15 Repetition Overview Dr Werner Van Belle wernersigtrans

15. Repetition / Overview Dr. Werner Van Belle werner@sigtrans. org 25/11/2009 - Pg. 1 v 1 © Van Belle Werner

1. Correlation Mathematical definition of Average, Variance, Standard deviation Mathematical definition of L. P. Correlation Graphical interpretation of correlation Implementing a correlation routine given the math. Definition What is numerical stability ? Give an example using the variance How to deal with missing numbers in a correlation calculation ? What does significance of a correlation value mean ? 25/11/2009 - Pg. 2 v 1 © Van Belle Werner
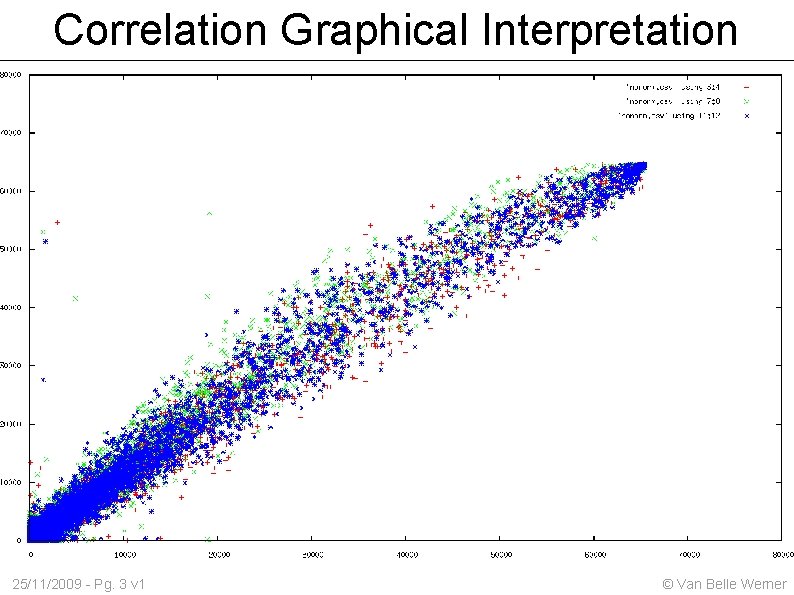
Correlation Graphical Interpretation 25/11/2009 - Pg. 3 v 1 © Van Belle Werner

Given a vector of n numbers What is the average is of this vector ? 25/11/2009 - Pg. 4 v 1 © Van Belle Werner
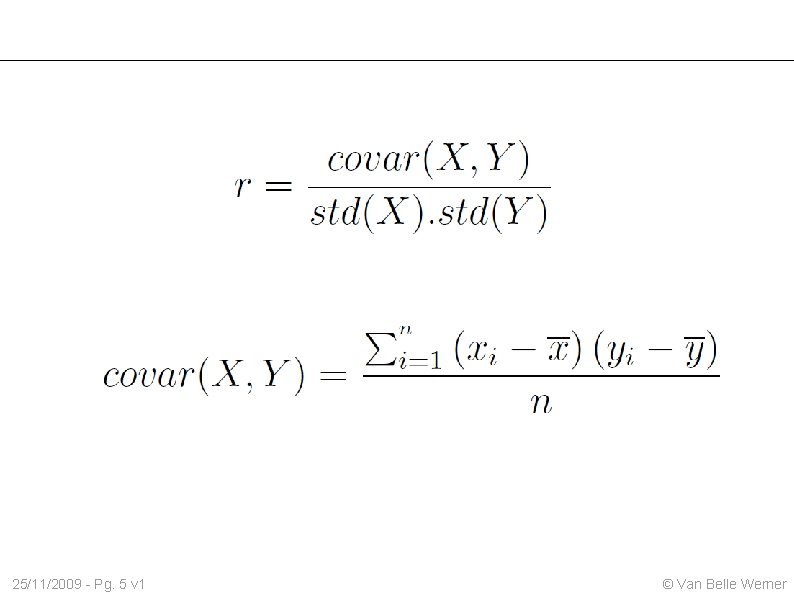
25/11/2009 - Pg. 5 v 1 © Van Belle Werner
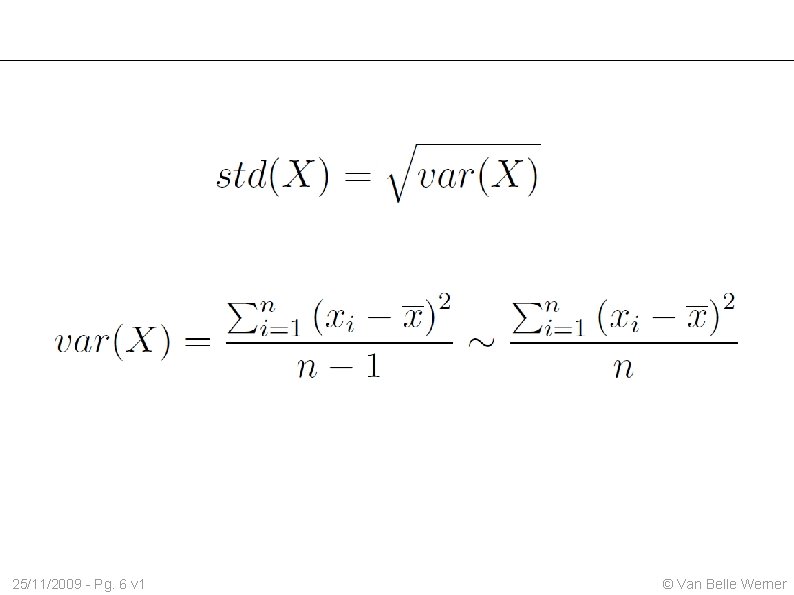
25/11/2009 - Pg. 6 v 1 © Van Belle Werner
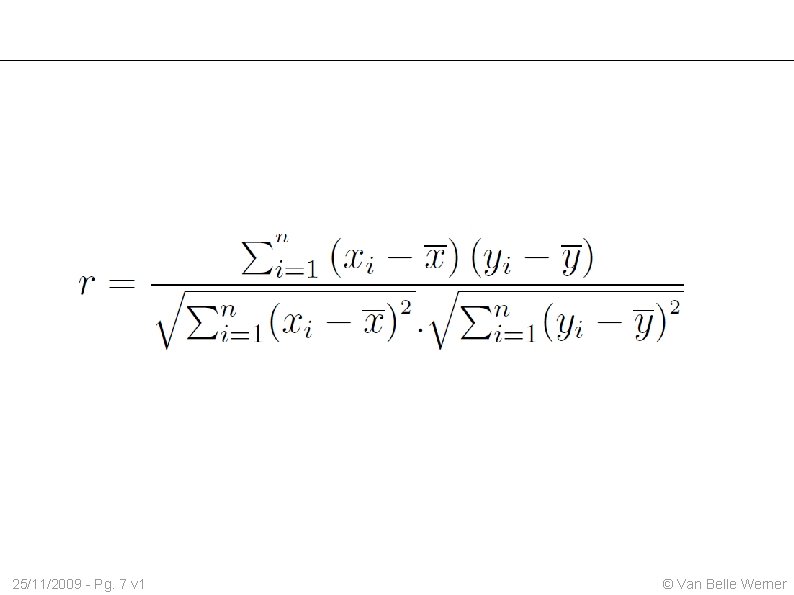
25/11/2009 - Pg. 7 v 1 © Van Belle Werner

Step 1. Translation 25/11/2009 - Pg. 8 v 1 © Van Belle Werner
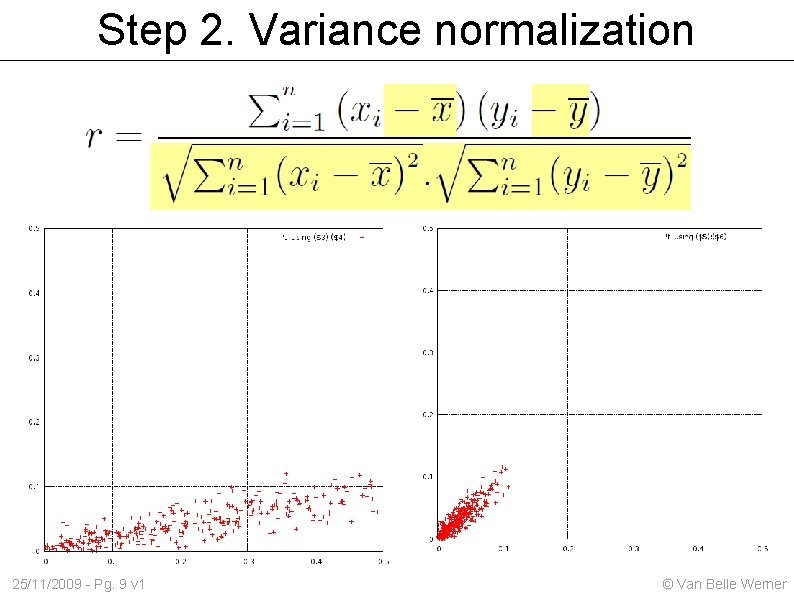
Step 2. Variance normalization 25/11/2009 - Pg. 9 v 1 © Van Belle Werner
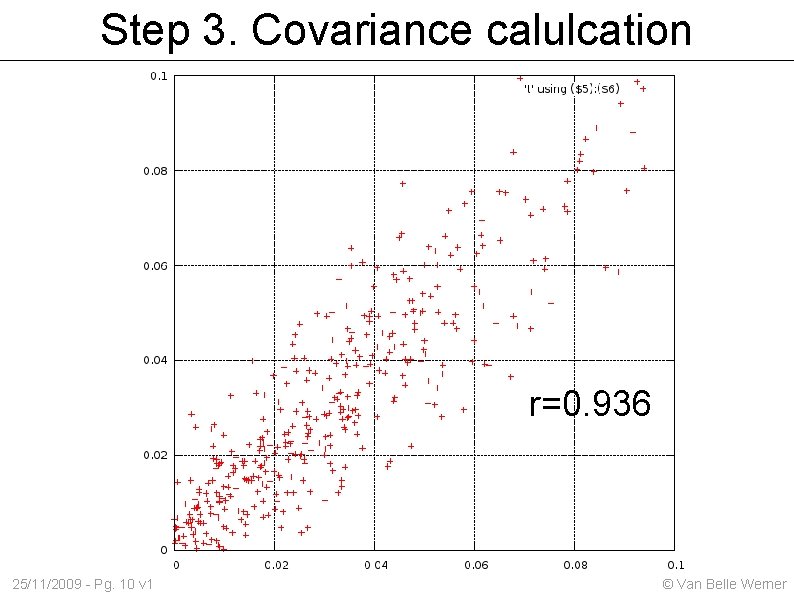
Step 3. Covariance calulcation r=0. 936 25/11/2009 - Pg. 10 v 1 © Van Belle Werner

1. Write a function that takes two inputs X and Y and returns the linear pearson correlation between these two vectors. 2. Write a routine which reads a binary file in which we have a consecutive sequence of double precision floats. 3. Modify your program to take 2 filenames as argument and let it report the correlation value 25/11/2009 - Pg. 11 v 1 © Van Belle Werner

4. Write a program that will generate two random vectors of size N and let it report the correlation between those two random vectors. Let this program repeat this action 100 times and report the average absolute correlation. Investigate the effect of the groupsize N on the average reported correlation. 5. We have 10 vectors stored in 10 different files. Write a program to read these files and report a correlation matrix. 25/11/2009 - Pg. 12 v 1 © Van Belle Werner

What should we do with missing numbers ? What is the significance of a correlation ? 25/11/2009 - Pg. 13 v 1 © Van Belle Werner

Numerical Stability Variance calculations are notorious for numerical instability 100'000 10**10 A Simple Algorithm for(int i=0; i < n; i++) var+=(x_i-avg)*(x_i-avg) var/=n; 25/11/2009 - Pg. 14 v 1 100'000 1 1 1 1 20**10 30*10 40**10 50**10+1 ? . . . © Van Belle Werner

Incremental online algorithm Note on a Method for Calculating Corrected Sums of Squares and Products B. P. Welford, technometrics, Vol. 4, No. 3 (Aug. , 1962), pp. 419 -420 25/11/2009 - Pg. 15 v 1 © Van Belle Werner

2. Multi Dimensional Correlations What is 2 D Gel electrophoresis ? What is a function/method declaration ? Exercise on converting a single dimensional routine to multiple dimensions Correlation is not causation Correlation is not linear regression Correlations can be accidental Both no correlations -as well as- high/lo correlations can be indicative 25/11/2009 - Pg. 16 v 1 © Van Belle Werner
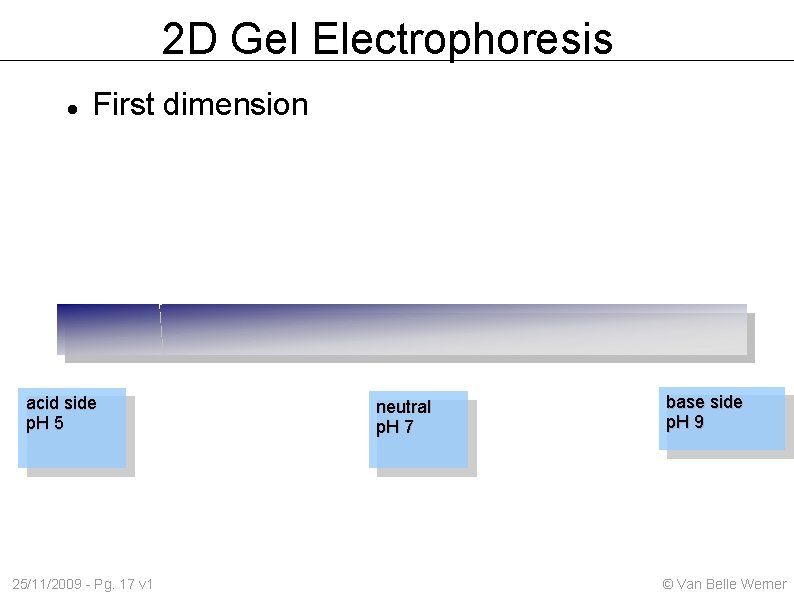
2 D Gel Electrophoresis First dimension acid side p. H 5 25/11/2009 - Pg. 17 v 1 neutral p. H 7 base side p. H 9 © Van Belle Werner
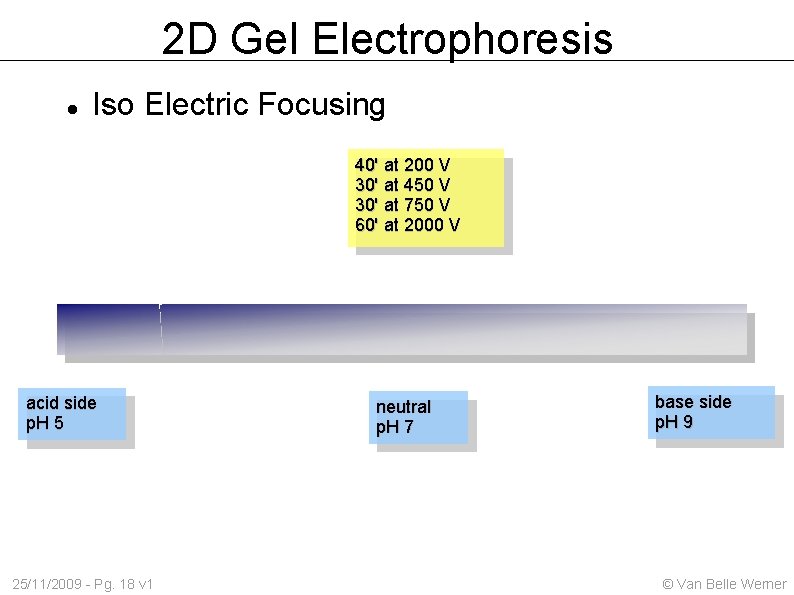
2 D Gel Electrophoresis Iso Electric Focusing 40' at 200 V 30' at 450 V 30' at 750 V 60' at 2000 V acid side p. H 5 25/11/2009 - Pg. 18 v 1 neutral p. H 7 base side p. H 9 © Van Belle Werner

