OSPREY Tutorial Ivelin Georgiev Bruce Donald Lab Duke

OSPREY Tutorial Ivelin Georgiev Bruce Donald Lab Duke University

Distribution of Structures Maximum Likelihood (pick most probable) Bayesian (average over all conformations) min ( ) Global Minimum Energy Conformation 1 Z ò Probability « Energy using Boltzmann distribution
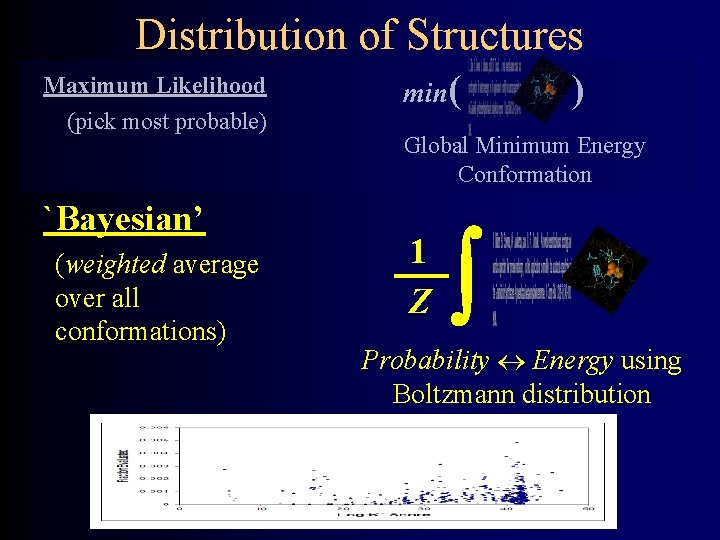
Distribution of Structures Maximum Likelihood (pick most probable) `Bayesian’ (weighted average over all conformations) min( ) Global Minimum Energy Conformation 1 Z ò Probability « Energy using Boltzmann distribution
traditional-DEE Min. DEE BD GMEC maximum likelihood
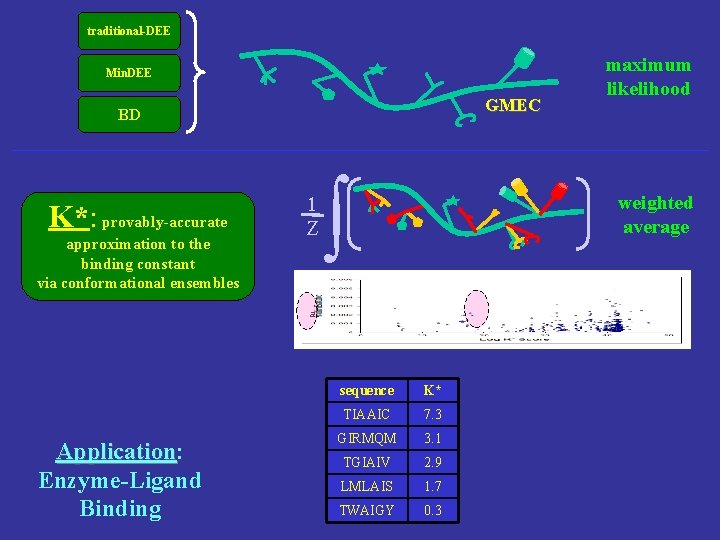
traditional-DEE Min. DEE GMEC BD K*: provably-accurate approximation to the binding constant via conformational ensembles 1 Z ∫ weighted average a Application: Application Enzyme-Ligand Binding maximum likelihood sequence K* TIAAIC 7. 3 GIRMQM 3. 1 TGIAIV 2. 9 LMLAIS 1. 7 TWAIGY 0. 3
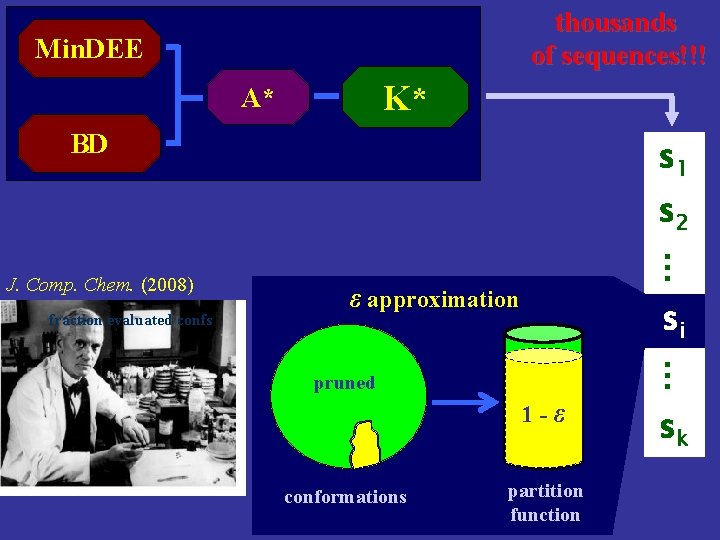
thousands of sequences!!! Min. DEE K* A* BD s 1 s 2 fraction evaluated confs … J. Comp. Chem. (2008) ε approximation si … pruned 1 -ε conformations partition function sk

