Genome Rearrangements Belle marquise vos beaux yeux me
Genome Rearrangements Belle marquise, vos beaux yeux me font mourir d'amour. Vos yeux beaux d'amour me font, belle marquise, mourir. Me font vos beaux yeux mourir, belle marquise, d'amour. Anne Bergeron, Comparative Genomics Laboratory Université du Québec à Montréal

1. General introduction to genome rearrangements Examples of rearranged genomes 2. Measures of distance Rearrangement operations The Hannenhalli-Pevzner distance equation 3. A unifying view of genome rearrangements The Double-Cut-and-Join operation The adjacency graph and the distance equation

1. General introduction to genome rearrangements Examples of rearranged genomes 2. Measures of distance Rearrangement operations The Hannenhalli-Pevzner distance equation 3. A unifying view of genome rearrangements The Double-Cut-and-Join operation The adjacency graph and the distance equation
Example of rearranged genomes : Mitochondrial Genomes Mitochondria are small, oval shaped organelles surrounded by two highly specialized membranes. Homo sapiens Bombyx mori Animal mitochondrial genomes are normally circular, ~16 k. B in length, and encode: 13 proteins 22 t. RNAs and 2 r. RNAs.

Example of rearranged genomes : Mitochondrial Genomes Here is an alignment of the cytochrome c oxidase I of, respectively, Homo sapiens and Bombyx mori. RWLFSTNHKDIGTLYLLFGAWAGVLGTALSLLIRAELGQPGNLLGNDHIYNVIVTAHAFVMIFFMVMPIMIGGFGNWLVPLMIGAPDMAFPRMNNM KWIYSTNHKDIGTLYFIFGIWSGMIGTSLSLLIRAELGNPGSLIGDDQIYNTIVTAHAFIMIFFMVMPIMIGGFGNWLVPLMLGAPDMAFPRMNNM : *: : ******: : ** *: *: : ******: **. *: *: *: *******: ***********: ******* : X: : XXXXXX: : XX X: X: : XXXXXX: XX. X: X: X: XXXXXXX: XXXXXXXXXXX: XXXXXXX SFWLLPPSLLLLLASAMVEAGAGTGWTVYPPLAGNYSHPGASVDLTIFSLHLAGVSSILGAINFITTIINMKPPAMTQYQTPLFVWSVLITAVLLLLSLP SFWLLPPSLMLLISSSIVENGAGTGWTVYPPLSSNIAHSGSSVDLAIFSLHLAGISSIMGAINFITTMINMRLNNMSFDQLPLFVWAVGITAFLLLLSLP *****: **: : ** ******: . * XXXXX: XX: : XX XXXXXX: . X : *. *: ********: ***: : X. X: XXXXXXXX: XXX: *: * *****: * ******* X: X XXXXX: X XXXXXXX VLAAGITMLLTDRNLNTTFFDPAGGGDPILYQHLFWFFGHPEVYILILPGFGMISHIVTYYSGKKEPFGYMGMVWAMMSIGFLGFIVWAHHMFTVGMDVD VLAGAITMLLTDRNLNTSFFDPAGGGDPILYQHLFWFFGHPEVYILILPGFGMISHIISQESGKKETFGCLGMIYAMLAIGLLGFIVWAHHMFTVGMDID ***. . ******: ********************: : *****. ** : **: : *********: * XXXXX. XX : XX: : XXXXXXXXX: X XXX. . XXXXXX: XXXXXXXXXXXXXXXXXXXX: : TRAYFTSATMIIAIPTGVKVFSWLATLHGSNMKWSAAVLWALGFIFLFTVGGLTGIVLANSSLDIVLHDTYYVVAHFHYVLSMGAVFAIMGGFIHWFPLF TRAYFTSATMIIAVPTGIKIFSWLATMHGTQINYNPNILWSLGFVFLFTVGGLTGVILANSSIDITLHDTYYVVAHFHYVLSMGAVFAIIGGFINWYPLF *******: *: ******: : : . . : ***: *****: **. ************: *: *** XXXXXXX: X: XXXXXX: : : . . : XXX: XXXXX: XX. XXXXXXXXXXXX: X: XXX SGYTLDQTYAKIHFTIMFIGVNLTFFPQHFLGLSGMPRRYSDYPDAYTTWNILSSVGSFISLTAVMLMIFMIWEAFASKRKVLMVEEPSMNLE TGLSLNSYMLKIQFFTMFIGVNMTFFPQHFLGLAGMPRRYSDYPDSYISWNMISSLGSYISLLSVMMMLIIIWESMINQRINLFSLNLPSSIE : *: . : X: . **: * XX: X ******: ***********: * XXXXXX: XXXXXXXXXXX: X : **: **: *** : **: *: : : ***: : : XX: XXX. : * *: : . . : * : XX: X: : : XXX: : . : X X: : . . : X 73% identity over more than 500 amino acids.