2 D Gel Electrophoresis Transfer onto 2 nd gel Transfer 25/11/2009 - Pg. 19 v 1 © Van Belle Werner

2 D Gel Electrophoresis Transfer onto 2 nd gel Time Based Mass Separation 25/11/2009 - Pg. 20 v 1 © Van Belle Werner

2 D Gel Electrophoresis Washing/Drying/Staining 25/11/2009 - Pg. 21 v 1 © Van Belle Werner
2 D Gel Electrophoresis Capturing 25/11/2009 - Pg. 22 v 1 © Van Belle Werner
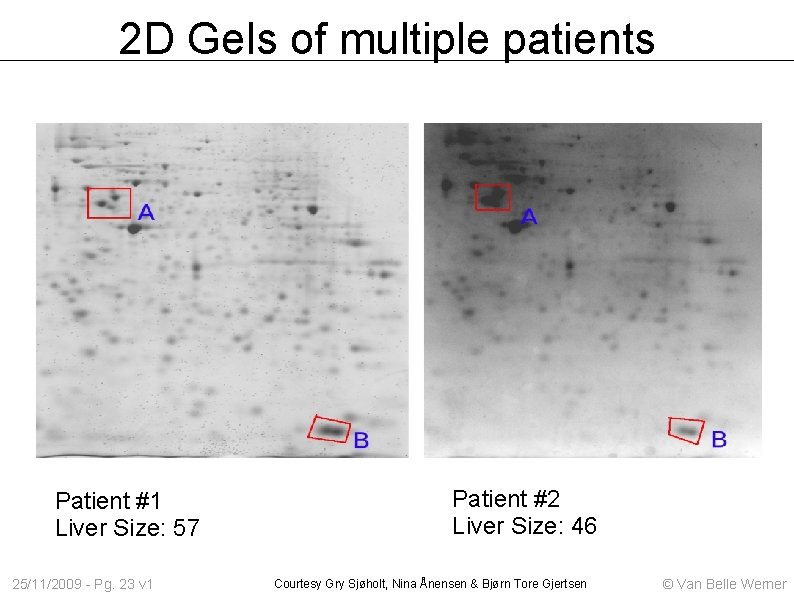
2 D Gels of multiple patients Patient #1 Liver Size: 57 25/11/2009 - Pg. 23 v 1 Patient #2 Liver Size: 46 Courtesy Gry Sjøholt, Nina Ånensen & Bjørn Tore Gjertsen © Van Belle Werner
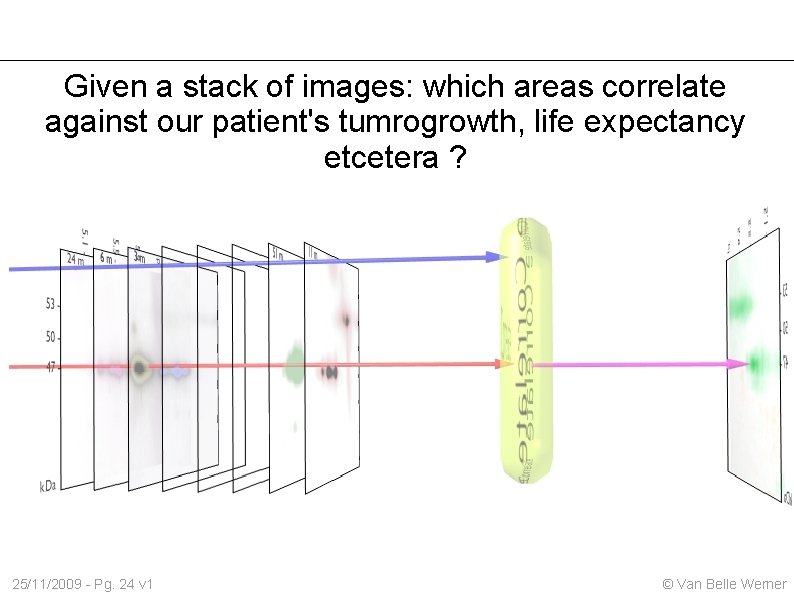
Given a stack of images: which areas correlate against our patient's tumrogrowth, life expectancy etcetera ? 25/11/2009 - Pg. 24 v 1 © Van Belle Werner
![Reading The Image int img_sx int img_sy byte[, ] read_image(String filename) Will read the Reading The Image int img_sx int img_sy byte[, ] read_image(String filename) Will read the](http://slidetodoc.com/presentation_image/a6c9df4e44cd58f5d0b83a887f4c2c97/image-25.jpg)
Reading The Image int img_sx int img_sy byte[, ] read_image(String filename) Will read the image from the provided filename When img_sx is not set, will set both of them to the size of the image being loaded When img_sx is set then the image must have the same size. Otherwise null is returned. If the image does not exist null is returned as well. The byte array is ordered as image[x][y] 25/11/2009 - Pg. 25 v 1 © Van Belle Werner

Exercise Import your correlation routine from last time. It should have the following declaration float correlate(float[] X, float[] Y, int n) 25/11/2009 - Pg. 26 v 1 © Van Belle Werner
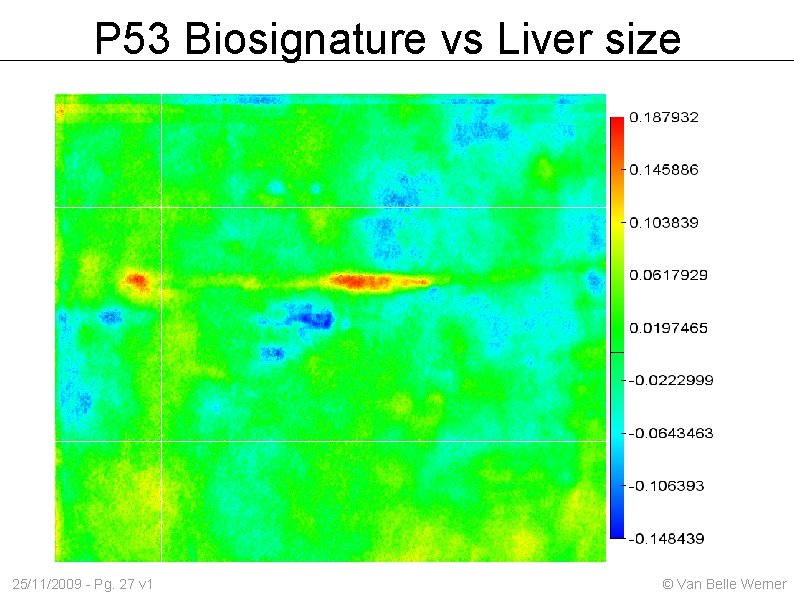
P 53 Biosignature vs Liver size 25/11/2009 - Pg. 27 v 1 © Van Belle Werner

Masking 25/11/2009 - Pg. 28 v 1 © Van Belle Werner

Significance 25/11/2009 - Pg. 29 v 1 © Van Belle Werner
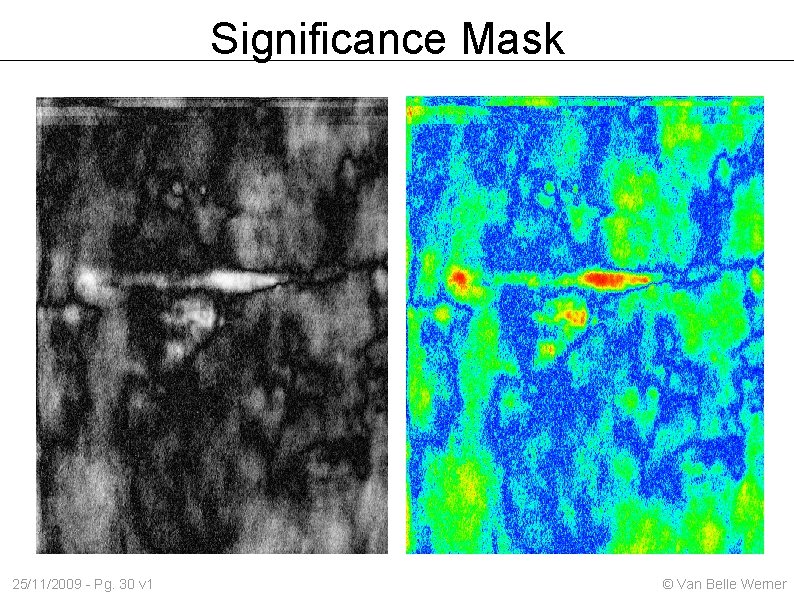
Significance Mask 25/11/2009 - Pg. 30 v 1 © Van Belle Werner

Variance 25/11/2009 - Pg. 31 v 1 © Van Belle Werner
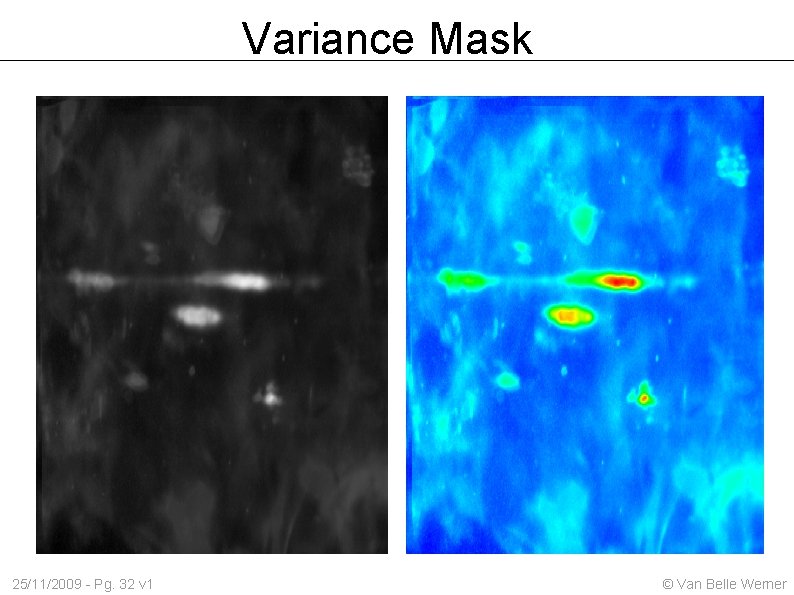
Variance Mask 25/11/2009 - Pg. 32 v 1 © Van Belle Werner

Overall Mask 25/11/2009 - Pg. 33 v 1 © Van Belle Werner
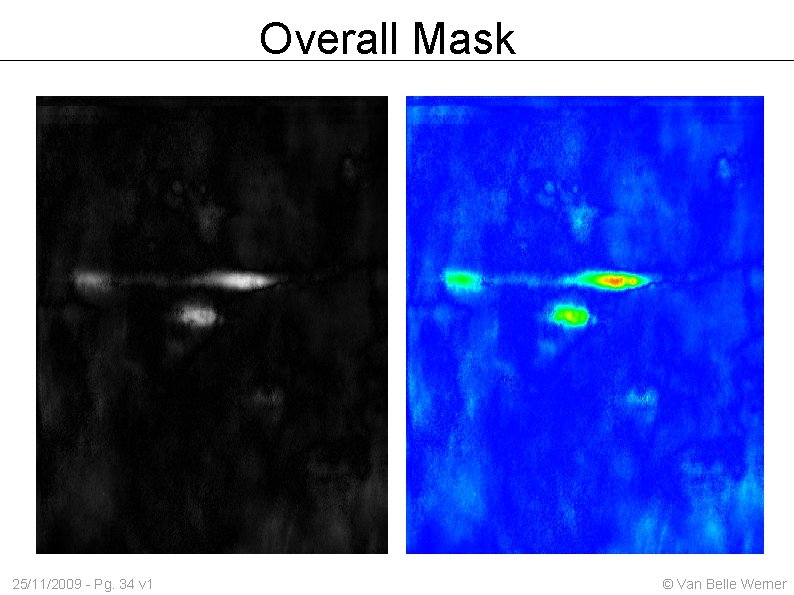
Overall Mask 25/11/2009 - Pg. 34 v 1 © Van Belle Werner
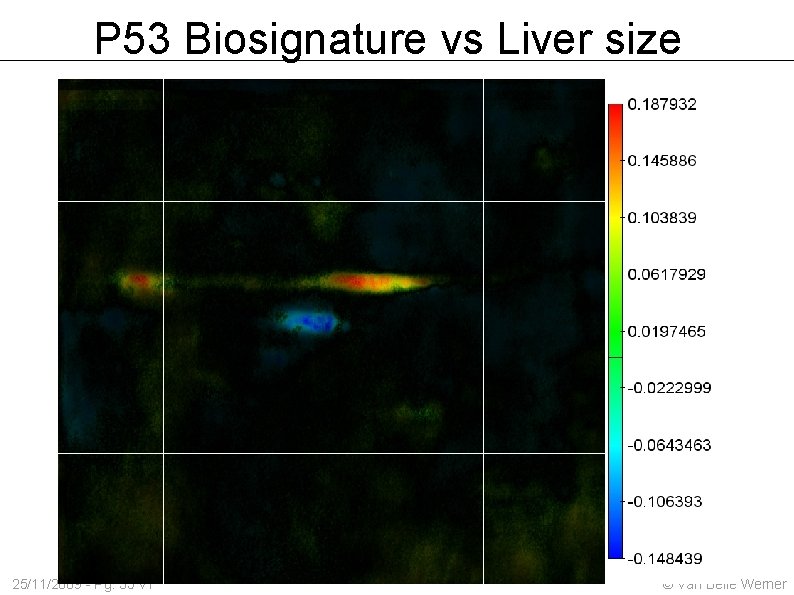
P 53 Biosignature vs Liver size 25/11/2009 - Pg. 35 v 1 © Van Belle Werner

3. Nucleotides to Amino Acides Various biological terms briefly explained Prokaryotes/Eukaryotes/Chromosomes/Chroma tide/Chromatine/Karyotyping/Diploid/Haploid/Ga metes Where is the genetic material stored ? Nucleotides / Amino Acids Complement, Reverse Sequence Proteins Translation Reading Frames, Open reading Frames 25/11/2009 - Pg. 36 v 1 © Van Belle Werner

Cells Prokaryotic - no nucleus (bacteria) Eukaryotic – with nucleus (plants/animals) The nucleus Contains the genetic material Genetic material can be in two states 25/11/2009 - Pg. 37 v 1 Heterochromatine / Euchromatine (a diffuse state which makes the DNA accessible) Chromosomes © Van Belle Werner
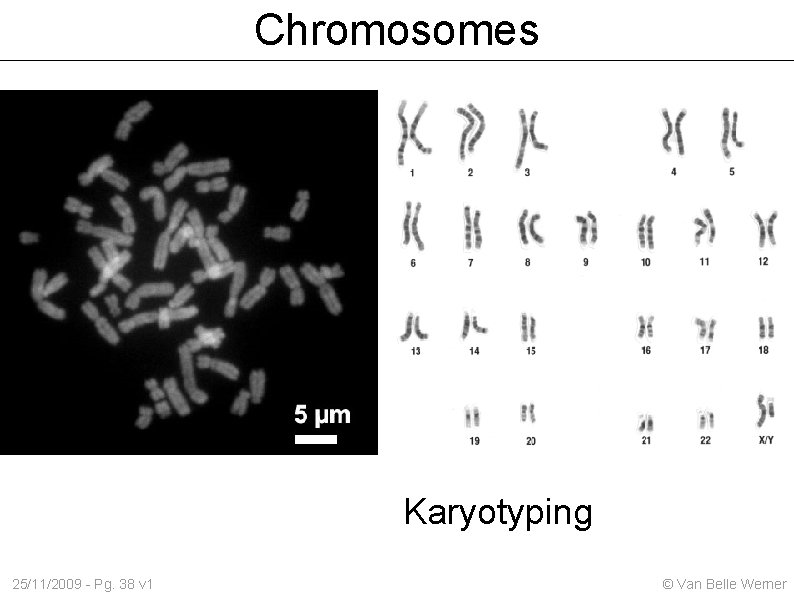
Chromosomes Karyotyping 25/11/2009 - Pg. 38 v 1 © Van Belle Werner

Humans 23 chromosomes pairs: Diploid Other organisms can have different layouts Tetraploid Hexaploid Octoploid 22 chromsome types 1 sex chromosome 25/11/2009 - Pg. 39 v 1 (autosomes) © Van Belle Werner