Example (PNAS, 2009) Cheng-Yu Chen Ivelin Georgiev Amy Anderson Bruce Donald
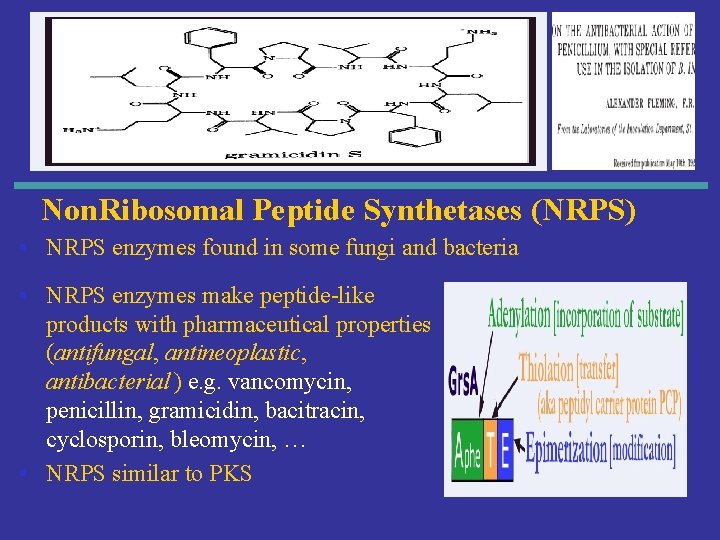
Non. Ribosomal Peptide Synthetases (NRPS) • NRPS enzymes found in some fungi and bacteria • NRPS enzymes make peptide-like products with pharmaceutical properties (antifungal, antineoplastic, antibacterial ) e. g. vancomycin, penicillin, gramicidin, bacitracin, cyclosporin, bleomycin, … • NRPS similar to PKS
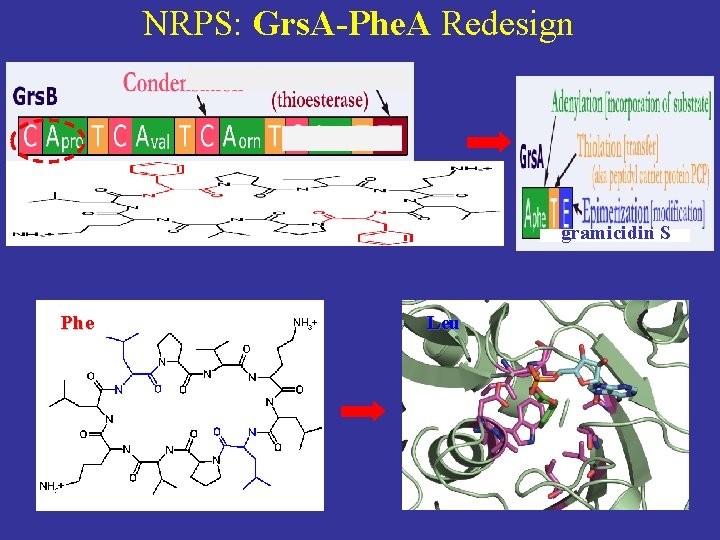
NRPS: Grs. A-Phe. A Redesign gramicidin S Phe Leu
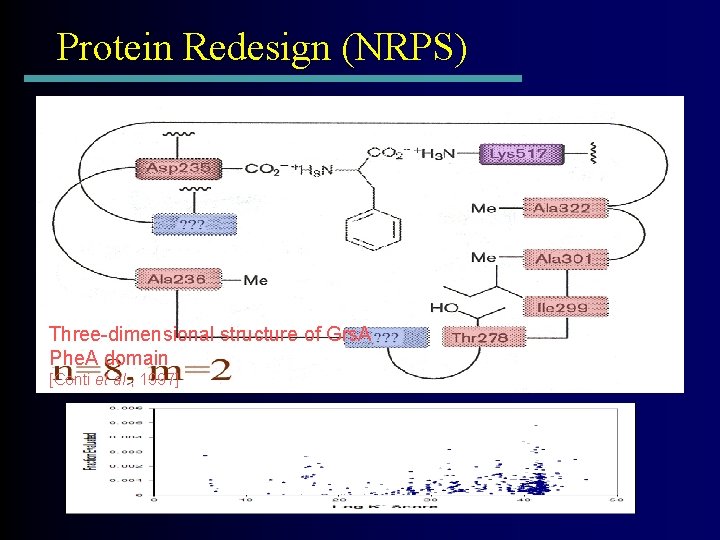
Protein Redesign (NRPS) Three-dimensional structure of Grs. A Phe. A domain [Conti et al. , 1997]

