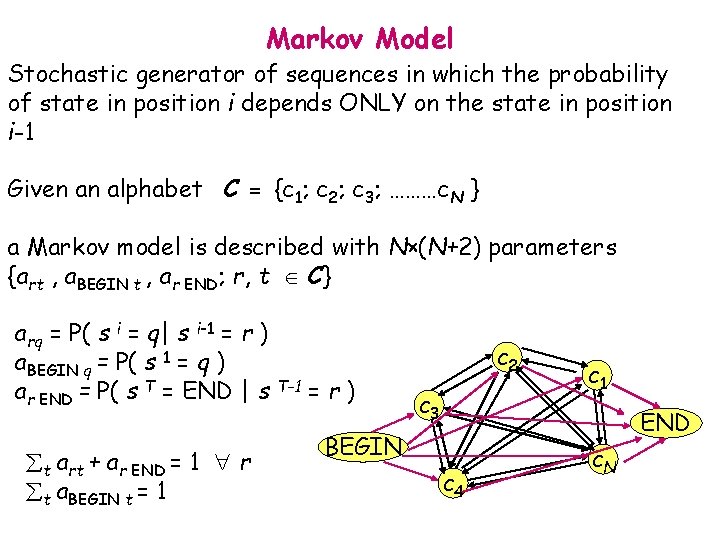
Markov Model Stochastic generator of sequences in which the probability of state in position i depends ONLY on the state in position i-1 Given an alphabet C = {c 1; c 2; c 3; ………c. N } a Markov model is described with N×(N+2) parameters {art , a. BEGIN t , ar END; r, t C} arq = P( s i = q| s i-1 = r ) a. BEGIN q = P( s 1 = q ) ar END = P( s T = END | s T-1 = r ) t art + ar END = 1 r t a. BEGIN t = 1 c 2 c 3 c 1 END BEGIN c 4 c. N

Markov Models Given the sequence: s = s 1 s 2 s 3 s 4 s 6 ……………s. T with si C = {c 1; c 2; c 3; ………c. N } P( s | M ) = P( s 1 ) i=2 P( s i | s i-1 ) = = a. BEGIN s 1 Ti=2 as i-1 s i as. T END P(“ALKALI”)= a. BEGIN A a. A L a. L K a. K A a. A L a. L I a. I END
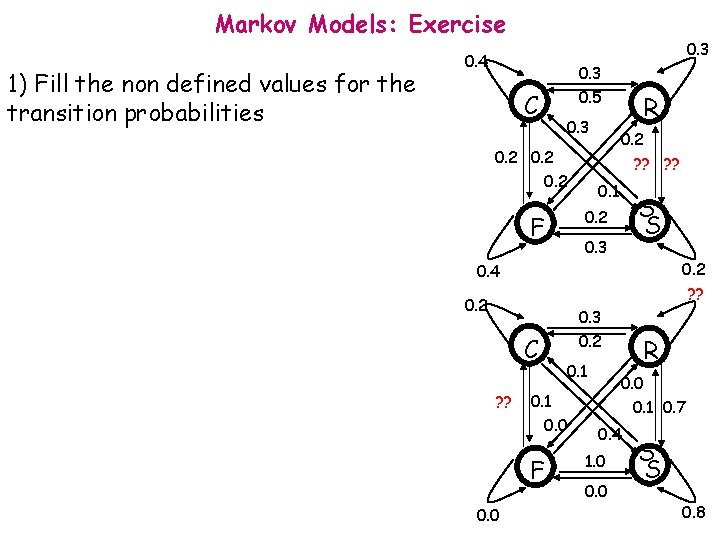
Markov Models: Exercise 1) Fill the non defined values for the transition probabilities 0. 3 0. 4 C 0. 3 0. 5 0. 3 0. 2 R 0. 2 ? ? 0. 1 S S 0. 2 F 0. 3 0. 2 ? ? 0. 4 0. 2 C ? ? 0. 1 0. 0 F 0. 0 0. 3 0. 2 R 0. 0 0. 1 0. 7 0. 4 1. 0 0. 0 S S 0. 8
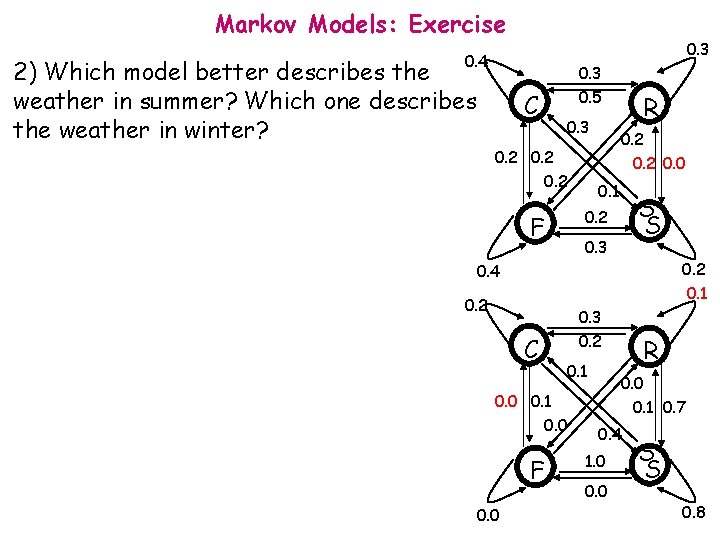
Markov Models: Exercise 0. 3 0. 4 2) Which model better describes the weather in summer? Which one describes the weather in winter? C 0. 3 0. 5 0. 3 0. 2 R 0. 2 0. 0 0. 1 S S 0. 2 F 0. 3 0. 2 0. 1 0. 4 0. 2 C 0. 3 0. 2 0. 1 0. 0 F 0. 0 R 0. 0 0. 1 0. 7 0. 4 1. 0 0. 0 S S 0. 8
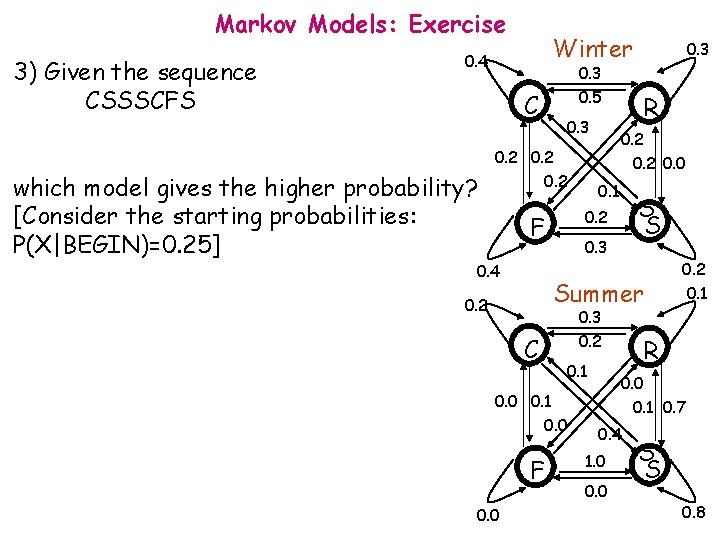
Markov Models: Exercise 3) Given the sequence CSSSCFS Winter 0. 4 C which model gives the higher probability? [Consider the starting probabilities: P(X|BEGIN)=0. 25] 0. 3 0. 5 0. 3 0. 2 0. 4 0. 2 0. 0 0. 1 S S 0. 3 Summer 0. 2 C 0. 3 0. 2 0. 1 0. 0 F 0. 0 R 0. 2 F 0. 3 0. 2 0. 1 R 0. 0 0. 1 0. 7 0. 4 1. 0 0. 0 S S 0. 8
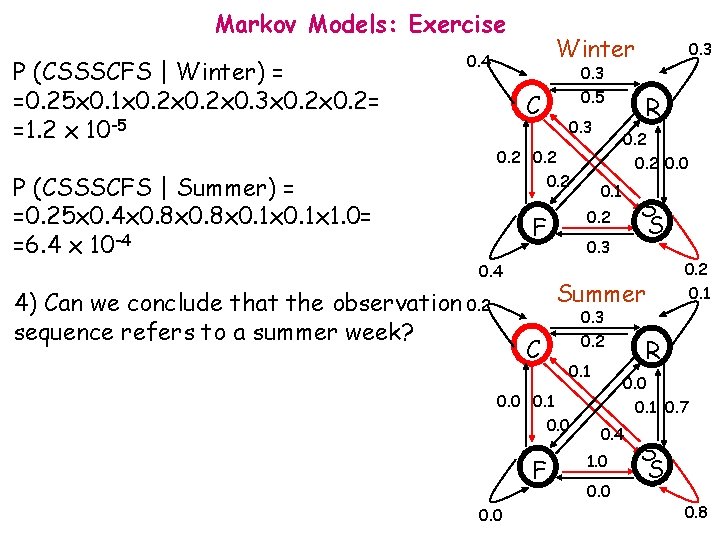
Markov Models: Exercise P (CSSSCFS | Winter) = =0. 25 x 0. 1 x 0. 2 x 0. 3 x 0. 2= =1. 2 x 10 -5 P (CSSSCFS | Summer) = =0. 25 x 0. 4 x 0. 8 x 0. 1 x 1. 0= =6. 4 x 10 -4 Winter 0. 4 0. 3 0. 5 C 0. 3 0. 2 0. 0 0. 1 Summer 0. 3 0. 2 C 0. 1 0. 0 F 0. 2 0. 1 R 0. 0 0. 1 0. 0 S S 0. 3 0. 4 4) Can we conclude that the observation 0. 2 sequence refers to a summer week? R 0. 2 F 0. 3 0. 1 0. 7 0. 4 1. 0 0. 0 S S 0. 8
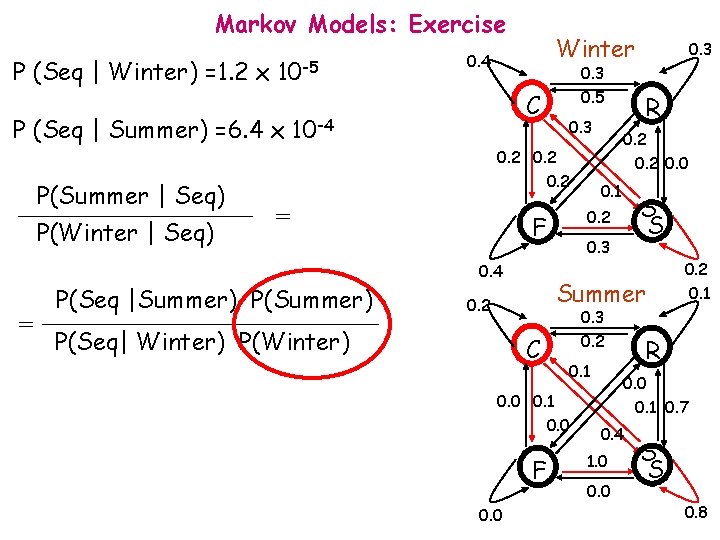
Markov Models: Exercise P (Seq | Winter) =1. 2 x 10 -5 P(Winter | Seq) 0. 3 0. 5 C P (Seq | Summer) =6. 4 x 10 -4 P(Summer | Seq) Winter 0. 4 0. 3 0. 2 = = 0. 2 0. 0 0. 1 0. 3 0. 2 C 0. 1 0. 0 F 0. 2 0. 1 R 0. 0 0. 1 0. 0 S S Summer 0. 2 P(Seq| Winter) P(Winter) 0. 2 0. 3 0. 4 P(Seq |Summer) P(Summer) R 0. 2 F 0. 3 0. 1 0. 7 0. 4 1. 0 0. 0 S S 0. 8
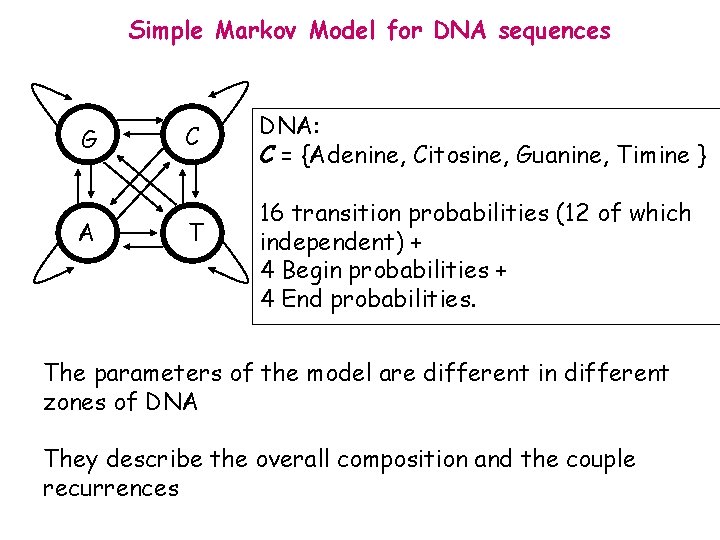
Simple Markov Model for DNA sequences G A C T DNA: C = {Adenine, Citosine, Guanine, Timine } 16 transition probabilities (12 of which independent) + 4 Begin probabilities + 4 End probabilities. The parameters of the model are different in different zones of DNA They describe the overall composition and the couple recurrences
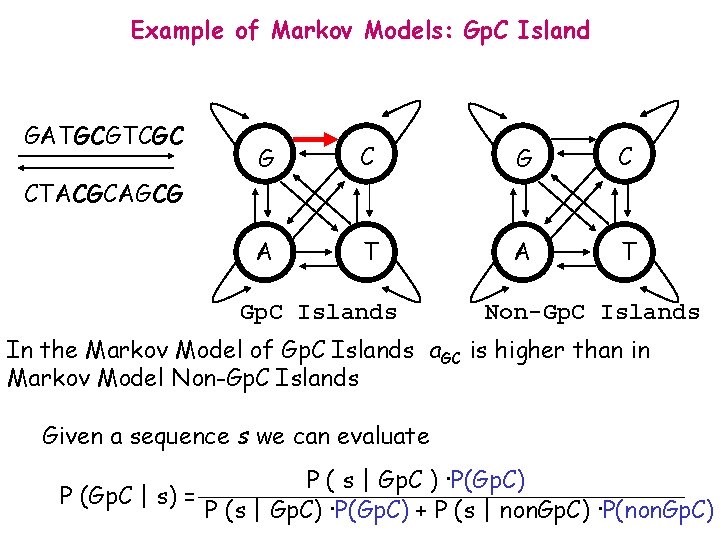
Example of Markov Models: Gp. C Island GATGCGTCGC G C A T CTACGCAGCG Gp. C Islands Non-Gp. C Islands In the Markov Model of Gp. C Islands a. GC is higher than in Markov Model Non-Gp. C Islands Given a sequence s we can evaluate P (Gp. C | s) = P ( s | Gp. C ) ·P(Gp. C) P (s | Gp. C) ·P(Gp. C) + P (s | non. Gp. C) ·P(non. Gp. C)

The simplest probabilistic models: Markov Models • Definition • Training

Training of Markov Models Let M be the set of parameters of model M. During the training phase, M parameters are estimated from the set of known data D Maximum Likelihood Extimation (ML) ML = argmax P( D | M, ) It can be proved that: nik Frequency of occurrence as counted in the aik = j nij data set D Maximum A Posteriori Extimation (MAP) MAP = argmax P( | M, D ) = argmax [P( D | M, ) P( ) ]

Maximum Likelihood training: Proof Given a sequence s contained in D: s = s 1 s 2 s 3 s 4 s 6 ……………s. T We can count the number of transitions between any to states j and k: njk Where states 0 and N+1 are BEGIN and END Normalisation contstraints are taken into account using the Lagrange multipliers lk

Hidden Markov Models • Preliminary examples

Loaded dice We have 99 regular dice (R) and 1 loaded die (L). R L P(1) 1/6 1/10 P(2) 1/6 1/10 P(3) 1/6 1/10 P(4) 1/6 1/10 P(5) 1/6 1/10 P(6) 1/6 1/2 Given a sequence: 4156266656321636543662152611536264162364261664616263 We don’t know the sequence of dice that generated it. RRRRRLRLRRRRRRRRRRRRLRLRRRRLRRRRLRR

Loaded dice Hypothesis: We chose a different die for each roll Two stochastic processes give origin to the sequence of observations. 1) Choosing the die ( R o L ). 2) Rolling the die The sequence of dice is hidden The first process is assumed to be Markovian (in this case a 0 -order MM) The outcome of the second process depends only on the state reached in the first process (that is the chosen die)
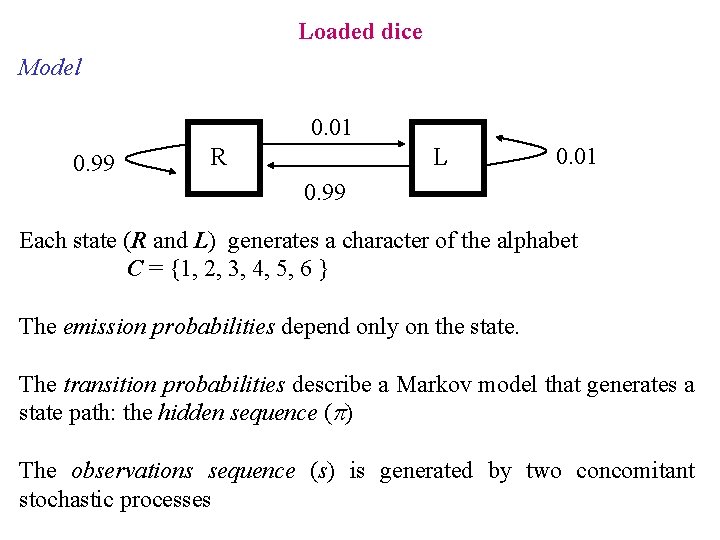
Loaded dice Model 0. 01 0. 99 R L 0. 01 0. 99 Each state (R and L) generates a character of the alphabet C = {1, 2, 3, 4, 5, 6 } The emission probabilities depend only on the state. The transition probabilities describe a Markov model that generates a state path: the hidden sequence (p) The observations sequence (s) is generated by two concomitant stochastic processes

Alcuni esempi semi-seri 1) DEMOGRAFICO Osservabile: Numero di nascite e/o morti in un anno in un luogo Variabile Nascosta: Stato economico (in prima istanza, se consideriamo la “fortuna” economica in un anno un processo casuale, la ricchezza accumulata è un prodesso markoviano ---> possiamo ricostruire lo stato economico a partire dai registri anagrafici? 2) DOCENTE METEOPATICO Osservabile: Media dei voti giornalieri registrati su un registro di un professore meteopatico Variabile Nascosta: Stato meteorologico ---> possiamo ricostruire lo stato meteorologico a partire dal registro del docente?

