Computational Protein Redesign and the NonRibosomal Code Ivelin

Computational Protein Redesign and the Non-Ribosomal Code Ivelin Georgiev Cheng-Yu Chen Bruce R. Donald Lab

Non. Ribosomal Peptide Synthetases (NRPS) • NRPS enzymes found in some fungi and bacteria • NRPS enzymes make peptide-like products with pharmaceutical properties (antifungal , antineoplastic , antibacterial ) e. g. vancomycin, penicillin, gramicidin, bacitracin, cyclosporin, bleomycin, … • NRPS similar to PKS
Walsh Group
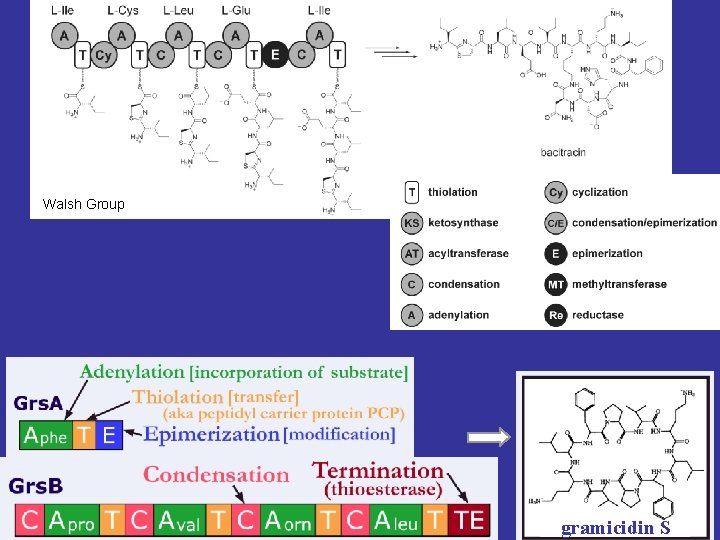
Walsh Group gramicidin S
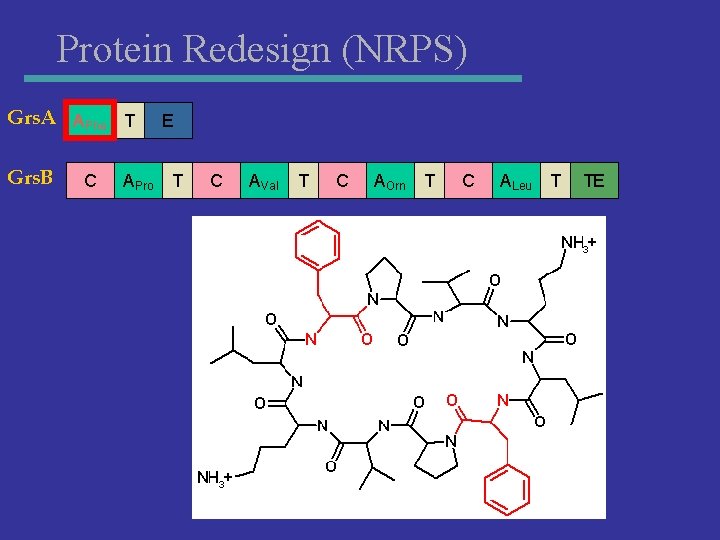
Protein Redesign (NRPS) Grs. A APhe T Grs. B C APro E T C AVal T C AOrn T C ALeu T TE
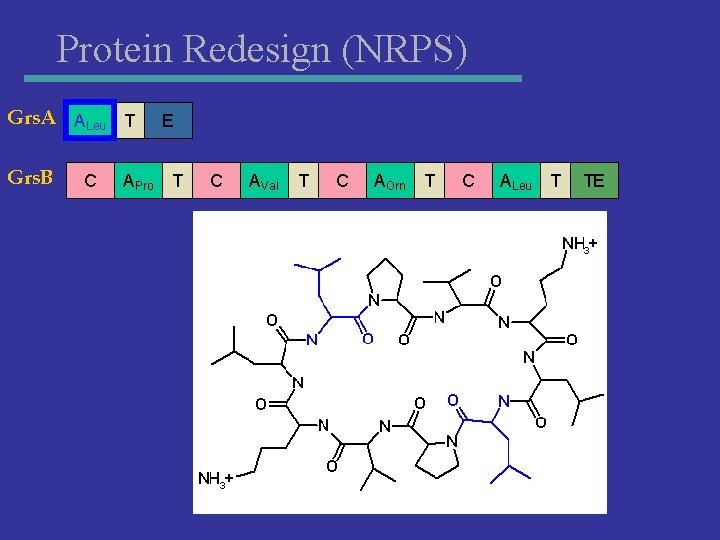
Protein Redesign (NRPS) Grs. A ALeu T Grs. B C APro E T C AVal T C AOrn T C ALeu T TE
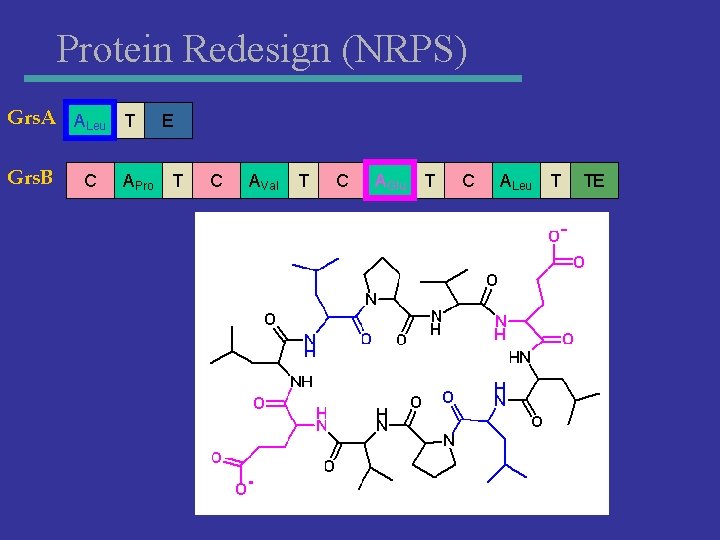
Protein Redesign (NRPS) Grs. A ALeu T Grs. B C APro E T C AVal T C AGlu T C ALeu T TE
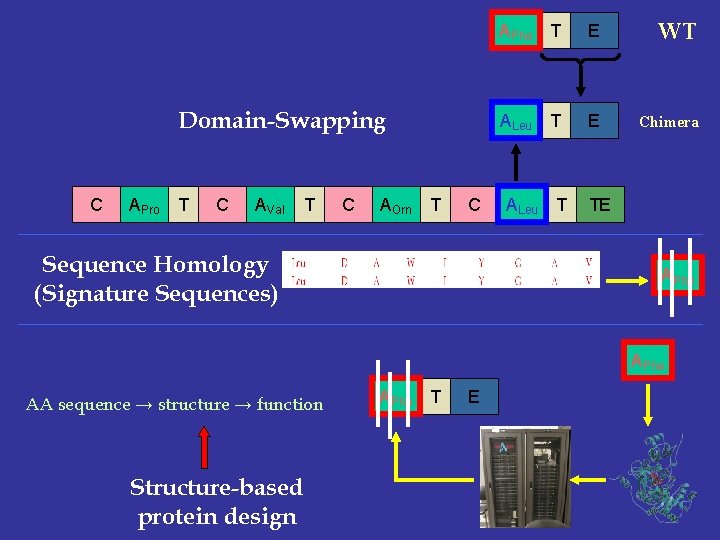
Domain-Swapping C APro T C AVal T C AOrn T C Sequence Homology (Signature Sequences) APhe T E WT ALeu T E Chimera ALeu T TE APhe AA sequence → structure → function Structure-based protein design APhe T E
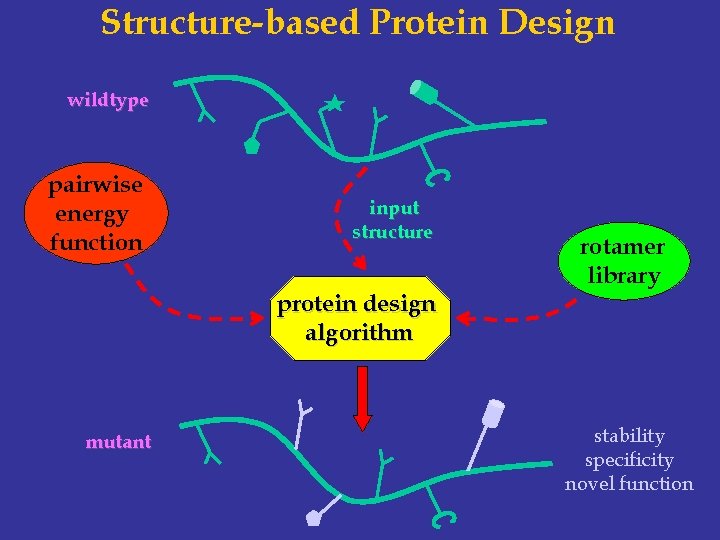
Structure-based Protein Design wildtype pairwise energy function input structure protein design algorithm mutant rotamer library stability specificity novel function