Example of rearranged genomes : Mitochondrial Genomes The 37 genes of animal mitochondria are highly conserved. Charles Darwin, 1809 - 1882 A lowly worm But the order of the genes differs from species to species.
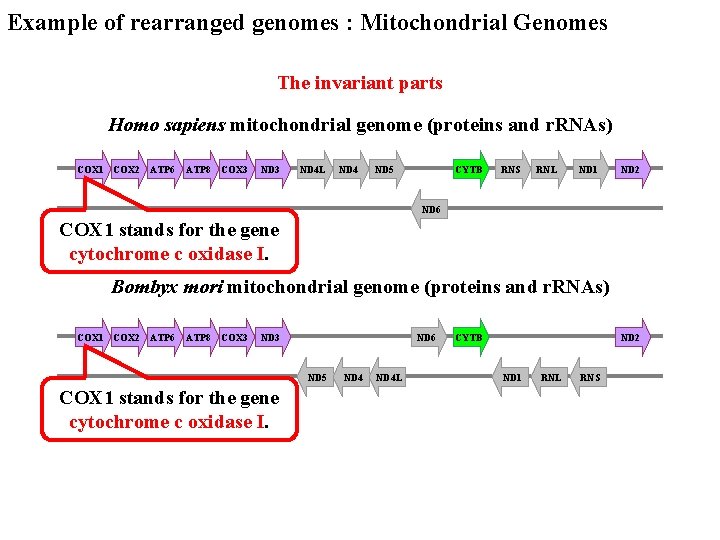
Example of rearranged genomes : Mitochondrial Genomes The invariant parts Homo sapiens mitochondrial genome (proteins and r. RNAs) COX 1 COX 2 ATP 6 ATP 8 COX 3 ND 4 L ND 4 ND 5 CYTB RNS RNL ND 1 ND 2 ND 6 COX 1 stands for the gene cytochrome c oxidase I. Bombyx mori mitochondrial genome (proteins and r. RNAs) COX 1 COX 2 ATP 6 ATP 8 COX 3 ND 6 ND 5 COX 1 stands for the gene cytochrome c oxidase I. ND 4 L CYTB ND 2 ND 1 RNL RNS
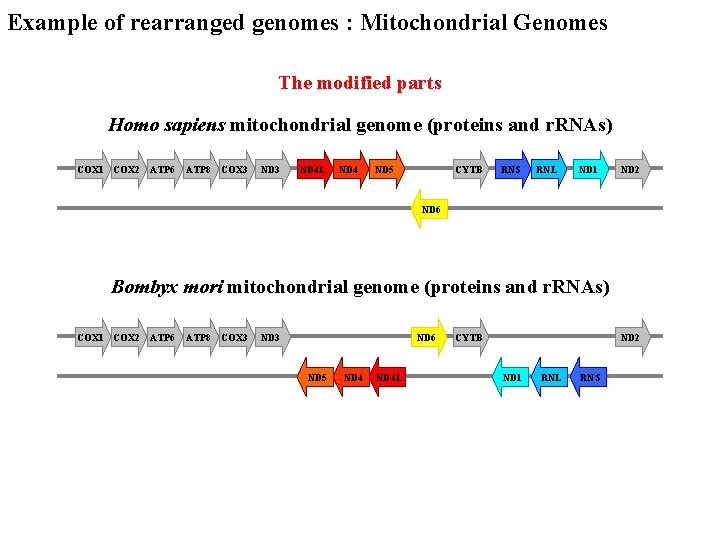
Example of rearranged genomes : Mitochondrial Genomes The modified parts Homo sapiens mitochondrial genome (proteins and r. RNAs) COX 1 COX 2 ATP 6 ATP 8 COX 3 ND 4 L ND 4 ND 5 CYTB RNS RNL ND 1 ND 2 ND 6 Bombyx mori mitochondrial genome (proteins and r. RNAs) COX 1 COX 2 ATP 6 ATP 8 COX 3 ND 6 ND 5 ND 4 L CYTB ND 2 ND 1 RNL RNS

Example of rearranged genomes : Mitochondrial Genomes of 6 Arthropoda Fruit Fly Mosquito Silkworm Locust Tick Centipede Identical ‘runs’ of genes have been grouped.

Example of rearranged genomes : mammal X chromosomes Sixteen large synteny blocks are ordered differently in the X chromosomes of the human, mouse and rat. Blocks have similar gene content and order. Note that the estimated number of genes in the X chromosome is 2000. (Art work by Guillaume Bourque, scientific work by Guillaume Bourque, Pavel Pevzner and Glenn Tesler, 2004)

Example of rearranged genomes : mammal X chromosomes (Art work by Guillaume Bourque, scientific work by Guillaume Bourque, Pavel Pevzner and Glenn Tesler, 2004)

Problem: Given two or more genomes, How do we measure their similarity and/or distance with respect to gene order and gene content? Sub-problem: How do we know that two genes or blocks are the "same" in two different species?

1. General introduction to genome rearrangements Examples of rearranged genomes 2. Measures of distance Rearrangement operations The Hannenhalli-Pevzner distance equation 3. A unifying view of genome rearrangements The Double-Cut-and-Join operation The adjacency graph and the distance equation