Alcuni esempi più seri 1) STRUTTURA SECONDARIA Osservabile: Sequenza proteica Variabile Nascosta: Struttura secondaria ---> possiamo predire la struttura secondaria a partire dalla sequenza? 2) ALLINEAMENTI Osservabile: Sequenza proteica Variabile Nascosta: Posizione di ogni residuo nell’allineamento di una famiglia proteica ---> possiamo ricostruire l’allineamento della sequenza alla famiglia proteica a partire dalla sequenza?
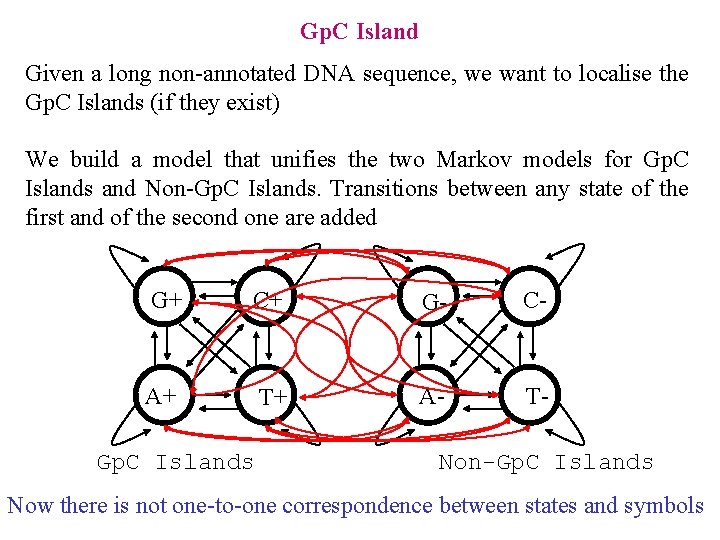
Gp. C Island Given a long non-annotated DNA sequence, we want to localise the Gp. C Islands (if they exist) We build a model that unifies the two Markov models for Gp. C Islands and Non-Gp. C Islands. Transitions between any state of the first and of the second one are added G+ C+ G- C- A+ T+ A- T- Gp. C Islands Non-Gp. C Islands Now there is not one-to-one correspondence between states and symbols
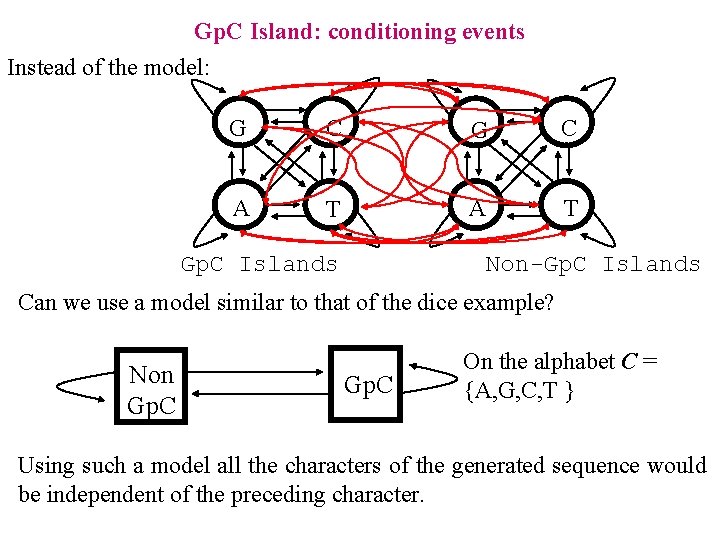
Gp. C Island: conditioning events Instead of the model: G C A T Non-Gp. C Islands Can we use a model similar to that of the dice example? Non Gp. C On the alphabet C = {A, G, C, T } Using such a model all the characters of the generated sequence would be independent of the preceding character.

Hidden Markov Models • Preliminary examples • Formal definition

Formal definition of Hidden Markov Models A HMM is a stochastic generator of sequences characterised by: · N states · A set of transition probabilities between two states {akj} akj = P( (i) = j | (i-1) = k ) · A set of starting probabilities {a 0 k} a 0 k = P( (1) = k ) · A set of ending probabilities {ak 0} ak 0 = P( (i) = END | (i-1) = k ) · An alphabet C with M characters. · A set of emission probabilities for each state {ek (c)} ek (c) = P( s i = c | (i) = k ) s: sequence ·Constraints: p: path through the states k a 0 k = 1 ak 0 + j ak j = 1 k c C ek (c) = 1 k
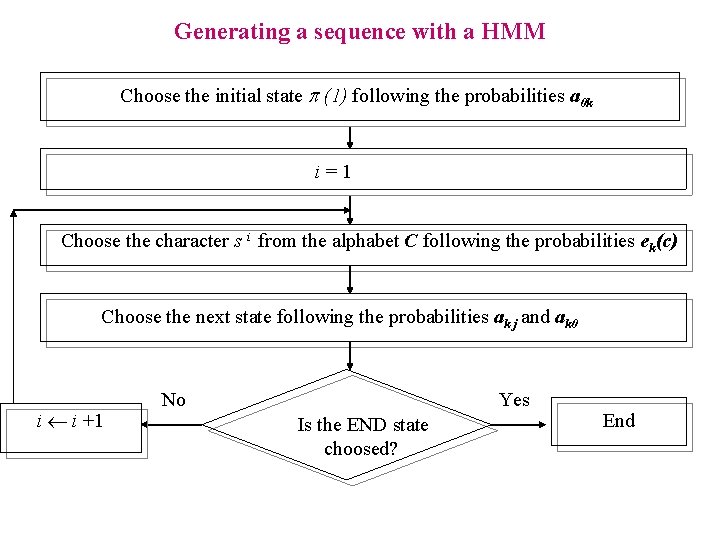
Generating a sequence with a HMM Choose the initial state p (1) following the probabilities a 0 k i=1 Choose the character s i from the alphabet C following the probabilities ek(c) Choose the next state following the probabilities ak j and ak 0 i i +1 No Yes Is the END state choosed? End
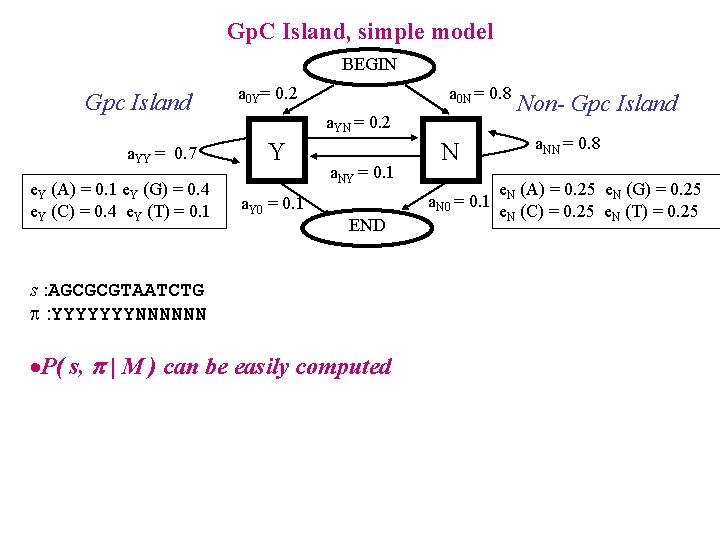
Gp. C Island, simple model BEGIN Gpc Island a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 a 0 Y= 0. 2 a 0 N = 0. 8 a. YN = 0. 2 Y a. Y 0 = 0. 1 a. NY = 0. 1 N a. N 0 = 0. 1 END s : AGCGCGTAATCTG : YYYYYYYNNNNNN ·P( s, | M ) can be easily computed Non- Gpc Island a. NN = 0. 8 e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25
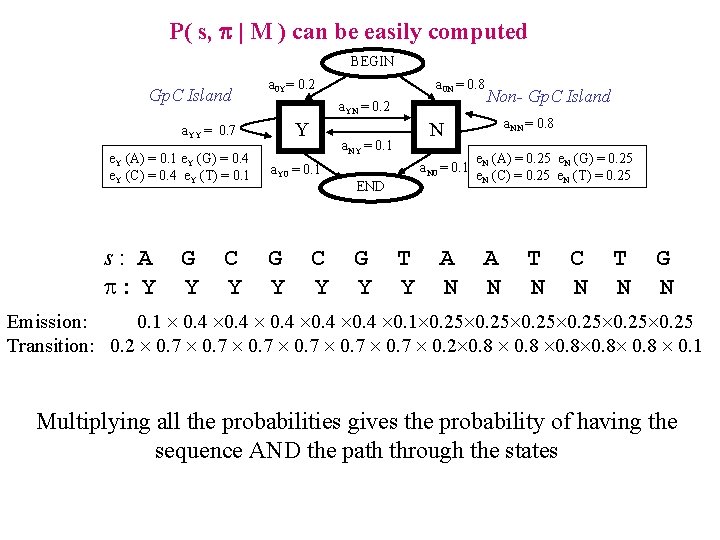
P( s, p | M ) can be easily computed BEGIN Gp. C Island a 0 Y= 0. 2 a. YN = 0. 2 Y a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 s: A : Y G Y C Y a 0 N = 0. 8 G Y C Y a. N 0 = 0. 1 END G Y a. NN = 0. 8 N a. NY = 0. 1 a. Y 0 = 0. 1 Non- Gp. C Island T Y A N e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 A N T N C N T N G N Emission: 0. 1 0. 4 0. 1 0. 25 Transition: 0. 2 0. 7 0. 2 0. 8 0. 8 0. 1 Multiplying all the probabilities gives the probability of having the sequence AND the path through the states

Evaluation of the joint probability of the sequence ad the path

Hidden Markov Models • Preliminary examples • Formal definition • Three questions
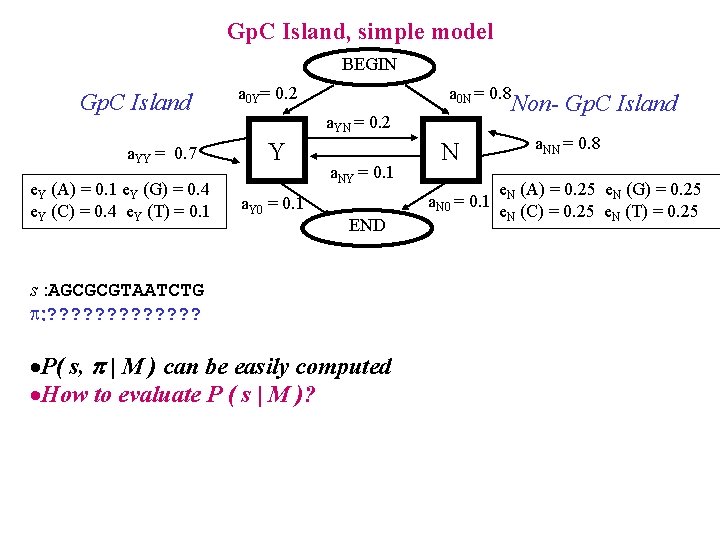
Gp. C Island, simple model BEGIN Gp. C Island a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 a 0 Y= 0. 2 a 0 N = 0. 8 Non- Gp. C Island a. YN = 0. 2 Y a. Y 0 = 0. 1 a. NY = 0. 1 N a. N 0 = 0. 1 END s : AGCGCGTAATCTG : ? ? ? ? ·P( s, | M ) can be easily computed ·How to evaluate P ( s | M )? a. NN = 0. 8 e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25
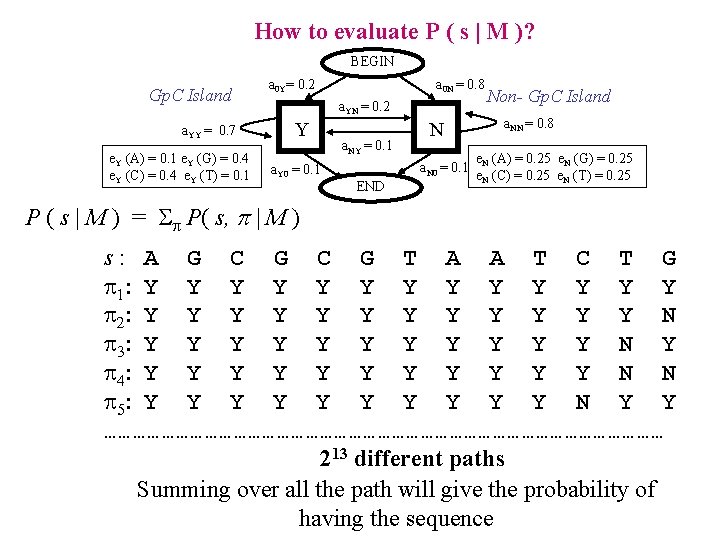
How to evaluate P ( s | M )? BEGIN Gp. C Island a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 a 0 Y= 0. 2 a 0 N = 0. 8 a. YN = 0. 2 Y a. Y 0 = 0. 1 a. NY = 0. 1 N a. N 0 = 0. 1 END Non- Gp. C Island a. NN = 0. 8 e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 P ( s | M ) = P( s, p | M ) s: A G C G T A A T C T G 1: Y Y Y Y 2: Y Y Y N 3: Y Y Y N Y 4: Y Y Y N N 5: Y Y Y Y Y N Y Y …………………………………………………… 213 different paths Summing over all the path will give the probability of having the sequence
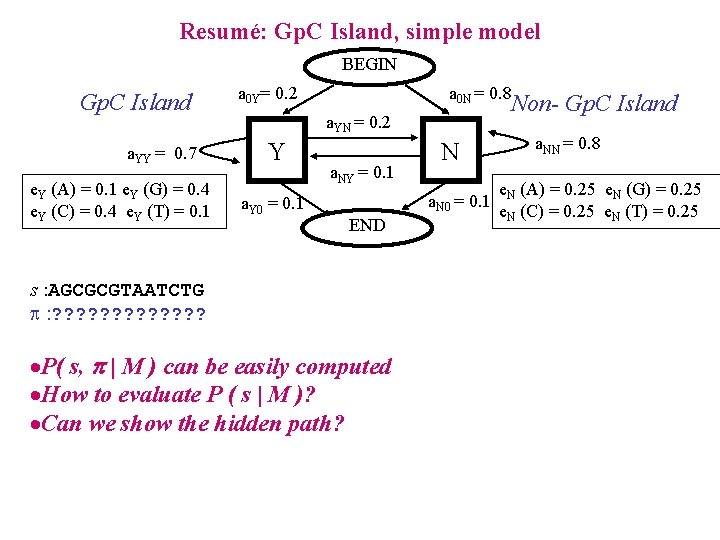
Resumé: Gp. C Island, simple model BEGIN Gp. C Island a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 a 0 Y= 0. 2 a 0 N = 0. 8 Non- Gp. C Island a. YN = 0. 2 Y a. Y 0 = 0. 1 a. NY = 0. 1 N a. N 0 = 0. 1 END s : AGCGCGTAATCTG : ? ? ? ? ·P( s, | M ) can be easily computed ·How to evaluate P ( s | M )? ·Can we show the hidden path? a. NN = 0. 8 e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25
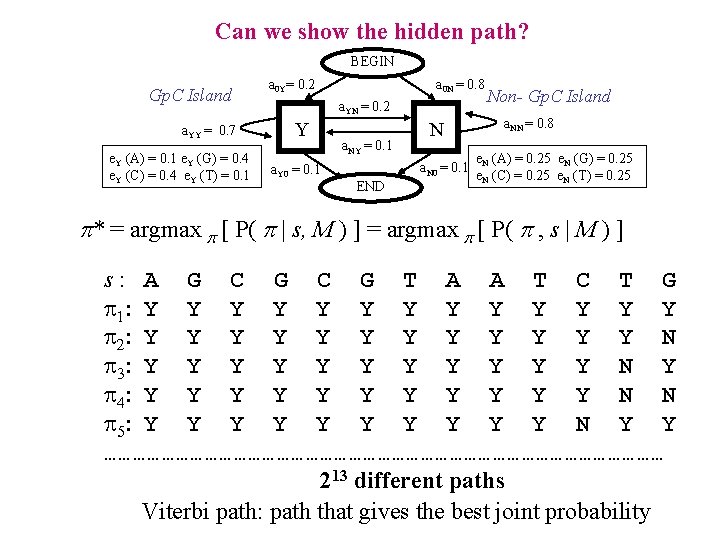
Can we show the hidden path? BEGIN Gp. C Island a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 a 0 Y= 0. 2 a 0 N = 0. 8 a. YN = 0. 2 Y a. Y 0 = 0. 1 a. NY = 0. 1 N a. N 0 = 0. 1 END Non- Gp. C Island a. NN = 0. 8 e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 p* = argmax p [ P( p | s, M ) ] = argmax p [ P( p , s | M ) ] s: A G C G T A A T C T G 1: Y Y Y Y 2: Y Y Y N 3: Y Y Y N Y 4: Y Y Y N N 5: Y Y Y Y Y N Y Y …………………………………………………… 213 different paths Viterbi path: path that gives the best joint probability