Redesign of Grs. A-Phe. A input structure 1 amu (1. 9Å)
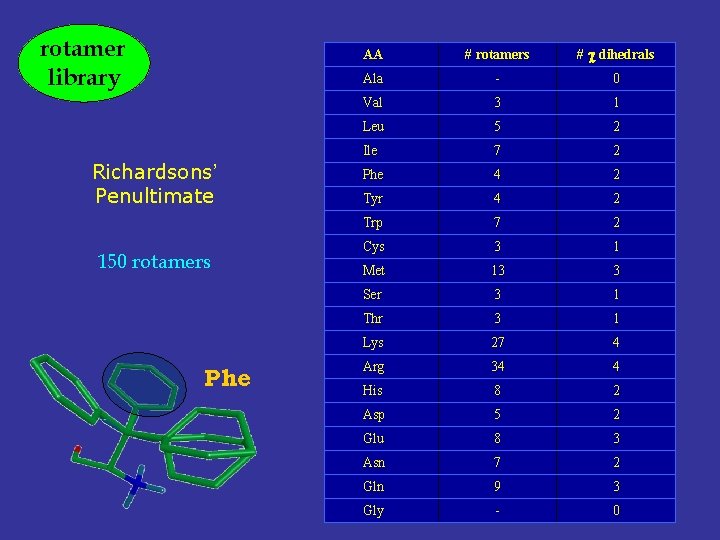
rotamer library Richardsons’ Penultimate 150 rotamers Phe AA # rotamers # dihedrals Ala - 0 Val 3 1 Leu 5 2 Ile 7 2 Phe 4 2 Tyr 4 2 Trp 7 2 Cys 3 1 Met 13 3 Ser 3 1 Thr 3 1 Lys 27 4 Arg 34 4 His 8 2 Asp 5 2 Glu 8 3 Asn 7 2 Gln 9 3 Gly - 0
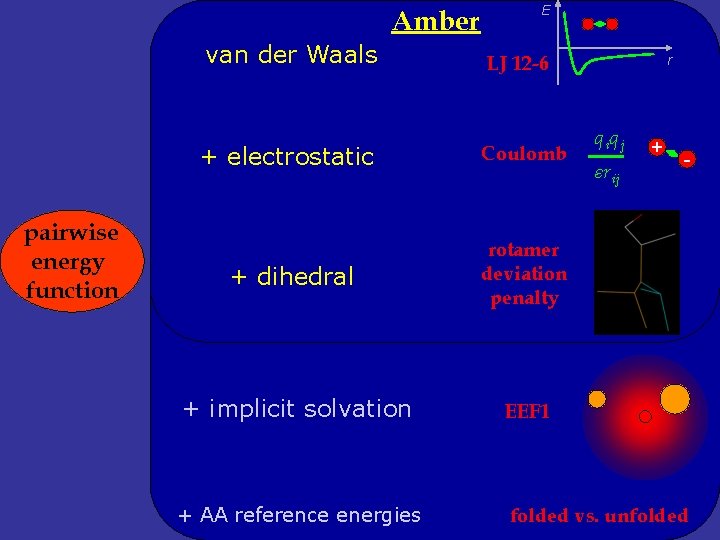
Amber van der Waals pairwise energy function E LJ 12 -6 r q iq j + + electrostatic Coulomb + dihedral rotamer deviation penalty + implicit solvation EEF 1 + AA reference energies folded vs. unfolded εrij -
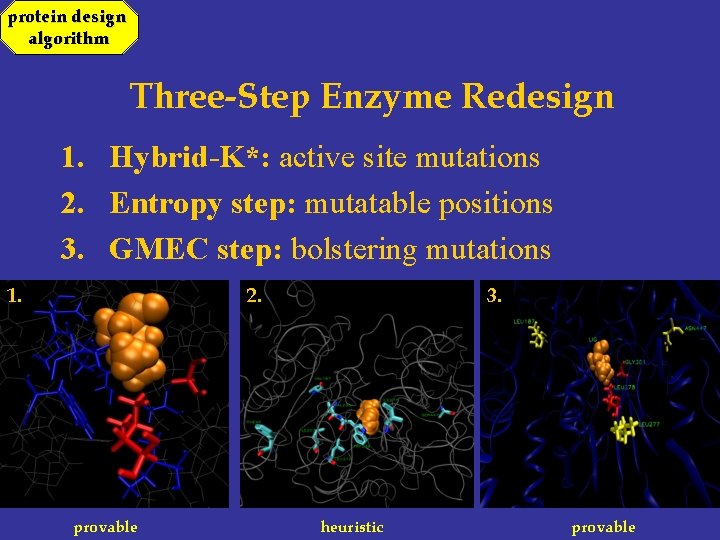
protein design algorithm Three-Step Enzyme Redesign 1. Hybrid-K*: active site mutations 2. Entropy step: mutatable positions 3. GMEC step: bolstering mutations 1. 2. provable 3. heuristic provable
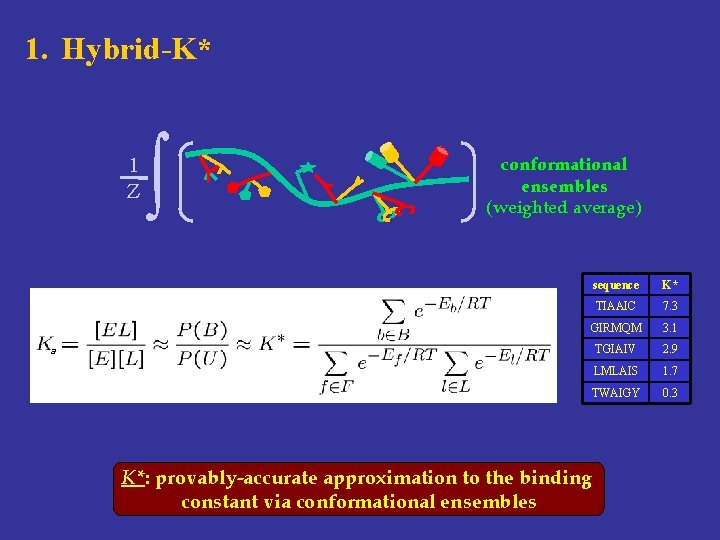