Change specificity from Phe to Leu by allowing any 2 (of 9) mutations Mutations to GAVLIFYWM Appx. 3000 Mutation Sequences = 680, 000 Conformations (78, 200 after pruning) +H 3 N Leu - CO 2 r=9 s =2
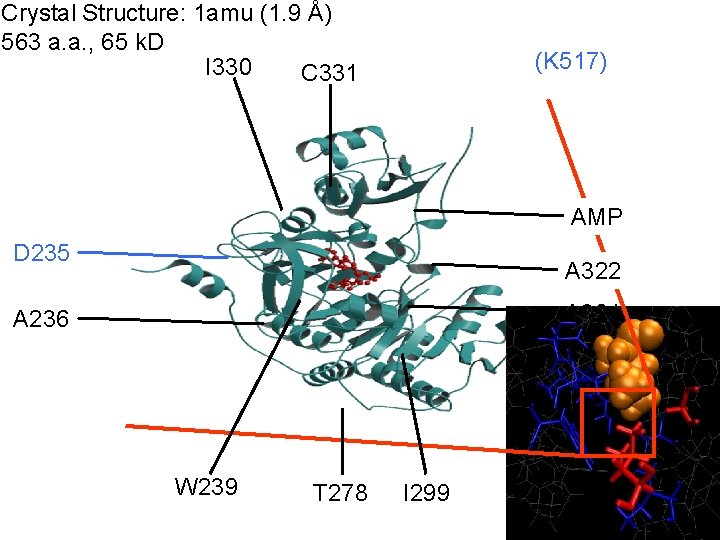
Crystal Structure: 1 amu (1. 9 Å) 563 a. a. , 65 k. D I 330 C 331 (K 517) AMP D 235 A 322 A 301 A 236 W 239 T 278 I 299
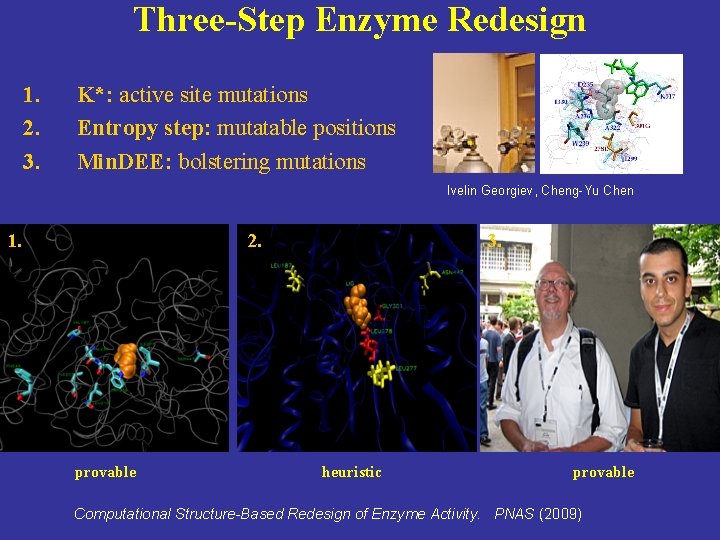
Three-Step Enzyme Redesign 1. 2. 3. K*: active site mutations Entropy step: mutatable positions Min. DEE: bolstering mutations Ivelin Georgiev, Cheng-Yu Chen 1. 2. provable 3. heuristic provable Computational Structure-Based Redesign of Enzyme Activity. PNAS (2009)
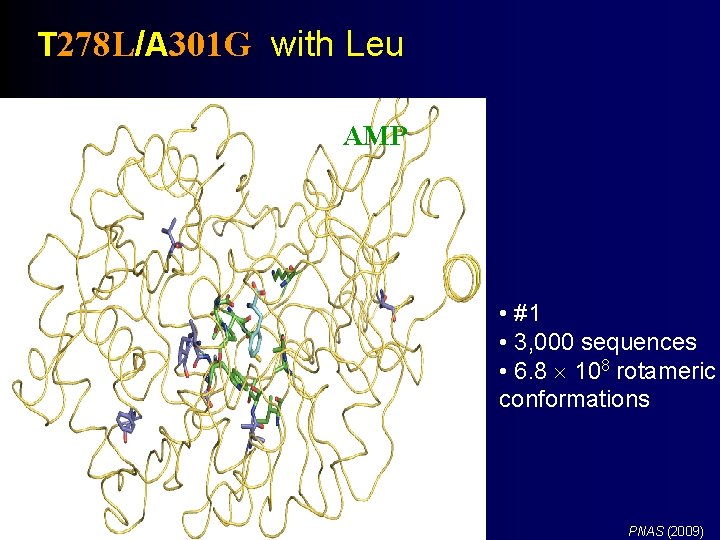
T 278 L/A 301 G with Leu AMP K 517 • #1 • 3, 000 sequences • 6. 8 108 rotameric conformations PNAS (2009)
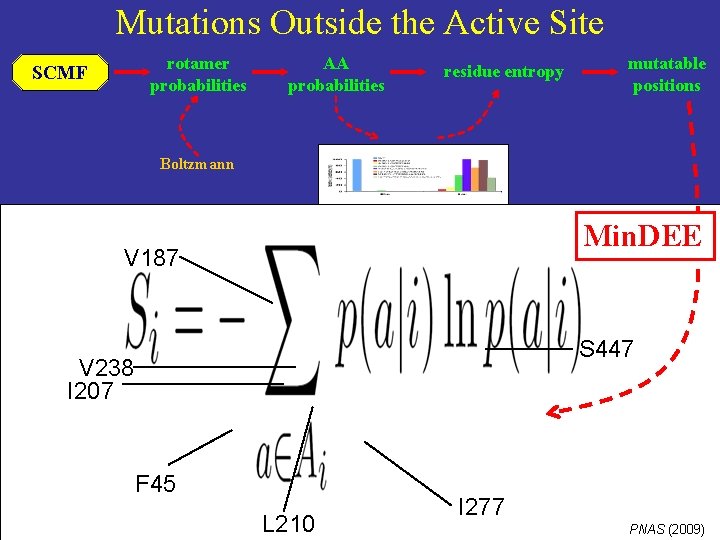
Mutations Outside the Active Site rotamer probabilities SCMF AA probabilities residue entropy mutatable positions Boltzmann Min. DEE V 187 S 447 V 238 I 207 F 45 L 210 I 277 PNAS (2009)