Can we show the hidden path? A Posteriori decoding For each position choose the state p (t) : p (i) = argmax k [ P( p (i) = k| s, M ) ] The contribution to this probability derives from all the paths that go through the state k at position i. The A posteriori path can be a non-sense path (it may not be a legitimate path if some transitions are not permitted in the model)
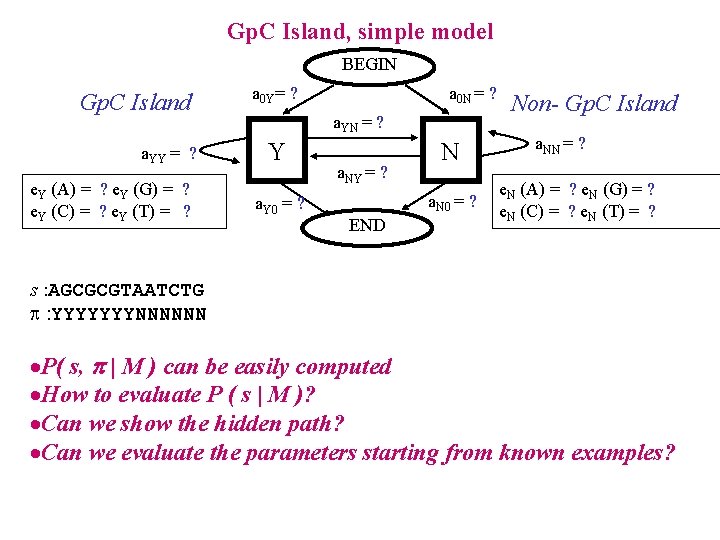
Gp. C Island, simple model BEGIN Gp. C Island a. YY = ? e. Y (A) = ? e. Y (G) = ? e. Y (C) = ? e. Y (T) = ? a 0 Y= ? a 0 N = ? a. YN = ? Y a. Y 0 = ? a. NY = ? N a. N 0 = ? END Non- Gp. C Island a. NN = ? e. N (A) = ? e. N (G) = ? e. N (C) = ? e. N (T) = ? s : AGCGCGTAATCTG : YYYYYYYNNNNNN ·P( s, | M ) can be easily computed ·How to evaluate P ( s | M )? ·Can we show the hidden path? ·Can we evaluate the parameters starting from known examples?
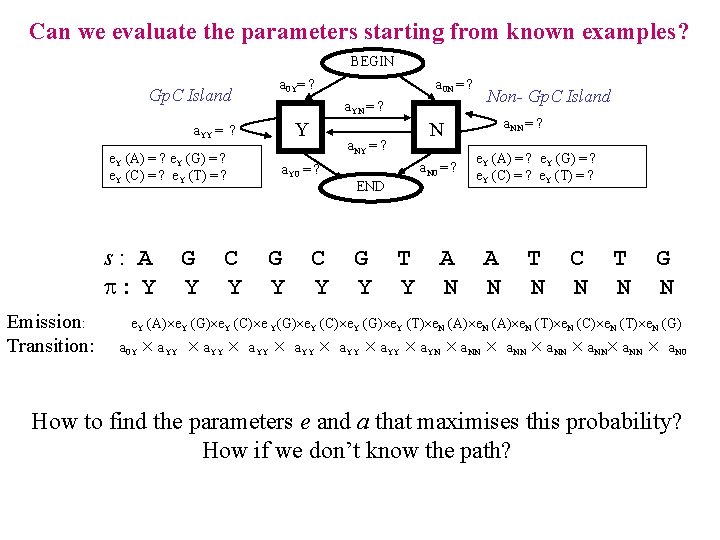
Can we evaluate the parameters starting from known examples? BEGIN a 0 Y= ? Gp. C Island a. YN = ? Y a. YY = ? e. Y (A) = ? e. Y (G) = ? e. Y (C) = ? e. Y (T) = ? s: A : Y Emission: Transition: G Y C Y a 0 N = ? G Y C Y a. N 0 = ? END G Y a. NN = ? N a. NY = ? a. Y 0 = ? Non- Gp. C Island T Y A N e. Y (A) = ? e. Y (G) = ? e. Y (C) = ? e. Y (T) = ? A N T N C N T N G N e. Y (A) e. Y (G) e. Y (C) e Y(G) e. Y (C) e. Y (G) e. Y (T) e. N (A) e. N (T) e. N (C) e. N (T) e. N (G) a 0 Y a. YY a. YY a. YN a. NN a. NN a. N 0 How to find the parameters e and a that maximises this probability? How if we don’t know the path?

Hidden Markov Models: Algorithms • Resumé • Evaluating P(s | M): Forward Algorithm
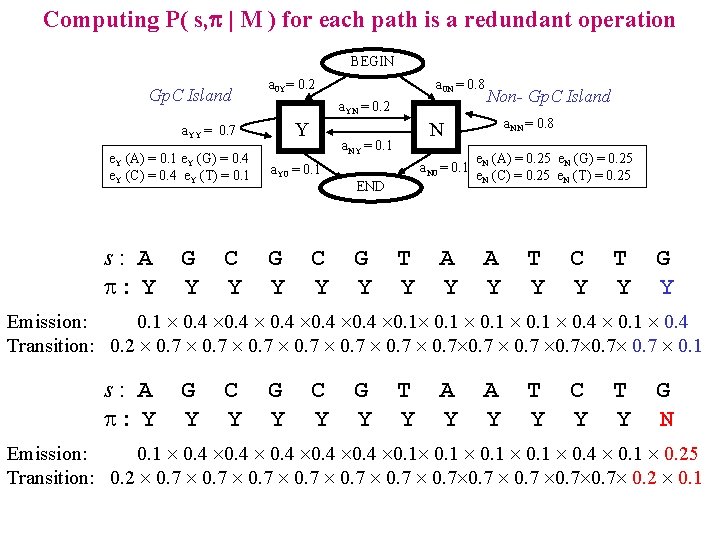
Computing P( s, p | M ) for each path is a redundant operation BEGIN Gp. C Island a 0 Y= 0. 2 a. YN = 0. 2 Y a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 s: A : Y G Y C Y a 0 N = 0. 8 G Y C Y a. N 0 = 0. 1 END G Y a. NN = 0. 8 N a. NY = 0. 1 a. Y 0 = 0. 1 Non- Gp. C Island T Y A Y e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 A Y T Y C Y T Y G Y Emission: 0. 1 0. 4 0. 1 0. 4 Transition: 0. 2 0. 7 0. 7 0. 7 0. 1 s: A : Y G Y C Y G Y T Y A Y T Y C Y T Y G N Emission: 0. 1 0. 4 0. 1 0. 4 0. 1 0. 25 Transition: 0. 2 0. 7 0. 7 0. 2 0. 1
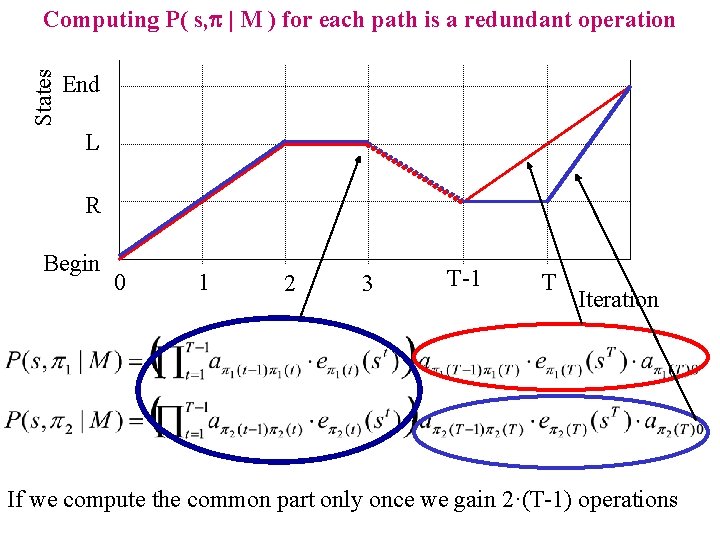
States Computing P( s, p | M ) for each path is a redundant operation End L R Begin 0 1 2 3 T-1 T Iteration If we compute the common part only once we gain 2·(T-1) operations
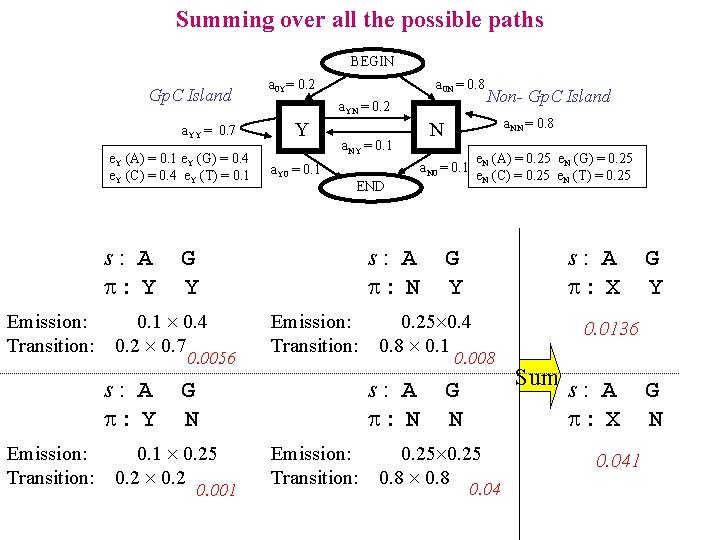
Summing over all the possible paths BEGIN Gp. C Island a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 s: A : Y Emission: Transition: a 0 N = 0. 8 a. YN = 0. 2 Y a. Y 0 = 0. 1 a. NY = 0. 1 0. 0056 Emission: Transition: 0. 001 e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 Emission: Transition: s: A : X G Y 0. 25 0. 4 0. 8 0. 1 0. 008 s: A : N G N 0. 1 0. 25 0. 2 a. NN = 0. 8 END s: A : N Non- Gp. C Island N a. N 0 = 0. 1 G Y 0. 1 0. 4 0. 2 0. 7 s: A : Y a 0 Y= 0. 2 G N 0. 25 0. 8 0. 04 G Y 0. 0136 Sum s: A : X 0. 041 G N
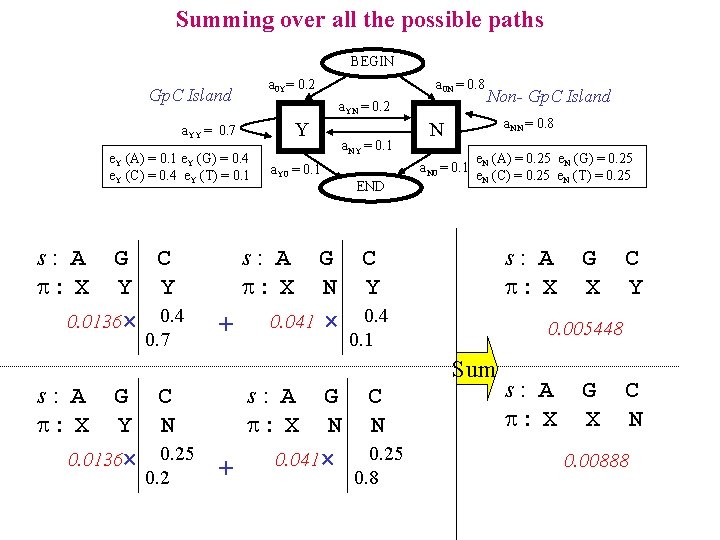
Summing over all the possible paths BEGIN a 0 Y= 0. 2 Gp. C Island a. YN = 0. 2 Y a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 s: A : X G C Y Y 0. 0136 0. 4 0. 7 s: A : X a 0 N = 0. 8 a. Y 0 = 0. 1 a. NY = 0. 1 a. NN = 0. 8 N a. N 0 = 0. 1 END Non- Gp. C Island e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 s: A : X + s: A : X G C N Y 0. 041 0. 4 C Y 0. 005448 0. 1 G C s: A G C Y N : X N N 0. 25 0. 0136 0. 25 0. 041 + 0. 2 0. 8 G X Sum s: A : X G X C N 0. 00888
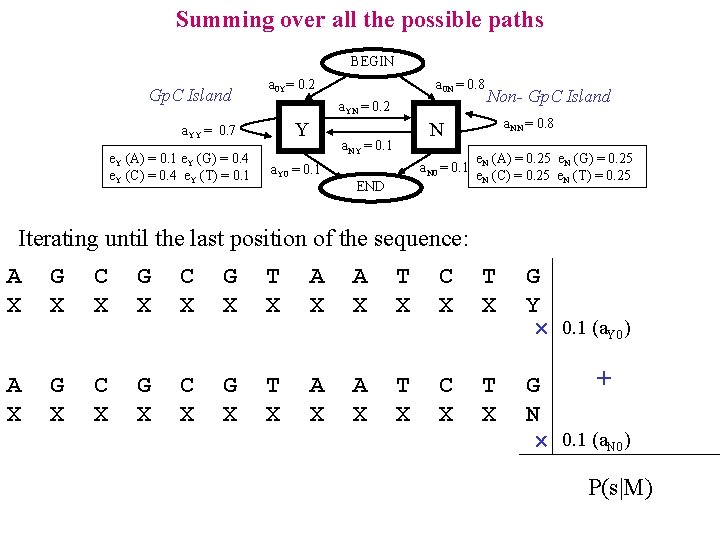
Summing over all the possible paths BEGIN Gp. C Island a 0 Y= 0. 2 a. YN = 0. 2 Y a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 a 0 N = 0. 8 a. NN = 0. 8 N a. NY = 0. 1 a. Y 0 = 0. 1 Non- Gp. C Island a. N 0 = 0. 1 END e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 Iterating until the last position of the sequence: A X G X C X G X T X A X T X C X T X G Y 0. 1 (a. Y 0) A X G X C X G X T X A X T X C X T X + G N 0. 1 (a. N 0) P(s|M)
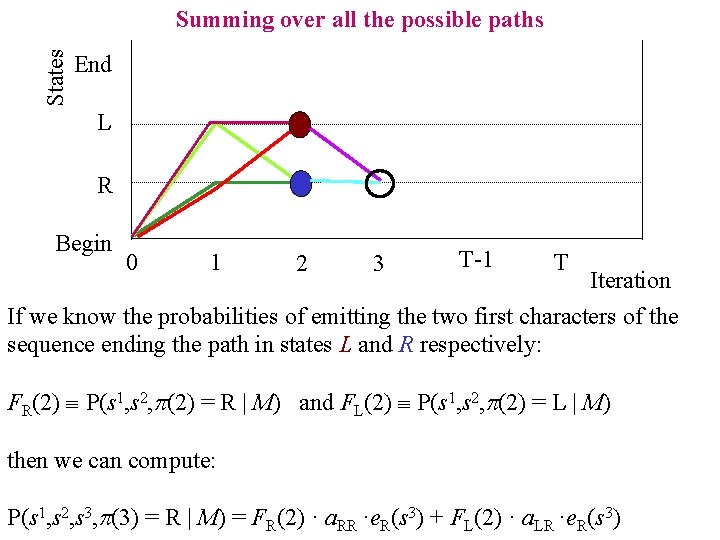
States Summing over all the possible paths End L R Begin 0 1 2 3 T-1 T Iteration If we know the probabilities of emitting the two first characters of the sequence ending the path in states L and R respectively: FR(2) P(s 1, s 2, p(2) = R | M) and FL(2) P(s 1, s 2, p(2) = L | M) then we can compute: P(s 1, s 2, s 3, p(3) = R | M) = FR(2) · a. RR ·e. R(s 3) + FL(2) · a. LR ·e. R(s 3)

