Towards Semantic Typing Support for Scientific Workflows Bertram

Towards Semantic Typing Support for Scientific Workflows Bertram Ludäscher Knowledge-Based Information Systems Lab San Diego Supercomputer Center University of California San Diego http: //seek. ecoinformatics. org http: //www. geongrid. org

Outline 1. Motivation: Traditional vs Scientific Data Integration 2. Semantic (a. k. a. Model-Based) Mediation 3. Scientific Workflows (a. k. a. Analysis Pipelines) 4. DB Theory Appetizer: Web Service Composition Through Declarative Queries B. Ludäscher – Scientific Data Management 2
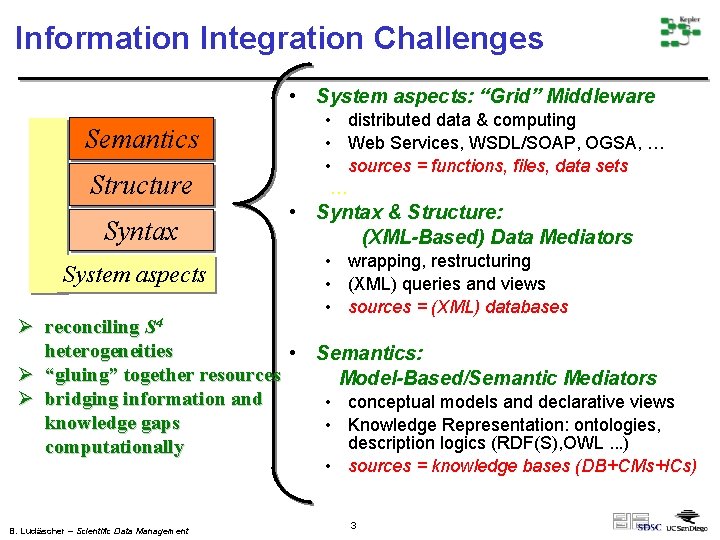
Information Integration Challenges • System aspects: “Grid” Middleware Semantics Structure Syntax System aspects • distributed data & computing • Web Services, WSDL/SOAP, OGSA, … • sources = functions, files, data sets … • Syntax & Structure: (XML-Based) Data Mediators • wrapping, restructuring • (XML) queries and views • sources = (XML) databases Ø reconciling S 4 heterogeneities • Semantics: Ø “gluing” together resources Model-Based/Semantic Mediators Ø bridging information and • conceptual models and declarative views knowledge gaps • Knowledge Representation: ontologies, description logics (RDF(S), OWL. . . ) computationally • sources = knowledge bases (DB+CMs+ICs) B. Ludäscher – Scientific Data Management 3