Rearrangement operations affect gene order and gene content. There are various types: • Inversions • Transpositions • Reverse transpositions • Translocations, fusions and fissions • Duplications and losses • Others. . . Any set of operations yields a distance between genomes, by counting the minimum number of operations needed to transform one genome into the other.
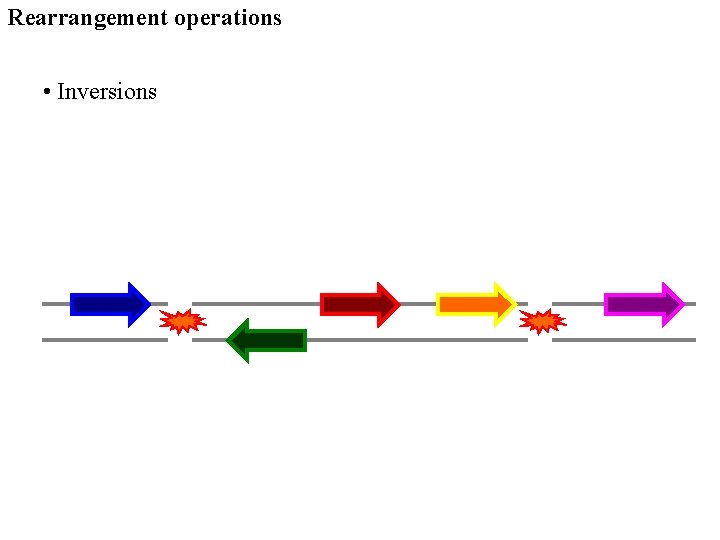
Rearrangement operations • Inversions
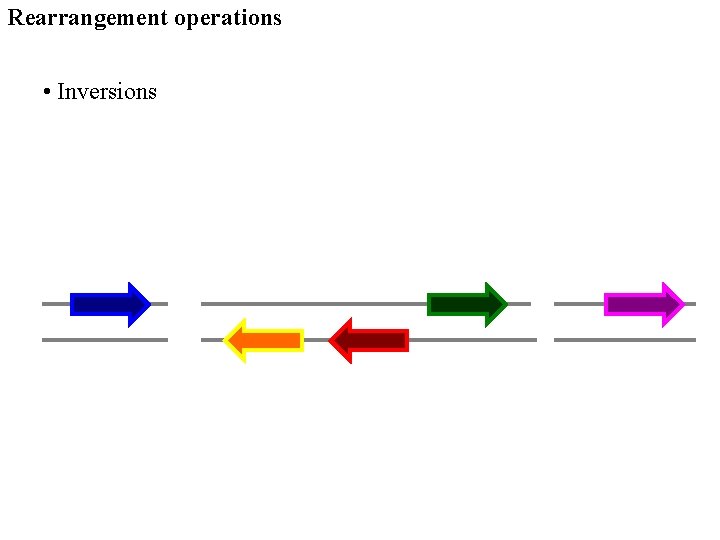
Rearrangement operations • Inversions
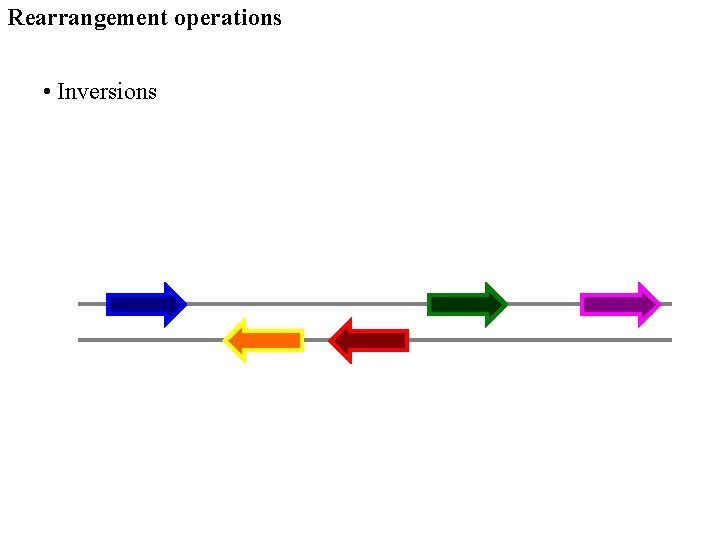
Rearrangement operations • Inversions
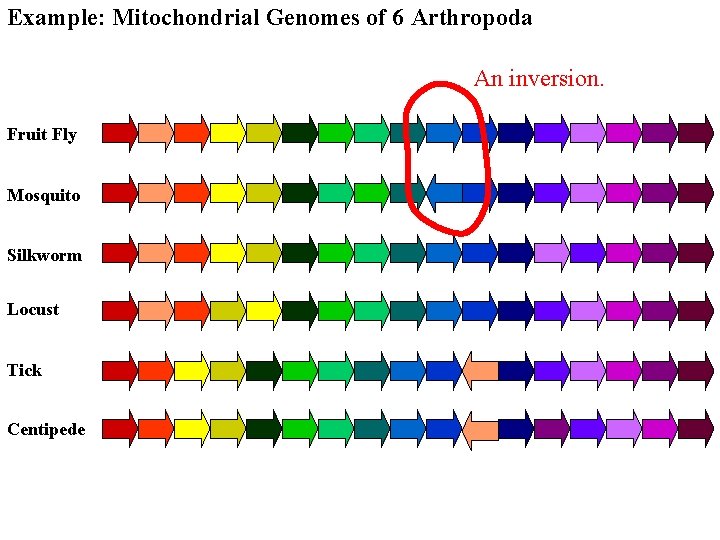
Example: Mitochondrial Genomes of 6 Arthropoda An inversion. Fruit Fly Mosquito Silkworm Locust Tick Centipede
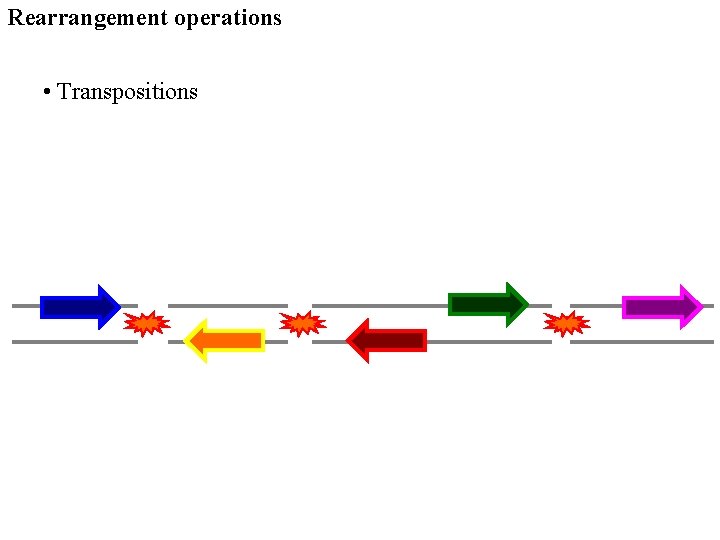
Rearrangement operations • Transpositions
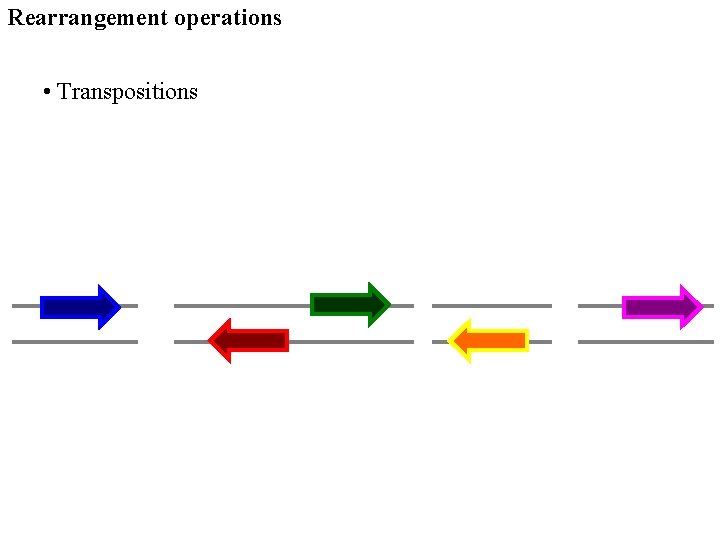
Rearrangement operations • Transpositions
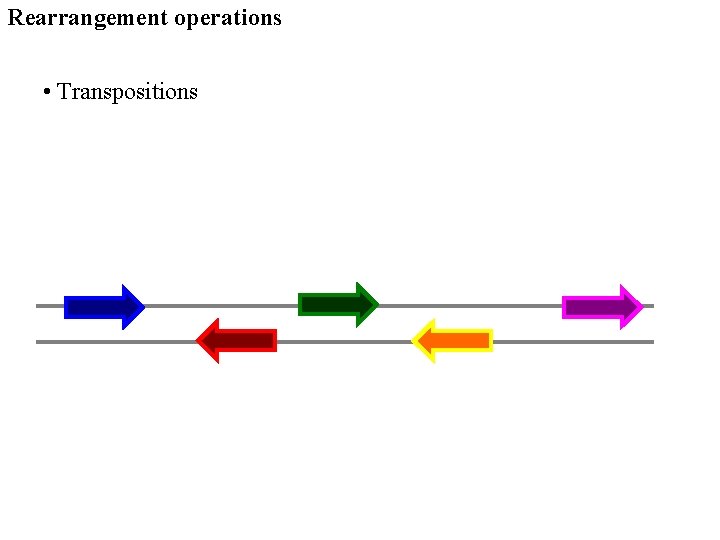
Rearrangement operations • Transpositions