Forward Algorithm On the basis of preceding observations the computation of P(s | M) can be decomposed in simplest problems For each state k and each position i in the sequence, we compute: Fk(i) = P( s 1 s 2 s 3……s i, Initialisation: FBEGIN (0) = 1 Recurrence: p (i) = k | M) Fi (0) = 0 Will be understood i BEGIN Fl ( i+1) = P( s 1 s 2…s is i+1, p (i + 1) = l ) = = k P( s 1 s 2 …s i, p (i) = k ) a k l e l ( s i+1 ) = = e l ( s i+1 ) k Fk ( i ) a k l Termination: P( s ) = P( s 1 s 2 s 3……s T, p (T + 1) = END ) = = k P( s 1 s 2 …s T , p (T) = k ) a k 0 = k Fk ( T ) a k 0
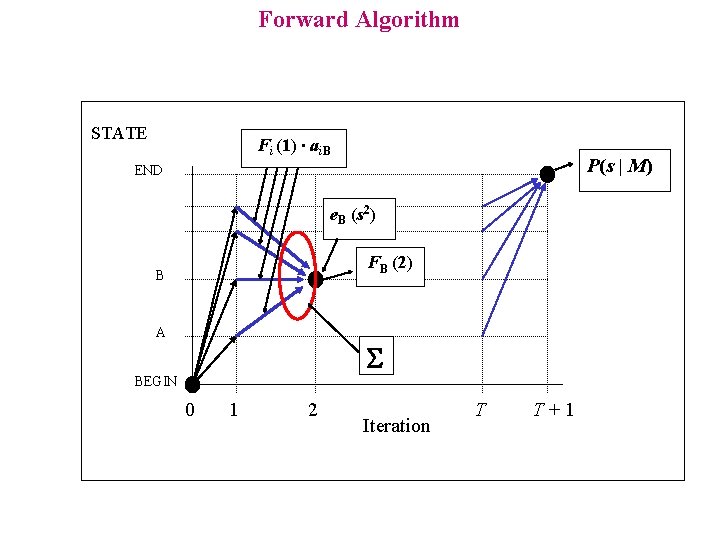
Forward Algorithm STATE Fi (1) ∙ ai. B P(s | M) END e. B (s 2) FB (2) B A S BEGIN 0 1 2 Iteration T T+1

Forward algorithm: computational complexity Naïf method P ( s | M ) = P( s, p | M ) There are N T possible paths. Each path requires about 2 T operations. The time for the computation is O( T N T ) s: A : Y G Y C Y G Y T Y A Y T Y C Y T Y G Y Emission: 0. 1 0. 4 0. 1 0. 4 Transition: 0. 2 0. 7 0. 7 0. 7 0. 1 s: A : Y G Y C Y G Y T Y A Y T Y C Y T Y G N Emission: 0. 1 0. 4 0. 1 0. 4 0. 1 0. 25 Transition: 0. 2 0. 7 0. 7 0. 2 0. 1

Forward algorithm: computational complexity ·Forward algorithm T positions, N values for each position Each element requires about 2 N product and 1 sum The time for the computation is O(T N 2) s: A : X G C Y Y 0. 0136 0. 4 0. 7 s: A : X + s: A : X G C N Y 0. 041 0. 4 C Y 0. 005448 0. 1 G C s: A G C Y N : X N N 0. 25 0. 0136 0. 25 0. 041 + 0. 2 0. 8 G X Sum s: A : X G X C N 0. 00888
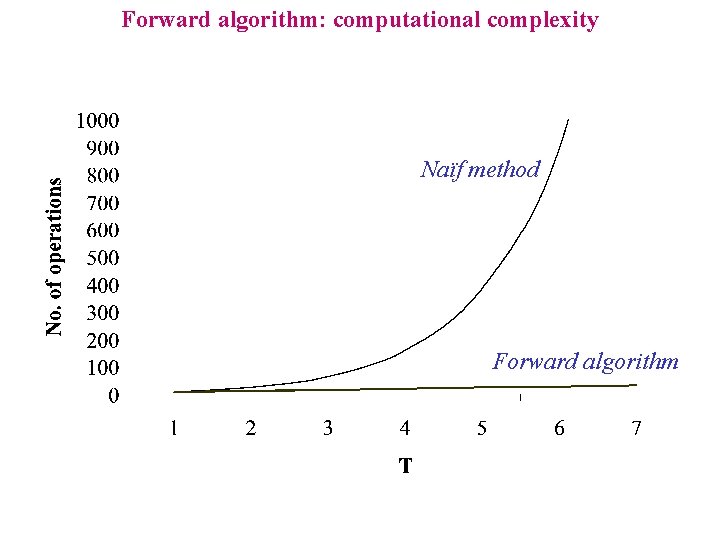
Forward algorithm: computational complexity Naïf method Forward algorithm

Hidden Markov Models: Algorithms • Resumé • Evaluating P(s | M): Forward Algorithm • Evaluating P(s | M): Backward Algorithm

Backward Algorithm Similar to the Forward algorithm: it computes P( s | M ), reconstructing the sequence from the end For each state k and each position i in the sequence, we compute: Bk(i) = P( s i+1 s i+2 s i+3……s T | p (i) = k ) Initialisation: Bk (T) = P(p (T+1) = END | p (T) = k ) = ak 0 Recurrence: Bl ( i-1) = P(s is i+1…s T | p (i - 1) = l ) = = k P(s i+1 s i+2…s T | p (i) = k) a l k e k (s i )= = k Bk ( i ) e k ( s i ) a l k Termination: P( s ) = P( s 1 s 2 s 3……s T | p (0) = BEGIN ) = = k P( s 2 …s T | p (1) = k ) a 0 k e k ( s 1 ) = = k Bk ( 1 ) a 0 k e k ( s 1 )
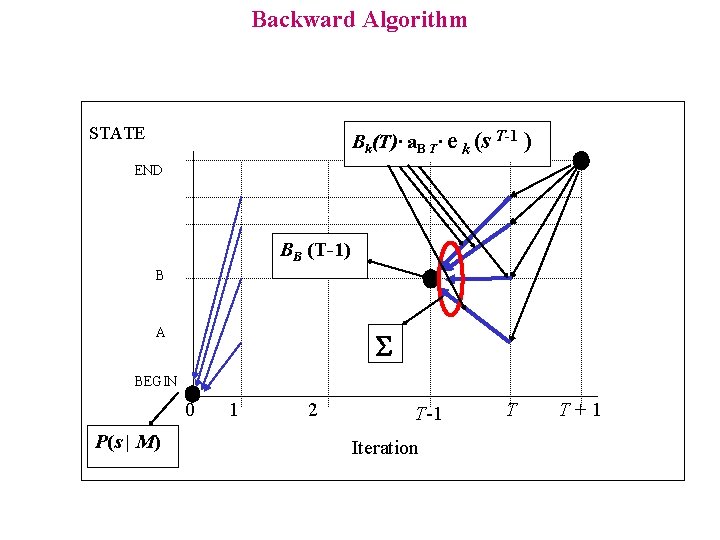
Backward Algorithm STATE Bk(T)· a. B T· e k (s T-1 ) END BB (T-1) B A S BEGIN 0 P(s | M) 1 2 T-1 Iteration T T+1

Hidden Markov Models: Algorithms • Resumé • Evaluating P(s | M): Forward Algorithm • Evaluating P(s | M): Backward Algorithm • Showing the path: Viterbi decoding
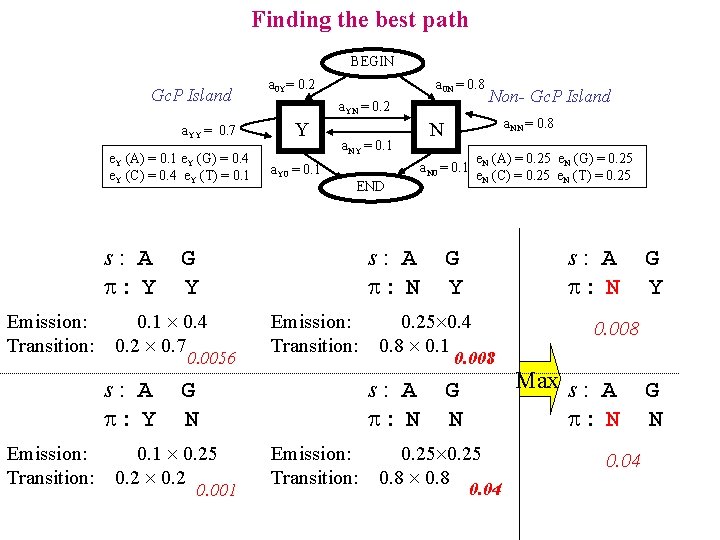
Finding the best path BEGIN Gc. P Island a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 s: A : Y Emission: Transition: a 0 N = 0. 8 a. YN = 0. 2 Y a. Y 0 = 0. 1 a. NY = 0. 1 0. 0056 Emission: Transition: 0. 001 e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 Emission: Transition: s: A : N G Y 0. 25 0. 4 0. 8 0. 1 0. 008 s: A : N G N 0. 1 0. 25 0. 2 a. NN = 0. 8 END s: A : N Non- Gc. P Island N a. N 0 = 0. 1 G Y 0. 1 0. 4 0. 2 0. 7 s: A : Y a 0 Y= 0. 2 G N 0. 25 0. 8 0. 04 G Y 0. 008 Max s : A : N 0. 04 G N
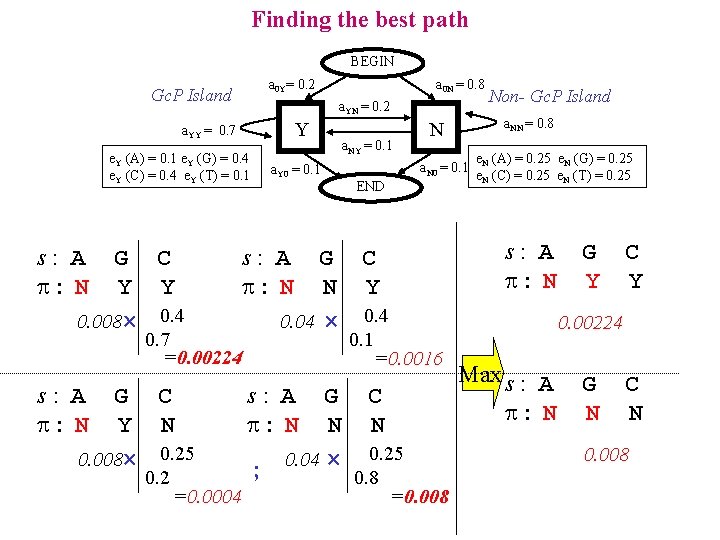
Finding the best path BEGIN a 0 Y= 0. 2 Gc. P Island a. YN = 0. 2 Y a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 s: A : N G C Y Y 0. 008 0. 4 0. 7 =0. 00224 s: A : N G C Y N 0. 008 0. 25 0. 2 =0. 0004 a 0 N = 0. 8 a. Y 0 = 0. 1 a. NY = 0. 1 a. NN = 0. 8 N a. N 0 = 0. 1 END e. N (A) = 0. 25 e. N (G) = 0. 25 e. N (C) = 0. 25 e. N (T) = 0. 25 s: A : N G C N Y 0. 04 0. 4 0. 1 Non- Gc. P Island G Y C Y 0. 00224 =0. 0016 s: A : N G C N N 0. 04 0. 25 ; 0. 8 =0. 008 Max s : A : N G N C N 0. 008
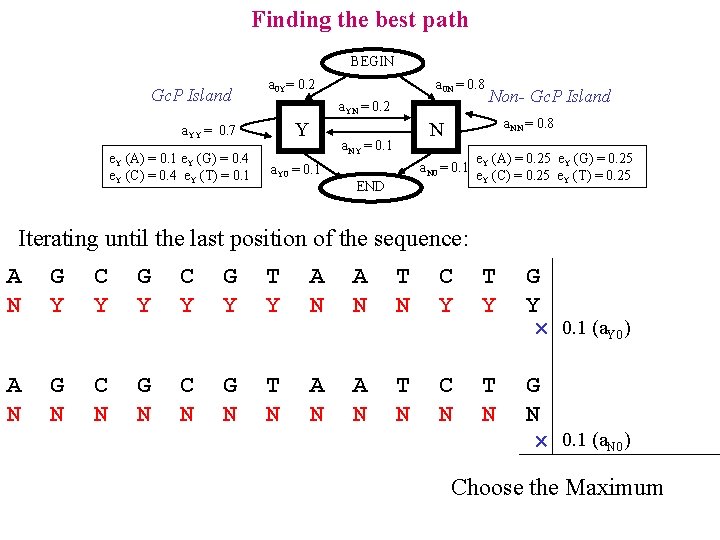
Finding the best path BEGIN Gc. P Island a 0 Y= 0. 2 a. YN = 0. 2 Y a. YY = 0. 7 e. Y (A) = 0. 1 e. Y (G) = 0. 4 e. Y (C) = 0. 4 e. Y (T) = 0. 1 a 0 N = 0. 8 a. NN = 0. 8 N a. NY = 0. 1 a. Y 0 = 0. 1 Non- Gc. P Island a. N 0 = 0. 1 END e. Y (A) = 0. 25 e. Y (G) = 0. 25 e. Y (C) = 0. 25 e. Y (T) = 0. 25 Iterating until the last position of the sequence: A N G Y C Y G Y T Y A N T N C Y T Y G Y 0. 1 (a. Y 0) A N G N C N G N T N A N T N C N T N G N 0. 1 (a. N 0) Choose the Maximum

Viterbi Algorithm p* = argmax p [ P( p , s | M ) ] The computation of P(s, p*| M) can be decomposed in simplest problems Let Vk(i) be the probability of the most probable path for generating the subsequence s 1 s 2 s 3……s i ending in the state k at iteration i Initialisation: VBEGIN (0) = 1 Recurrence: Vi (0) = 0 i BEGIN Vl ( i+1) = e l ( s i+1 ) Max k ( Vk ( i ) a k l ) ptr i ( l ) = argmax k ( Vk ( i ) a k l ) Termination: P( s, p* ) =Maxk (Vk ( T ) a k 0 ) Traceback: p* ( T ) = argmax k (Vk ( T ) a k 0 ) p* ( i-1 ) = ptr i (p* ( i ))
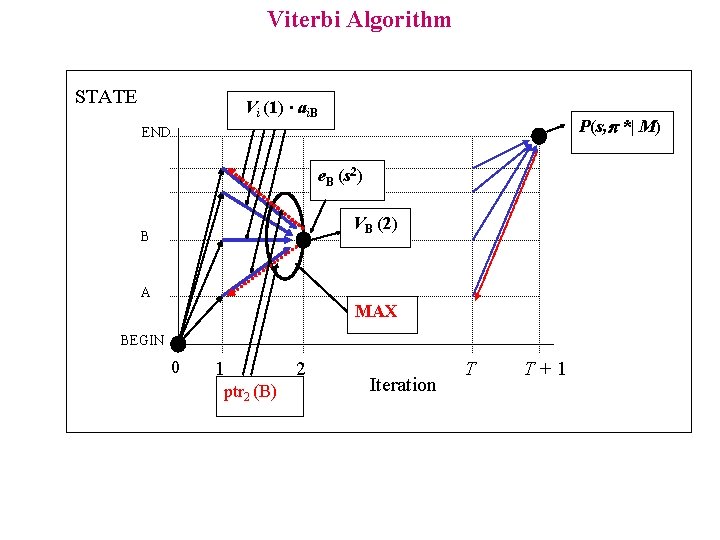
Viterbi Algorithm STATE Vi (1) ∙ ai. B P(s, *| M) END e. B (s 2) VB (2) B A MAX BEGIN 0 1 ptr 2 (B) 2 Iteration T T+1
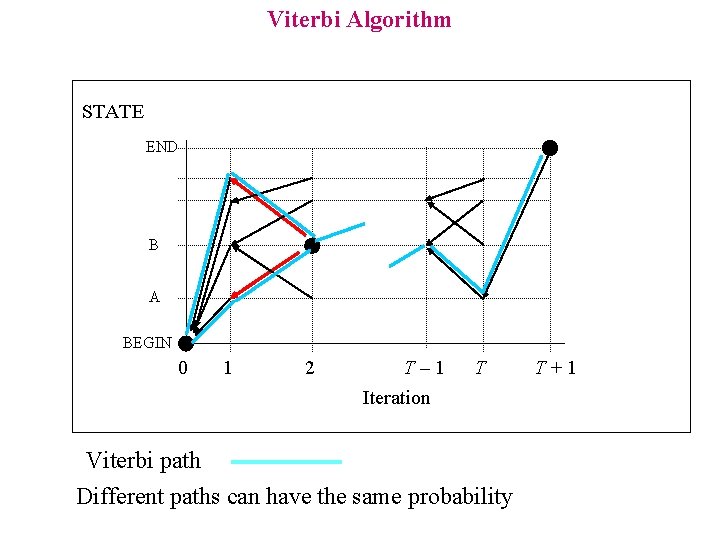
Viterbi Algorithm STATE END B A BEGIN 0 1 2 T– 1 T Iteration Viterbi path Different paths can have the same probability T+1