1. Hybrid-K* 1 Z ∫ conformational ensembles (weighted average) sequence K* TIAAIC 7. 3 GIRMQM 3. 1 TGIAIV 2. 9 LMLAIS 1. 7 TWAIGY 0. 3 a K*: provably-accurate approximation to the binding constant via conformational ensembles
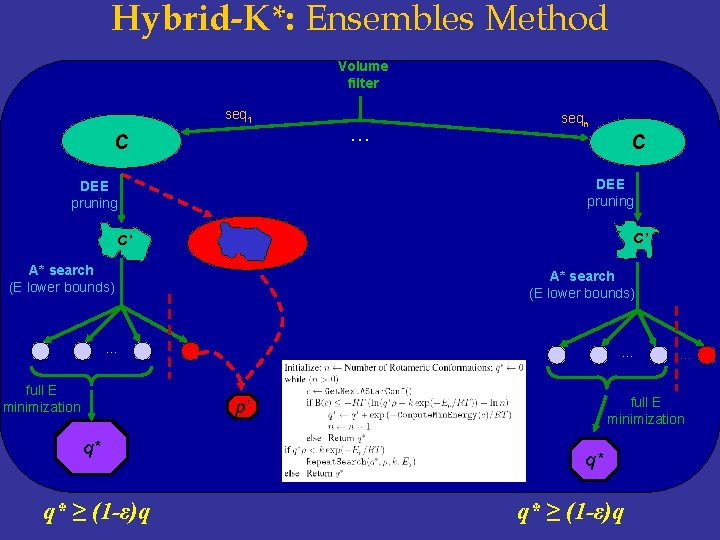
Hybrid-K*: Ensembles Method Volume filter seq 1 C … seqn C DEE pruning C’ C’ A* search (E lower bounds) … full E minimization p’ q* q* ≥ (1 -ε)q … q* q* ≥ (1 -ε)q
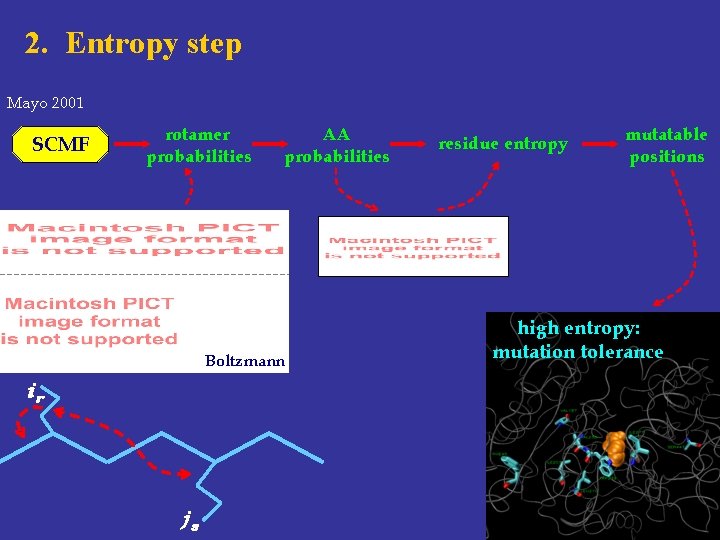
2. Entropy step Mayo 2001 SCMF rotamer probabilities AA probabilities Boltzmann ir js residue entropy mutatable positions high entropy: mutation tolerance
3. GMEC step min single lowest-energy conformation
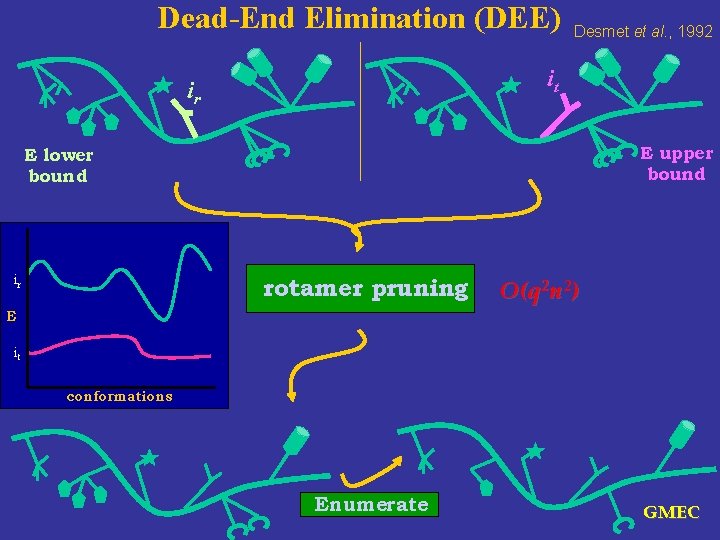
Dead-End Elimination (DEE) Desmet et al. , 1992 it ir E upper bound E lower bound ir rotamer pruning O(q 2 n 2) E it conformations Enumerate GMEC
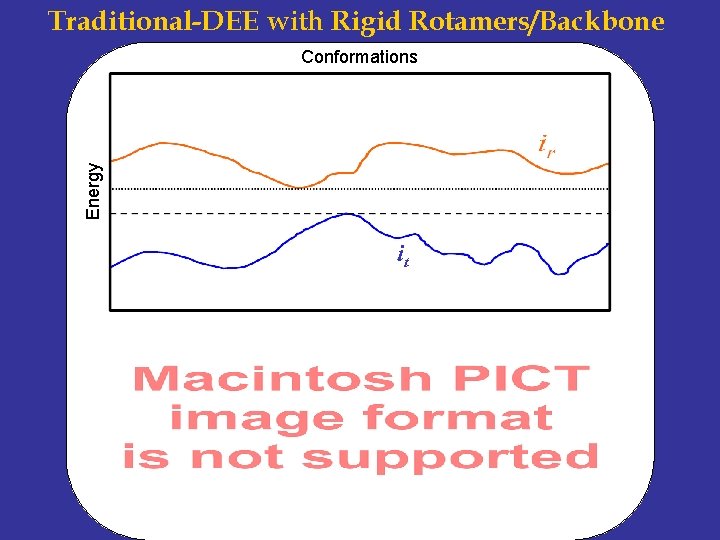
Traditional-DEE with Rigid Rotamers/Backbone Energy Conformations it