Chromosome Various chromosome layouts Somatic cells – diploid Gametes – haploid One set from mother One set from father Mother -or. Father Gametes do not have the same genetic code Autosomes in diploid cells are not strictly identical. Although 99% is the same 25/11/2009 - Pg. 40 v 1 © Van Belle Werner
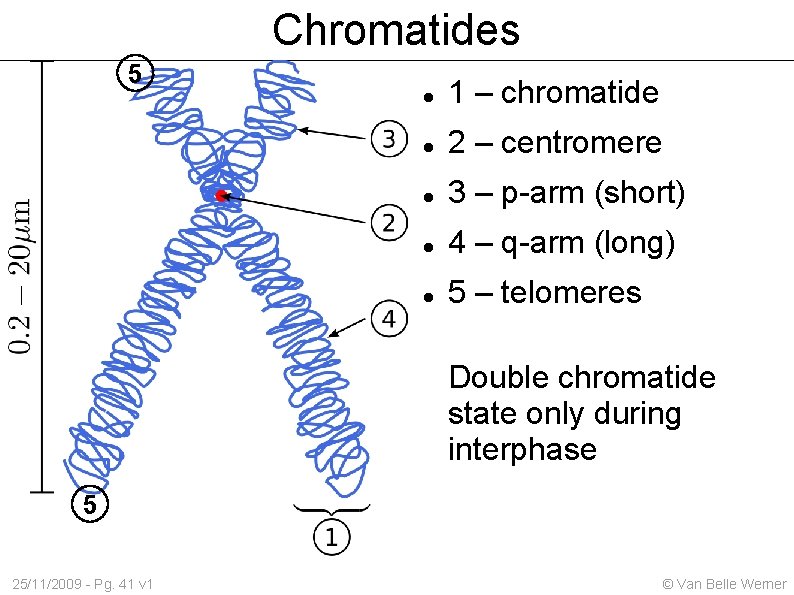
Chromatides 5 1 – chromatide 2 – centromere 3 – p-arm (short) 4 – q-arm (long) 5 – telomeres Double chromatide state only during interphase 5 25/11/2009 - Pg. 41 v 1 © Van Belle Werner
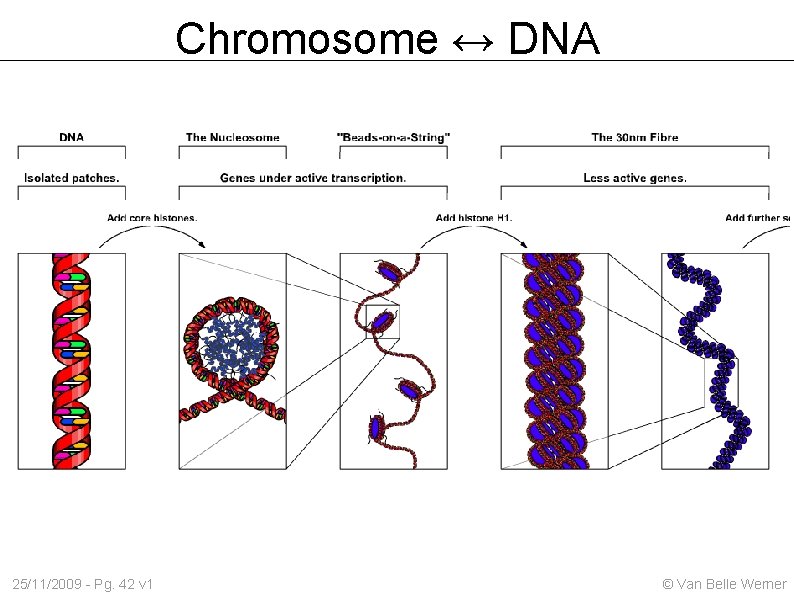
Chromosome ↔ DNA 25/11/2009 - Pg. 42 v 1 © Van Belle Werner
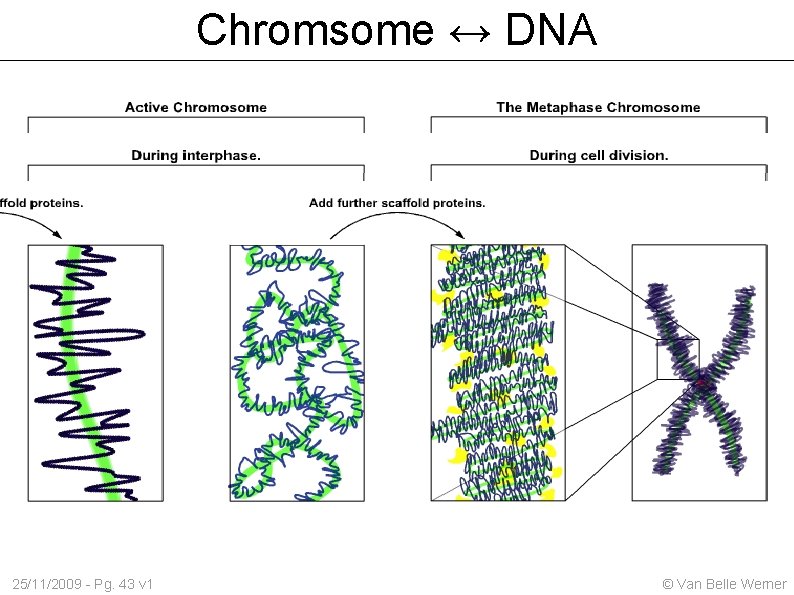
Chromsome ↔ DNA 25/11/2009 - Pg. 43 v 1 © Van Belle Werner
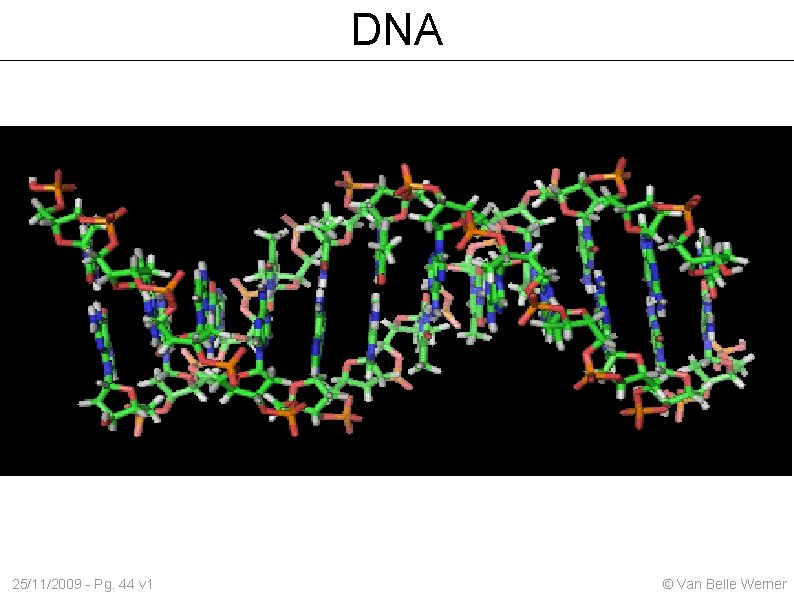
DNA 25/11/2009 - Pg. 44 v 1 © Van Belle Werner
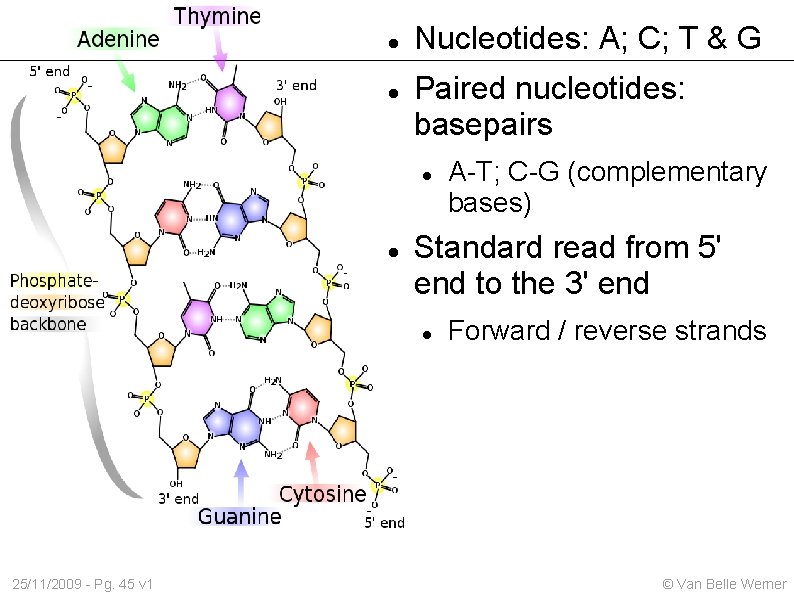
Nucleotides: A; C; T & G Paired nucleotides: basepairs Standard read from 5' end to the 3' end 25/11/2009 - Pg. 45 v 1 A-T; C-G (complementary bases) Forward / reverse strands © Van Belle Werner

Genes Gene identification is problematic Position identifiers are not unique Sequences are not completely unique A biologists' agreement on terminology 'somewhere around this area' having 'largely' this sequence. Similar sequences across species 25/11/2009 - Pg. 46 v 1 © Van Belle Werner

Genes Specific areas in the genome (loci) have meaning and translate to proteins afterward Number of bases in the genome ? Number of genes in the genome ? 25/11/2009 - Pg. 47 v 1 © Van Belle Werner
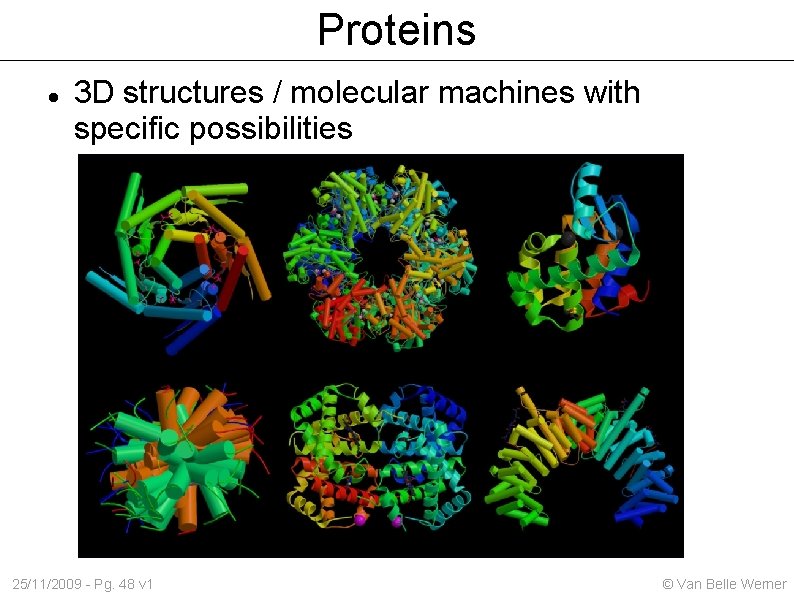
Proteins 3 D structures / molecular machines with specific possibilities 25/11/2009 - Pg. 48 v 1 © Van Belle Werner

Amino Acids Consist of a sequence of 20++ amino acids Alanine (Ala, A) Cysteine (Cys, C), Aspartic Acid (Asp, D), Glutamic Acid (Glu, E), Phenylalanine (Phe, F), Glycine (Gly, G), Histidine (His, H), Isoleucine (Ile, I), Lysine (Lys, K), Leucine (Leu, L), Methionine (Met, M), Asparagine (Asn, N), Proline (Pro, P), Glutamine (Gln, Q), Arginine (Arg, R), Serine (Ser, S), Threonine (Thr, T), Valine (Val, V), Tryptophan (Trp, W), Tyrosine (Tyr, Y) Selenocysteine, pyrrolysine (rare) 25/11/2009 - Pg. 49 v 1 © Van Belle Werner

Essential Amino Acids Consist of a sequence of 20 amino acids Alanine (Ala, A) Cysteine (Cys, C), Aspartic Acid (Asp, D), Glutamic Acid (Glu, E), Phenylalanine (Phe, F), Glycine (Gly, G), Histidine (His, H), Isoleucine (Ile, I), Lysine (Lys, K), Leucine (Leu, L), Methionine (Met, M), Asparagine (Asn, N), Proline (Pro, P), Glutamine (Gln, Q), Arginine (Arg, R), Serine (Ser, S), Threonine (Thr, T), Valine (Val, V), Tryptophan (Trp, W), Tyrosine (Tyr, Y) 25/11/2009 - Pg. 50 v 1 © Van Belle Werner

DNA → Protein Codons (3 nucleotide sequence) translate to Amino acid DNA copies to RNA, which is moved out of the nucleus (T → U) Polymerases convert the sequence to proteins Multiple translations possible. Most common is RNA Polymerase-II 25/11/2009 - Pg. 51 v 1 © Van Belle Werner
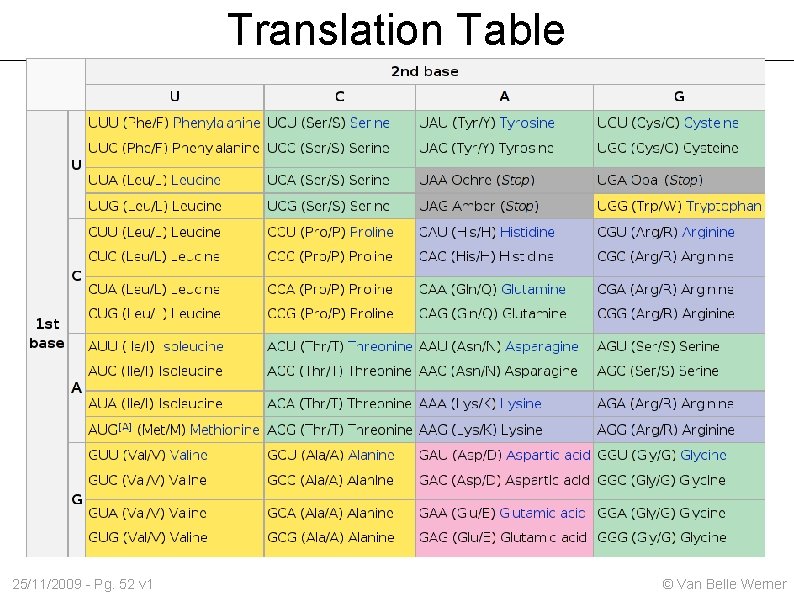
Translation Table 25/11/2009 - Pg. 52 v 1 © Van Belle Werner

Reading frames UCU AAA AUG GGU GAC . . . CUA AAA UGG GUG AC . . . UAA AAU GGG UGA C An open reading frame (ORF) is a reading frame that contains a start codon a subsequent region which usually has a length which is a multiple of 3 nucleotides a stop codon at its end. 25/11/2009 - Pg. 53 v 1 © Van Belle Werner

Exercise Create a routine that calculates the complement of a DNA sequence Create a routine that calculates the reverse of a DNA sequence Create a routine that translates a DNA sequence into an amino acid sequence Let the program try each reading frame and report the sequence with the longest distance to the first stop codon Include now complement, reverse and reverse complement sequences. 25/11/2009 - Pg. 54 v 1 © Van Belle Werner

4. Exons, Introns, Splice Variants Translation mechanisms in eukaryotes Splice variants; Exons / Introns Ensembl Browsing of this type of information Designing probes to detect specific splice variants 25/11/2009 - Pg. 55 v 1 © Van Belle Werner

Transcription / Translation Prokaryotes Transcription: Polymerase copies the gene into an RNA strand: m. RNA Translation: The m. RNA is then used to generate proteins These peptide chains then fold into Proteins Problem for Eukaryotes DNA stays in the nucleus Proteins are mainly in the cytoplasm 25/11/2009 - Pg. 56 v 1 © Van Belle Werner
Eukaryotic Translation Translate DNA to pre-m. RNA Process pre-m. RNA to m. RNA Adding caps (5' cap, polyadenylation) Splicing (select certain parts of the pre-m. RNA) Editing (nucleotide modifications) Transport m. RNA to cytoplasm Translate m. RNA to proteins. 25/11/2009 - Pg. 57 v 1 © Van Belle Werner
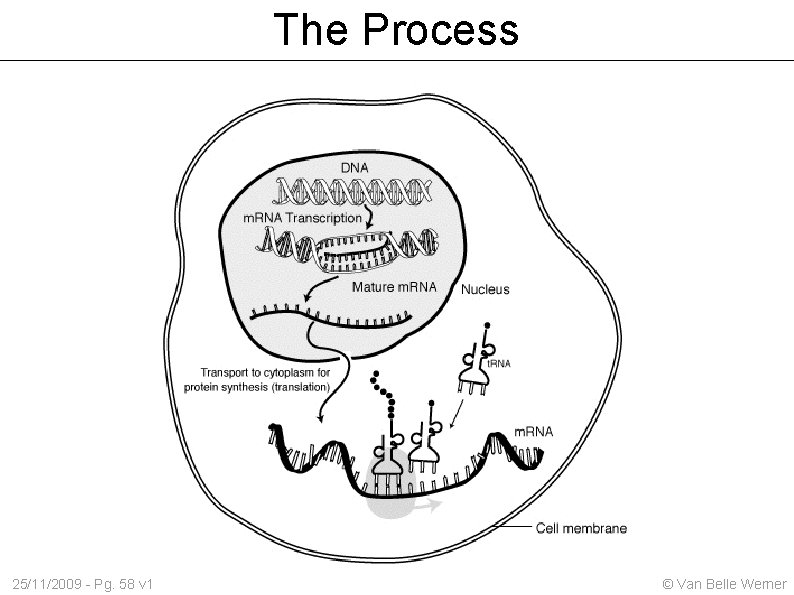
The Process 25/11/2009 - Pg. 58 v 1 © Van Belle Werner
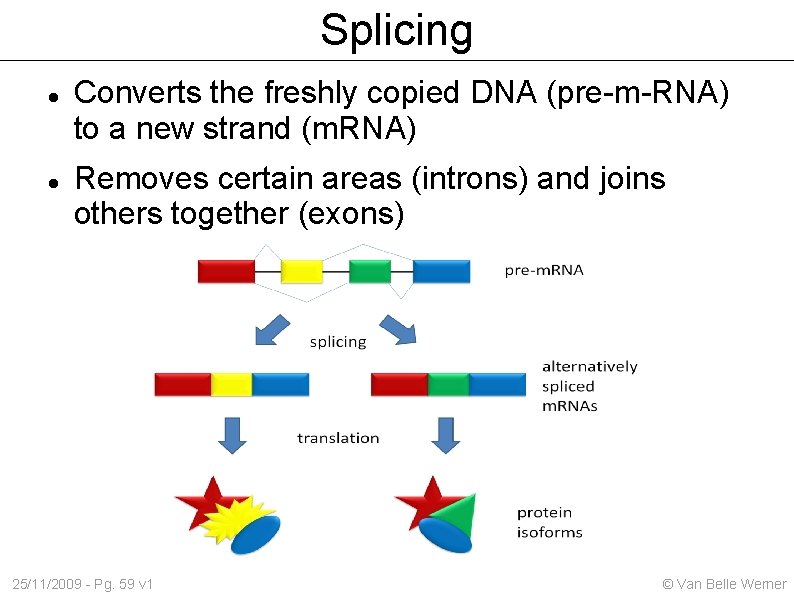
Splicing Converts the freshly copied DNA (pre-m-RNA) to a new strand (m. RNA) Removes certain areas (introns) and joins others together (exons) 25/11/2009 - Pg. 59 v 1 © Van Belle Werner
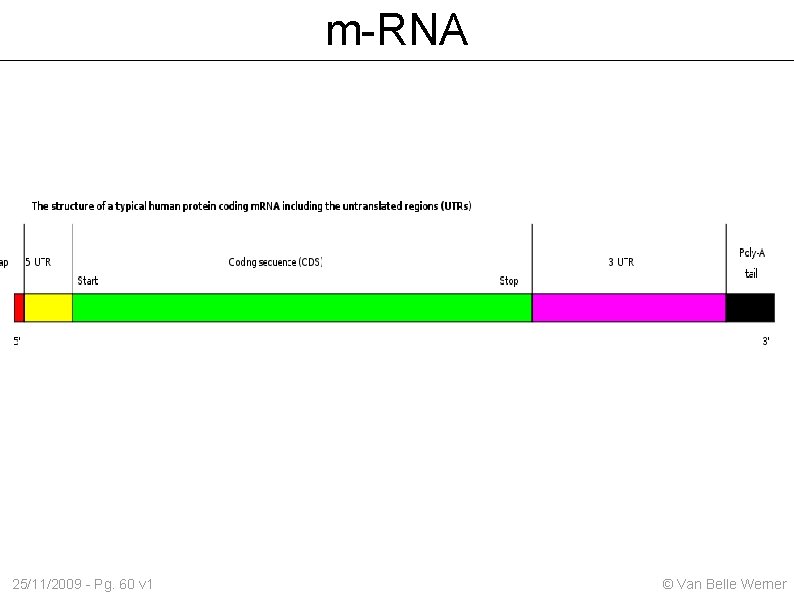
m-RNA 25/11/2009 - Pg. 60 v 1 © Van Belle Werner