Hidden Markov Models: Algorithms • Resumé • Evaluating P(s | M): Forward Algorithm • Evaluating P(s | M): Backward Algorithm • Showing the path: Viterbi decoding • Showing the path: A posteriori decoding • Training a model: EM algorithm
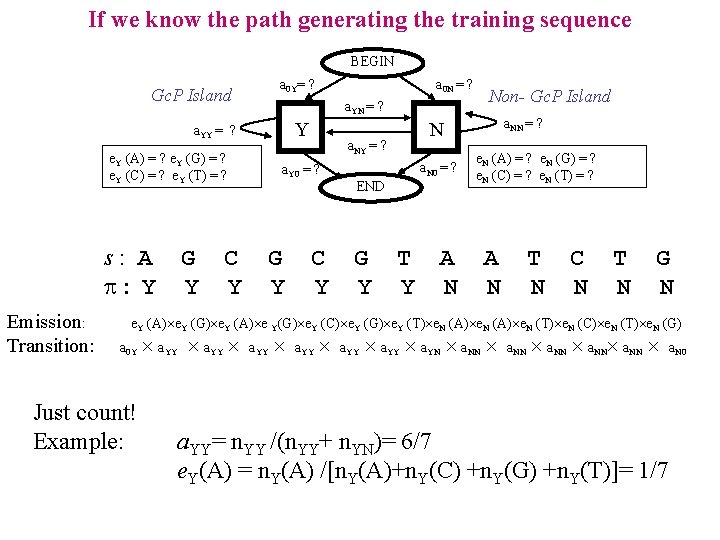
If we know the path generating the training sequence BEGIN a 0 Y= ? Gc. P Island a. YN = ? Y a. YY = ? e. Y (A) = ? e. Y (G) = ? e. Y (C) = ? e. Y (T) = ? s: A : Y Emission: Transition: G Y C Y a 0 N = ? G Y C Y a. N 0 = ? END G Y a. NN = ? N a. NY = ? a. Y 0 = ? Non- Gc. P Island T Y A N e. N (A) = ? e. N (G) = ? e. N (C) = ? e. N (T) = ? A N T N C N T N G N e. Y (A) e. Y (G) e. Y (A) e Y(G) e. Y (C) e. Y (G) e. Y (T) e. N (A) e. N (T) e. N (C) e. N (T) e. N (G) a 0 Y Just count! Example: a. YY a. YY a. YN a. NN a. NN a. N 0 a. YY= n. YY /(n. YY+ n. YN)= 6/7 e. Y(A) = n. Y(A) /[n. Y(A)+n. Y(C) +n. Y(G) +n. Y(T)]= 1/7

Expectation-Maximisation algorithm We need to estimate the Maximum Likelihood parameters when the paths generating the training sequences are unknown ML = argmax [P ( s | , M)] Given a model with parameters 0 the EM algorithm finds new parameters that increase the likelihood of the model: P( s | ) > P( s| 0 )

Expectation-Maximisation algorithm s: A G C G T A A T C T G 1: Y Y Y Y 2: Y Y Y N 3: Y Y Y N Y 4: Y Y Y N N ………………………………. . . Given a path p we can count the number of transition between states k and l: Ak, l(p) the number on emissions of character c from state k: Ek (c, p) We can compute the expected values over all the paths Ak, l = P(p | s, 0) · Ak, l(p) Ek (c) = P(p | s, 0) · Ek (c, p) The updated parameters are: ak, l = Ak, l N m=1 Ak, m ek(c) = Ek (c) c Ek (c)

Expectation-Maximisation algorithm We need to estimate the Maximum Likelihood parameters when the paths generating the training sequences are unknown ML = argmax [P ( s | , M)] Given a model with parameters 0 the EM algorithm finds new parameters that increase the likelihood of the model: P( s | ) > P( s| 0 ) or equivalentely log P( s | ) > log P( s| 0 )

Expectation-Maximisation algorithm log P( s | ) = log P(s, p | ) - log P(p | s, ) Multiplying for P(p | s, 0) and summing over all the possible paths: log P( s | ) = = P(p | s, 0) ·log P(s, p | ) - P(p | s, 0) · log P(p | s, ) Q( | 0) : Expectation value of log P(s, p | ) over all the “current” paths log P( s | ) - log P(s | 0) = = Q( | 0) - Q( 0| 0) + Q( | 0) - Q( 0| 0) 0

Expectation-Maximisation algorithm The EM algorithm is an iterative process Each iteration performs two steps: E-step: evaluation of Q( | 0) = P(p | s, 0) ·log P(s, p | ) M-step: Maximisation of Q( | 0) over all It does NOT assure to converge to the GLOBAL Maximum Likelihood

Baum-Welch implementation of the EM algorithm E-step: Q( | 0) = P(p | s, 0) ·log P(s, p | ) = a 0, (1) · P N T i = 1 a (i), (i+1) N Ak. l ( ) = Pk = 0 Pl = 1 ak, l ·e (i)(si) = N · Pk = 1 Pc C ek (c) Ek (c, ) Ak, l(p): number of transitions between the states k and l in path Ek (c, p): number of emissions of character c in path Ak, l = P(p | s, 0) · Ak, l(p) Ek (c) = P(p | s, 0) · Ek (c, p) So: N N Expected values over all the “actual” paths N Q( | = k = 0 l = 1 Ak, l · log ak, l + k = 1 c C Ek (c) ·log ek (c) 0)

Baum-Welch implementation of the EM algorithm M-step: For any state k and l, with l ak, l = 1 For any state k and character c, with c ek(c) = 1 By means of Lagrange’s multipliers techniques, we can solve the system ak, l = ek(c) = Ak, l N m=1 Ak, m Ek (c) N m=1 Em (c)

Baum-Welch implementation of the EM algorithm How to compute the expected number of transitions and emissions over all the paths Fk(i) = P( s 1 s 2 s 3……s i, p (i) = k ) Bk(i) = P( s i+1 s i+2 s i+3……s T | p (i) = k ) Ak, l i +1) B (i + 1) F (i ) a e (s kl l l = P(p (i ) = k , p ( i +1) = l | s, ) = i k P (s ) Ek (c) = P( s = c , p (i ) = k | s, ) = i s = c Fk(i ) Bl(i) i P (s )
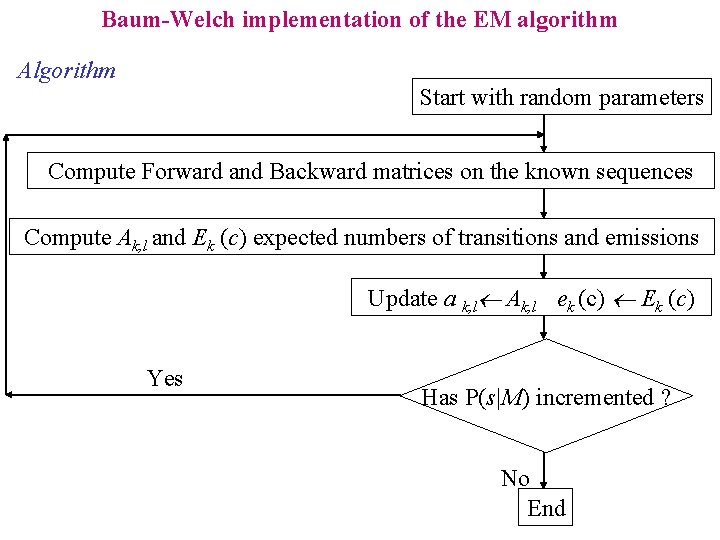
Baum-Welch implementation of the EM algorithm Algorithm Start with random parameters Compute Forward and Backward matrices on the known sequences Compute Ak, l and Ek (c) expected numbers of transitions and emissions Update a k, l Ak, l ek (c) Ek (c) Yes Has P(s|M) incremented ? No End

Profile HMMs • HMMs for alignments

How to align? M 0 M 1 M 2 M 3 M 5 M 4 Each state represent a position in the alignment. A M 0 C M 1 G M 2 G M 3 T M 4 A M 5 A M 0 C M 1 G M 2 A M 3 T M 4 C M 5 A M 0 ?
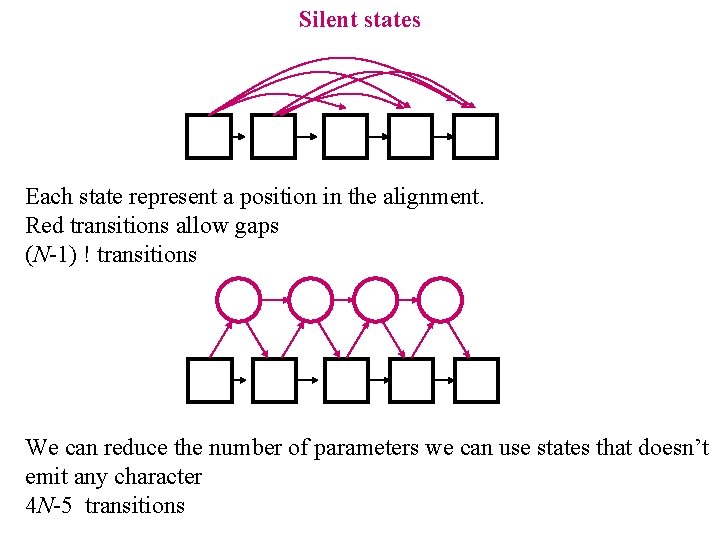
Silent states Each state represent a position in the alignment. Red transitions allow gaps (N-1) ! transitions We can reduce the number of parameters we can use states that doesn’t emit any character 4 N-5 transitions
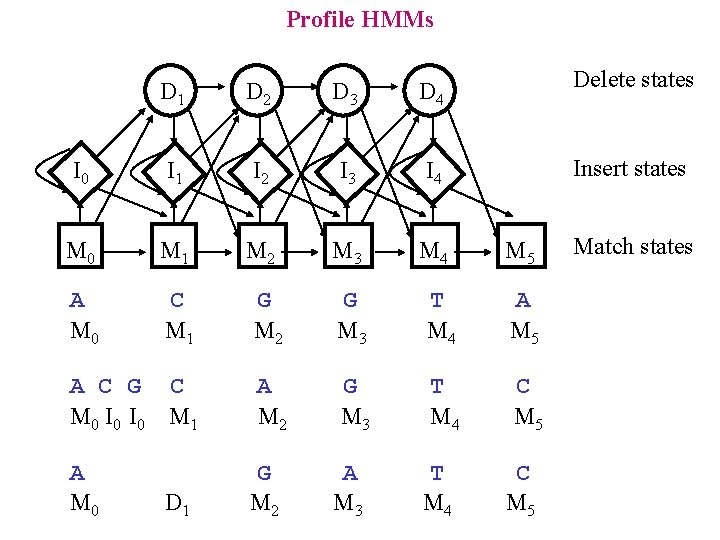
Profile HMMs Delete states D 1 D 2 D 3 D 4 I 0 I 1 I 2 I 3 I 4 M 0 M 1 M 2 M 3 M 4 A M 0 C M 1 G M 2 G M 3 T M 4 A M 5 A C G C M 0 I 0 M 1 A M 2 G M 3 T M 4 C M 5 A M 0 G M 2 D 1 A M 3 T M 4 Insert states M 5 C M 5 Match states
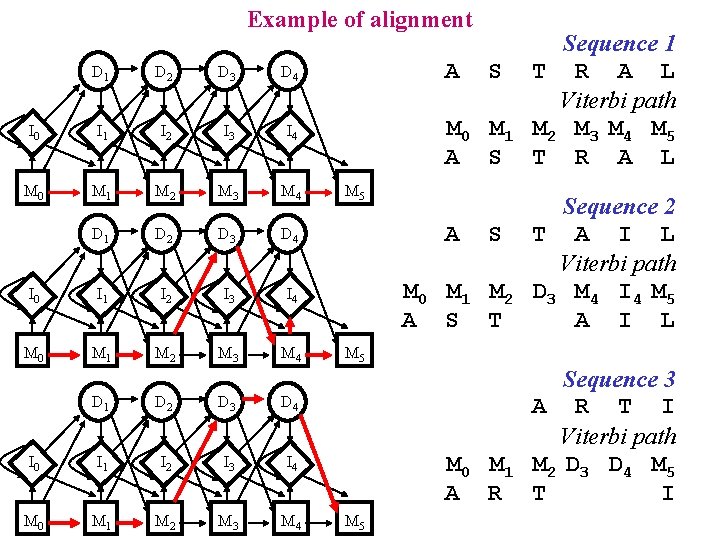
Example of alignment D 1 D 2 D 3 D 4 I 0 I 1 I 2 I 3 I 4 M 0 M 1 M 2 M 3 M 4 A S M 0 M 1 A S M 5 Sequence 1 T R A L Viterbi path M 2 M 3 M 4 M 5 T R A L M 0 M 1 M 2 A S T Sequence 2 T A I L Viterbi path D 3 M 4 I 4 M 5 A I L M 0 M 1 A R Sequence 3 A R T I Viterbi path M 2 D 3 D 4 M 5 T I A S M 5
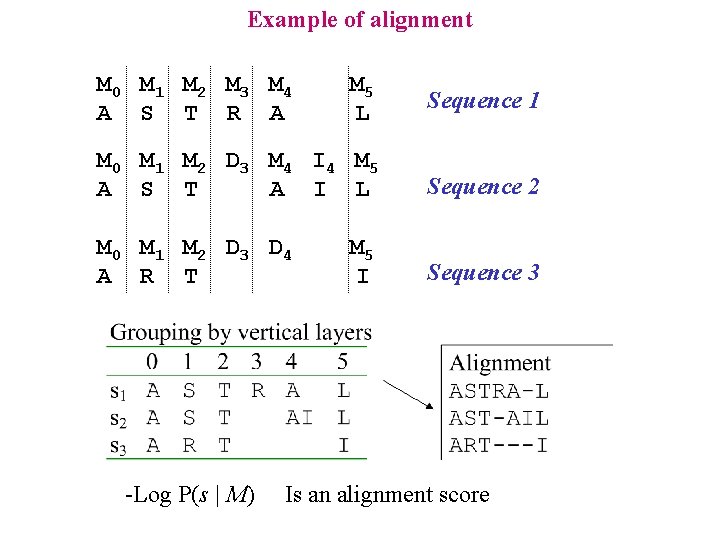
Example of alignment M 0 M 1 M 2 M 3 M 4 A S T R A M 5 L Sequence 1 M 0 M 1 M 2 D 3 M 4 I 4 M 5 A S T A I L Sequence 2 M 0 M 1 M 2 D 3 D 4 A R T Sequence 3 -Log P(s | M) M 5 I Is an alignment score

Searching for a structural/functional pattern in protein sequence Zn binding loop: C C C D C C H H H C C C I L I L C C D C R K S T S K I I L I I I Cysteines can be replaced by an Aspartic Acid, but only ONCE for each sequence C C C
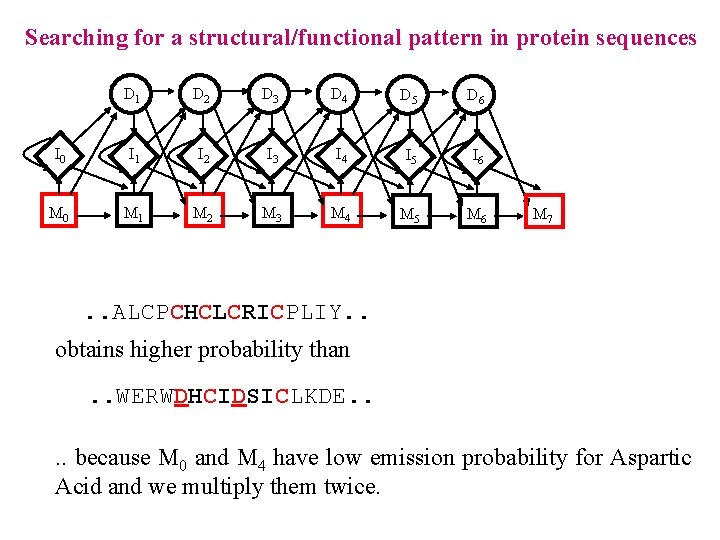
Searching for a structural/functional pattern in protein sequences D 1 D 2 D 3 D 4 D 5 D 6 I 0 I 1 I 2 I 3 I 4 I 5 I 6 M 0 M 1 M 2 M 3 M 4 M 5 M 6 M 7 . . ALCPCHCLCRICPLIY. . obtains higher probability than. . WERWDHCIDSICLKDE. . because M 0 and M 4 have low emission probability for Aspartic Acid and we multiply them twice.