• All top 10 • 3, 000 sequences • 6. 8 108 rotameric conformations A 301 G/T 278 L [L-Leu] m. M Phe Normalized kcat/ KM Leu PNAS (2009)
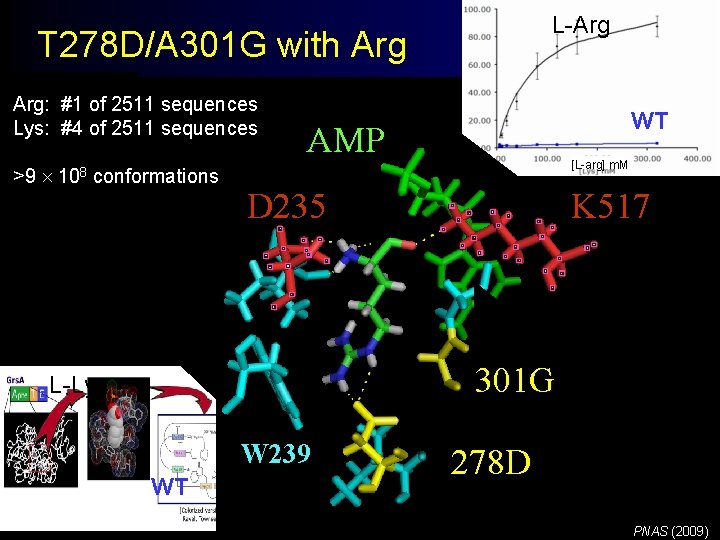
L-Arg T 278 D/A 301 G with Arg: #1 of 2511 sequences Lys: #4 of 2511 sequences >9 108 conformations WT AMP [L-arg] m. M D 235 K 517 301 G L-Lys W 239 WT 278 D PNAS (2009)