Example: Mitochondrial Genomes of 6 Arthropoda Fruit Fly Mosquito Silkworm Locust Tick A transposition Centipede
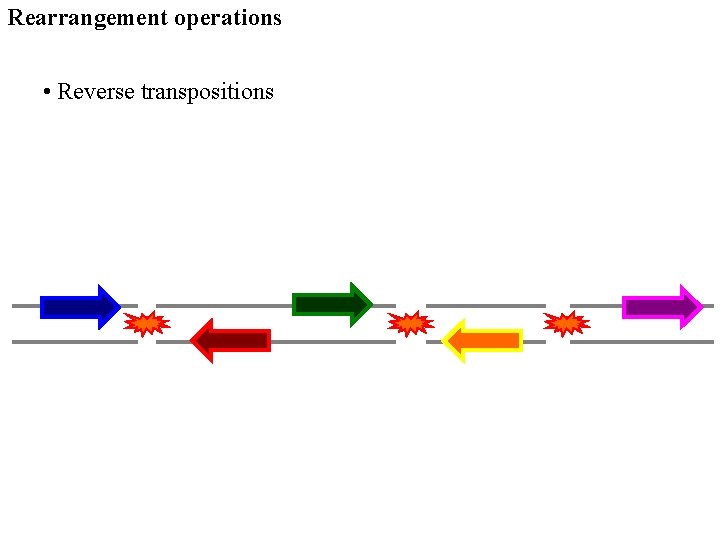
Rearrangement operations • Reverse transpositions
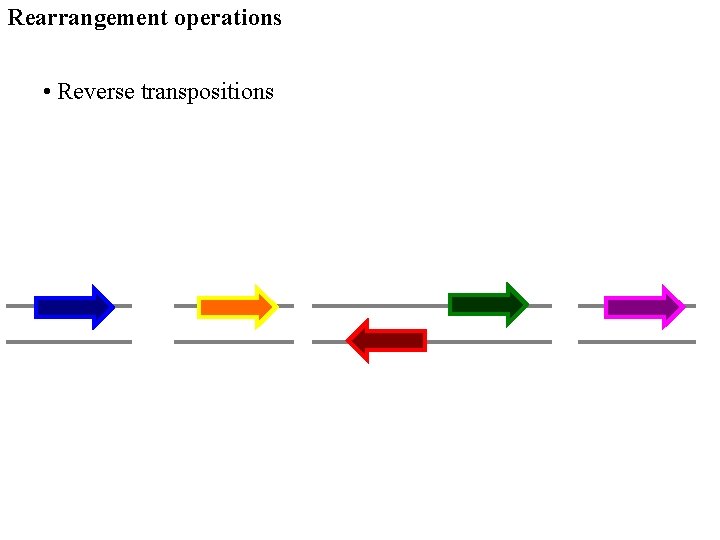
Rearrangement operations • Reverse transpositions
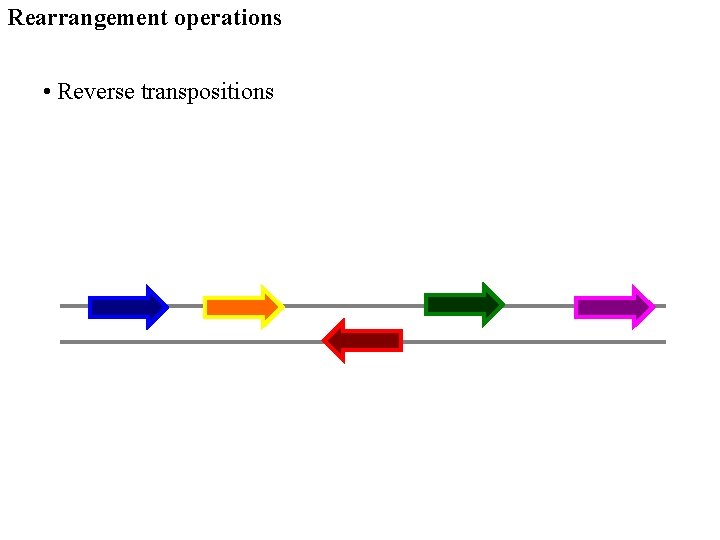
Rearrangement operations • Reverse transpositions