Profile HMMs • HMMs for alignments • Example on globins

Structural alignment of globins Bashdorf D, Chothia C & Lesk AM, (1987) Determinants of a protein fold: unique features of the globin amino sequence. J. Mol. Biol. 196, 199 -216

Alignment of globins reconstructed with profile HMMs Krogh A, Brown M, Mian IS, Sjolander K & Haussler D (1994) Hidden Markov Models in computational biology: applications to protein modelling. J. Mol. Biol. 235, 1501 -1531
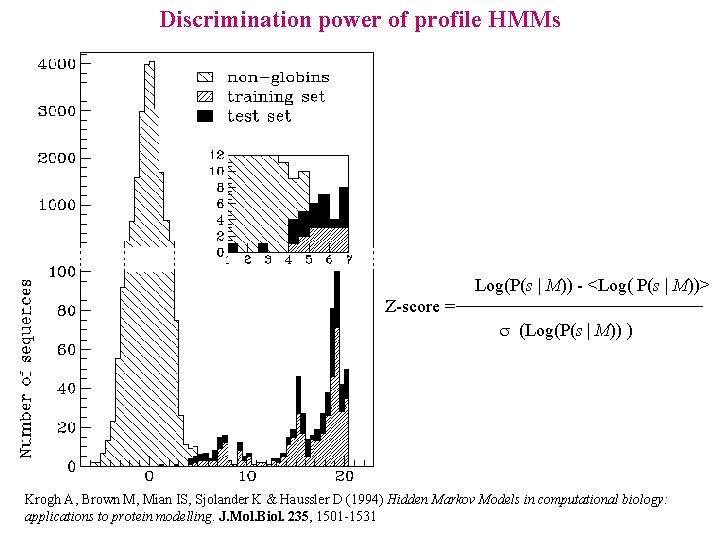
Discrimination power of profile HMMs Log(P(s | M)) - <Log( P(s | M))> Z-score = s (Log(P(s | M)) ) Krogh A, Brown M, Mian IS, Sjolander K & Haussler D (1994) Hidden Markov Models in computational biology: applications to protein modelling. J. Mol. Biol. 235, 1501 -1531

Profile HMMs • HMMs for alignments • Example on globins • Other applications
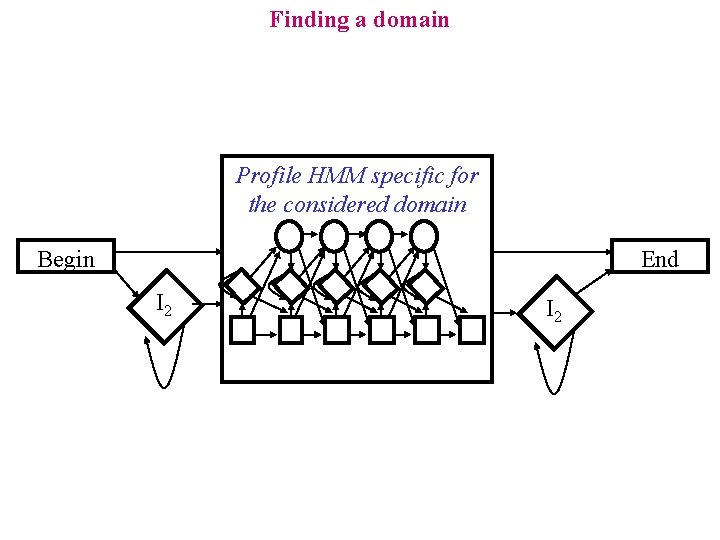
Finding a domain Profile HMM specific for the considered domain Begin End I 2
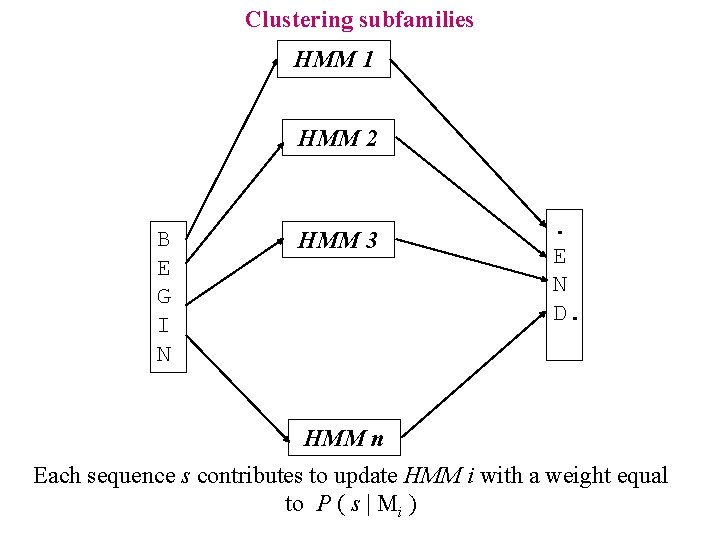
Clustering subfamilies HMM 1 HMM 2 B E G I N HMM 3 . E N D. HMM n Each sequence s contributes to update HMM i with a weight equal to P ( s | Mi )

Profile HMMs • HMMs for alignments • Example on globins • Other applications • Available codes and servers
HMMER at WUSTL: http: //hmmer. wustl. edu/ Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14: 755 -763
HMMER applications: http: //www. sanger. ac. uk/Software/Pfam/
SAM at UCSD: http: //www. soe. ucsc. edu/research/compbio/sam. html Krogh A, Brown M, Mian IS, Sjolander K & Haussler D (1994) Hidden Markov Models in computational biology: applications to protein modelling. J. Mol. Biol. 235, 1501 -1531
SAM applications: http: //www. cse. ucsc. edu/research/compbio/HMM-apps/T 02 -query. html
HMMPRO: http: //www. netid. com/html/hmmpro. html Pierre Baldi, Net-ID

HMMs for Mapping problems • Mapping problems in protein prediction
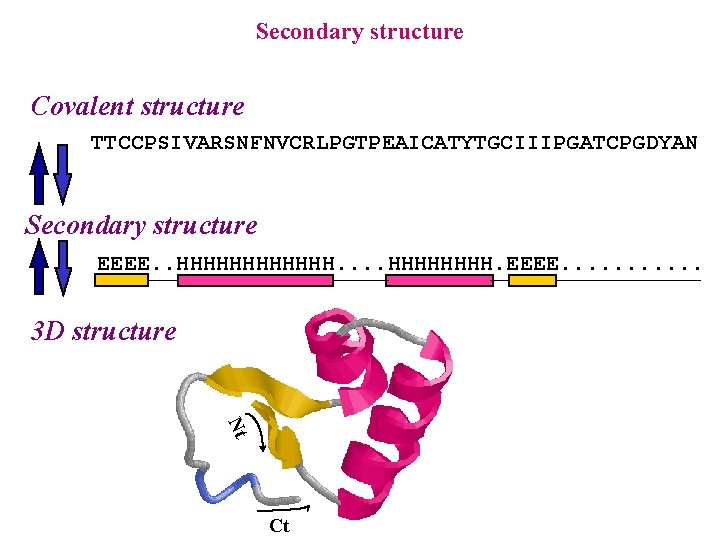
Secondary structure Covalent structure TTCCPSIVARSNFNVCRLPGTPEAICATYTGCIIIPGATCPGDYAN Secondary structure EEEE. . HHHHHH. EEEE. . . 3 D structure Nt Ct
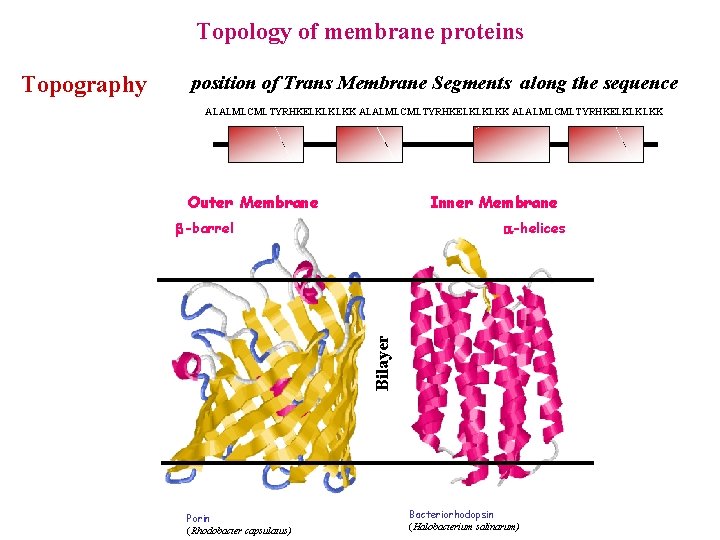
Topology of membrane proteins position of Trans Membrane Segments along the sequence ALALMLCMLTYRHKELKLKLKK Outer Membrane Inner Membrane -barrel -helices Bilayer Topography Porin (Rhodobacter capsulatus) Bacteriorhodopsin (Halobacterium salinarum)

HMMs for Mapping problems • Mapping problems in protein prediction • Labelled HMMs
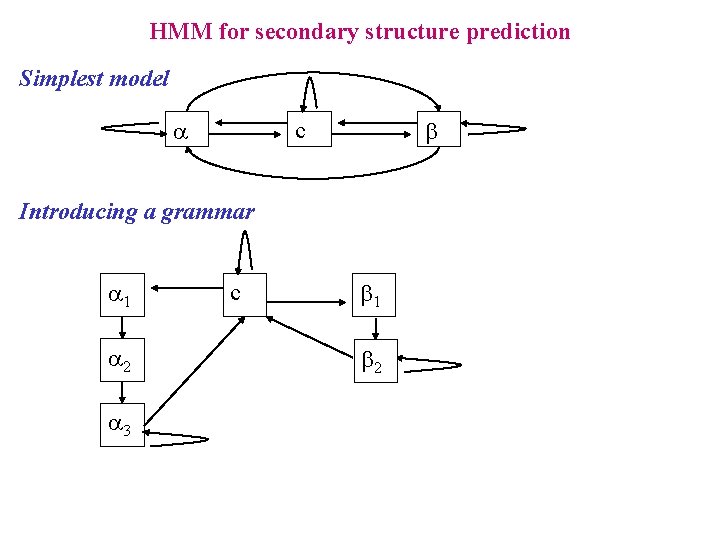
HMM for secondary structure prediction Simplest model a b c Introducing a grammar a 1 a 2 a 3 c b 1 b 2
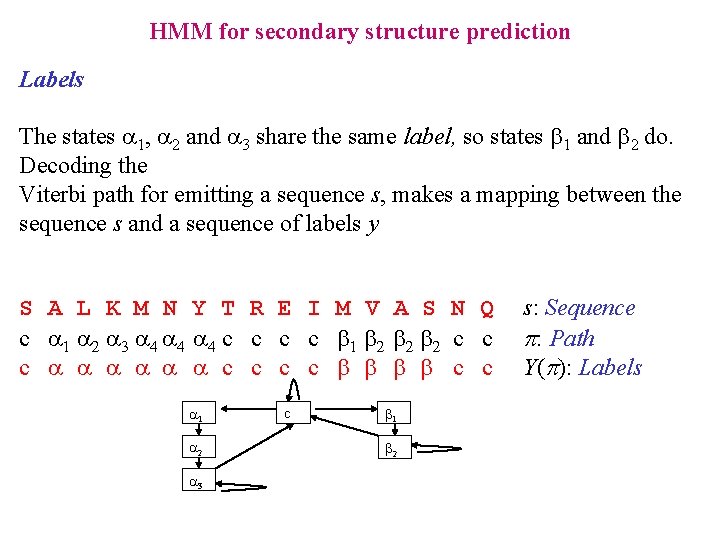
HMM for secondary structure prediction Labels The states a 1, a 2 and a 3 share the same label, so states b 1 and b 2 do. Decoding the Viterbi path for emitting a sequence s, makes a mapping between the sequence s and a sequence of labels y S A L K M N Y T R E I M V A S N Q c a 1 a 2 a 3 a 4 a 4 c c b 1 b 2 b 2 c c c a a a c c b b c c a 1 a 2 a 3 c b 1 b 2 s: Sequence p: Path Y(p): Labels

Computing P(s, y | M) Only the path whose labelling is y have to be considered in the sum In Forward and Backward algorithms it means to set Fk(i) = 0, Bk(i) = 0 if Y(k) yi S A L K M N Y T R E I M V A S N Q s: Sequence c a a a c c b b c c y: Labels 1 2 3 4 1 2 c a a b b c States Labelling

Baum-Welch training algorithm for labelled HMMs Given a set of known labelled sequences (e. g. amino acid sequences and their native secondary structure) we want to find the parameters of the model, without knowing the generating paths: ML = argmax [P ( s, y | , M)] The algorithm is the same as in the non-labelled case if we use the Forward and Backward matrices defined in the last slide. Supervised learning of the mapping

HMMs for Mapping problems • Mapping problems in protein prediction • Labelled HMMs • Duration modelling

Self loops and geometric decay p Begin 1 -p End P(l) p=0. 9 P(l) = p l-1 ·( 1 -p ) p=0. 5 p=0. 1 l The length distribution of the generated segments is always exp-like
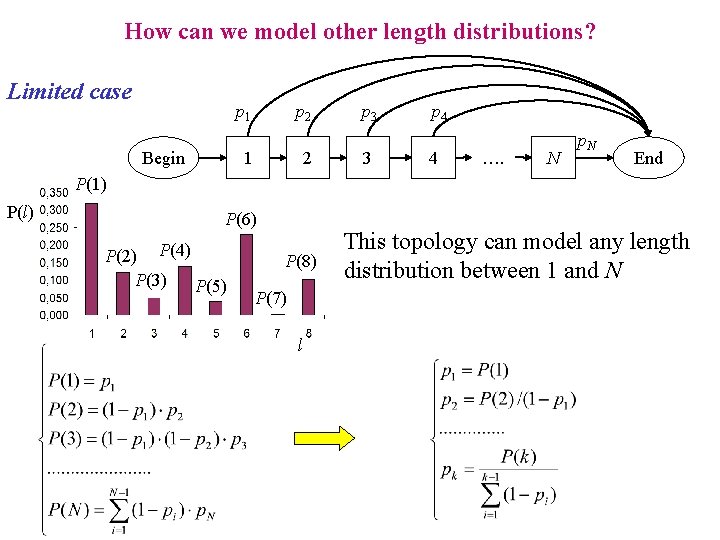
How can we model other length distributions? Limited case p 1 Begin p 2 1 2 p 3 3 p 4 4 …. N p. N End P(1) P(l) P(6) P(2) P(4) P(3) P(5) P(8) P(7) l This topology can model any length distribution between 1 and N
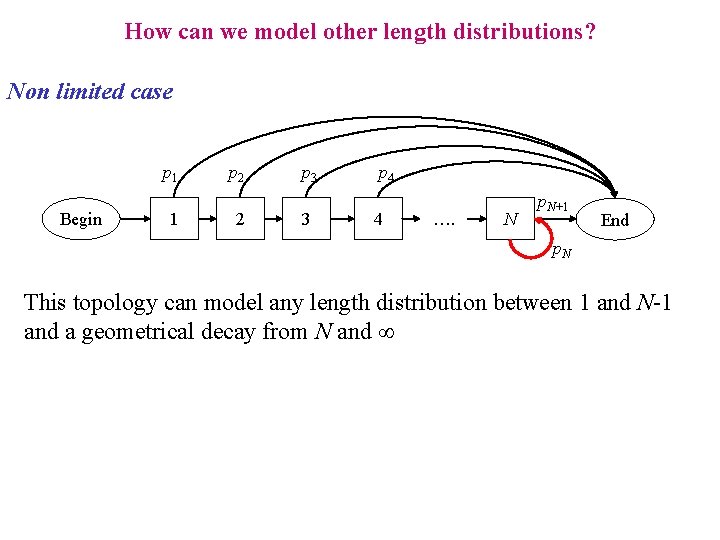
How can we model other length distributions? Non limited case p 1 Begin 1 p 2 2 p 3 3 p 4 4 …. N p. N+1 End p. N This topology can model any length distribution between 1 and N-1 and a geometrical decay from N and
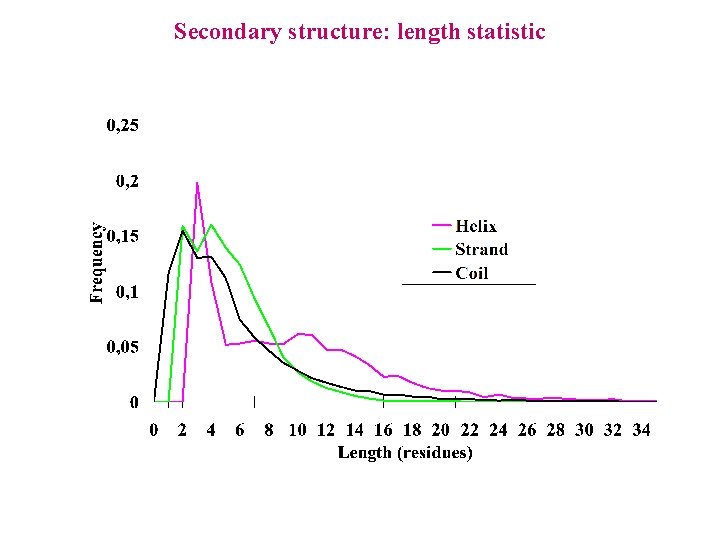
Secondary structure: length statistic
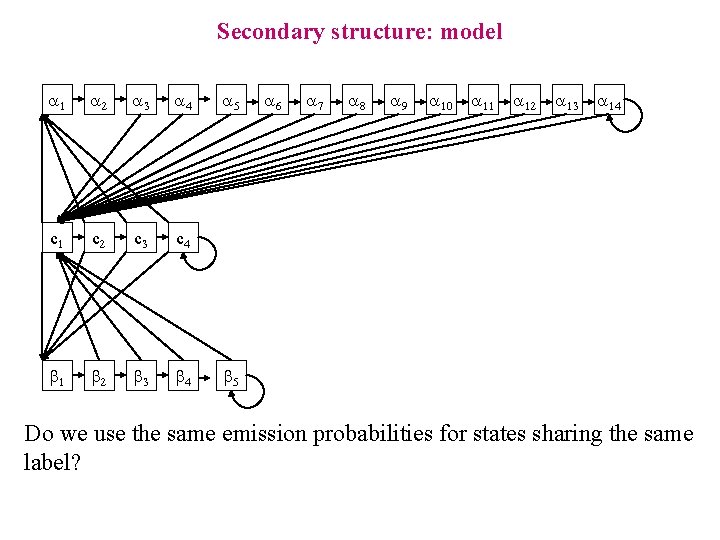
Secondary structure: model a 1 a 2 a 3 a 4 c 1 c 2 c 3 c 4 b 1 b 2 b 3 b 4 a 5 a 6 a 7 a 8 a 9 a 10 a 11 a 12 a 13 a 14 b 5 Do we use the same emission probabilities for states sharing the same label?