Tutorial

Installation Setup Running OSPREY
Installation Java mpi. Java 32 -bit √ MPICH 2 64 -bit may require special instructions

Setup Compute Nodes Input Structure Rotamer Library Energy Function
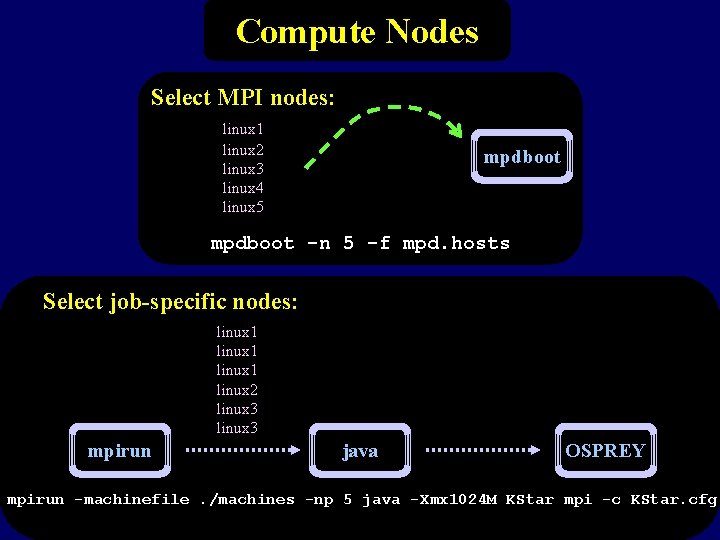
Compute Nodes Select MPI nodes: linux 1 linux 2 linux 3 linux 4 linux 5 mpdboot -n 5 -f mpd. hosts Select job-specific nodes: linux 1 linux 2 linux 3 mpirun java OSPREY mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi -c KStar. cfg
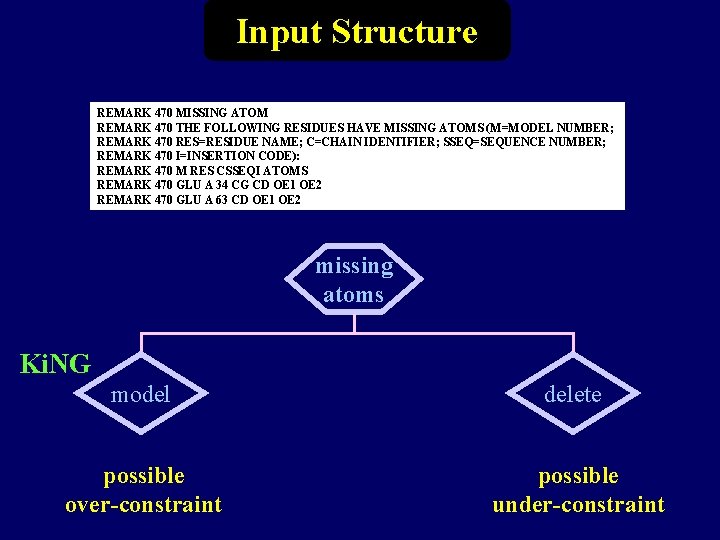
Input Structure REMARK 470 MISSING ATOM REMARK 470 THE FOLLOWING RESIDUES HAVE MISSING ATOMS (M=MODEL NUMBER; REMARK 470 RES=RESIDUE NAME; C=CHAIN IDENTIFIER; SSEQ=SEQUENCE NUMBER; REMARK 470 I=INSERTION CODE): REMARK 470 M RES CSSEQI ATOMS REMARK 470 GLU A 34 CG CD OE 1 OE 2 REMARK 470 GLU A 63 CD OE 1 OE 2 missing atoms Ki. NG model delete possible over-constraint possible under-constraint
Input Structure adding hydrogens proteins general compounds recommended: Mol. Probity recommended: Accelrys DS Visualizer Check: protonation states missing protons
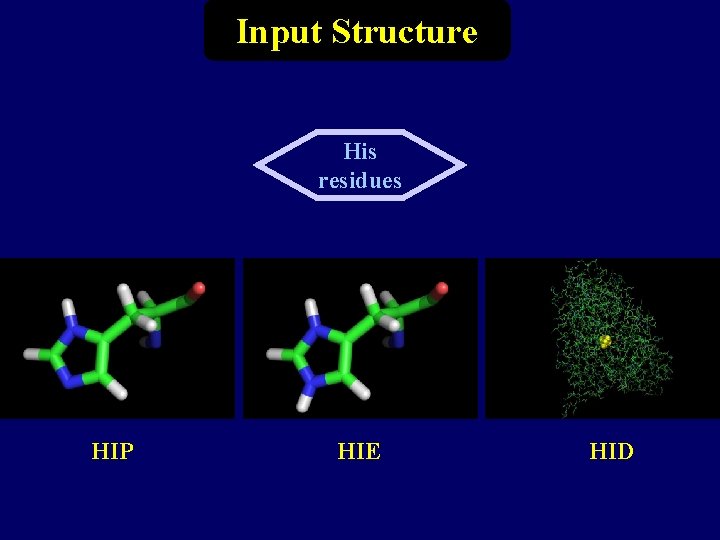
Input Structure His residues HIP HIE HID
Input Structure steric shell • close to design site • significant speedup