Information Integration from a DB Perspective • Information Integration Problem – Given: data sources S 1, . . . , Sk (DBMS, web sites, . . . ) and user questions Q 1, . . . , Qn that can be answered using the Si – Find: the answers to Q 1, . . . , Qn • The Database Perspective: source = “database” Þ Si has a schema (relational, XML, OO, . . . ) Þ Si can be queried Þ define virtual (or materialized) integrated/global view G over S 1 , . . . , Sk using database query languages (SQL, XQuery, . . . ) Þ questions become queries Qi against G(S 1, . . . , Sk) B. Ludäscher – Scientific Data Management 4
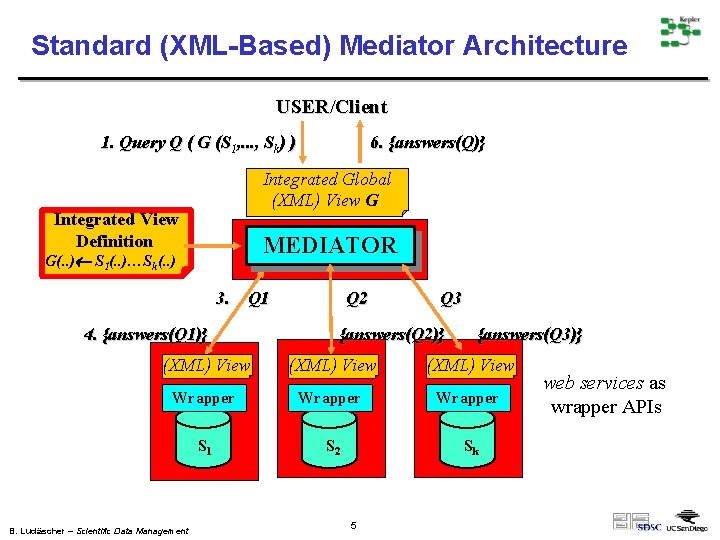
Standard (XML-Based) Mediator Architecture USER/Client 1. Query Q ( G (S 1, . . . , Sk) ) 6. {answers(Q)} Integrated Global (XML) View G Integrated View Definition MEDIATOR G(. . ) S 1(. . )…Sk(. . ) 3. Q 1 4. {answers(Q 1)} Q 2 Q 3 {answers(Q 2)} {answers(Q 3)} (XML) View Wrapper S 1 S 2 Sk B. Ludäscher – Scientific Data Management 5 web services as wrapper APIs