Example: Mitochondrial Genomes of 6 Arthropoda Fruit Fly Mosquito Silkworm Locust A reverse transposition Tick Centipede
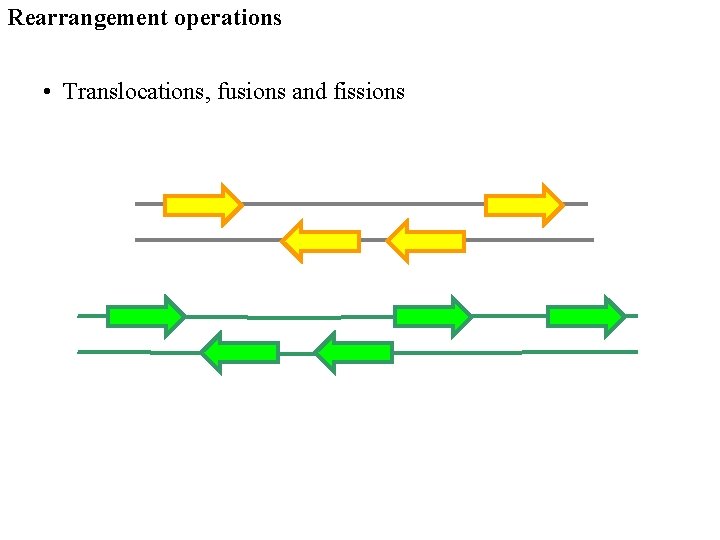
Rearrangement operations • Translocations, fusions and fissions
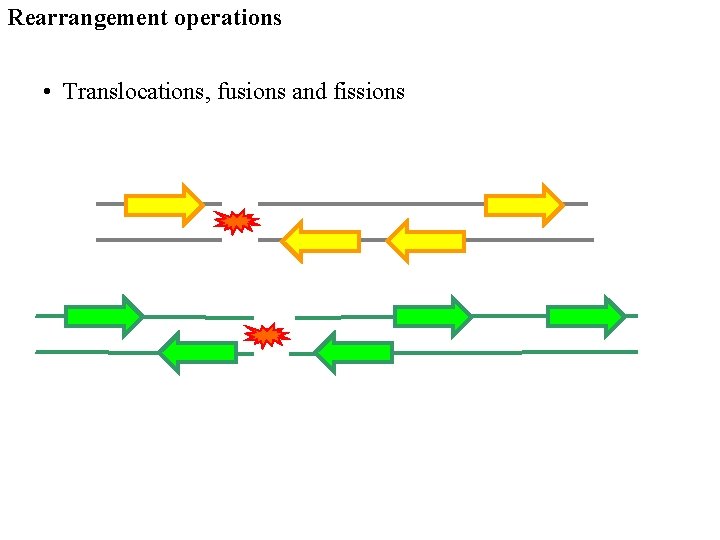
Rearrangement operations • Translocations, fusions and fissions
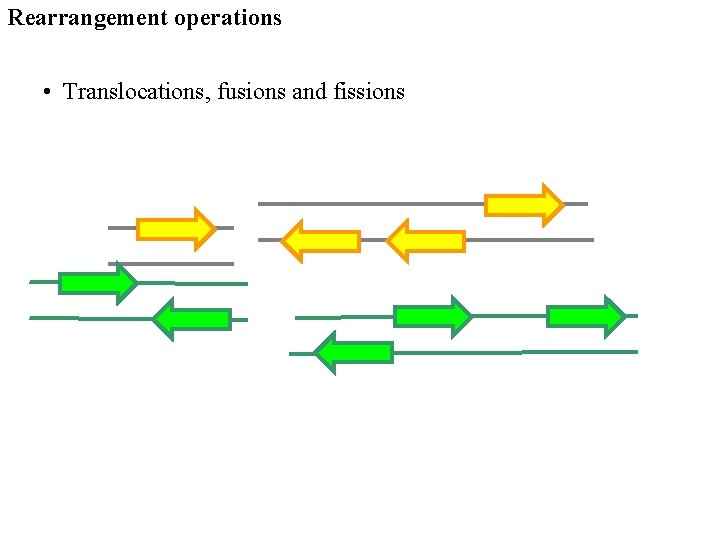
Rearrangement operations • Translocations, fusions and fissions
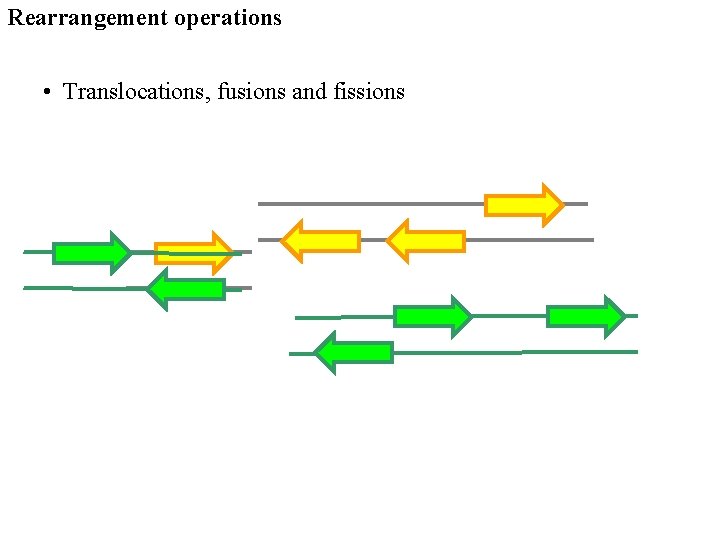
Rearrangement operations • Translocations, fusions and fissions
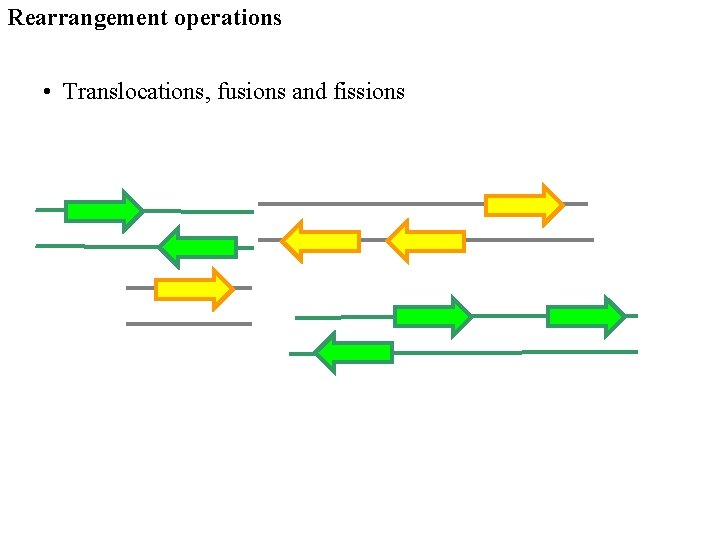
Rearrangement operations • Translocations, fusions and fissions
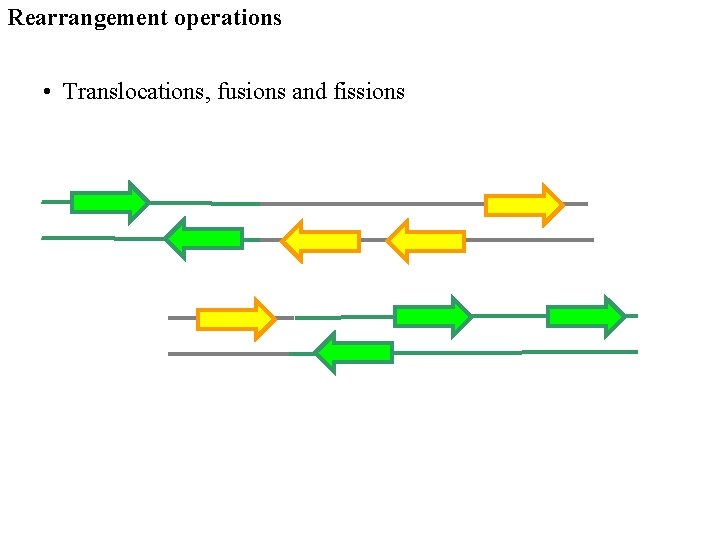
Rearrangement operations • Translocations, fusions and fissions
![From 24 chromosomes To 21 chromosomes [Source: Linda Ashworth, LLNL] DOE Human Genome Program From 24 chromosomes To 21 chromosomes [Source: Linda Ashworth, LLNL] DOE Human Genome Program](http://slidetodoc.com/presentation_image_h/0f35cd793190ec9a461ec20cbdeb7a4b/image-33.jpg)
From 24 chromosomes To 21 chromosomes [Source: Linda Ashworth, LLNL] DOE Human Genome Program Report