Min. DEE Energy Conformations min max it
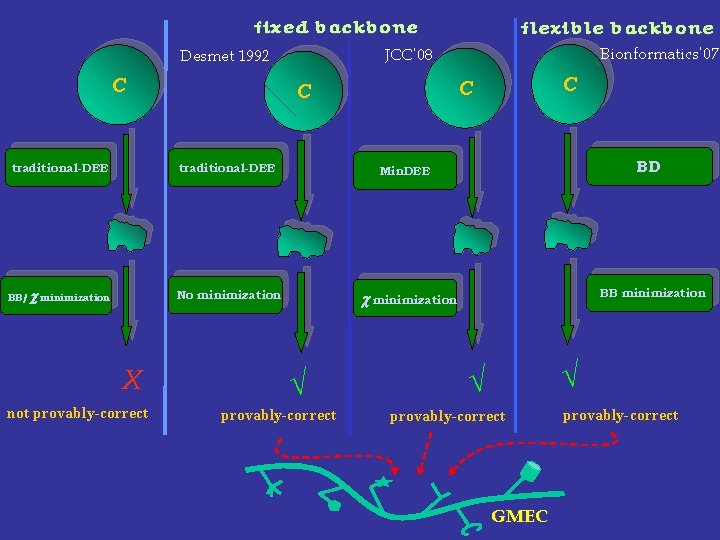
fixed backbone traditional-DEE Min. DEE BB/ c minimization No minimization c minimization not provably-correct √ provably-correct C C C traditional-DEE X Bionformatics’ 07 JCC’ 08 Desmet 1992 C flexible backbone BD BB minimization √ √ provably-correct GMEC provably-correct
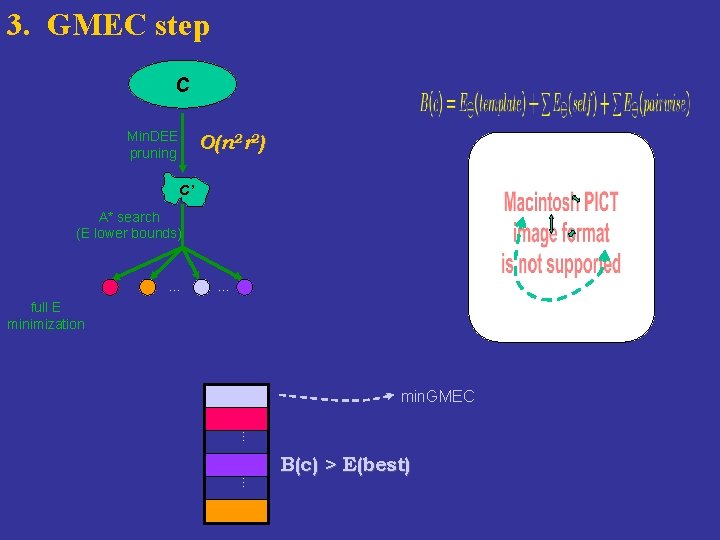
3. GMEC step C Min. DEE pruning O(n 2 r 2) C’ A* search (E lower bounds) … … full E minimization min. GMEC … … B(c) > E(best)
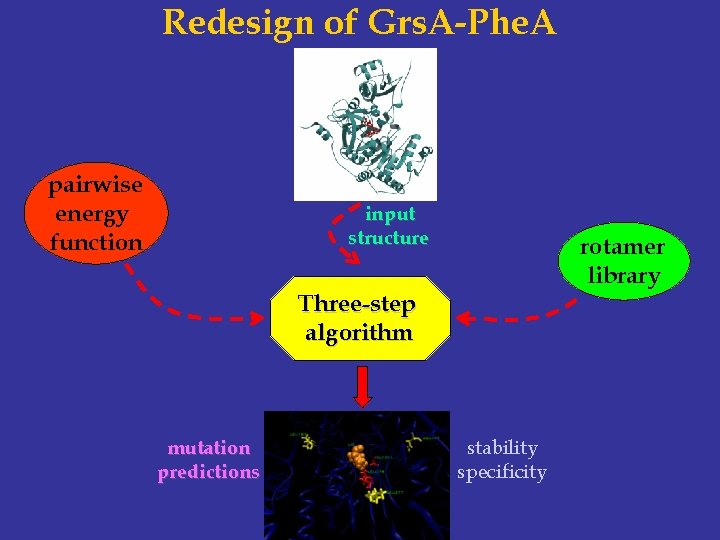
Redesign of Grs. A-Phe. A pairwise energy function input structure rotamer library Three-step algorithm mutation predictions stability specificity