Query Planning for Mediators • Given: – User query Q: answer(…) …G. . . – …&{G …S…} global-as-view (GAV) – …&{S …G…} local-as-view (LAV) – … & { false … S … G… } integrity constraints (ICs) • Find: – equivalent (or min. containing, max. contained) query plan Q’: answer(…) … S … • Results: – A variety of results/algorithms; depending on classes of queries, views, and ICs: P, NP, …, undecidable – many variants still open B. Ludäscher – Scientific Data Management 6
From Scientific Data Integration to Process & Application Integration (and back…) • Data Integration – Database mediation + Knowledge-based extension Query rewriting w/ GAV, LAV, ICs, access patterns • “Process/Application”Integration – Scientific models (ocean, atmosphere, ecology, …), assimilation models (e. g. , real-time data feeds), … – Data sets – Legacy tools Components = web services Applications = composite components (“workflows”) Need for semantic type extensions B. Ludäscher – Scientific Data Management 7

Geologic Map Integration • Given: – Geologic maps from different state geological surveys (shapefiles w/ different data schemas) – Different ontologies: • • Geologic age ontology Rock type ontologies: – – • Multiple hierarchies (chemical, fabric, texture, genesis) from Geological Survey of Canada (GSC) Single hierarchy from British Geological Survey (BGS) Problem – Support uniform queries against the multiple geologic maps using different ontologies – Support registration w/ ontology A, querying w/ ontology B B. Ludäscher – Scientific Data Management 8
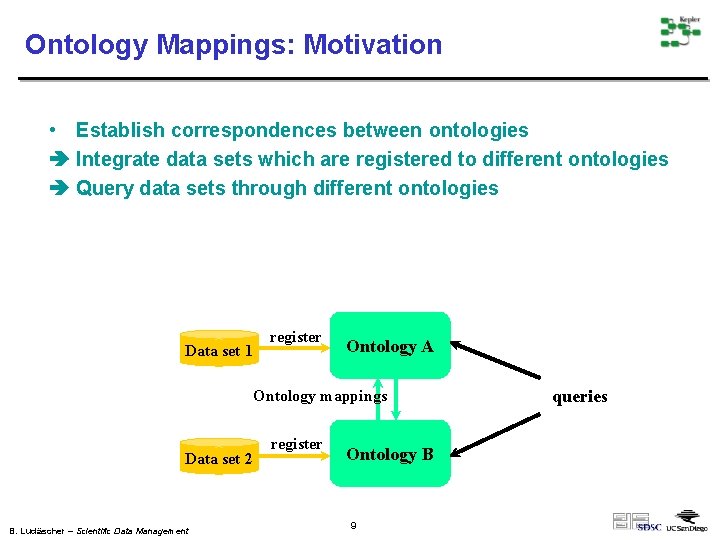
Ontology Mappings: Motivation • Establish correspondences between ontologies Integrate data sets which are registered to different ontologies Query data sets through different ontologies Data set 1 register Ontology A Ontology mappings Data set 2 B. Ludäscher – Scientific Data Management register Ontology B 9 queries
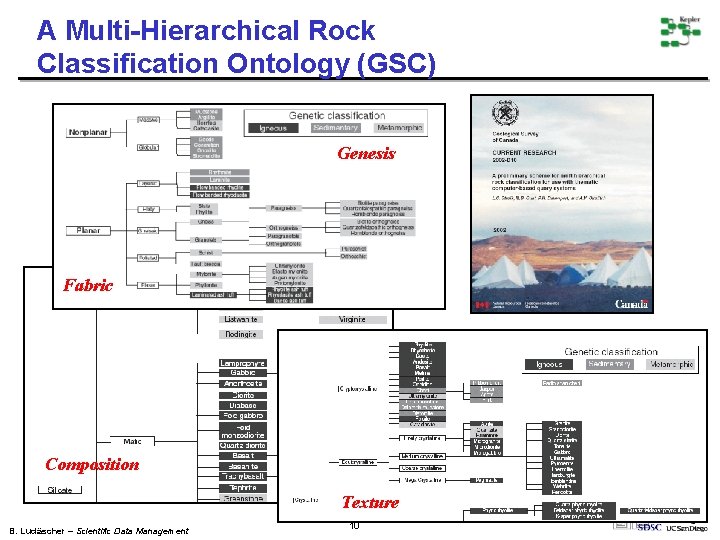
A Multi-Hierarchical Rock Classification Ontology (GSC) Genesis Fabric Composition Texture B. Ludäscher – Scientific Data Management 10
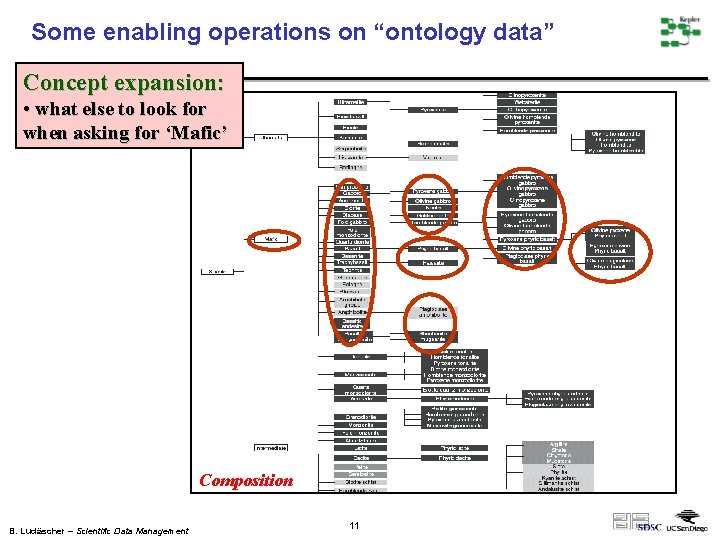
Some enabling operations on “ontology data” Concept expansion: • what else to look for when asking for ‘Mafic’ Composition B. Ludäscher – Scientific Data Management 11
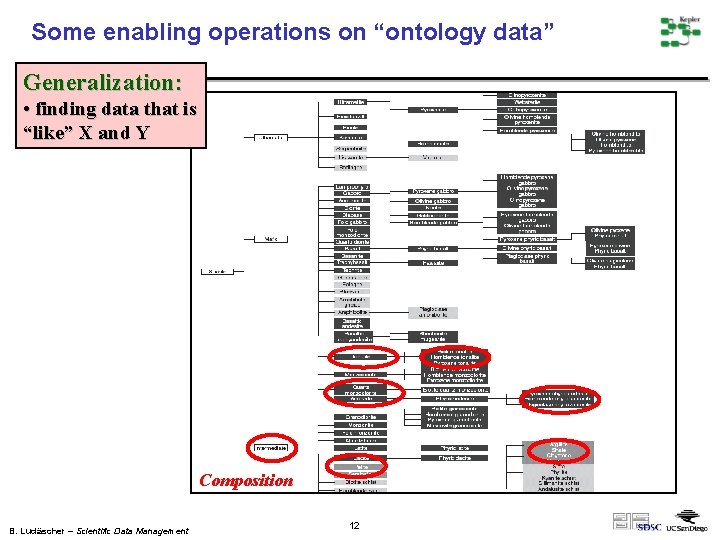