One Gene, One Protein ? Non-coding genes Alternative splicing: one gene can have multiple splice variants leading to different proteins Monocistronic m. RNA: when the m. RNA codes for only one protein Polycistronic m. RNA: codes for multiple genes (operon) 25/11/2009 - Pg. 61 v 1 © Van Belle Werner

TP 53 In human: on which chromosome is it located ? Does this area in the genome also overlap with other genes ? How many splice variants does the TP 53 gene have ? How many bases is the gene long ? How many exons does the gene have ? Is there a transcript which includes all exons ? What is the sequence of the shortest transcript ? [TP 53 -205] 25/11/2009 - Pg. 62 v 1 © Van Belle Werner

Some other Genes: 5 HTT, BRCA 2, Wingless Alternative names ? Chromosome location ? Overlapping genes at this position ? Splice variants ? Do all exons transcribe in the same direction ? 25/11/2009 - Pg. 63 v 1 © Van Belle Werner

Exercise We want to design a probe that will uniquely detect a specific splice variant (target) We have a nice large table of all existing splice variants in human together with their gene name and variant number (ENSTxxx) Given a length L, we now want to find the first subsequence of the target that does not match any of the other existing splice variant. The subsequence is of length L The splice variant table will nopt contain the target itself We also want the shortest sufficient probe to detect 25/11/2009 - Pg. 64 v 1 the target © Van Belle Werner

5. Ensembl SQL Part 1 What is a database, schema, table, row, column, attribute, value Ensembl Stable ID's Ensembl Genes, mapipng to stable ids Transcripts, Translations, Exons One-Many relationships across tables 25/11/2009 - Pg. 65 v 1 © Van Belle Werner
![Relational Databases Database Server Database aka [Database] Schema Tables Columns with specific types Rows Relational Databases Database Server Database aka [Database] Schema Tables Columns with specific types Rows](http://slidetodoc.com/presentation_image/a6c9df4e44cd58f5d0b83a887f4c2c97/image-66.jpg)
Relational Databases Database Server Database aka [Database] Schema Tables Columns with specific types Rows Values can be NULL or real values 25/11/2009 - Pg. 66 v 1 © Van Belle Werner
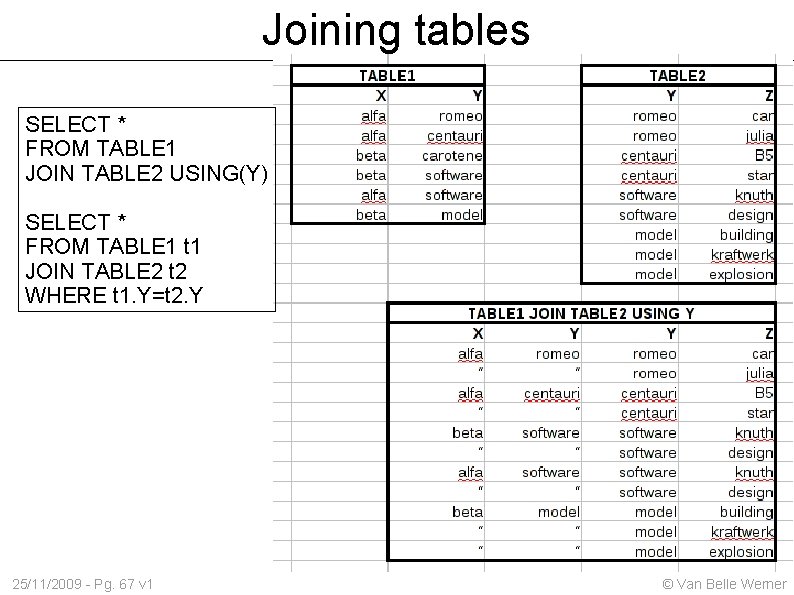
Joining tables SELECT * FROM TABLE 1 JOIN TABLE 2 USING(Y) SELECT * FROM TABLE 1 t 1 JOIN TABLE 2 t 2 WHERE t 1. Y=t 2. Y 25/11/2009 - Pg. 67 v 1 © Van Belle Werner

Ensembl Provides very structured biological information Integrates many different data sources Cares about metadata Keeps track of different versions, releases Keeps track where the data came from Keeps track how a specific analysis was performed Access using Mysql-query-browser Host: ensembldb. ensembl. org Port: 3306 (up to #47) or 5306 (#48 onwards) Login: anonymous 25/11/2009 - Pg. 68 v 1 © Van Belle Werner

Schemata Each organism has its own collection of databases homo_sapiens_core_47_36 i homo_sapiens_cdna_<x>_<y> homo_sapiens_core_expression_est_<x>_<y> homo_sapiens_core_expression_gnf_<x>_<y> homo_sapiens_disease_<x>_<y> homo_sapiens_estgene_<x>_<y> homo_sapiens_funcgen_<x>_<y> homo_sapiens_haplotype_<x>_<y> homo_sapiens_lite_<x>_<y> homo_sapiens_otherfeatures_<x>_<y> 25/11/2009 - Pg. 69 v 1 © Van Belle Werner

Stable Gene Identifiers Table gene_stable_id gene_id – the current gene identification, acts as (part of the) primary key in many tables – a number stable_id – the publicly visible ENSG<xxx> identifier creation_date – when was this gene introduced ? version – what is the current version of the gene modified_date – when was the last change ? 25/11/2009 - Pg. 70 v 1 © Van Belle Werner

Genes Table gene_id biotype: proteincoding or not seq_region_id seq_region_start seq_region_end seq_region_strand display_xref_id source: where did it come from status: KNOWN/NOVEL description: human readable 25/11/2009 - Pg. 71 v 1 © Van Belle Werner

Gene to Stable id mapping Write a query that will map a gene to its stable id. The output should contain gene_id, Biotype, seq_region_id, seq_region_start, seq_region_end, seq_region_strand, display_xref_id, source, status, description and of course the stable_id 25/11/2009 - Pg. 72 v 1 © Van Belle Werner

Transcript Table transcript_id – these ids can coincide with gene_ids. Do not mix them ! gene_id seq_region_id / seq_region_start / seq_region_end / seq_region_strand display_xref_id biotype status Description Table transcript_stable_id – something like ENST 000. . . 25/11/2009 - Pg. 73 v 1 © Van Belle Werner

Translation Table translation_id transcript_id seq_start_exon_id seq_end end_exon_id Table translation_stable_id – something like ENSP 0000. . translation_id 25/11/2009 - Pg. 74 v 1 © Van Belle Werner

Exon Table exon exon_id seq_region_id / seq_region_start / seq_region_end / seq_region_strand phase end_phase Table exon_transcript maps the transcript to something ? exon_id transcript_id Rank – exon number (1 to 10, 13, 170) 25/11/2009 - Pg. 75 v 1 © Van Belle Werner
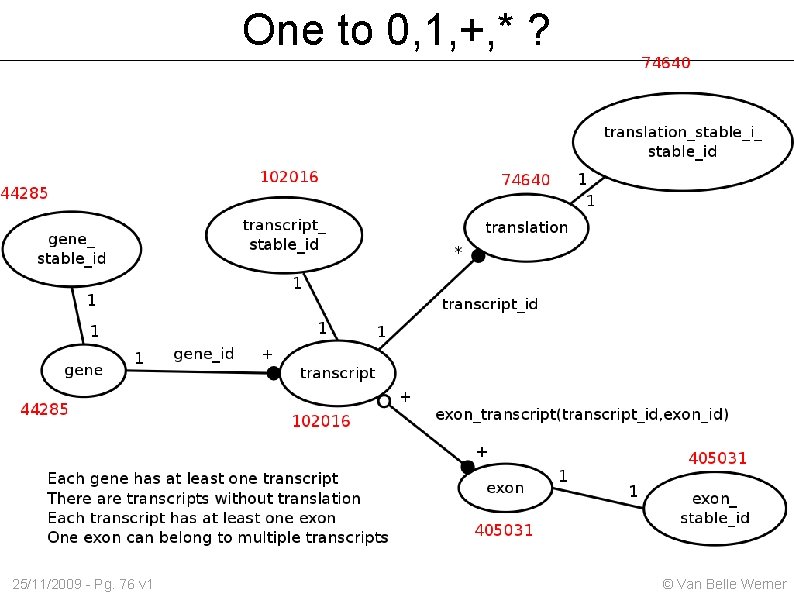
One to 0, 1, +, * ? 25/11/2009 - Pg. 76 v 1 © Van Belle Werner

Genes ↔ Transcript ↔ Exon Obtain a table with 3 columns gene_id transcript_id exon_id Start out with a table that lists all gene_id transcript_id Then extend the table with exons belonging to that transcript 25/11/2009 - Pg. 77 v 1 © Van Belle Werner
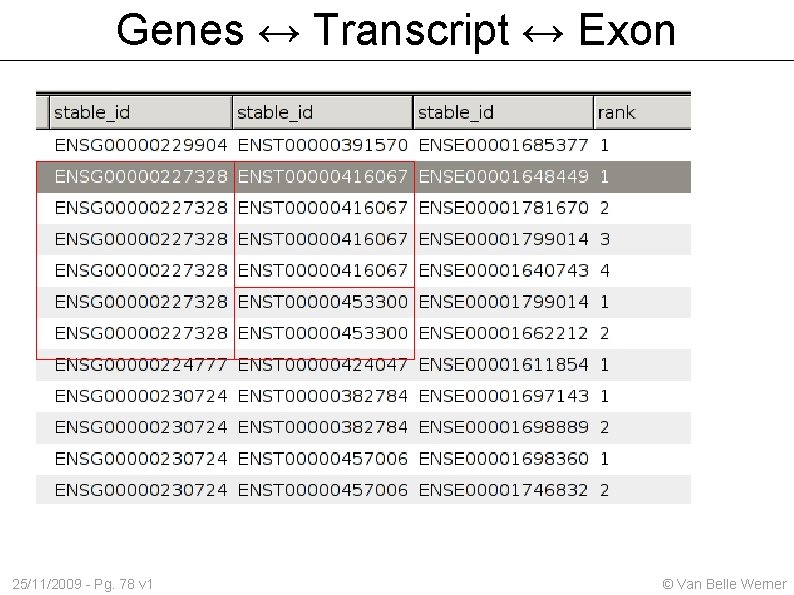
Genes ↔ Transcript ↔ Exon 25/11/2009 - Pg. 78 v 1 © Van Belle Werner

6. Ensembl SQL Part 2 Various mappings: Genes to Proteins Various grouping operations: average, maxima, minima, countings etcetera Ensembl Regions and Chromosome information 25/11/2009 - Pg. 79 v 1 © Van Belle Werner

Genes ↔ Protein Mapping Write a query to map genes to potential proteins. The output table should contain A stable gene identifier A stable protein identifier 25/11/2009 - Pg. 80 v 1 © Van Belle Werner

Genes ↔ Protein Mapping Write a query to map genes to potential proteins. The output table should contain A stable gene identifier A stable protein identifier SELECT G. stable_id as gen, T. stable_id as protein FROM gene JOIN transcript USING (gene_id) JOIN translation USING(transcript_id) JOIN translation_stable_id T USING (translation_id) JOIN gene_stable_id G USING (gene_id) LIMIT 10 25/11/2009 - Pg. 81 v 1 © Van Belle Werner

Genes ↔ Protein Mapping 25/11/2009 - Pg. 82 v 1 © Van Belle Werner

Averages What is the average number of transcripts per gene ? What is the average number of exons per transcript ? Based on #47 of the database 33761 unique genes 57365 unique transcripts 503655 unique exon/transcript combinations 288309 unique exons Which gives 1. 7 transcript/gene And 8. 79 exons per transcript But only 8. 5 unique exons per gene 25/11/2009 - Pg. 83 v 1 © Van Belle Werner

Largest Gene Which gene has the most transcripts ? SELECT COUNT(DISTINCT t. transcript_id) as tcount, si. stable_id, g. gene_id FROM gene g JOIN transcript t USING(gene_id) JOIN gene_stable_id si USING (gene_id) GROUP BY gene_id ORDER BY tcount DESC LIMIT 10 ENSG 00000154556 with 44 transcripts 25/11/2009 - Pg. 84 v 1 © Van Belle Werner

Transcript with the most exons Which transcript has the most exons ? SELECT MAX(rank) ecount, si. stable_id, t. transcript_id FROM transcript t JOIN exon_transcript et USING (transcript_id) JOIN transcript_stable_id si USING (transcript_id) GROUP BY transcript_id ORDER BY ecount DESC LIMIT 10 ENST 00000356127 with 313 exons 25/11/2009 - Pg. 85 v 1 © Van Belle Werner

Regions The region codes in Ensembl can be a variety of things. Table seq_region_id name – can be a chromosome name coord_system_id length 25/11/2009 - Pg. 86 v 1 © Van Belle Werner

Largest area covered Which gen covers the largest area in the genome ? On which chromosome ? SELECT seq_region_end-seq_region_start as L, stable_id, gene_id, name FROM gene JOIN gene_stable_id USING (gene_id) JOIN seq_region USING (seq_region_id) ORDER BY L desc LIMIT 10 Answer: ENSG 00000174469 with 2'304'637 bases 25/11/2009 - Pg. 87 v 1 © Van Belle Werner
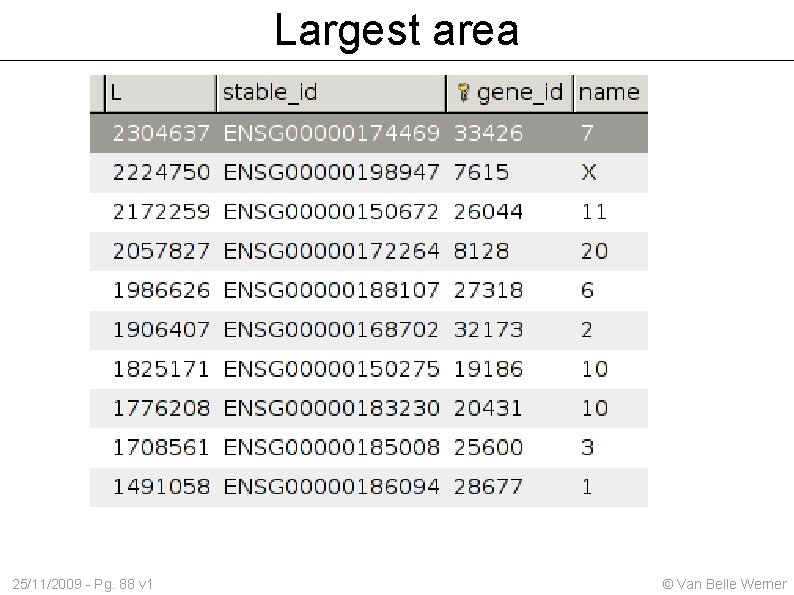
Largest area 25/11/2009 - Pg. 88 v 1 © Van Belle Werner

Create a table of TSS Retrieve a list of potential transcription start sites and to which gene they belong SELECT t. seq_region_start, t. seq_region_strand, stable_id FROM transcript t JOIN gene USING (gene_id) JOIN gene_stable_id USING (gene_id) 25/11/2009 - Pg. 89 v 1 © Van Belle Werner

7. Ensembl Identifiers How are external identifiers represented in Ensembl ? Object_xref – links Ensembl objects to external names Xref – remembers the external object name External_db – keeps track of a variety of different databases How could we add a new nomenclature Map one nomenclature to ensembl identifiers Mapping exercices from one nomenclature to another 25/11/2009 - Pg. 90 v 1 © Van Belle Werner