Hybrid-K* Conf. Remaining Pruning Factor (%) Initial 6. 8 x 108 - Volume Filter 2. 04 x 108 3. 33 (70. 0) Min. DEE Filter 4. 13 x 106 49. 43 (98. 0) Steric Filter 3. 86 x 106 1. 07 (6. 5) A* Filter 7. 82 x 104 49. 41 (98. 0) all 7. 82 x 104 8685. 65 (99. 99) Computational • 9 hrs. on 24 processors • K* w/o filters: ≈ 3, 263 days Top 40 Mutations – Hybrid-K* Predictions • Experimental validation • T 278 M/A 301 G (Stachelhaus et al. , 1999) ranked 3 rd • G 301 in all known natural Leu adenylation domains

Entropy mutatable residues: 45, 187, 207, 210, 238, 277, 447

Entropy (SCMF) Sequence Alignment of 402 Sequences from AMP-binding Domains (Pfam)

Entropy (SCMF) Min. DEE G N T F L N I I L G I T F L I I L V V L V A L L A F Allowed 45 ala 187 ala 207 ala 210 ala 238 ala 277 ala 447 ala AA types (DEE input) leu ile phe asn gly val leu ile ser thr asp glu asn gly leu ile ser thr asn gly leu ile phe tyr asn gly leu ile ser thr asn gly val leu ile ser thr asn gly V I L V I S

Sequence Alignment of 402 Sequences from AMP-binding Domains (Pfam) Min. DEE G N I N T F L I I I V L I T L L V L L A F V I L V I S L V A
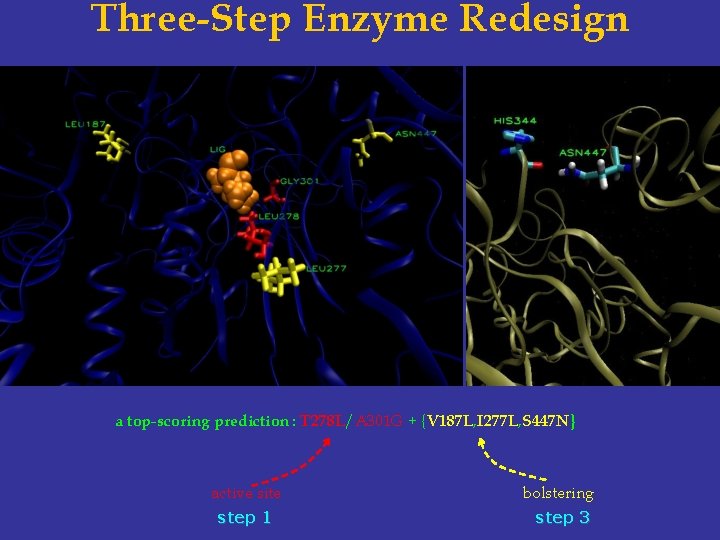
Three-Step Enzyme Redesign a top-scoring prediction : T 278 L/A 301 G + {V 187 L, I 277 L, S 447 N} active site bolstering step 1 step 3

Binding Pocket of Phe. A I 330 S 331 A 322 I 299 A 301 D 235 A 236 K 517 W 239 T 278

K* Prediction: Mutants to bind L-Leu D A W T I A A I S

Enzymes coupled continuous assay E + ATP + a. a. UDPG + PPi KM E • a. a • ATP UDPG Pyrophosphorylase Glucose 1 -Phosphate Glucose 6 -P + NAD kcat E • a. a-AMP + PPi UTP + Glucose 1 -Phosphate Phosphoglucomutase G-6 -PDH Glucose 6 -Phosphate 6 -Phosphogluconate + NADH 340 nm

Steady-state Kinetics E + ATP + a. a. KM E • a. a • ATP kcat E • a. a-AMP + PPi Vmax KM