Input Structure Other considerations: • protein, ligand, cofactor • ligand: natural AA, small molecule • water molecules • no chain ID’s • unique residue numbers • protein-peptide, protein-protein • connectivity (good input structures)

Input Structure Check and double-check!!!
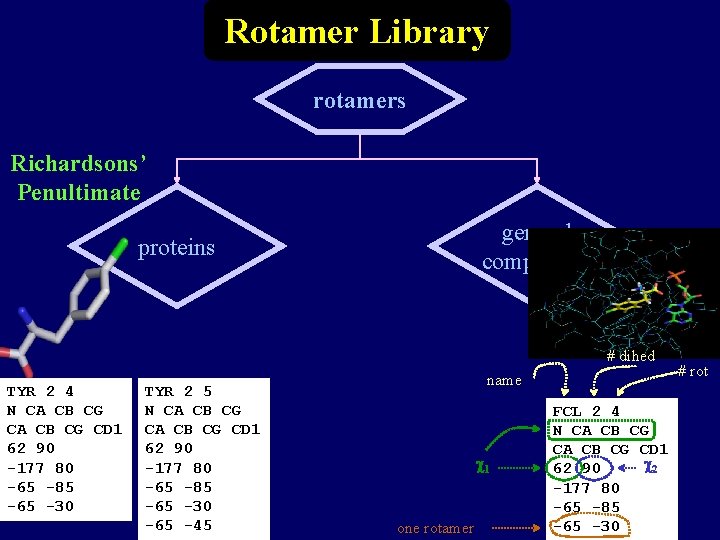
Rotamer Library rotamers Richardsons’ Penultimate general compounds proteins # dihed TYR 2 4 N CA CB CG CD 1 62 90 -177 80 -65 -85 -65 -30 TYR 2 5 N CA CB CG CD 1 62 90 -177 80 -65 -85 -65 -30 -65 -45 name 1 one rotamer FCL 2 4 N CA CB CG CD 1 2 62 90 -177 80 -65 -85 -65 -30 # rot

Energy Function parm 96 a. dat • atom types • dihedral parameters • vd. W parameters add params for new atom types antechamber all_amino 94 X. in all_nuc 94_and_gr. in • amino acids • partial charges • connectivity • general compounds • partial charges • connectivity typically no changes can modify partial charges add params for new compounds antechamber user control: distance-dependent dielectric, dielectric value, vd. W radii scaling, solvation energy scaling, dihedral energies switch

Running OSPREY GMEC-based Ensemble-based Residue entropy

GMEC-based mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi -c KStar. cfg do. DEE System. cfg DEE. cfg input structure rotamer library energy function mutation search parameters do. DEE energy minimization (Min. DEE, BD, BRDEE) DACS 1 MET GLY ASP ARG FCL 6 0 2 18 3 un. Min. E: -273. 75 min. E: -273. 75 best. E: -273. 75 2 MET GLY ASP MET FCL 6 0 2 6 3 un. Min. E: -271. 96 min. E: -271. 96 best. E: -273. 75 3 MET GLY ASP ARG FCL 6 0 2 18 3 un. Min. E: -271. 78 min. E: -271. 78 best. E: -273. 75 1 MET GLY SER ARG FCL 6 3 2 18 2 un. Min. E: -276. 50 min. E: -276. 50 best. E: -276. 50 2 MET GLY SER ARG FCL 6 3 1 18 2 un. Min. E: -276. 42 min. E: -276. 42 best. E: -276. 50