Some enabling operations on “ontology data” Generalization: • finding data that is “like” X and Y Composition B. Ludäscher – Scientific Data Management 12
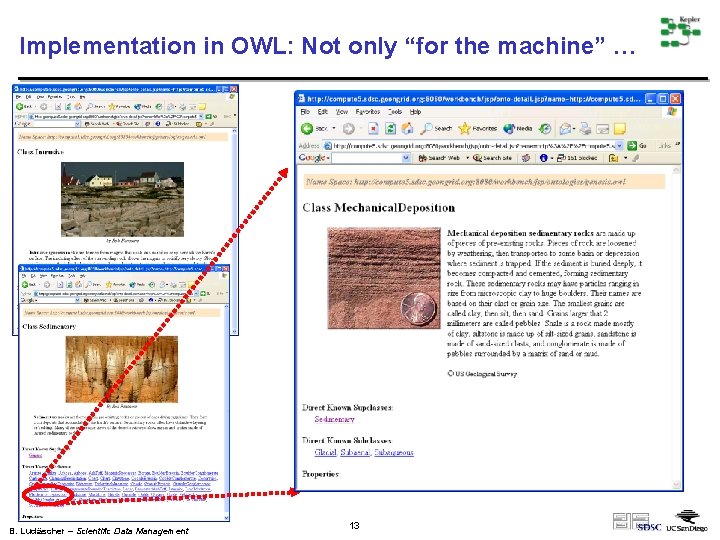
Implementation in OWL: Not only “for the machine” … B. Ludäscher – Scientific Data Management 13
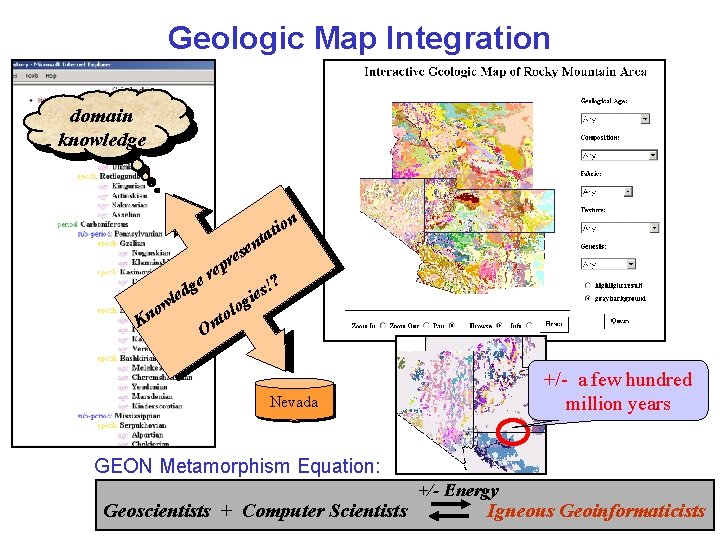
Geologic Map Integration domain knowledge io tat n n se e r p e er g led w ? ! s gie o l nto o n K O +/- a few hundred million years Nevada GEON Metamorphism Equation: Geoscientists + Computer Scientists +/- Energy Igneous Geoinformaticists
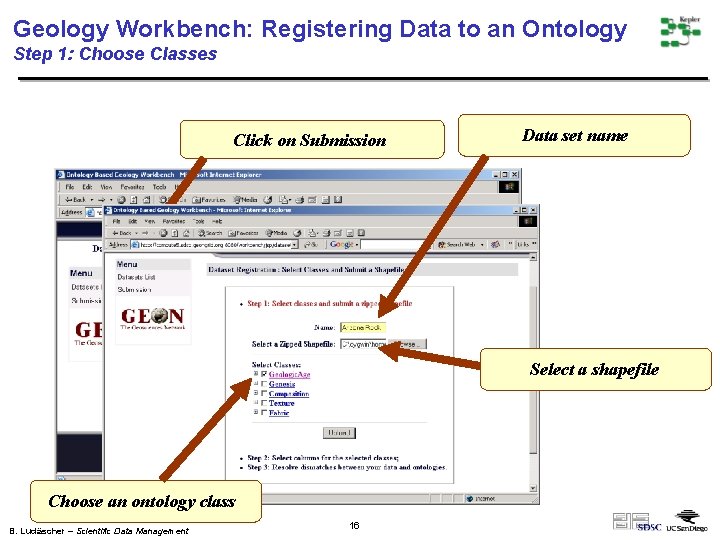
Geology Workbench: Registering Data to an Ontology Step 1: Choose Classes Click on Submission Data set name Select a shapefile Choose an ontology class B. Ludäscher – Scientific Data Management 16
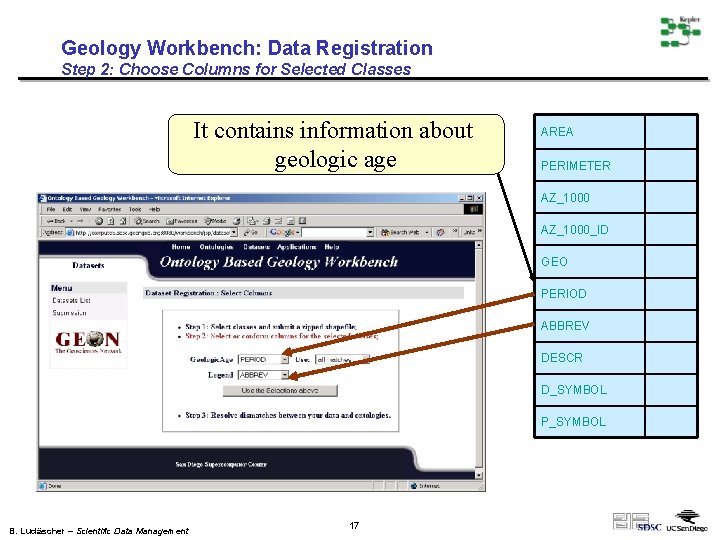
Geology Workbench: Data Registration Step 2: Choose Columns for Selected Classes It contains information about geologic age AREA PERIMETER AZ_1000_ID GEO PERIOD ABBREV DESCR D_SYMBOL P_SYMBOL B. Ludäscher – Scientific Data Management 17
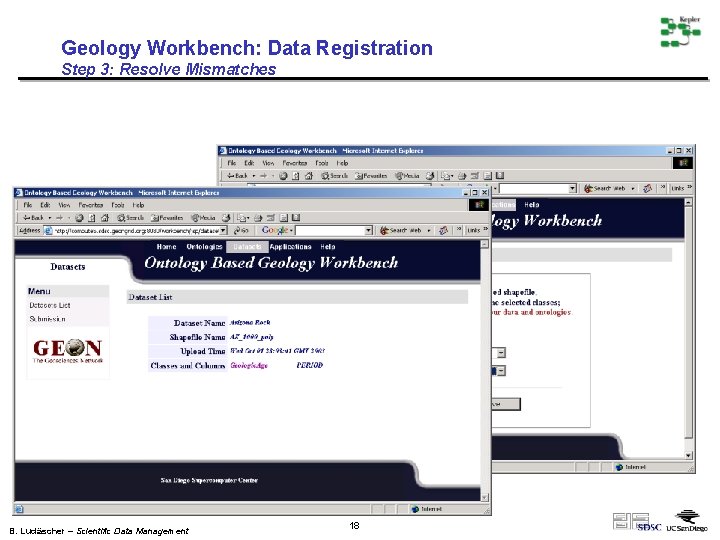
Geology Workbench: Data Registration Step 3: Resolve Mismatches Two terms are not matched any ontology terms Manually mapping algonkian into the ontology B. Ludäscher – Scientific Data Management 18