Specificity switched from L-Phe to L-Leu

Bolstering mutants L-Phe L-Leu

The Reaction Pathway KM E + ATP + a. a. Katp E·ATP E • a. a • ATP E·ATP·a. a k 3[ATP] Kaa E·a. a. E • a. a-AMP + PPi k 1[a. a] k-1 E kcat k 2 k-3 Random sequential order E·a. a-AMP·PPi k 4 k-4[PPi] E·AMP-a. a + PPi

Stopped-Flow Kinetics • Dead time (Before measurement) = 1. 2 ms • Total reaction time = 0. 5 ~ 5 s • Sampling points = 1000 • # of exponential terms = # of steps

E • ATP + L-Phe k 1 k-1 E • a. a • ATP k 2 E* • a. a • ATP kobs 1 = k 1[L-Phe] + k-1 A*e-kobs 1*t+B*e-kobs 2*t+C A*e-kobs 1*t + C

E • ATP + L-Leu k 1 k-1 Wild-type E • Leu • ATP A 301 G/T 278 L A 301 G/T 278 M kobs 1 = k 1[L-Leu] + k-1
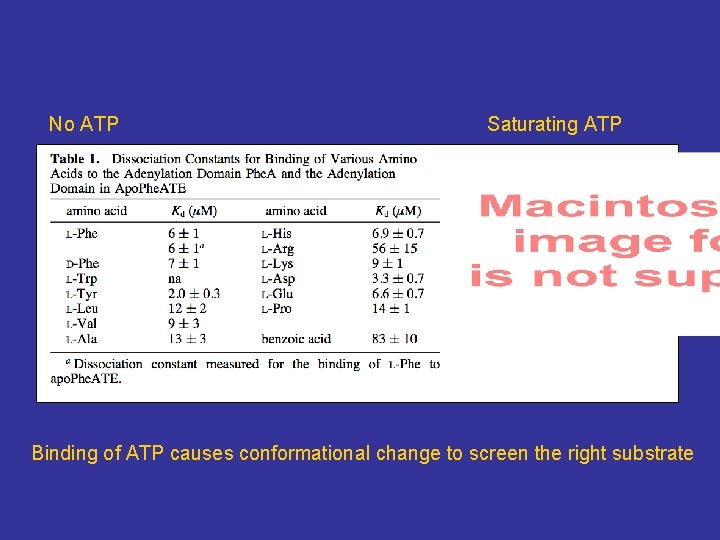
No ATP Saturating ATP Binding of ATP causes conformational change to screen the right substrate

Free Energy Profile (kcal/mole) E ‡S +ATP ∆G‡ ES‡ ∆G ES ∆G‡ = -RT[ln(k 1)-ln(k. B/Th)]

Stability of Wild-type and Mutant Phe. A

Conclusion • Switching specificity not only improves activity but also affects binding • Binding of ATP causes conformational change for the enzyme to select its right substrate • Active site mutation lowers the energy barrier (activation energy) for the enzyme to bind different substrate.
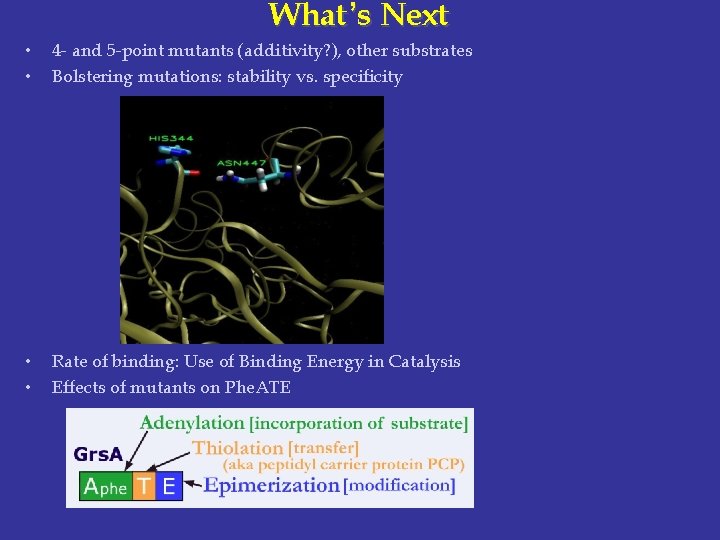
What ’s Next • • 4 - and 5 -point mutants (additivity? ), other substrates Bolstering mutations: stability vs. specificity • • Rate of binding: Use of Binding Energy in Catalysis Effects of mutants on Phe. ATE

What ’s Next • NMR structures and dynamics of WT Grs. A-Phe. A and mutants John Mac. Master 800 m. Hz [1 H, 15 N]-TROSY-HSQC spectrum of perdeuterated, 15 N-labeled WT Grs. A-Phe. A

Acknowledgments • • John Mac. Master Tony Yan Dan Keedy All members of Donald Lab Funding: • NIH