GMEC-based java -Xmx 1024 M KStar -c KStar. cfg gen. Struct. DEE System. cfg Gen. Struct. cfg input structure rotamer library energy function struct generation parameters gen. Struct. DEE energy minimization (Min. DEE, BD, BRDEE) 1 MET GLY SER ARG FCL 6 3 2 18 2 un. Min. E: -276. 50 min. E: -276. 50 best. E: -276. 50 2 MET GLY SER ARG FCL 6 3 1 18 2 un. Min. E: -276. 42 min. E: -276. 42 best. E: -276. 50 3 MET GLY ASP ARG FCL 6 0 2 18 3 un. Min. E: -273. 75 min. E: -273. 75 best. E: -273. 75 rank 1 MET GLY ASP ARG FCL 6 0 2 18 3 un. Min. E: -273. 75 min. E: -273. 75 best. E: -273. 75 2 MET GLY ASP MET FCL 6 0 2 6 3 un. Min. E: -271. 96 min. E: -271. 96 best. E: -273. 75 3 MET GLY ASP ARG FCL 6 0 2 18 3 un. Min. E: -271. 78 min. E: -271. 78 best. E: -273. 75 1 MET GLY SER ARG FCL 6 3 2 18 2 un. Min. E: -276. 50 min. E: -276. 50 best. E: -276. 50 2 MET GLY SER ARG FCL 6 3 1 18 2 un. Min. E: -276. 42 min. E: -276. 42 best. E: -276. 50
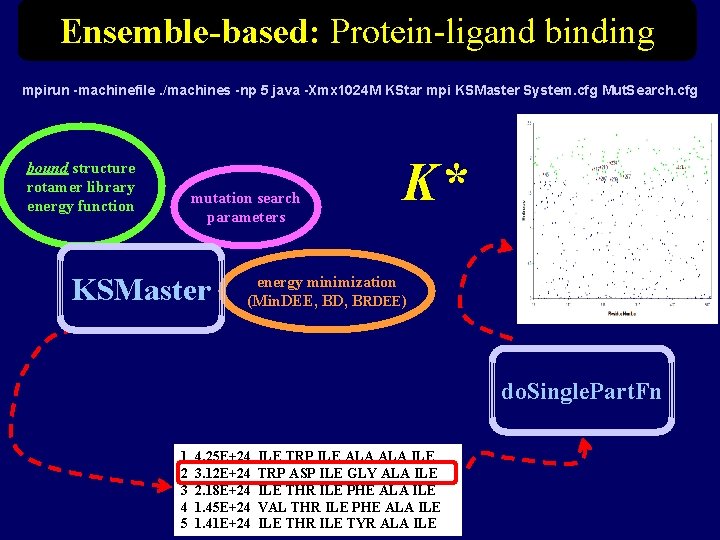
Ensemble-based: Protein-ligand binding mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi KSMaster System. cfg Mut. Search. cfg bound structure rotamer library energy function mutation search parameters KSMaster K* energy minimization (Min. DEE, BD, BRDEE) do. Single. Part. Fn 1 2 3 4 5 4. 25 E+24 3. 12 E+24 2. 18 E+24 1. 45 E+24 1. 41 E+24 ILE TRP ILE ALA ILE TRP ASP ILE GLY ALA ILE THR ILE PHE ALA ILE VAL THR ILE PHE ALA ILE THR ILE TYR ALA ILE
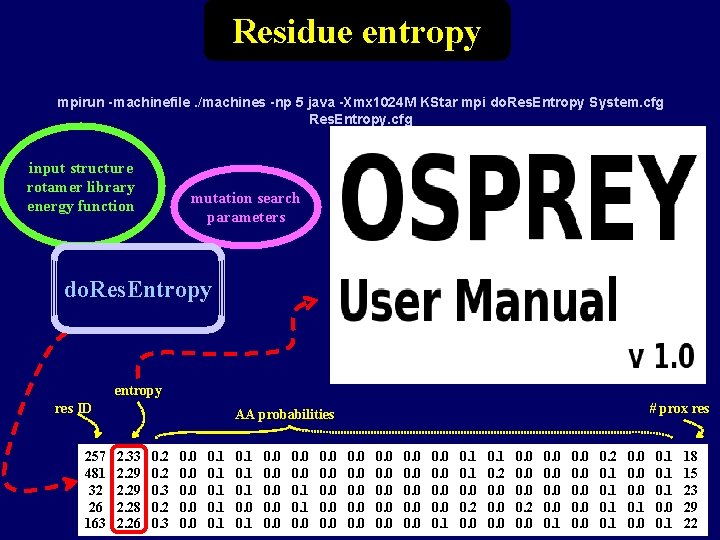
Residue entropy mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi do. Res. Entropy System. cfg Res. Entropy. cfg input structure rotamer library energy function mutation search parameters do. Res. Entropy entropy res ID 257 481 32 26 163 # prox res AA probabilities 2. 33 2. 29 2. 28 2. 26 0. 2 0. 3 0. 0 0. 1 0. 0 0. 0 0. 0 0. 1 0. 0 0. 2 0. 0 0. 1 0. 2 0. 0 0. 1 0. 0 0. 2 0. 1 0. 0 0. 1 18 15 23 29 22