1. General introduction to genome rearrangements Examples of rearranged genomes 2. Measures of distance Rearrangement operations The Hannenhalli-Pevzner distance equation 3. A unifying view of genome rearrangements The Double-Cut-and-Join operation The adjacency graph and the distance equation

The Hannenhalli-Pevzner distance equation In 1995, Hannenhalli and Pevzner found a formula to compute the minimum number of inversions, translocations, fusions or fissions necessary to transform a multichromosomal genome into another. Sketch of the approach: • Cap the chromosomes • Concatenate all the chromosomes • Sort the resulting genome by inversions

1. General introduction to genome rearrangements Examples of rearranged genomes 2. Measures of distance Rearrangement operations The Hannenhalli-Pevzner distance equation 3. A unifying view of genome rearrangements The Double-Cut-and-Join operation The adjacency graph and the distance equation
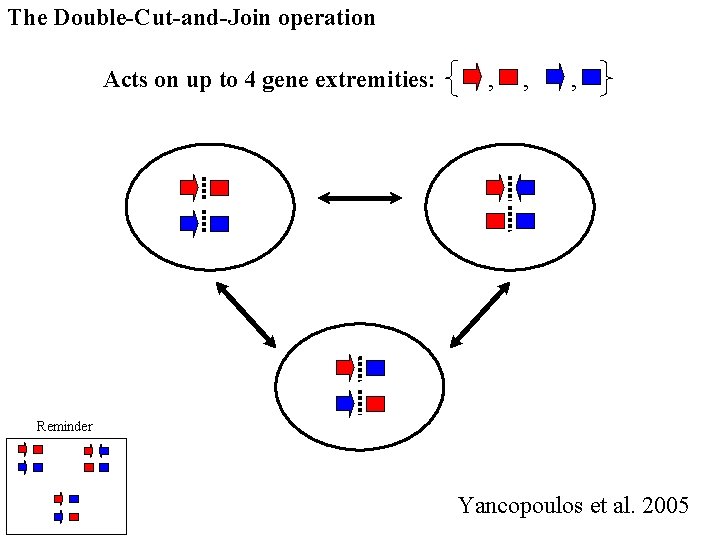
The Double-Cut-and-Join operation Acts on up to 4 gene extremities: , , , Reminder Yancopoulos et al. 2005
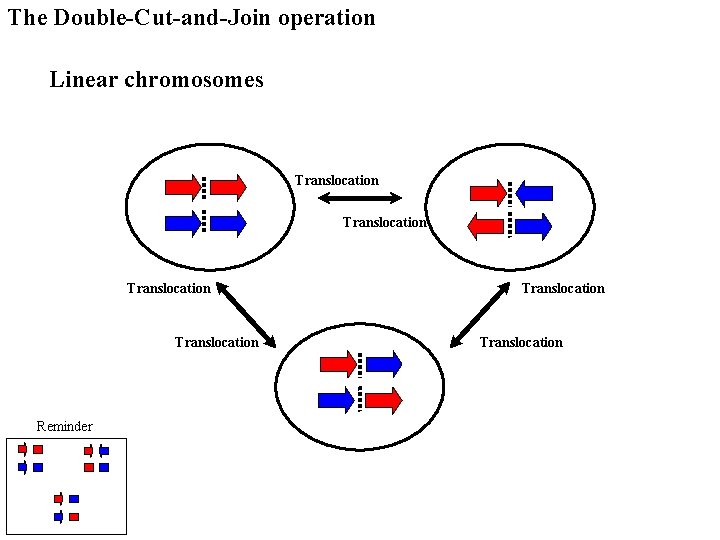
The Double-Cut-and-Join operation Linear chromosomes Translocation Reminder Translocation
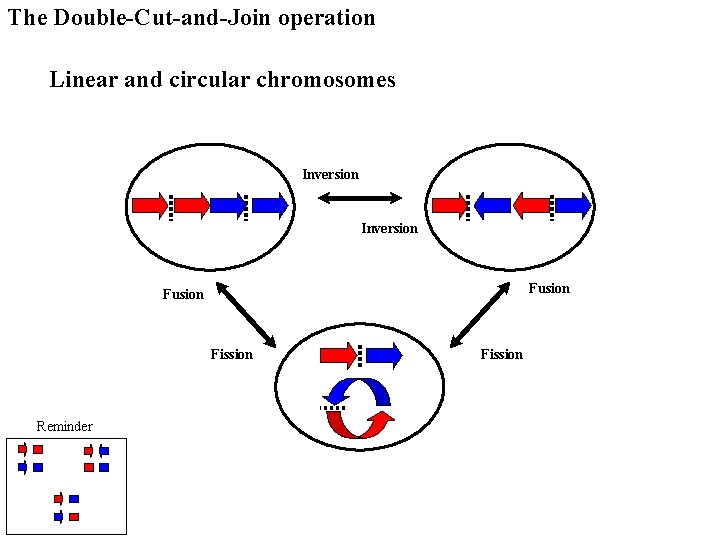
The Double-Cut-and-Join operation Linear and circular chromosomes Inversion Fusion Fission Reminder Fission
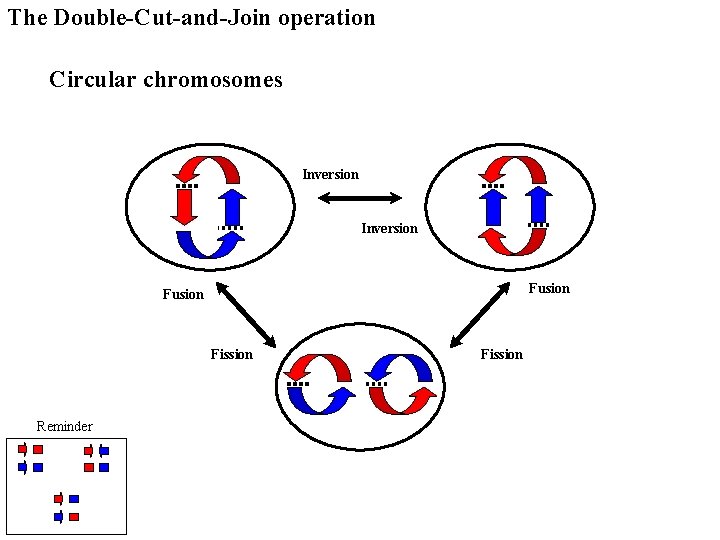
The Double-Cut-and-Join operation Circular chromosomes Inversion Fusion Fission Reminder Fission