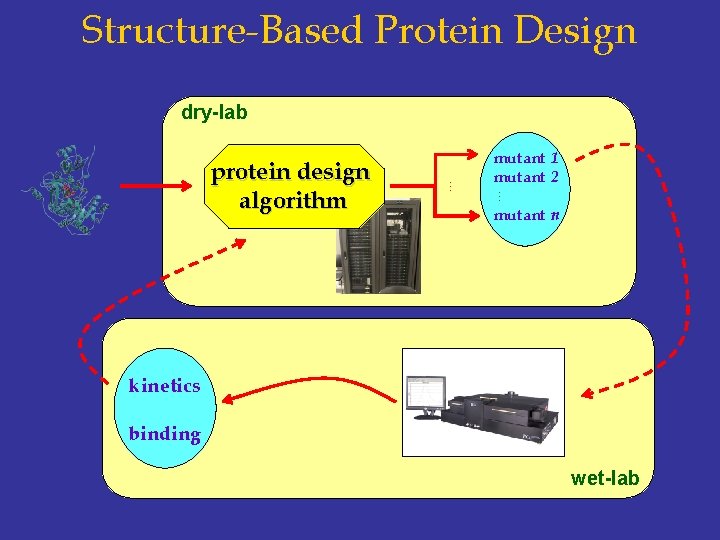
Structure-Based Protein Design dry-lab … … protein design algorithm mutant 1 mutant 2 mutant n kinetics binding wet-lab
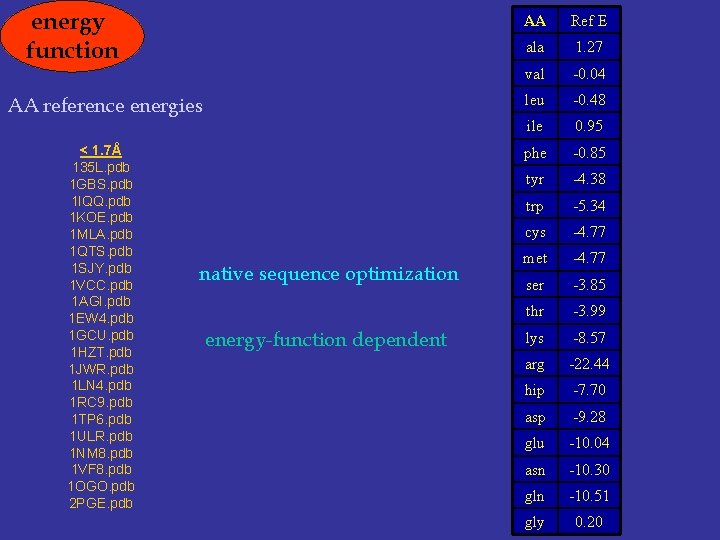
energy function AA reference energies < 1. 7Å 135 L. pdb 1 GBS. pdb 1 IQQ. pdb 1 KOE. pdb 1 MLA. pdb 1 QTS. pdb 1 SJY. pdb 1 VCC. pdb 1 AGI. pdb 1 EW 4. pdb 1 GCU. pdb 1 HZT. pdb 1 JWR. pdb 1 LN 4. pdb 1 RC 9. pdb 1 TP 6. pdb 1 ULR. pdb 1 NM 8. pdb 1 VF 8. pdb 1 OGO. pdb 2 PGE. pdb native sequence optimization energy-function dependent AA Ref E ala 1. 27 val -0. 04 leu -0. 48 ile 0. 95 phe -0. 85 tyr -4. 38 trp -5. 34 cys -4. 77 met -4. 77 ser -3. 85 thr -3. 99 lys -8. 57 arg -22. 44 hip -7. 70 asp -9. 28 glu -10. 04 asn -10. 30 gln -10. 51 gly 0. 20

energy function
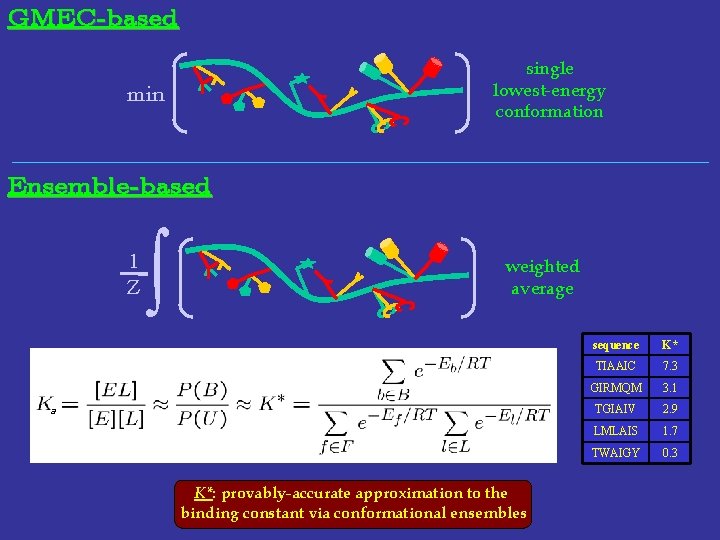
GMEC-based single lowest-energy conformation min Ensemble-based 1 Z ∫ weighted average a K*: provably-accurate approximation to the binding constant via conformational ensembles sequence K* TIAAIC 7. 3 GIRMQM 3. 1 TGIAIV 2. 9 LMLAIS 1. 7 TWAIGY 0. 3
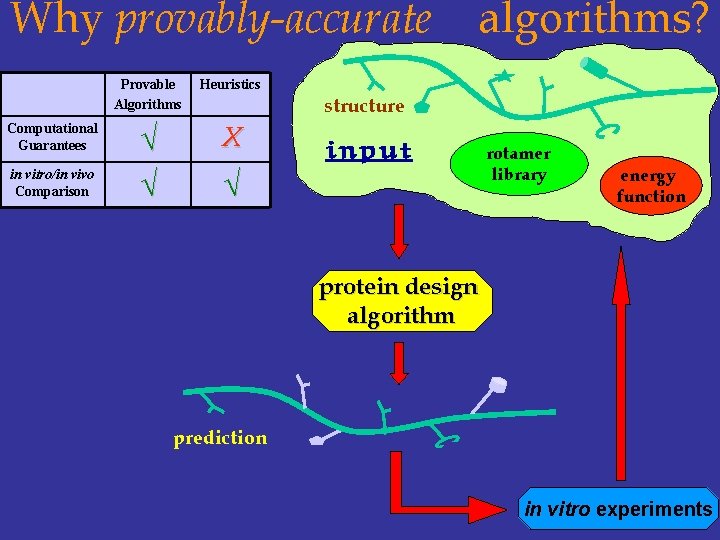
Why provably-accurate Computational Guarantees in vitro/in vivo Comparison Provable Algorithms Heuristics √ √ X algorithms? structure input √ rotamer library energy function protein design algorithm prediction in vitro experiments
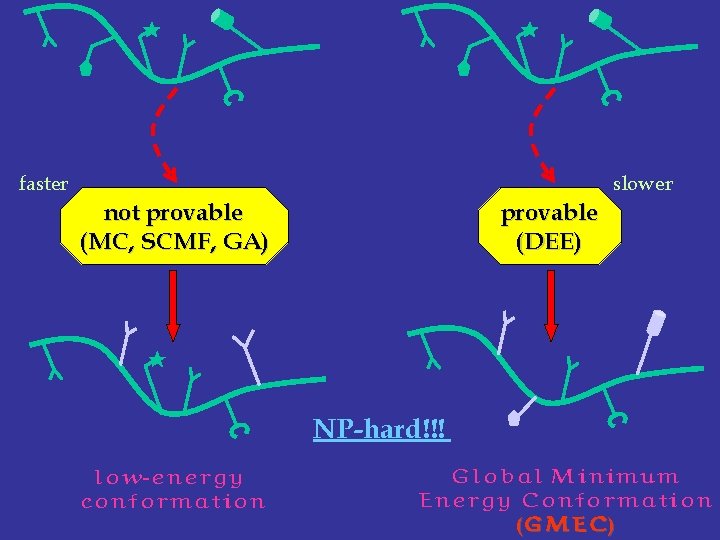
faster slower not provable (MC, SCMF, GA) provable (DEE) NP-hard!!! low-energy conformation Global Minimum Energy Conformation (GMEC)