HMMs for Mapping problems • Mapping problems in protein prediction • Labelled HMMs • Duration modelling • Models for membrane proteins
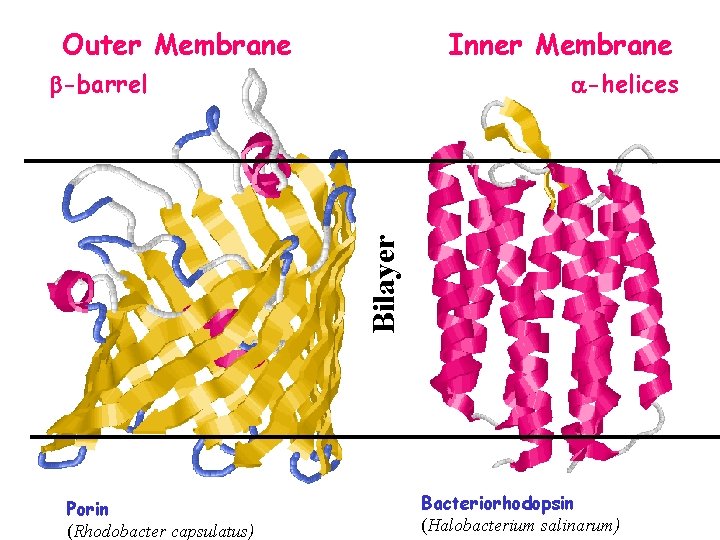
Outer Membrane Inner Membrane -helices Bilayer -barrel Porin (Rhodobacter capsulatus) Bacteriorhodopsin (Halobacterium salinarum)
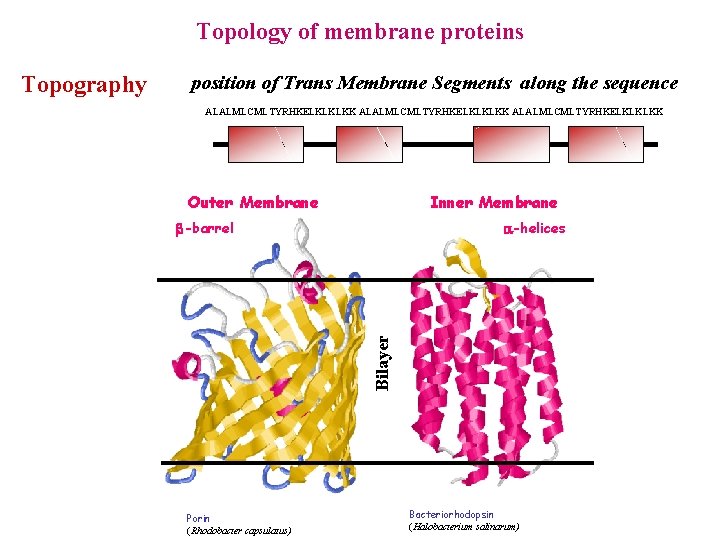
Topology of membrane proteins position of Trans Membrane Segments along the sequence ALALMLCMLTYRHKELKLKLKK Outer Membrane Inner Membrane -barrel -helices Bilayer Topography Porin (Rhodobacter capsulatus) Bacteriorhodopsin (Halobacterium salinarum)
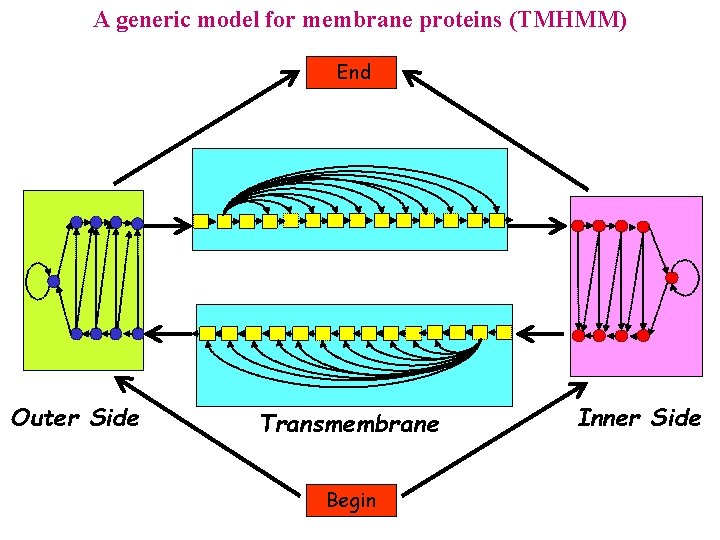
A generic model for membrane proteins (TMHMM) End Outer Side Transmembrane Begin Inner Side
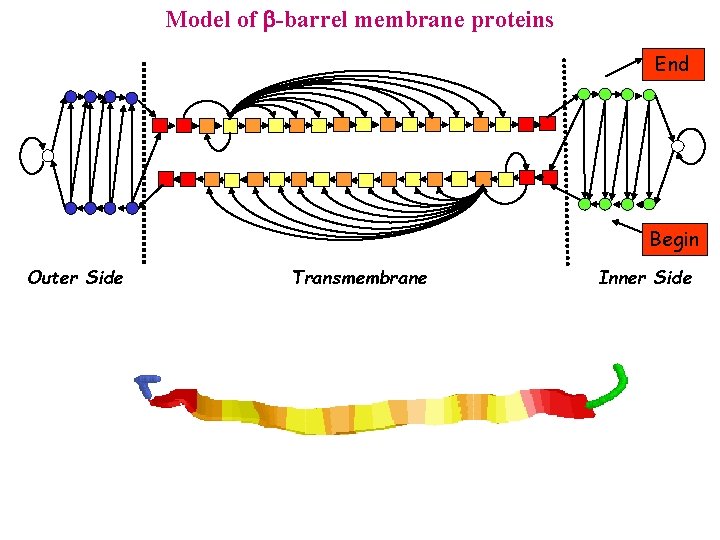
Model of -barrel membrane proteins End Begin Outer Side Transmembrane Inner Side
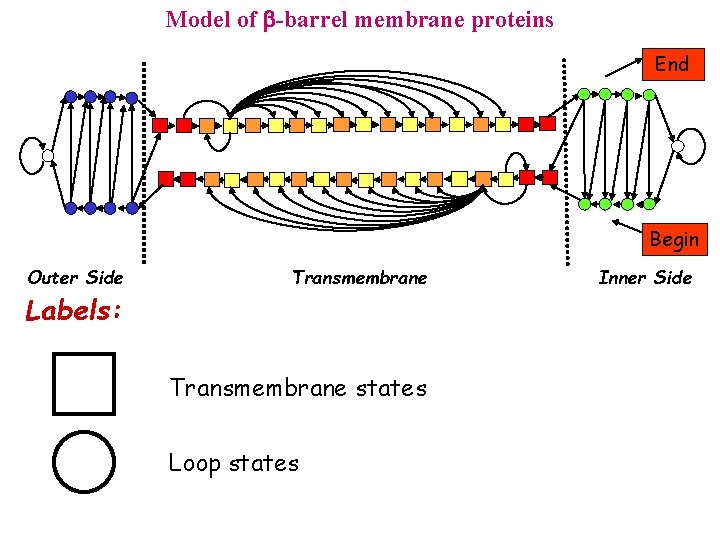
Model of -barrel membrane proteins End Begin Outer Side Labels: Transmembrane states Loop states Inner Side
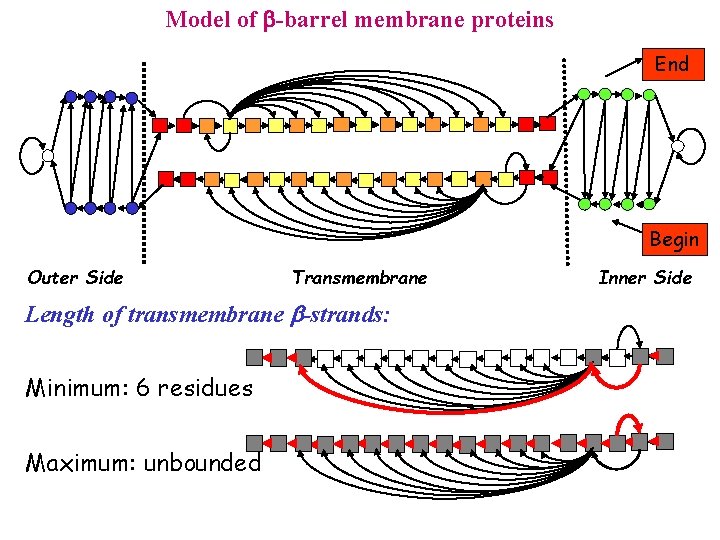
Model of -barrel membrane proteins End Begin Outer Side Transmembrane Length of transmembrane b-strands: Minimum: 6 residues Maximum: unbounded Inner Side
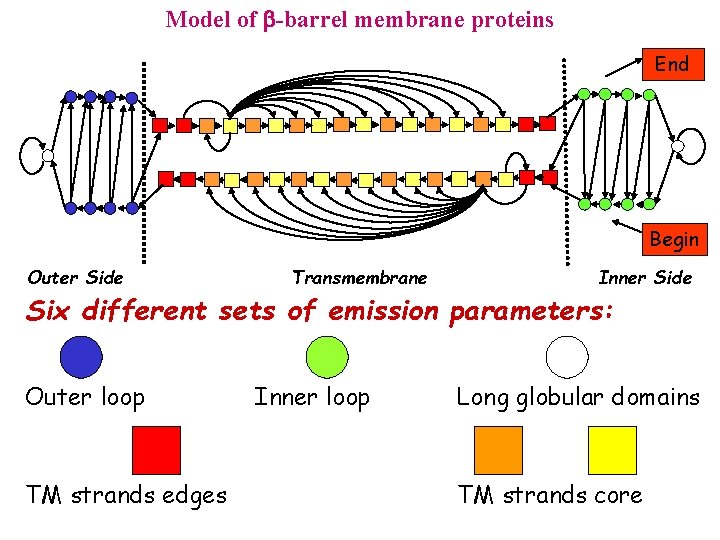
Model of -barrel membrane proteins End Begin Outer Side Transmembrane Inner Side Six different sets of emission parameters: Outer loop TM strands edges Inner loop Long globular domains TM strands core
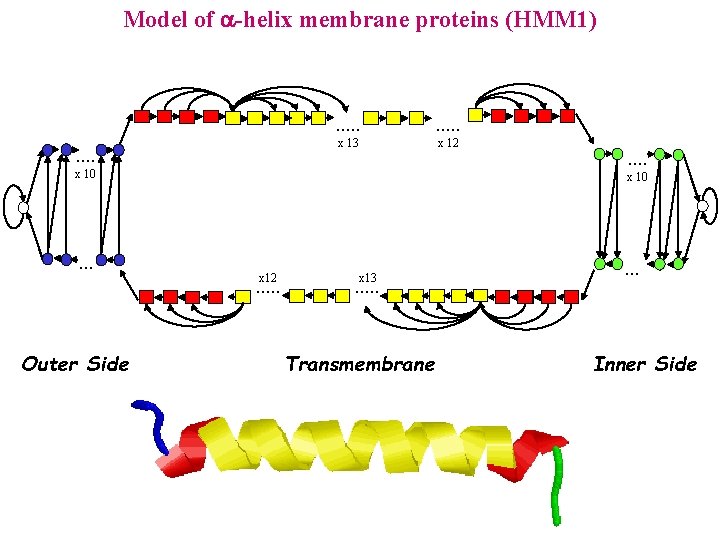
Model of -helix membrane proteins (HMM 1) . . . x 13 x 12 x 10 . . . x 10 x 12 . . . Outer Side . . x 13 . . . Transmembrane . . . Inner Side
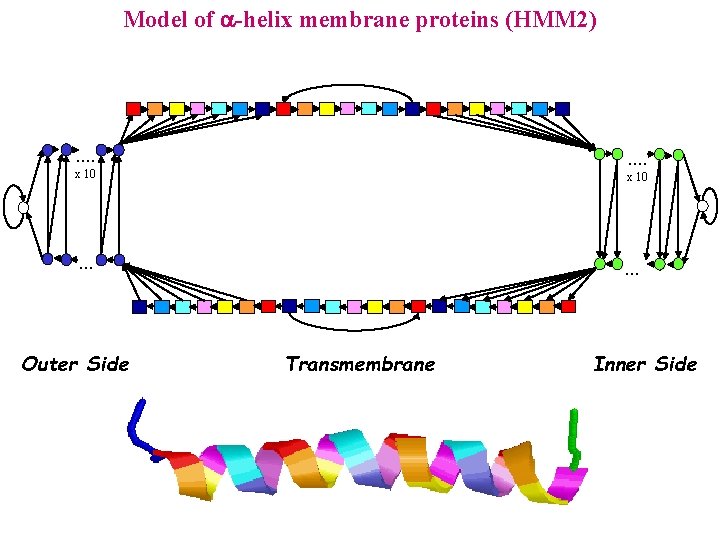
Model of -helix membrane proteins (HMM 2) . . . . x 10 . . . Outer Side . . . Transmembrane Inner Side
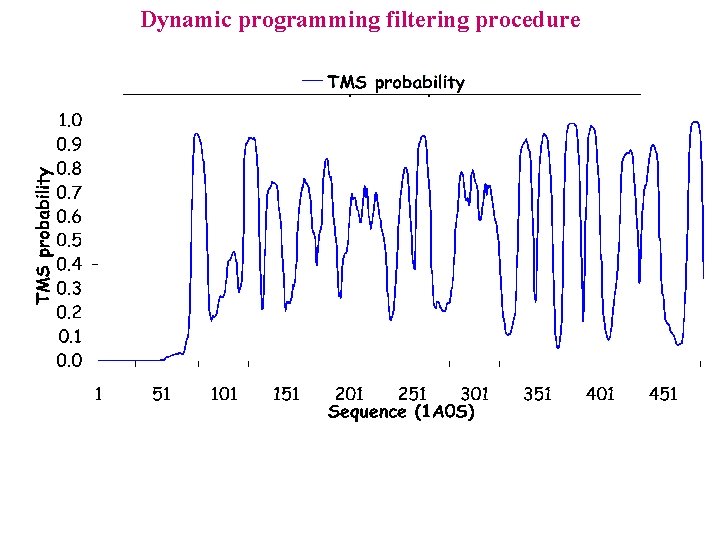
Dynamic programming filtering procedure
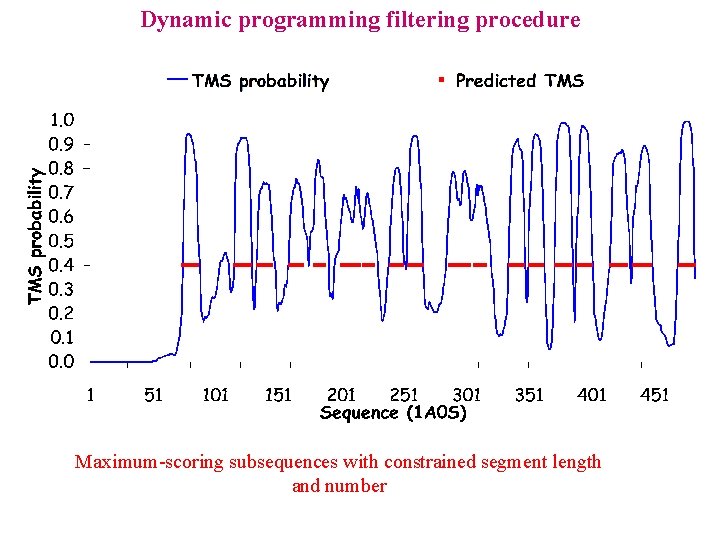
Dynamic programming filtering procedure Maximum-scoring subsequences with constrained segment length and number
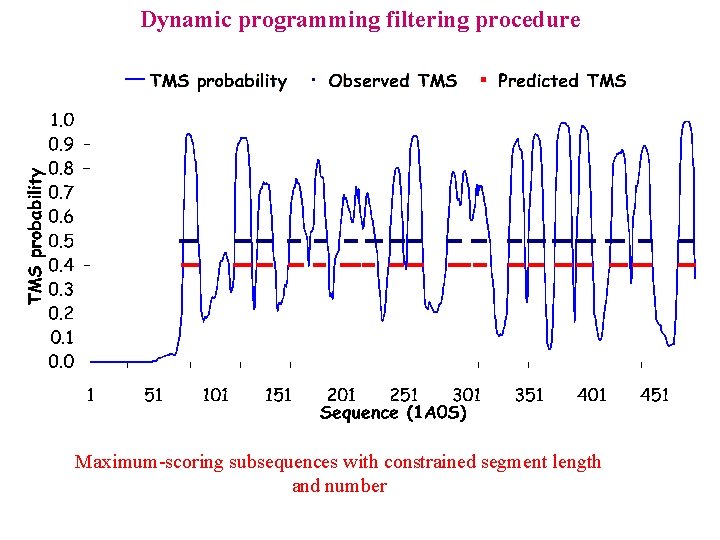
Dynamic programming filtering procedure Maximum-scoring subsequences with constrained segment length and number