External Databases Table external_db_id – primary key db_name – database name db_release – version status – predicted, known, cross referenced, orthologue mapped etcetera type - misc, array etcetera db_display_name – how to print this database name 25/11/2009 - Pg. 91 v 1 © Van Belle Werner

External Databases Table external_db_id – primary key db_name – database name type - misc, array etcetera db_display_name – how to print this database name SELECT * FROM external_db WHERE db_name=”HGNC” 25/11/2009 - Pg. 92 v 1 external_db_id: 1100 db_name: HGNC db_display_name: HGNC symbol Type: primary_db_synonym © Van Belle Werner

External Names Table xref_id – the cross reference primary key external_db_id – the external database key db_primary_acc – the 'primary key' in the external database display_label – how to print this gene identifier description – a description according to the external database 25/11/2009 - Pg. 93 v 1 © Van Belle Werner

External Names to Stable Ids General purpose table object_xref ensembl_id: the ensemble internal id (gene_id for instance) ensembl_object_type: translation, transcript, gene xref_id: id from the xref table linkage_annotation 25/11/2009 - Pg. 94 v 1 © Van Belle Werner

Adding new nomenclature Create external_db entry For each gene <A> in the nomenclature, allocate the name in the database xref Gene A → xref_id 764 Gene B → xref_id 987 . . . For each external gene A, B, . . . in the nomenclature, map it to the database A → ENSG 78646 → gene_id=76689 B → ENSG 98768 → gene_id=7577 Link A, B to ENSG. . . through an object_xref of type Gene 25/11/2009 - Pg. 95 v 1 © Van Belle Werner

External Names to Stable Ids SELECT * FROM xref x JOIN object_xref o WHERE external_db_id=1100 and x. xref_id=o. xref_id LIMIT 0, 1000 xref_id: 1793295 external_db_id: 1100 dbprimary_acc 21076 display_labe: TMEM 14 A Description: transmembrane protein 14 A info_type: Dependent info_text: Generated via NP_054770 ensembl_id: 18971 ensembl_object_type: Gene xref_id: 1793295 linkage_annotation: NULL 25/11/2009 - Pg. 96 v 1 © Van Belle Werner

HGNC to Ensembl SELECT * We obtain the wrong results ! FROM xref x Be aware that ensembl_id cannot always JOIN object_xref o Be mapped to a gene_id JOIN gene_stable_id g WHERE external_db_id=1100 and x. xref_id=o. xref_id and ensembl_id=g. gene_id display_label: CXYorf 1 [and display_label="CXYorf 1"] ensembl_id: 7888 LIMIT 0, 1000 ensembl_object_type: Transcript display_label: CXYorf 1 ensembl_id: 4373 emsembl_object_type: Gene 25/11/2009 - Pg. 97 v 1 © Van Belle Werner
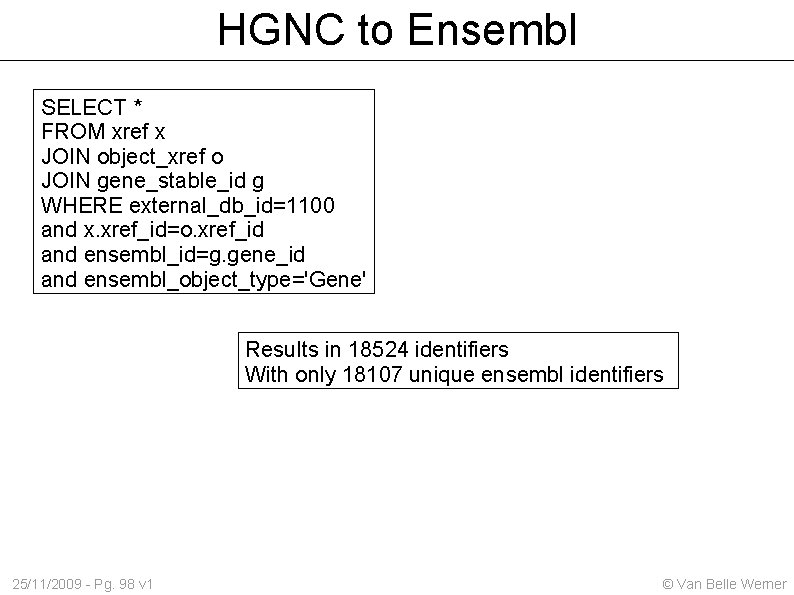
HGNC to Ensembl SELECT * FROM xref x JOIN object_xref o JOIN gene_stable_id g WHERE external_db_id=1100 and x. xref_id=o. xref_id and ensembl_id=g. gene_id and ensembl_object_type='Gene' Results in 18524 identifiers With only 18107 unique ensembl identifiers 25/11/2009 - Pg. 98 v 1 © Van Belle Werner

Ensembl to HGNC Find all ensembl genes that have no existing HGNC name First: map all the stable_ids through the object_xref table to the xref identities SELECT * FROM gene_stable_id g JOIN object_xref xr JOIN xref x USING(xref_id) WHERE xr. ensembl_object_type='Gene' AND xr. ensembl_id=g. gene_id LIMIT 100 25/11/2009 - Pg. 99 v 1 © Van Belle Werner

Ensembl to HGNC Find all ensembl genes that have no existing HGNC name Second: take only those that belong to the HGNC nomenclature (1100) SELECT * FROM gene_stable_id g JOIN object_xref xr JOIN xref x USING(xref_id) WHERE xr. ensembl_object_type='Gene' AND xr. ensembl_id=g. gene_id AND external_db_id=1100 LIMIT 100 25/11/2009 - Pg. 100 v 1 © Van Belle Werner

Ensembl to HGNC Find all ensembl genes that have no existing HGNC name Third: modify the query to only list the existing identifiers SELECT DISTINCT g. stable_id FROM gene_stable_id g JOIN object_xref xr JOIN xref x USING(xref_id) WHERE xr. ensembl_object_type='Gene' AND xr. ensembl_id=g. gene_id AND external_db_id=1100 25/11/2009 - Pg. 101 v 1 © Van Belle Werner

Ensembl to HGNC Find all ensembl genes that have no existing HGNC name Fourth: get rid of all the existing identifiers from the full stable_id list. SELECT stable_id FROM gene_stable_id WHERE stable_id NOT IN (SELECT DISTINCT g. stable_id FROM gene_stable_id g JOIN object_xref xr JOIN xref x USING(xref_id) WHERE xr. ensembl_object_type='Gene' AND xr. ensembl_id=g. gene_id AND external_db_id=1100) 15654 genes have no HGNC identifier 25/11/2009 - Pg. 102 v 1 © Van Belle Werner

HGNC to Uniprot Mapping Write a query that will map each known HGNC identifier to a Uniprot identifier Problem HGNC deals with genes Uniprot deals with proteins ('Translation') 25/11/2009 - Pg. 103 v 1 © Van Belle Werner

HGNC → Uniprot Map each HGNC identifier that is a gene to a Ensembl Translation identifier Map each HGNC identifier that is a transcript to a Ensembl Translation identifier Map each of those identifiers to an Uniprot identifier 25/11/2009 - Pg. 104 v 1 © Van Belle Werner

8. Simulating Realtime PCR What is the PCR ? Wrote a small simulation with a limit on the material copied How to calculate the volume after x cycles when the initial amount was I ? CT/CP Values – how to go back from a CT value to the initial amount Simulated the effect of less than 100% efficiency 25/11/2009 - Pg. 105 v 1 © Van Belle Werner

PCR Polymerase Chain reaction Denaturate DNA → single DNA strands Anneal primer – attaches only to complementary DNA Synthesize the rest of the strand – Polymerase Consumes d. NTPs (deoxynucleoside triphosphates) Enzyme + Reagents + DNA → 2 DNA + somewhat less reagents + enzyme 25/11/2009 - Pg. 106 v 1 © Van Belle Werner

RT-PCR / q. PCR Beware Reverse Transcription PCR Realtime PCR (= q. PCR) q-PCR Repetitive cycles (20 -40 cycles) Includes oligonucletodies that emit light when bound 25/11/2009 - Pg. 107 v 1 © Van Belle Werner

Simulating a RT-PCR reaction We start off with a specific volume of DNA material: amount With each cycle we increment amount by the DNA we copied (copied_dna) 25/11/2009 - Pg. 108 v 1 © Van Belle Werner
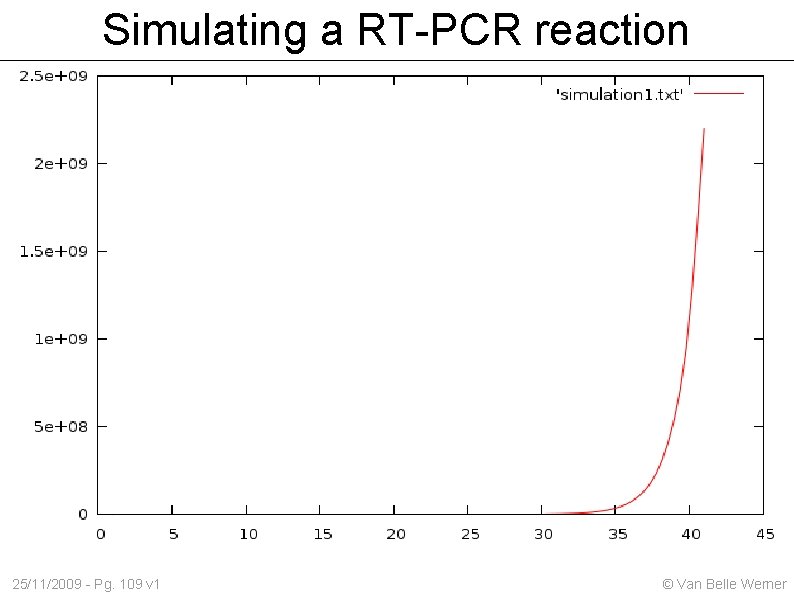
Simulating a RT-PCR reaction 25/11/2009 - Pg. 109 v 1 © Van Belle Werner

Simulating a RT-PCR reaction The cell volume is not infinite. We must observe how much reagentia is left for the reaction 25/11/2009 - Pg. 110 v 1 © Van Belle Werner
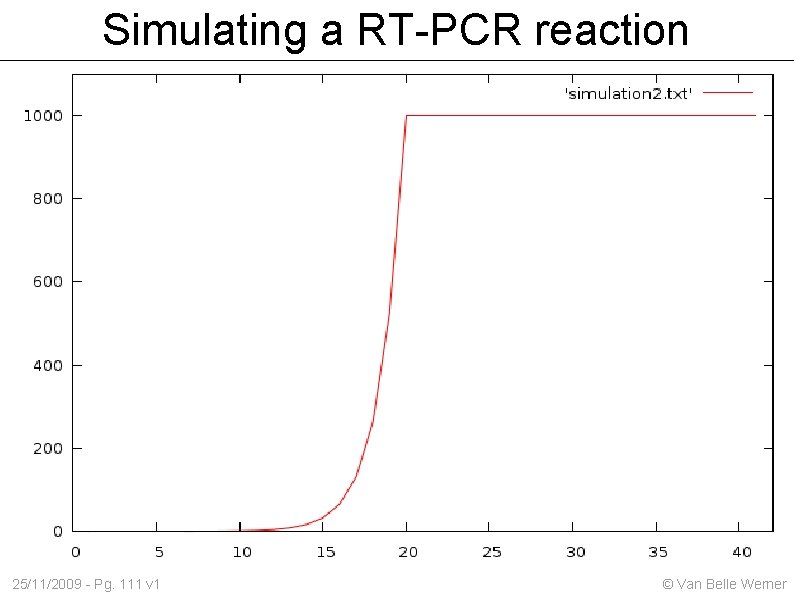
Simulating a RT-PCR reaction 25/11/2009 - Pg. 111 v 1 © Van Belle Werner

Simulating a RT-PCR reaction The usable reagents are only part of the remaining volume 25/11/2009 - Pg. 112 v 1 © Van Belle Werner
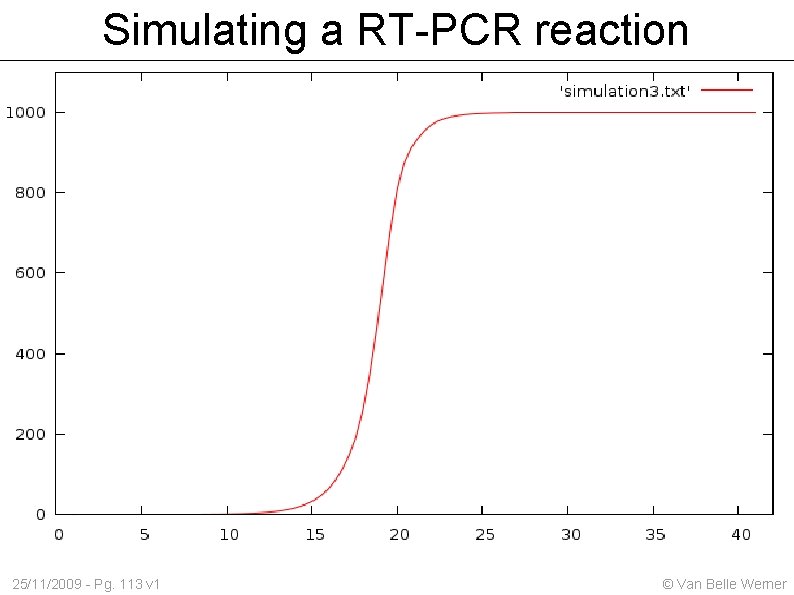
Simulating a RT-PCR reaction 25/11/2009 - Pg. 113 v 1 © Van Belle Werner
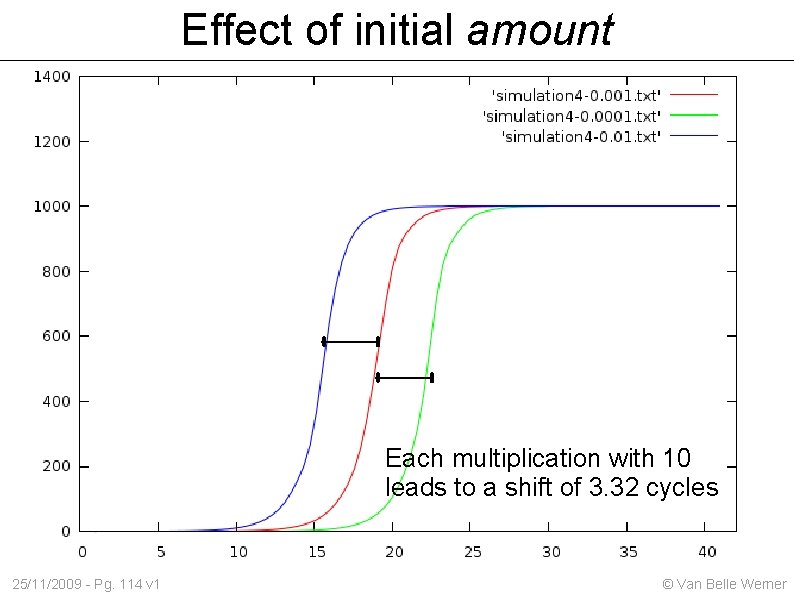
Effect of initial amount Each multiplication with 10 leads to a shift of 3. 32 cycles 25/11/2009 - Pg. 114 v 1 © Van Belle Werner
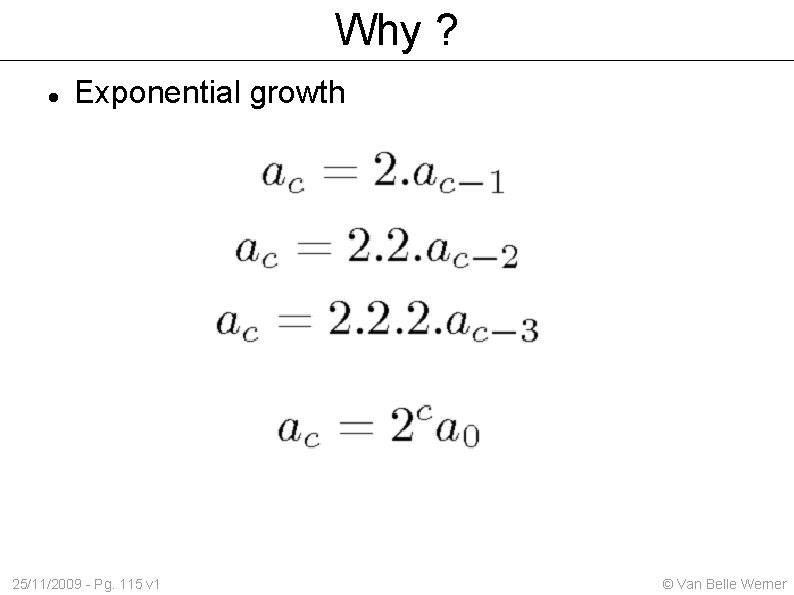
Why ? Exponential growth 25/11/2009 - Pg. 115 v 1 © Van Belle Werner
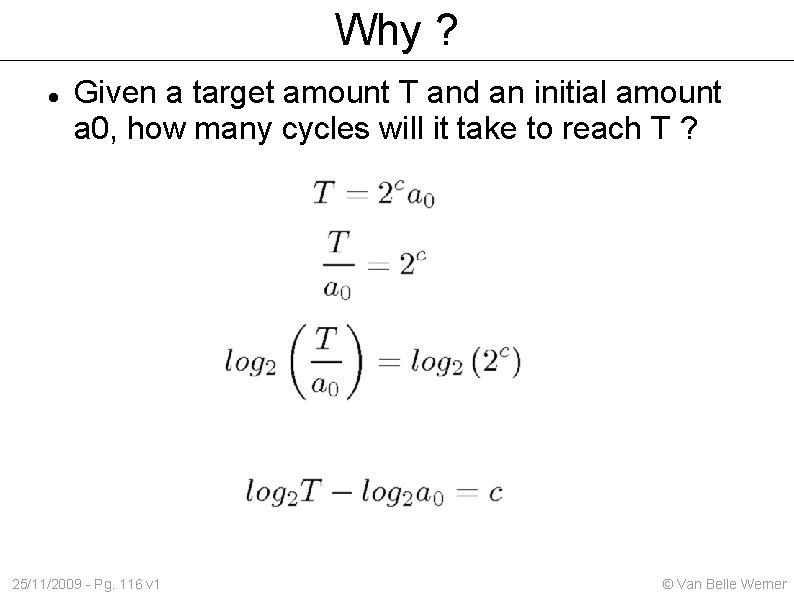
Why ? Given a target amount T and an initial amount a 0, how many cycles will it take to reach T ? 25/11/2009 - Pg. 116 v 1 © Van Belle Werner