Activity Assays Cheng-Yu Chen • mutants: ↓ specificity for Phe ↑ specificity for Leu

Grs. A-Phe. A Redesign Cheng-Yu Chen wt A 301 G/T 278 L A 301 G/T 278 M A 301 G/A 322 V T 278 L/A 301 G/S 447 N I 277 L/T 278 L/A 301 G V 187 L/T 278 L/A 301 G • mutants: switch of specificity
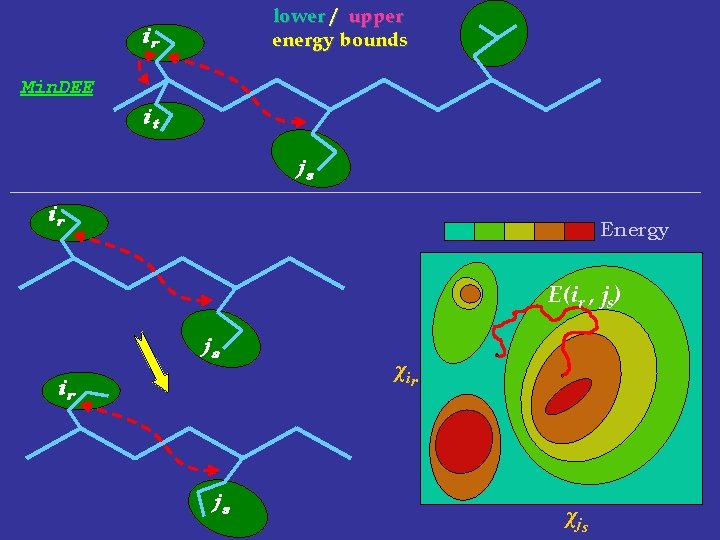
lower / upper energy bounds ir Min. DEE it js ir Energy E(ir , js) js ir js χ ir χ js
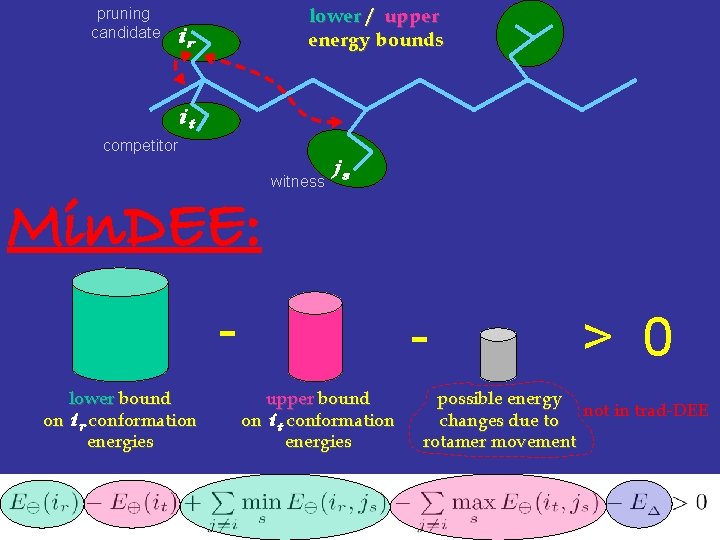
pruning candidate lower / upper energy bounds ir it competitor Min. DEE: witness js lower bound on ir conformation energies upper bound on it conformation energies > 0 possible energy not in trad-DEE changes due to rotamer movement
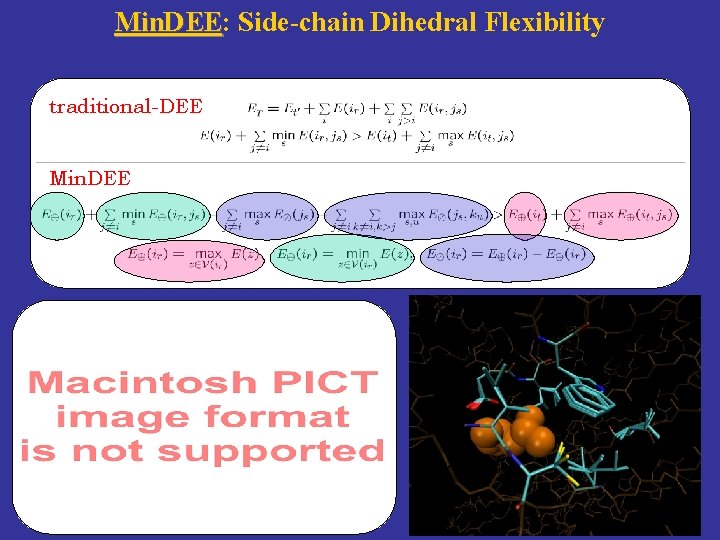
Min. DEE: Min. DEE Side-chain Dihedral Flexibility traditional-DEE Min. DEE

Energy Lower Bounds
- Slides: 59