1. General introduction to genome rearrangements Examples of rearranged genomes 2. Measures of distance Rearrangement operations The Hannenhalli-Pevzner distance equation 3. A unifying view of genome rearrangements The Double-Cut-and-Join operation The adjacency graph and the distance equation 4. Breakpoint reuse estimates Minimizing breakpoint reuse
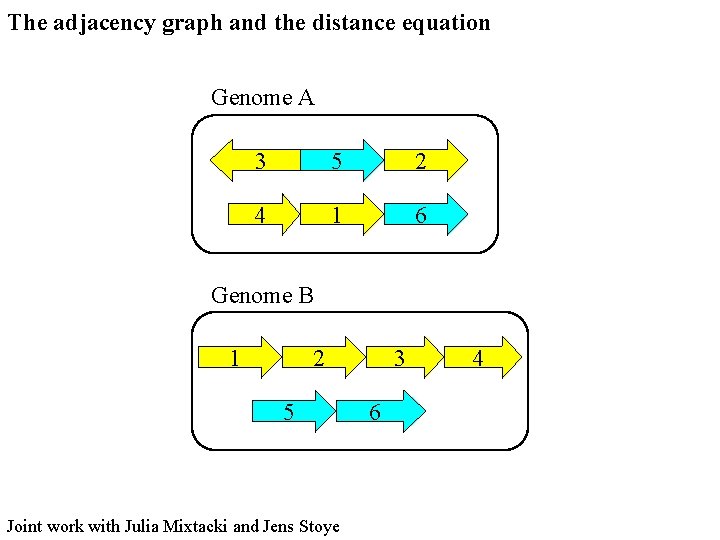
The adjacency graph and the distance equation Genome A 3 5 2 4 1 6 Genome B 1 2 5 Joint work with Julia Mixtacki and Jens Stoye 3 6 4
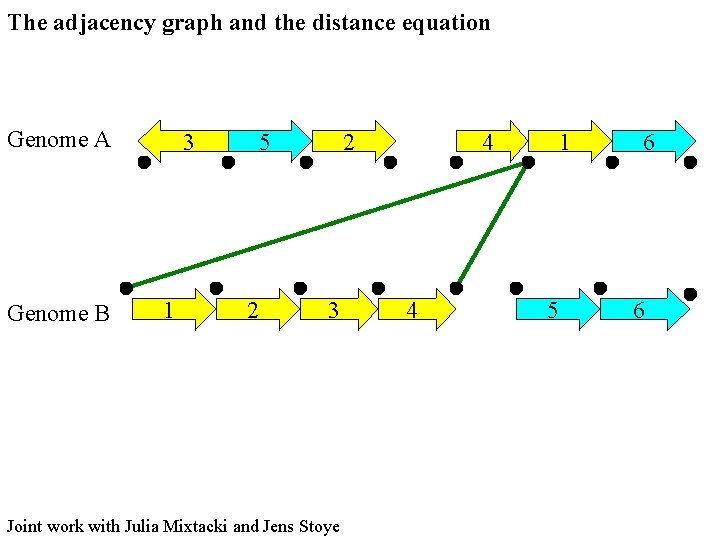
The adjacency graph and the distance equation Genome A Genome B 3 1 5 2 2 3 Joint work with Julia Mixtacki and Jens Stoye 4 4 1 5 6 6
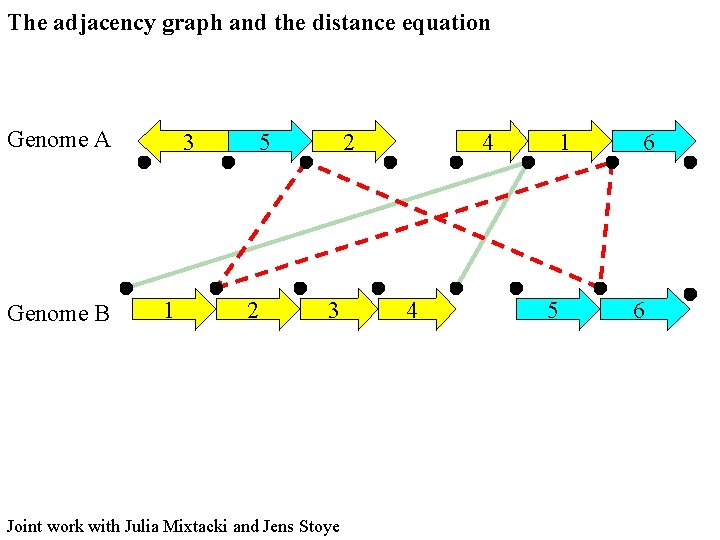
The adjacency graph and the distance equation Genome A Genome B 3 1 5 2 2 3 Joint work with Julia Mixtacki and Jens Stoye 4 4 1 5 6 6