Why ? Suppose now that the initial amount a 0 is multiplied with a factor 10, what effect does this have on the cyclecount ? 25/11/2009 - Pg. 117 v 1 © Van Belle Werner

CT / CP values Based on the 'cycles (c) to a certain threshold (T)' one can estimate the initial amount. Problem 1: we measurement after each cycle. There exists no such thing as a 3. 3 cycles. Solution: fit an exponential curve to the points we did measure. Problem 2: At a certain point the exponential growth tapers off. Solution: find the best point Still within 'exponential growth' Easy recognizable Useful 25/11/2009 - Pg. 118 v 1 © Van Belle Werner
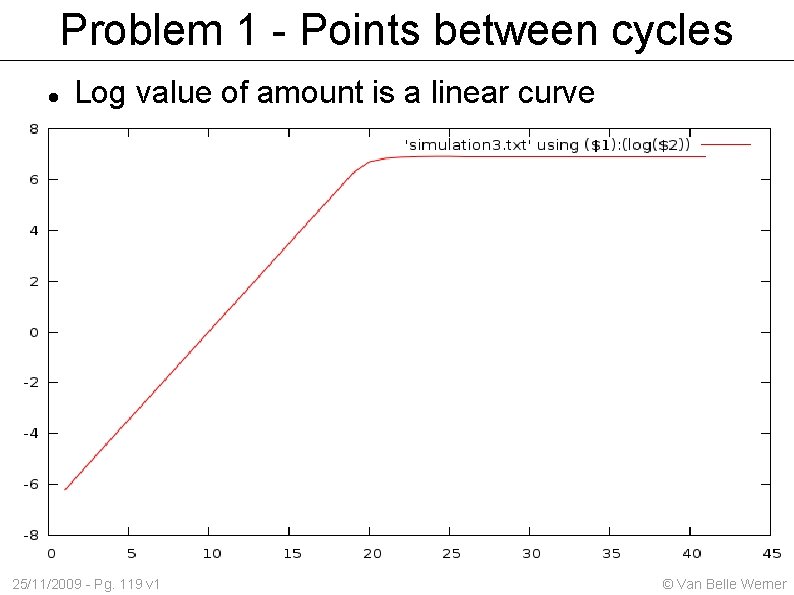
Problem 1 - Points between cycles Log value of amount is a linear curve 25/11/2009 - Pg. 119 v 1 © Van Belle Werner
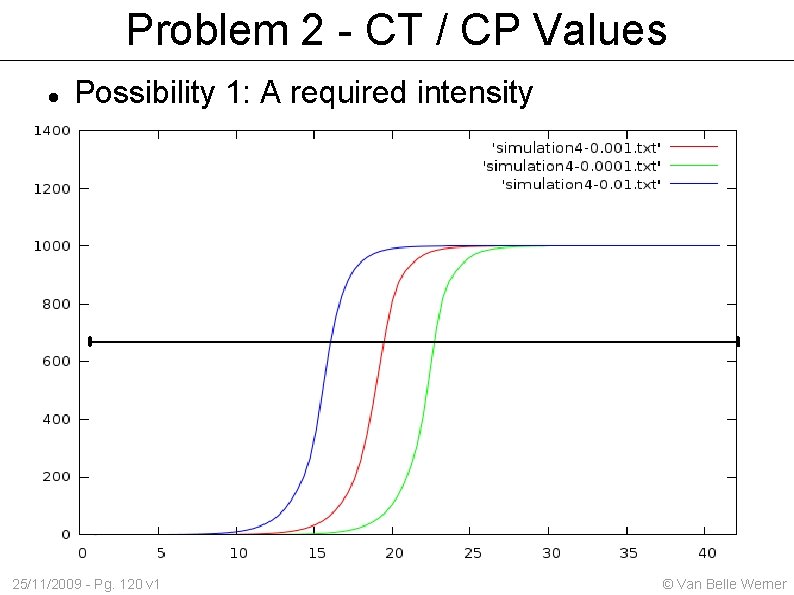
Problem 2 - CT / CP Values Possibility 1: A required intensity 25/11/2009 - Pg. 120 v 1 © Van Belle Werner
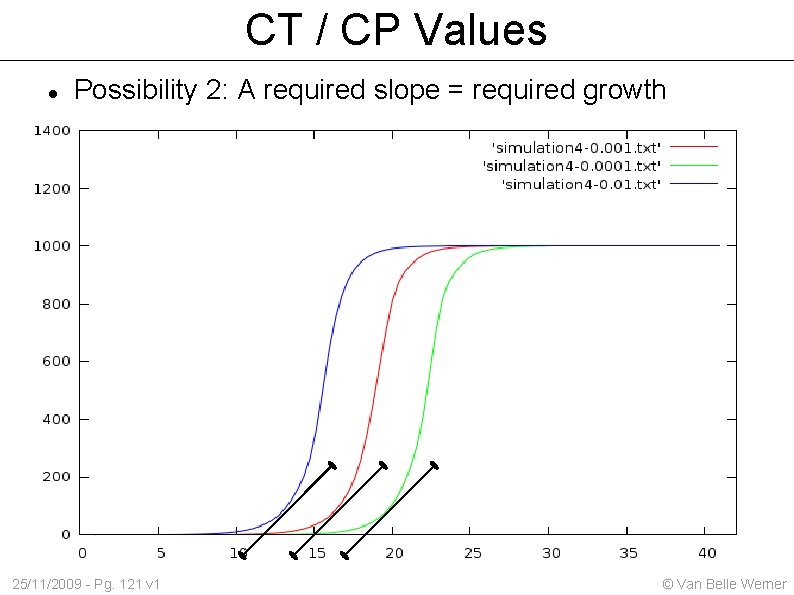
CT / CP Values Possibility 2: A required slope = required growth 25/11/2009 - Pg. 121 v 1 © Van Belle Werner
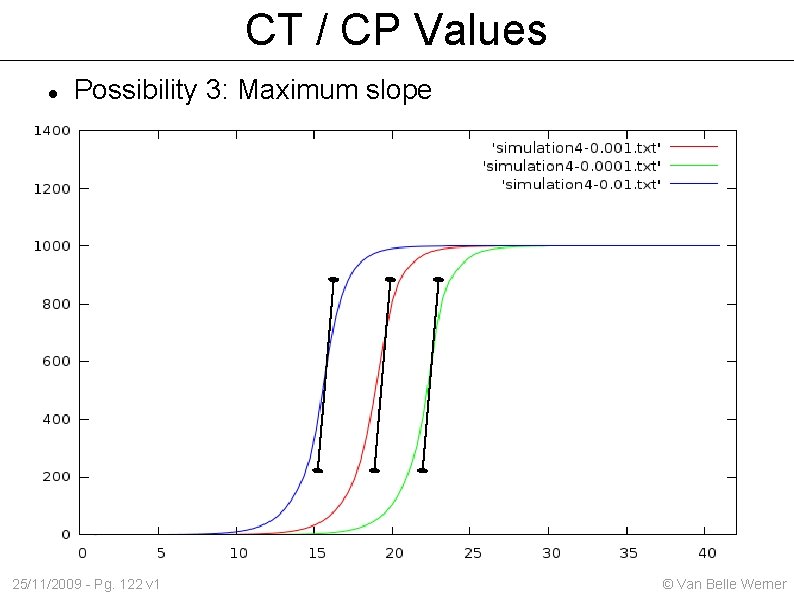
CT / CP Values Possibility 3: Maximum slope 25/11/2009 - Pg. 122 v 1 © Van Belle Werner
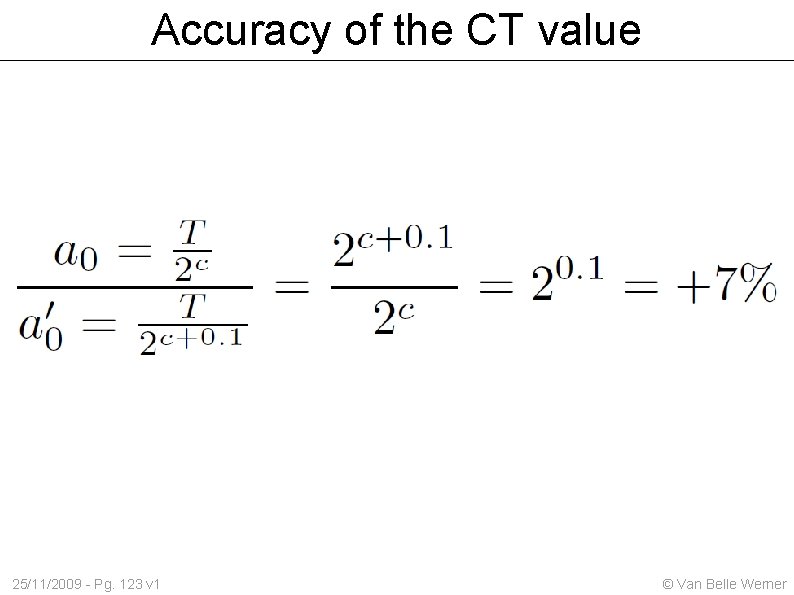
Accuracy of the CT value 25/11/2009 - Pg. 123 v 1 © Van Belle Werner

Cycle Variances Assume that with each cycle not everything is copied, but only something between 99% and 100% of the available amount, what effect will this have on our CT values ? To understand this Our algorithm needs to report its own CT value. We must modify the stepsize, instead of calculates cycle by cycle we will do it for every 1/1000 th of a cycle; this brings the error due to CT positioning down to 0. 07 % (= 0. 00069) 25/11/2009 - Pg. 124 v 1 © Van Belle Werner

Multi-step Beware off linear interpolation 25/11/2009 - Pg. 125 v 1 © Van Belle Werner

Exercise 1. Modify the simulate routine to return the cycle value before it reaches a volume of 500 2. Modify the routine such that it will decrease the efficiency of the copy process at random What is your reported CT value ? Before adding the dna_to_copy multiply it with a random number between 0. 99 and 1 3. Modify your routine to generate 1000 simulations and calculate the average reported CT value What is your result ? 25/11/2009 - Pg. 126 v 1 © Van Belle Werner

Results: 99% - 100% efficiency Initial amount: 0. 001 Without decreasing the efficiency: 18. 907 With decreasing the efficiency: 18. 9988 Difference: 0. 0918; effect on initial amount estimation: 6% underestimated Initial amount: 0. 00001 Without decreased efficiency: 25. 49 With decreased efficiency: 25. 588 Difference: 0. 098; effect on initial amount estimation: 7% underestimated 25/11/2009 - Pg. 127 v 1 © Van Belle Werner

Results 95%-100% efficiency Initial amount: 0. 001 Without decreasing the efficiency: 18. 907 With decreasing the efficiency: 19. 2524 Difference: 0. 3454; effect on initial amount estimation: 21% underestimated Initial amount: 0. 00001 Without decrease: 25. 49 With decrease: 26. 0581 Difference: 0. 5681; effect on initial amount estimation: 33% underestimated. 25/11/2009 - Pg. 128 v 1 © Van Belle Werner

9. Data Grouping Understanding questions Grouping data chunks together Across or foreach gene/plate etcera ? Layout of a PCR experiment and examples 25/11/2009 - Pg. 129 v 1 © Van Belle Werner

Unstructured Questions Calculate the up or down regulation between cell types For all or for each gene ? Including the different replicas ? Calculate the average expression in each cell line Averaged per gene after resolving replicates (each gene will have the same weight afterward) -or- directly across replicas ? Is there an effect between the cell line and the cell type Such unstructured questions can be understood and implemented differently and produce highly different results 25/11/2009 - Pg. 130 v 1 © Van Belle Werner

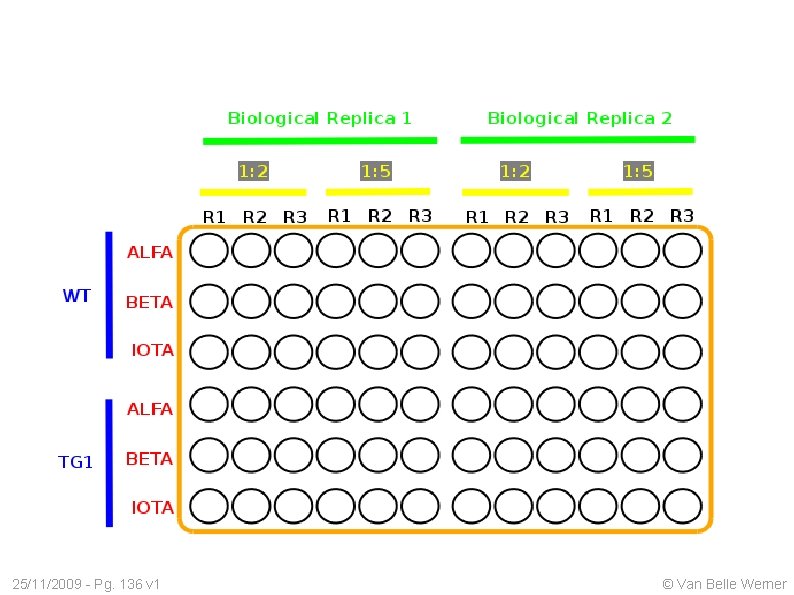
q. PCR A plate: 96 wells Different probes/gene: ALFA, BETA, IOTA A cell type: WT, TG Different dilutions: 1: 2, 1: 5, 1: 20, 1: 50 Technical Replicas: R 1, R 2 A cell line: He. La, SK-N-DZ Biological Replicas: B 1, B 2 25/11/2009 - Pg. 131 v 1 © Van Belle Werner

q. PCR: A Common Layout 25/11/2009 - Pg. 132 v 1 © Van Belle Werner

Why think about groups ? Group information is often implicit. If it is implicit: assume foreach. Groups can help to resolve missing data-points Groups determine the control flow in an analysis Calculate everything on technical replicates, then average things out over the biological replicates -or. Pool all technical and biological replicates together before continuing with the analysis Not all potential groups make sense Calculating the average of all dilutions is only possible if we have the same number of elements in each replica → dangerous to do 25/11/2009 - Pg. 133 v 1 © Van Belle Werner

Why think about groups ? A group of data tends to be smaller than the full dataset (we do not need to load other groups) Can make streaming possible Requires less RAM E. g: calculate an exon overlap map for each chromosome Can allow parallel execution Dependencies between data groups Recalculate only necessary groups 25/11/2009 - Pg. 134 v 1 © Van Belle Werner