Predictors of alpha-transmembrane topology www. cbs. dtu. dk/services/TMHMM

Hybrid systems: Basics • Sequence profile based HMMs

Sequence profile MSA Sequence profiles 1 2 3 4 5 6 7 8 9 10 A C D E F G H K I L M N P Q R S T V W Y Y Y Y G T T K R R K R Y K K D D D D G G G Y Y Y Y F Y Y H Q Q V Q N Q G G G 0 0 0 10 0 0 20 0 0 70 0 0 0 40 0 0 0 50 0 10 0 70 0 0 30 0 0 0 0 10 0 0 0 90 0 0 30 10 0 40 0 10 0 0 S T S S F T T F F G G D D D H D L L L 0 0 0 0 33 0 0 100 0 0 0 0 33 0 0 0 0 0 60 0 10 0 0 30 0 0 K Q H H Q Q H I I I K K K K K N N N G G G N A T T T E D E E S E D E E E 0 0 0 30 0 10 100 30 0 0 0 0 0 0 0 70 0 30 0 0 0 0 10 0 0 50 0 0 10 0 0 30 0 0 20 70 0 0 10 0 0 L L L S L T T T sequence position T T T K K K 0 0 0 0 0 0 100 0 0 0 60 0 0 0 10 0 0 30 100 0 0
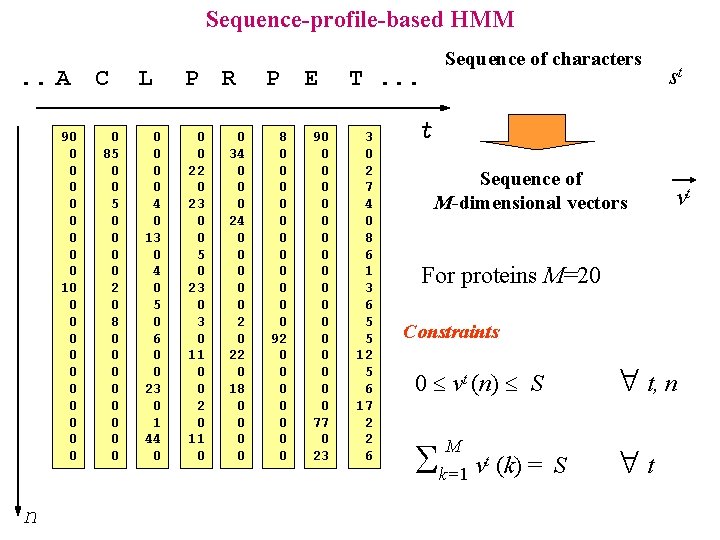
Sequence-profile-based HMM. . A C 90 0 0 0 0 10 0 0 n 0 85 0 0 0 0 2 0 8 0 0 0 0 L 0 0 4 0 13 0 4 0 5 0 6 0 0 23 0 1 44 0 P R 0 0 22 0 23 0 0 5 0 23 0 11 0 0 2 0 11 0 0 34 0 0 0 24 0 0 0 22 0 18 0 0 P E 8 0 0 0 92 0 0 0 0 90 0 0 0 0 77 0 23 Sequence of characters T. . . 3 0 2 7 4 0 8 6 1 3 6 5 5 12 5 6 17 2 2 6 st t Sequence of M-dimensional vectors vt For proteins M=20 Constraints t, n 0 vt (n) S k=1 v (k) = M t S t
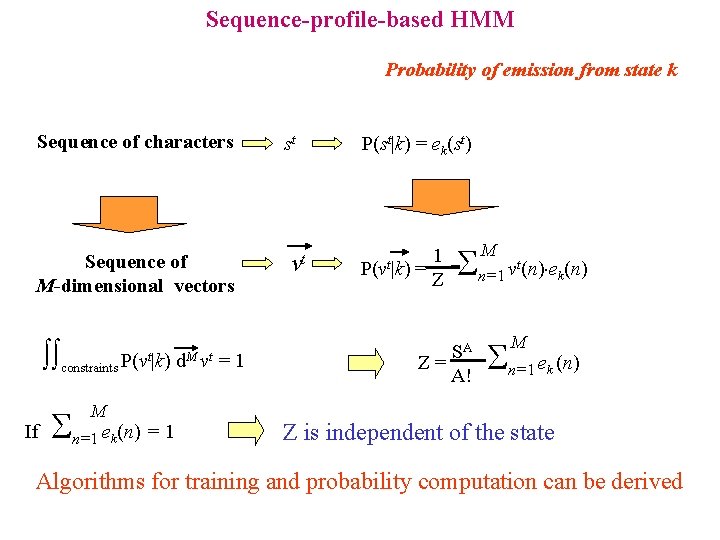
Sequence-profile-based HMM Probability of emission from state k Sequence of characters Sequence of M-dimensional vectors constraints P(vt|k) d. M vt = 1 If M n=1 e (n) = 1 k st vt P(st|k) = ek(st) 1 P(vt|k) = Z n=1 v (n) e (n) SA Z= A! M t k M n=1 e (n) k Z is independent of the state Algorithms for training and probability computation can be derived

Hybrid systems: Basics • Sequence profile based HMMs • Membrane protein topology

Scoring the prediction 1) Accuracy: Q 2=P/N where P is the total number of correctly predicted residues and N is the total number of residues 2) Correlation coefficient C: C(s)=(p(s)*n(s) - u(s)*o(s))/[(p(s)+u(s))(p(s)+o(s))(n(s)+u(s))(n(s)+o(s))]1/2 where, for each class s, p(s) and n(s) are respectively the total number of correct predictions and correctly rejected assignments while u(s) and o(s) are the numbers of under and over predictions 3) Accuracy for each discriminated structure s: Q(s)=p(s)/[p(s)+u(s)] where p(s) and u(s) are the same as in equation 2 4) Probability of correct predictions P(s) : P(s)=p(s)/[p(s)+o(s)] where p(s) and o(s) are the same as in equation 2 5) Segment-based measure (Sov):

Topology of -barrel membrane proteins Performance of sequence-profile-based HMM Q 2 83% 76% 78% QTMS Qloop 83% 77% 74% 82% 76% 82% PTMS Ploop 79% 72% 81% 85% 80% 76% Corr 0. 65 0. 53 0. 56 Sov 0. 83 0. 64 0. 79 HMM based on Multiple Sequence Alignment Standard HMM based on Single Sequence NN based on Multiple Sequence Alignment Martelli PL, Fariselli P, Krogh A, Casadio R -A sequence-profile-based HMM for predicting and discriminating beta barrel membrane proteins- Bioinformatics 18: S 46 -S 53 (2002)
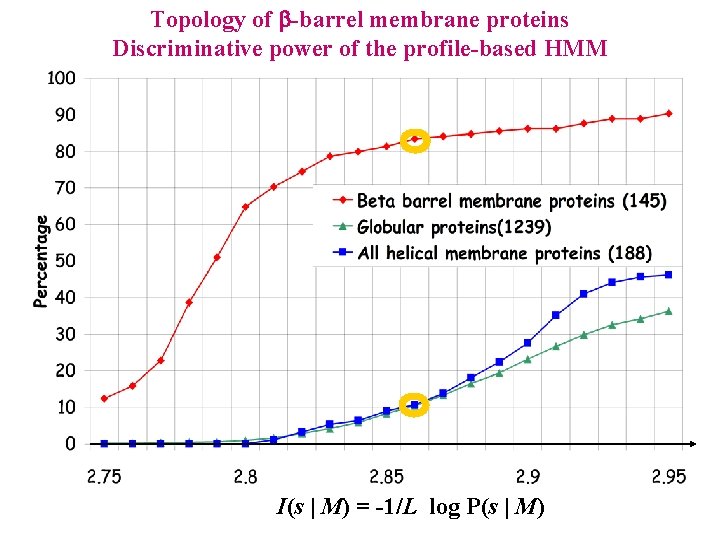
Topology of -barrel membrane proteins Discriminative power of the profile-based HMM I(s | M) = -1/L log P(s | M)
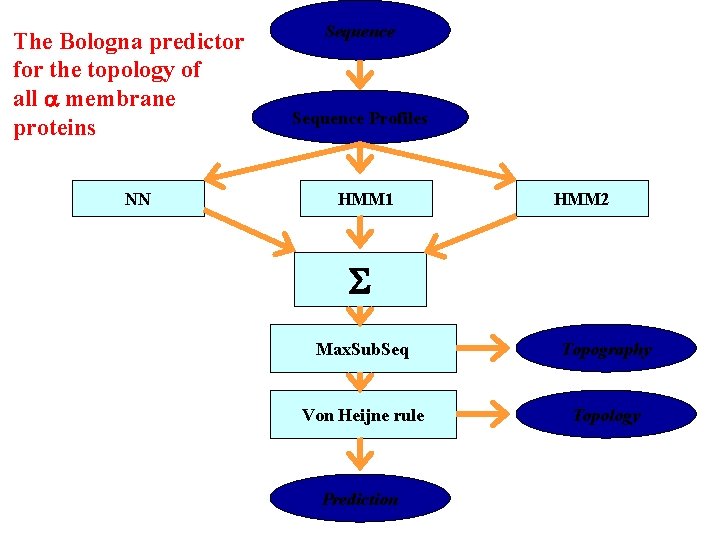
The Bologna predictor for the topology of all membrane proteins NN Sequence Profiles HMM 1 HMM 2 S Max. Sub. Seq Topography Von Heijne rule Topology Prediction
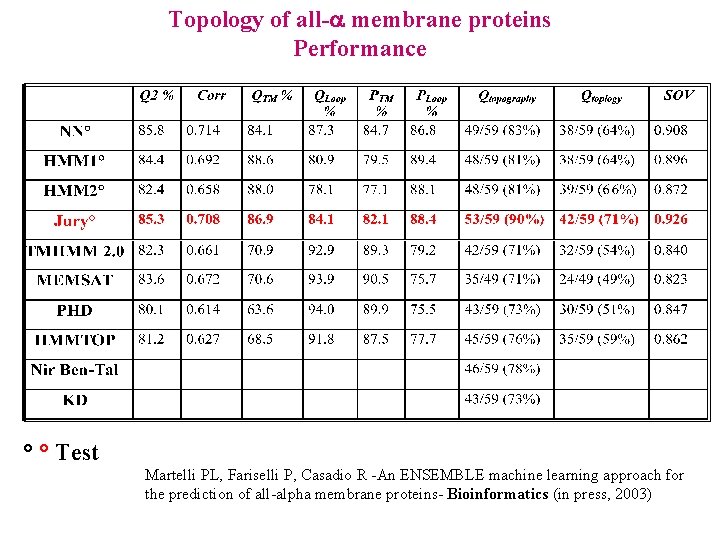
Topology of all- membrane proteins Performance ° ° Test Martelli PL, Fariselli P, Casadio R -An ENSEMBLE machine learning approach for the prediction of all-alpha membrane proteins- Bioinformatics (in press, 2003)

HMM: Application in gene finding • Basics
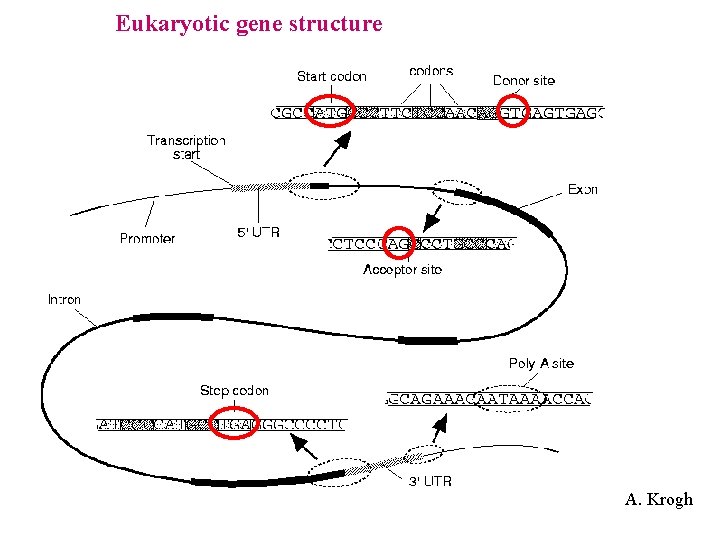
Eukaryotic gene structure A. Krogh
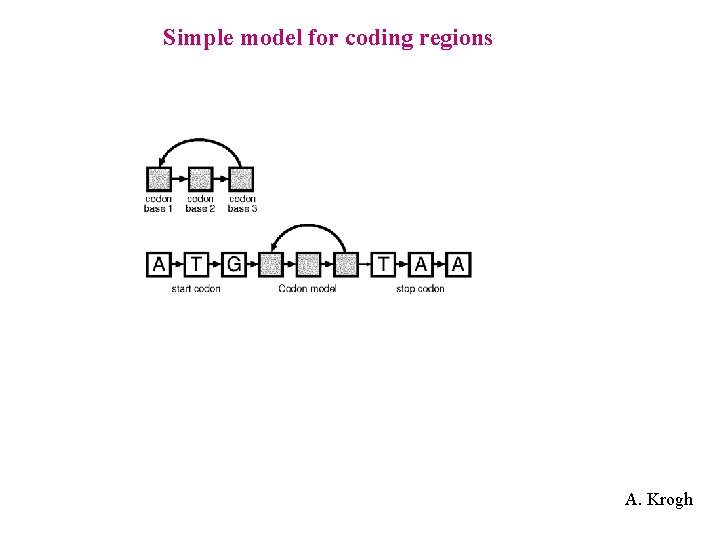
Simple model for coding regions A. Krogh
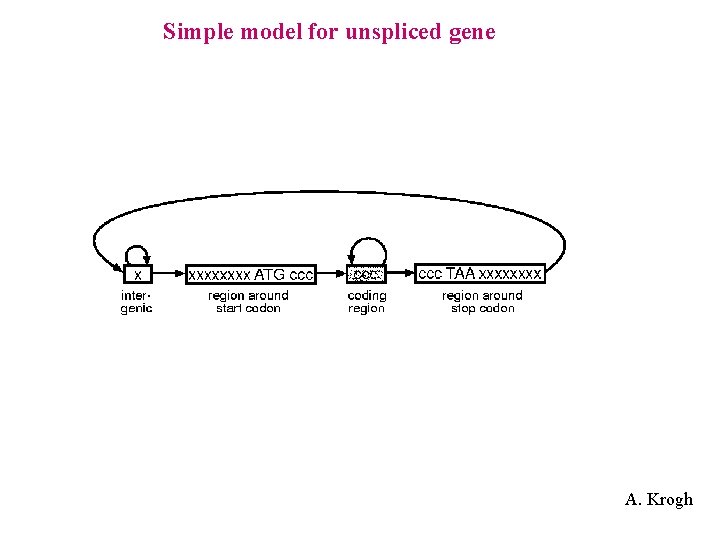
Simple model for unspliced gene A. Krogh
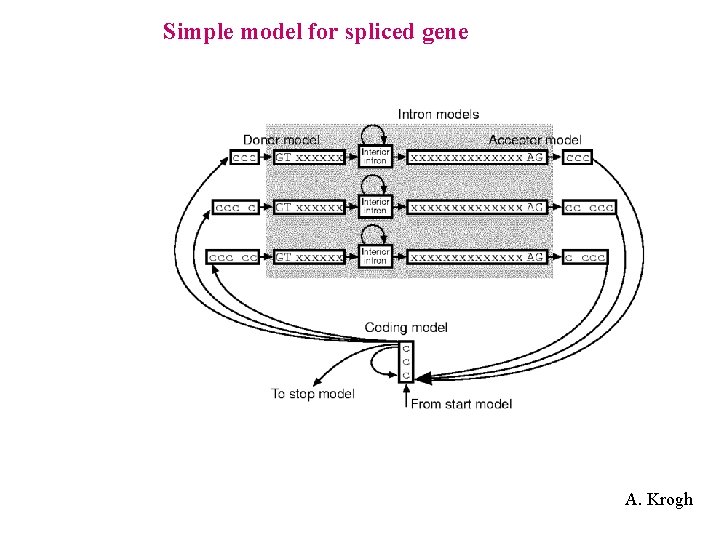
Simple model for spliced gene A. Krogh
- Slides: 154