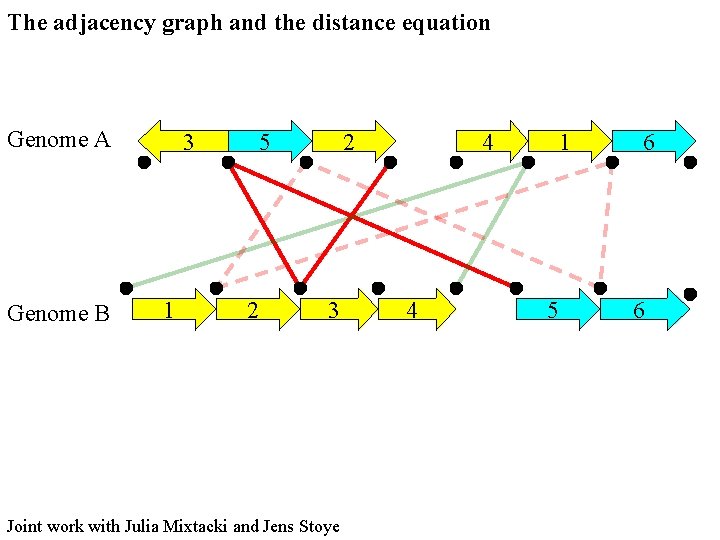
The adjacency graph and the distance equation Genome A Genome B 3 1 5 2 2 3 Joint work with Julia Mixtacki and Jens Stoye 4 4 1 5 6 6
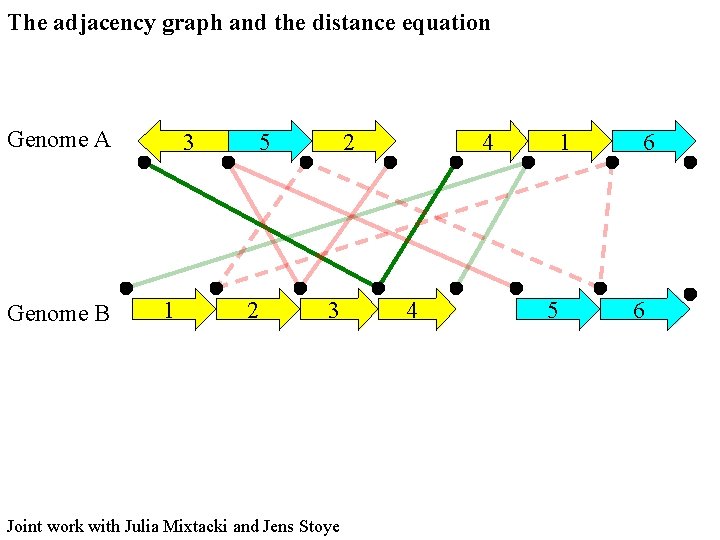
The adjacency graph and the distance equation Genome A Genome B 3 1 5 2 2 3 Joint work with Julia Mixtacki and Jens Stoye 4 4 1 5 6 6
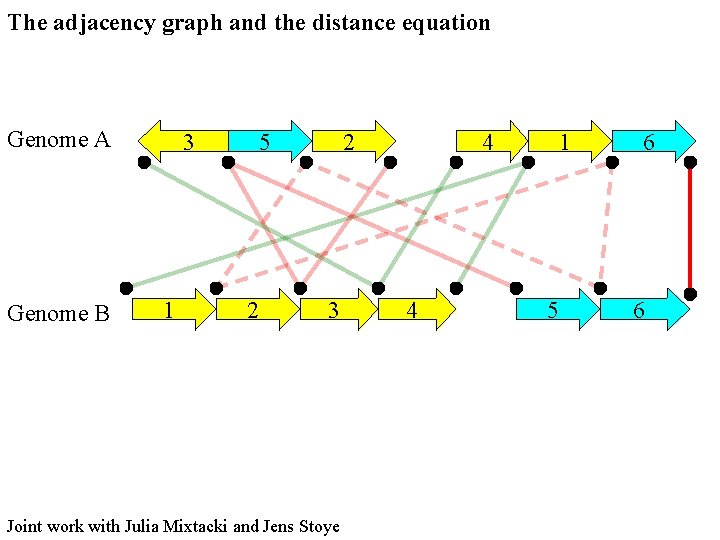
The adjacency graph and the distance equation Genome A Genome B 3 1 5 2 2 3 Joint work with Julia Mixtacki and Jens Stoye 4 4 1 5 6 6
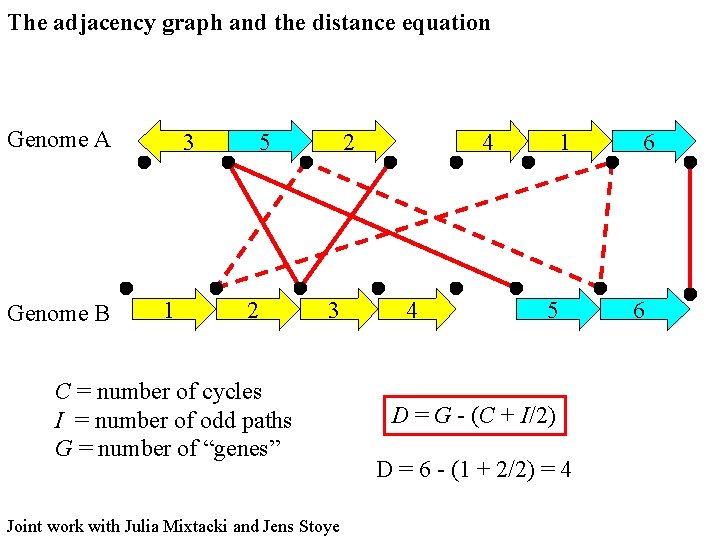
The adjacency graph and the distance equation Genome A Genome B 3 1 5 2 2 3 C = number of cycles I = number of odd paths G = number of “genes” Joint work with Julia Mixtacki and Jens Stoye 4 4 1 5 D = G - (C + I/2) D = 6 - (1 + 2/2) = 4 6 6
- Slides: 48