Language Issues Foreach, Per, (Forall) Foreach gene means that each group will only deal with one gene at a time. Forall, Across, Ignoring, (Pooled, Grouped), Aggregate. . . denotes a separation between groups. Denotes an aggregation of data independent of this particular variable Forall genes means that ALFA, BETA, THETA etc can all be included in each individual group. Pairs, Couples, Combinations, Multiples, Between Denotes subgroups within larger groups 25/11/2009 - Pg. 135 v 1 © Van Belle Werner
25/11/2009 - Pg. 136 v 1 © Van Belle Werner
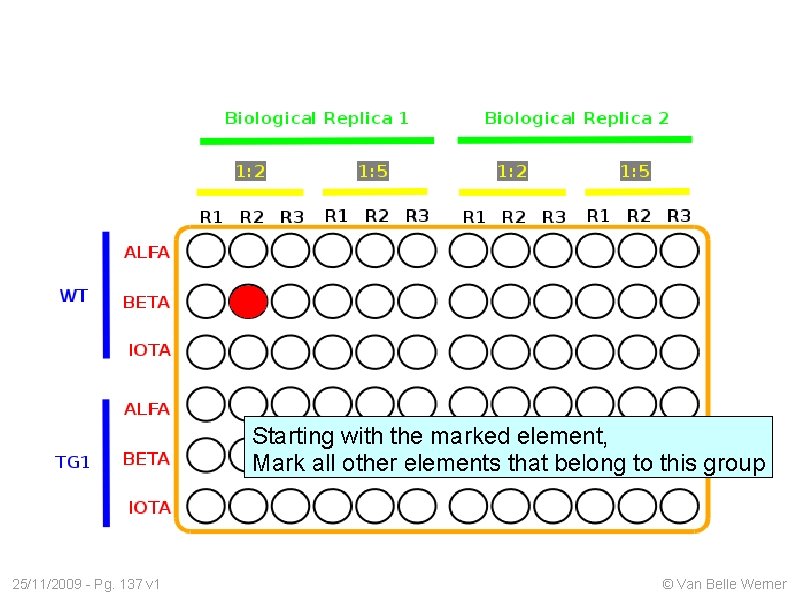
Starting with the marked element, Mark all other elements that belong to this group 25/11/2009 - Pg. 137 v 1 © Van Belle Werner
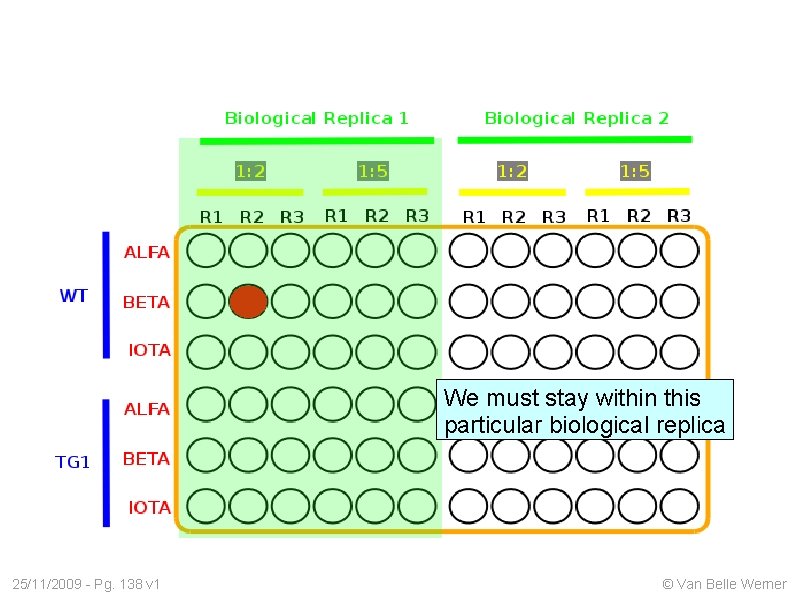
We must stay within this particular biological replica 25/11/2009 - Pg. 138 v 1 © Van Belle Werner
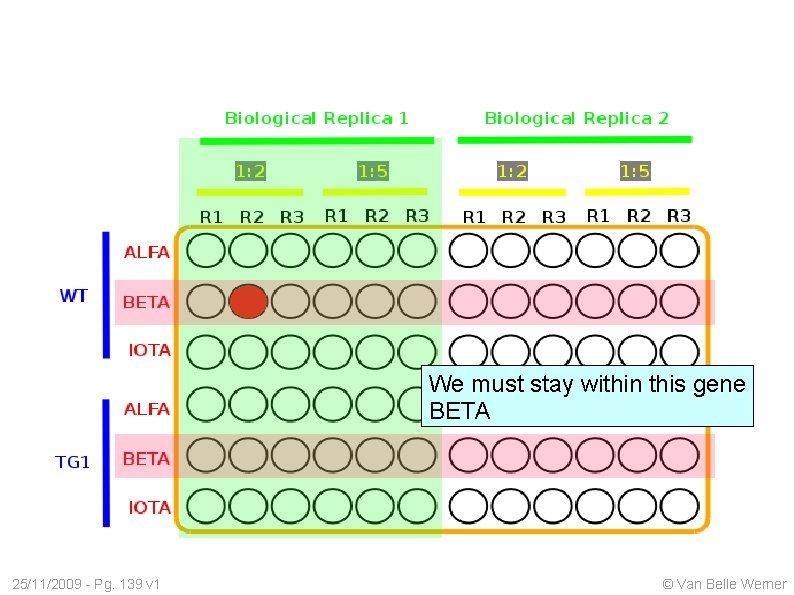
We must stay within this gene BETA 25/11/2009 - Pg. 139 v 1 © Van Belle Werner
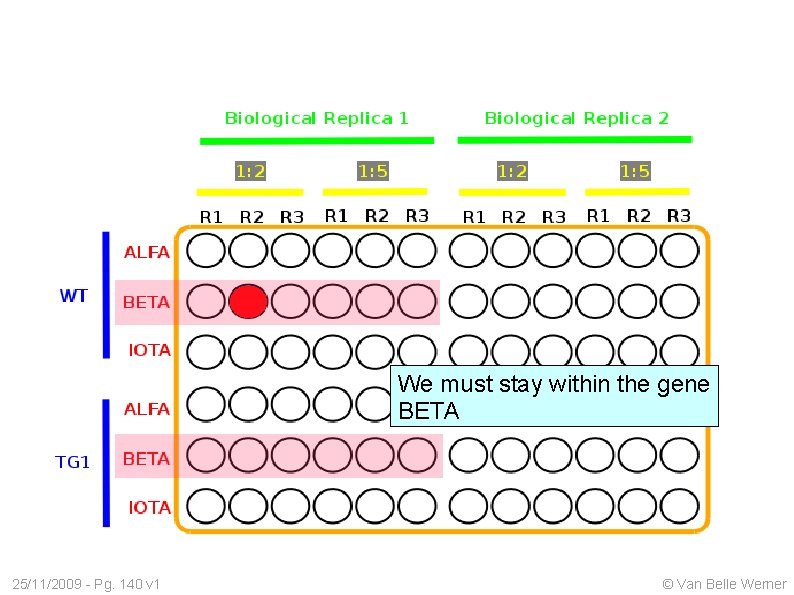
We must stay within the gene BETA 25/11/2009 - Pg. 140 v 1 © Van Belle Werner
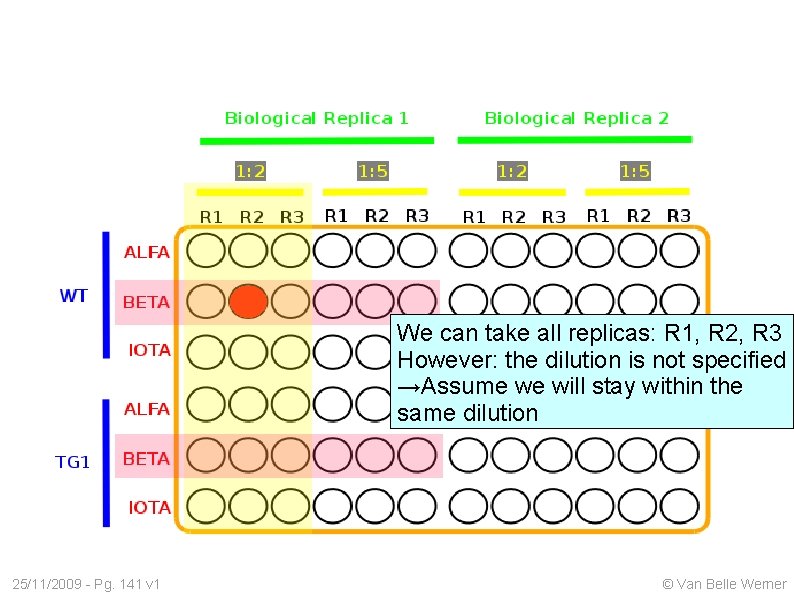
We can take all replicas: R 1, R 2, R 3 However: the dilution is not specified →Assume we will stay within the same dilution 25/11/2009 - Pg. 141 v 1 © Van Belle Werner
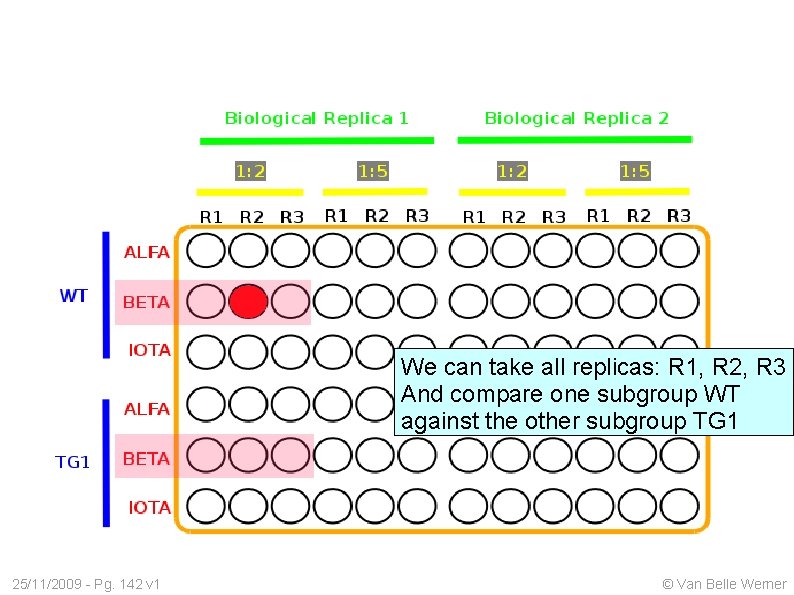
We can take all replicas: R 1, R 2, R 3 And compare one subgroup WT against the other subgroup TG 1 25/11/2009 - Pg. 142 v 1 © Van Belle Werner
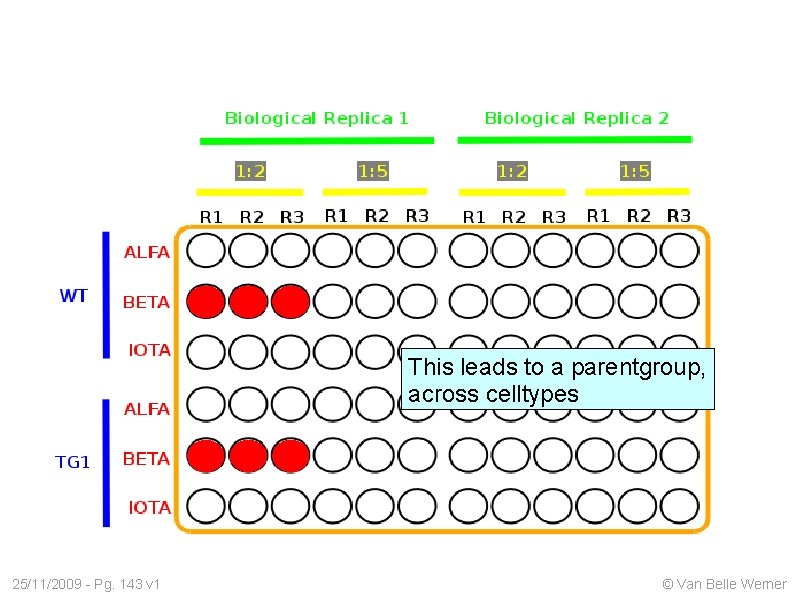
This leads to a parentgroup, across celltypes 25/11/2009 - Pg. 143 v 1 © Van Belle Werner
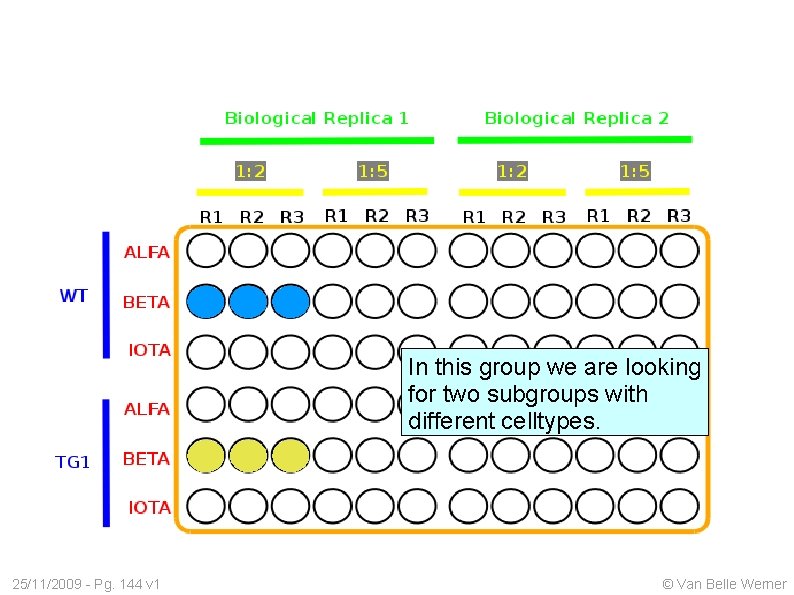
In this group we are looking for two subgroups with different celltypes. 25/11/2009 - Pg. 144 v 1 © Van Belle Werner

Dealing with combinations If variable X is listed as a 'combination' A celltype combination, or a dilution combination First create the parentgroup that assumes X is a group variable. Celltype is treated as a group Dilution is treated as a group From this parentgroup one can select a subgroup identified by a value of X. A subgroup where Celltype = WT A subgroup where Dilution = 1: 2 This subgroup is then the first group of the combination 25/11/2009 - Pg. 145 v 1 © Van Belle Werner
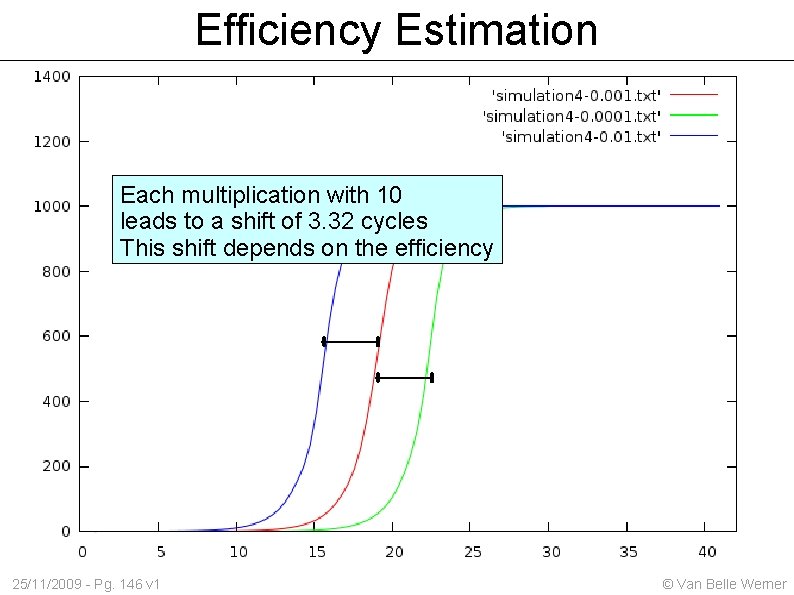
Efficiency Estimation Each multiplication with 10 leads to a shift of 3. 32 cycles This shift depends on the efficiency 25/11/2009 - Pg. 146 v 1 © Van Belle Werner

Efficiency Estimation How do we want to estimate the PCR efficiency ? For each plate For each probe/gene For each celltype (wildtype, modified) For each dilution combination For each replica Exercise Extend the group provided to you to include all elements of that group Color a second group belonging to the 'dilution combination' Belle Werner [Write down an object hierarchy to access the© Van data 25/11/2009 - Pg. 147 v 1
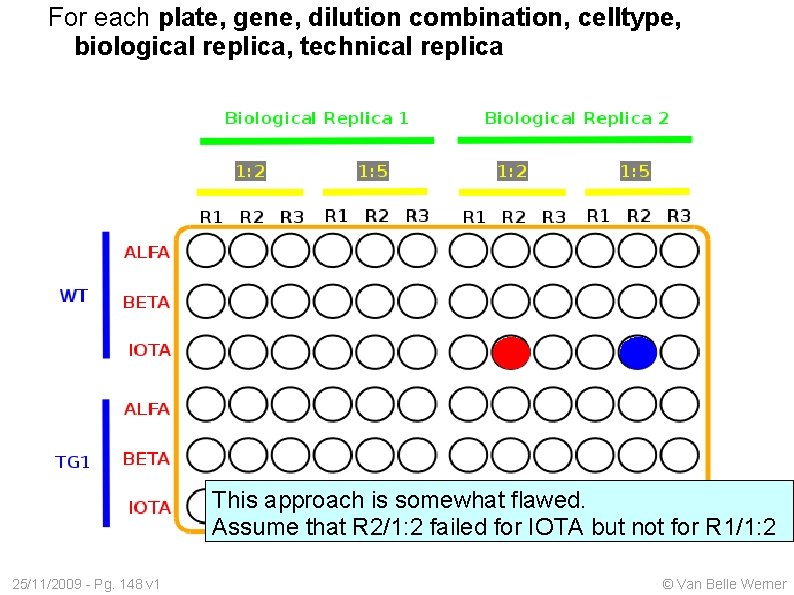
For each plate, gene, dilution combination, celltype, biological replica, technical replica This approach is somewhat flawed. Assume that R 2/1: 2 failed for IOTA but not for R 1/1: 2 25/11/2009 - Pg. 148 v 1 © Van Belle Werner
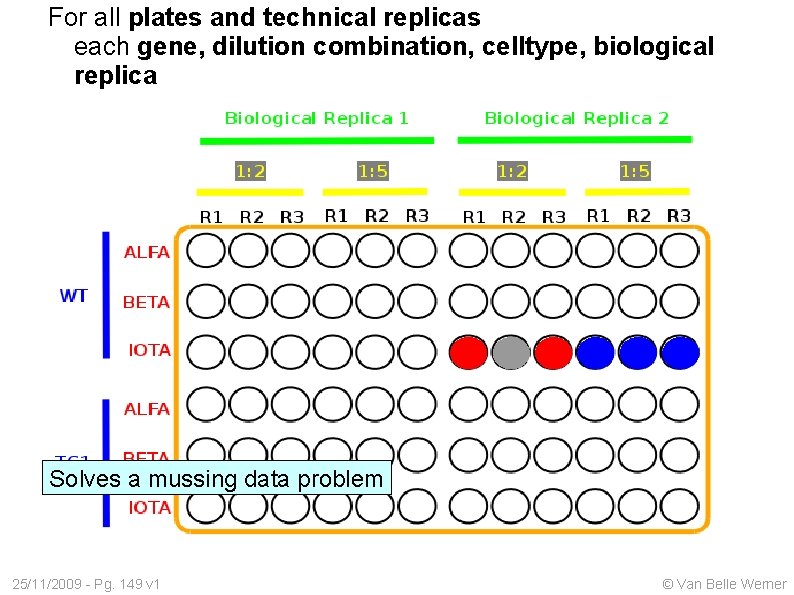
For all plates and technical replicas each gene, dilution combination, celltype, biological replica Solves a mussing data problem 25/11/2009 - Pg. 149 v 1 © Van Belle Werner