Some important parameters mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi -c KStar. cfg do. DEE System. cfg DEE. cfg KStar. cfg: h. Elect true h. VDW false energy function h. Steric false dist. Dep. Dielect true dielect. Const 6. 0 vdw. Mult 0. 95 do. Dihed. E true do. Solvation. E true solv. Scale 0. 8 steric filter rotamer libraries volume filter steric. Thresh 0. 4 soft. Steric. Thresh 1. 5 rot. File Lovell. Rotamer. dat grot. File Generic. Rotamers. dat vol. File AAVolumes. dat
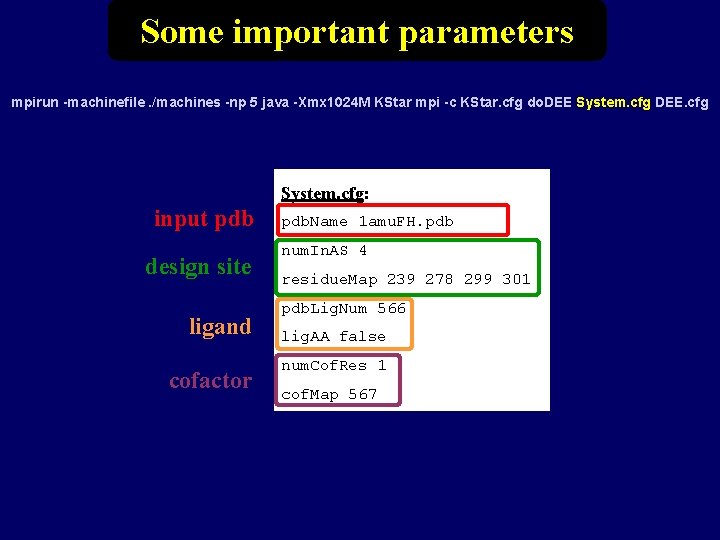
Some important parameters mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi -c KStar. cfg do. DEE System. cfg DEE. cfg System. cfg: input pdb design site ligand cofactor pdb. Name 1 amu. FH. pdb num. In. AS 4 residue. Map 239 278 299 301 pdb. Lig. Num 566 lig. AA false num. Cof. Res 1 cof. Map 567

Some important parameters mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi -c KStar. cfg do. DEE System. cfg DEE. cfg (partial): do. DACS true distr. DACS false DACS init. Depth 2 sub. Depth 1 diff. Fact 6 do. Minimize false minimization minimize. BB false do. Backrubs false backrub. File none reference energies ligand in search allowed mutations use. Eref true lig. Present false lig. Type none res. Allowed 0 gly ala val leu ile tyr phe trp met … res. Allowed 3 gly ala val leu ile tyr phe trp met resuming resume. Search false resume. Filename run. Info. out. partial

Some important parameters mpirun -machinefile. /machines -np 5 java -Xmx 1024 M KStar mpi KSMaster System. cfg Mut. Search. cfg (partial): mut. File. Name 1 amu. FCL_2 MUT. mut volume filter/ candidate mutants num. Mutations 2 target. Volume 620. 0 volume. Window 10000. 0 do. Minimize false minimization (1 -ε) accuracy inter-mutation at most 1 repeat unbound struct allowed mutations resuming minimize. BB false do. Backrubs false backrub. File none epsilon 0. 03 gamma 0. 01 repeat. Search true use. Unbound. Struct false unbound. Pdb. Name none res. Allowed 0 gly ala val leu ile tyr phe trp met resume. Search false resume. Filename 1 amu. FCL_Mut. Search. partial

Citing OSPREY General citation: K* and Min. DEE: BD: BRDEE: DACS: Original K* publication:

OSPREY is open source!!!

Acknowledgements Bruce Donald Ryan Lilien Funding: • NIH Faisal Reza Kyle Roberts Daniel Keedy Pablo Gainza Donald Lab

- Slides: 44