10. Accessing Data Groups An educational API to explore data groups Standard object hierarchies are difficult Data grouping enables concurrent access Helps with optimalisation (don't calculate what didn't change) Cleaned up the output of a q. PCR experiment 25/11/2009 - Pg. 150 v 1 © Van Belle Werner

Object Hierarchies Calculate the median across replicas and across plates, remove bad measurements first Calculate the copyratio per gene based on all dilution pairs Gene → Dilution → Cell Type → Plate* → average replicas Gene → Dilution → Cell Type → average replicas Gene → Replica → Dilution → Cell Type Gene → Dilution → [Cell Type] → CP Gene → Cell Type → Dilution → CP Calculate the up down regulation between two genes Gene → Dilution → Cell Type → Concentration → Gene 25/11/2009 - Pg. 151→ v 1 same Dilution → same Cell Type → Concentration © Van Belle Werner

Data Grouping Hard coding a data representation/object hierarchy often interferes with different data views / rotations of the data. XML/SQL/OODB SQL can group data for you Unsuitable to perform an analysis loading group by group from a database can be highly time consuming (latency) Group identification can be problematic. Requires a high level API that can be used to deal with data: data slices. 25/11/2009 - Pg. 152 v 1 © Van Belle Werner
A Table Interface A Table represents a table with attributes, records and values Retrieve all unique keys given an attribute list Retrieve all records associated with a specific key Differently stated: retrieve the group associated with (or identified by) key Retrieve a value from a record Retrieve all values for an attribute in a table (a column) Iterate over all records in a table Iterate over all groups in a table 25/11/2009 - Pg. 153 v 1 © Van Belle Werner

Table – loading in a tsv Table table=new Table("qpcr. tsv"); Will load the tab seperated value file qpcr. tsv in memory Console. Out. Write(table) Will print out the table content, record by record 25/11/2009 - Pg. 154 v 1 © Van Belle Werner

Records Each record maps an attribute (String) to a IComparable object (String, Double, …) Record r=new Record(); r[“Averaged CP”]=average; Records can be shared between tables ! Records should be treated read-only after creation and filling them with data. Records can be copied; the content of the record (the values that is) are not copied: r. copy(); 25/11/2009 - Pg. 155 v 1 © Van Belle Werner
![Adding a record to a table Table table=new Table(); Record record=new Record(); record[“test”]=60000; table. Adding a record to a table Table table=new Table(); Record record=new Record(); record[“test”]=60000; table.](http://slidetodoc.com/presentation_image/a6c9df4e44cd58f5d0b83a887f4c2c97/image-156.jpg)
Adding a record to a table Table table=new Table(); Record record=new Record(); record[“test”]=60000; table. add(record); Console. Out. Write(table); Records should be added only after their creation and initialization. 25/11/2009 - Pg. 156 v 1 © Van Belle Werner

Retrieving a set of keys Table table=new Table(“qpcr. txt”); table. keys(“Cell. Type”); Returns a list of unique Records, with only the attribute 'Cell. Type' table. keys(“Cell. Type”, ”Gene”); Returns a list of unique records with two attributes: 'Cell. Type and Gene' The return value is a List<Record> 25/11/2009 - Pg. 157 v 1 © Van Belle Werner

Retrieving values belonging to a key table. group(key) Key is a record (Cell. Type: WT; Gene: ALFA) The returned value is again a new Table. Remember the records are shared between the returned subtable and the parent table. Do not modify records after they have been created and initialized. If a record is added to the parent table it will not automatically appear in potential subtables. 25/11/2009 - Pg. 158 v 1 © Van Belle Werner
![Retrieving values from an attribute record[“Cell. Type”] returns the IComparable content in this record Retrieving values from an attribute record[“Cell. Type”] returns the IComparable content in this record](http://slidetodoc.com/presentation_image/a6c9df4e44cd58f5d0b83a887f4c2c97/image-159.jpg)
Retrieving values from an attribute record[“Cell. Type”] returns the IComparable content in this record table[“Cell. Type”] returns an Array. List of values linked to that attribute. 25/11/2009 - Pg. 159 v 1 © Van Belle Werner
![Iterating over the records in a table foreach(Record tr in table) { String str=(String)tr["CP"]; Iterating over the records in a table foreach(Record tr in table) { String str=(String)tr["CP"];](http://slidetodoc.com/presentation_image/a6c9df4e44cd58f5d0b83a887f4c2c97/image-160.jpg)
Iterating over the records in a table foreach(Record tr in table) { String str=(String)tr["CP"]; … } 25/11/2009 - Pg. 160 v 1 © Van Belle Werner

Iterating over all groups in a table List<Table> groups=table. groups(“A”, ”B”, . . . ); foreach(Table group in groups) { group. key → the current group identification … } 25/11/2009 - Pg. 161 v 1 (contains A, B, . . ) © Van Belle Werner

Advantages No need for an objects structure representing the data Flexible with regard to different regrouping (data rotations) Foreach group Flexible wrt new data fields The inner loop can theoretically be executed in parallel If necessary an SQL backend can be placed in the Table interface 25/11/2009 - Pg. 162 v 1 © Van Belle Werner

Exercise: Data Cleanup Document which attributes exist in qpcr. tsv Write a routine that will run through the dataset qpcr. tsv and clean up the data. All technical replicas should be averaged (for each of the other attributes) Replicas with useless data should be omitted cycle numbers >40 marked as bad [34. 56] without a value [outlier replicas (median)] 25/11/2009 - Pg. 163 v 1 © Van Belle Werner

11. Normalizing Efficiencies How to estimate the efficiency of a q. PCR experiment using dilution series How to normalize the CT values based on an estimated efficiency Created a normalized table 25/11/2009 - Pg. 164 v 1 © Van Belle Werner

Creating combinations Write a routine that will for each plate, cell line, cell type and probe report all dilution combinations of the averaged cp-values of the technical replicas 1. Reuse the table you created in the previous exercise 2. Foreach (plate, cellline, celltype, probe) obtain the associated group. Assume we want to have all dilutions included 3. Print out each combination of different dilutions 25/11/2009 - Pg. 165 v 1 © Van Belle Werner
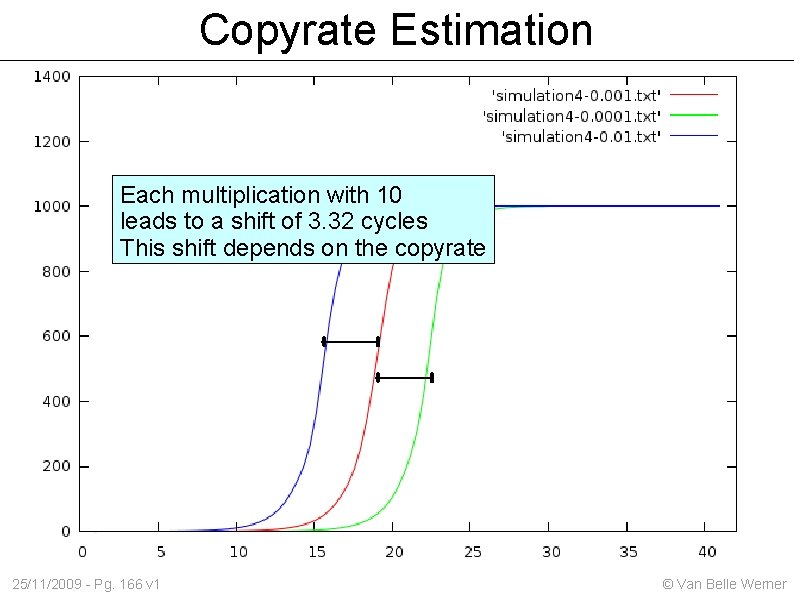
Copyrate Estimation Each multiplication with 10 leads to a shift of 3. 32 cycles This shift depends on the copyrate 25/11/2009 - Pg. 166 v 1 © Van Belle Werner

Normalization through dilution series If amplification were 100% efficient then halving the initial concentration would shift the measurement 3. 32 cycles to the right. In reality it isn't and we will see shifts larger/smaller than 3. 32 cycles. By creating a dilution series, one can estimate the copyrate / efficiency 25/11/2009 - Pg. 167 v 1 © Van Belle Werner

Copyrate Calculation If we have a dilution of factor 10 (becomes stronger) And we have a shift (to the left) of x cycles, then the copy ratio (r) is 25/11/2009 - Pg. 168 v 1 © Van Belle Werner

Examples After diluting a sample a factor 10 we observe a shift of 3. 33 → ratio = 2 3. 36 → ratio = 1. 98 3. 5 → ratio = 1. 93 3. 6 → ratio = 1. 89 3. 7 → ratio = 1. 86 25/11/2009 - Pg. 169 v 1 © Van Belle Werner

Routine to Calc. Eff. Based on the formula given before; write a routine Double estimate_r(Double x, Double y, Double ratio_x 2 y); To estimate the average multiplication factor for each cycle. To test your routine use the following inputs X=10; Y=13. 32; ratio_x 2 y=0. 1 → copyratio 2 X=10; Y=6. 68; ratio_x 2 y=10 → copyratio 2 25/11/2009 - Pg. 170 v 1 © Van Belle Werner

Estimating the copyratio Plug in your routine into Ex. 2 such that the average copy-ratio is calculated for each probe, celline, celltype and plate. All combinations should be taken into account. Use the function Double estimate_r(Double x, Double y, Double ratio_x 2 y)to return the cycleratio. 25/11/2009 - Pg. 171 v 1 © Van Belle Werner

Remarks on Efficiency One could also look at the material copied with each cycle. It should double. Based on that we have a direct measurement of the efficiency. In low areas our sensititve is not sufficient to estimate In High areas before the last cycle we have such a position Only one measurement Depends somewhat on the software provided with the machines 25/11/2009 - Pg. 172 v 1 © Van Belle Werner

Remarks on Efficiency Often the efficiency is expressed as the relative amount of input material that was used to create new DNA: efficiency = r-1. 0. (between 80% and 140%) Double estimate_efficiency (Double x, Double y, Double ratio_x 2 y) { return 100. 0*(estimate_r(x, y, ratio_x 2 y)-1. 0); } 25/11/2009 - Pg. 173 v 1 © Van Belle Werner

Normalizing CT Values Assume the reached volume was V, after x cycles at a copyrate of r What would be the number of cycles if the copyrate were 2 ? 25/11/2009 - Pg. 174 v 1 © Van Belle Werner

Normalizing CT Values It could also be possible to go back to the initial concentration instead of relying on normalizing the CT values 25/11/2009 - Pg. 175 v 1 © Van Belle Werner

Normalize the CT Value Use your estimated copyratio to normalize the CT value Implement the normalization equation Place the normalized CT values in a new table 25/11/2009 - Pg. 176 v 1 © Van Belle Werner

12. Dct Values Normalizing the results towards a houshold gene 25/11/2009 - Pg. 177 v 1 © Van Belle Werner

Normalization ? Cell. Type might affect the baseline gene expression in the cell. A WT cell might be less active than a TG cell Or vice versa To account for this problem one can compare a gene expression against a 'household' gene The household gene is supposed to be non related to the measured gene As we know from microarrays, using one gene to normalize various expressions is highly errorprone 25/11/2009 - Pg. 178 v 1 © Van Belle Werner

Normalizing against a known gene If CP_A is the CP value for gene of interest A And CP_H is the CT value for the householdgene H The concentrations [A] and [H] are given by 25/11/2009 - Pg. 179 v 1 © Van Belle Werner

Normalizing against a known gene The ratio of the concentrations is then given by 25/11/2009 - Pg. 180 v 1 © Van Belle Werner

Example A has a CT value of 15. 786 H has a CT value of 18. 875 DCT = 18. 875 -15. 786 The concentration ratio is 8. 51 This is an upregulation of a factor 8. 51 (against the household gene concentration) 25/11/2009 - Pg. 181 v 1 © Van Belle Werner

Example A has a CT value of 19. 6 H has a CT value of 12. 4 DCT = 12. 4 -19. 6 = -7. 2 The concentration ratio is 0. 068011 Which is a down regulation of a factor 147. 03 25/11/2009 - Pg. 182 v 1 © Van Belle Werner

Calculating DCT Using GADPH as a houehold gene, Calculate for any other gene the DCT value and report it in a new table 25/11/2009 - Pg. 183 v 1 © Van Belle Werner

13. Delta-Delta CT Calculating DDCT values from one celltype to another. 25/11/2009 - Pg. 184 v 1 © Van Belle Werner

Calculating up/down regulations Up/down regulations are typically calculated between celltypes. E. g: the relative expression of gene A in WT condition against the relative expression of gene A in TG condition. 25/11/2009 - Pg. 185 v 1 © Van Belle Werner

Example WT: A has a CT value of 15. 786 H has a CT value of 18. 875 DCT = 18. 875 -15. 786=3. 089 TG: A has a CT value of 19. 6 H has a CT value of 12. 4 DCT = 12. 4 -19. 6 = -7. 2 DDCT = -7. 2 -3. 089=-10. 289 WT/TG=0. 000799286 or a down regulation of a factor 1251. 12 25/11/2009 - Pg. 186 v 1 © Van Belle Werner

14. Reporting Regulations Reporting regulations as Log values Ratios larger than 1 25/11/2009 - Pg. 187 v 1 © Van Belle Werner

Reporting up-down regulations Can be reported as a ratio x 10 x 0. 01 x 0 Can be reported as a log value x 10 → log_10 value of 1 x 0. 1 → og_10 value of -1 x 1 → log_10 value of 0 x 0 → has no log_10 value 25/11/2009 - Pg. 188 v 1 © Van Belle Werner

Reporting up/down regulations Can be reported as a ratio and a direction x 10 → 10 times upregulated x 0. 1 → 10 times downregulated Exercise: Based on your DDCT table, create a report for the up / down regulations of each measured gene from the WT to the TG 25/11/2009 - Pg. 189 v 1 © Van Belle Werner
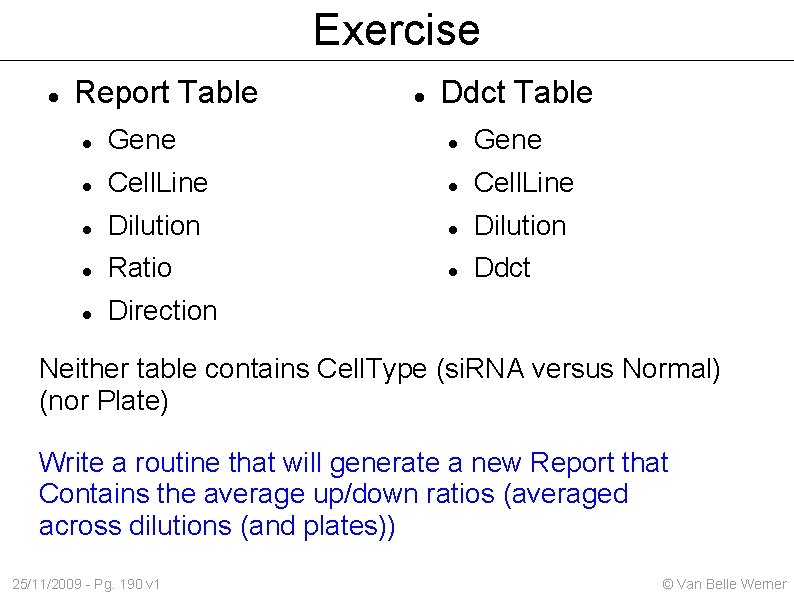
Exercise Report Table Ddct Table Gene Cell. Line Dilution Ratio Ddct Direction Neither table contains Cell. Type (si. RNA versus Normal) (nor Plate) Write a routine that will generate a new Report that Contains the average up/down ratios (averaged across dilutions (and plates)) 25/11/2009 - Pg. 190 v 1 © Van Belle Werner
- Slides: 190