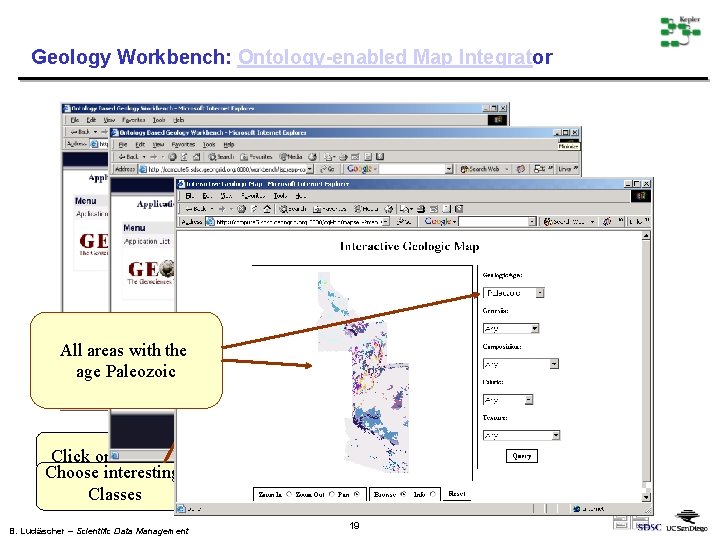
Geology Workbench: Ontology-enabled Map Integrator All areas with the age Paleozoic Click on the name Choose interesting Classes B. Ludäscher – Scientific Data Management 19
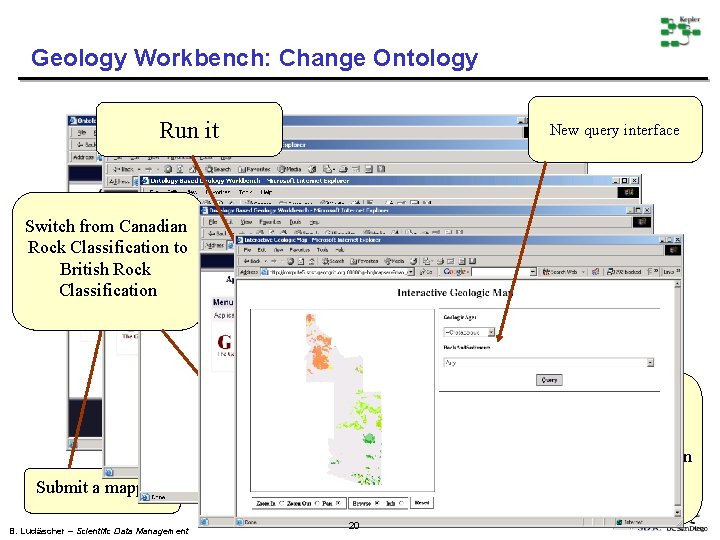
Geology Workbench: Change Ontology Run it New query interface Switch from Canadian Rock Classification to British Rock Classification Ontology mapping between British Rock Classification and Canadian Rock Classification Submit a mapping B. Ludäscher – Scientific Data Management 20
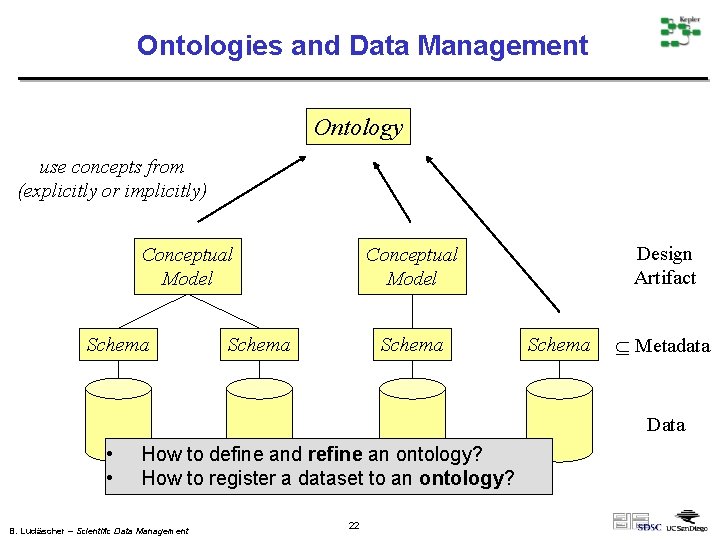
Ontologies and Data Management Ontology use concepts from (explicitly or implicitly) Conceptual Model Schema Design Artifact Conceptual Model Schema Metadata Data • • How to define and refine an ontology? How to register a dataset to an ontology? B. Ludäscher – Scientific Data Management 22
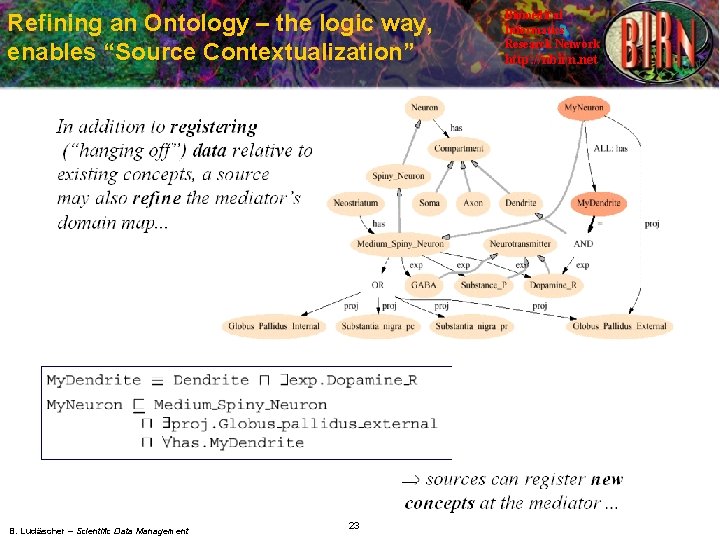
Refining an Ontology – the logic way, enables “Source Contextualization” B. Ludäscher – Scientific Data Management 23 Biomedical Informatics Research Network http: //nbirn. net
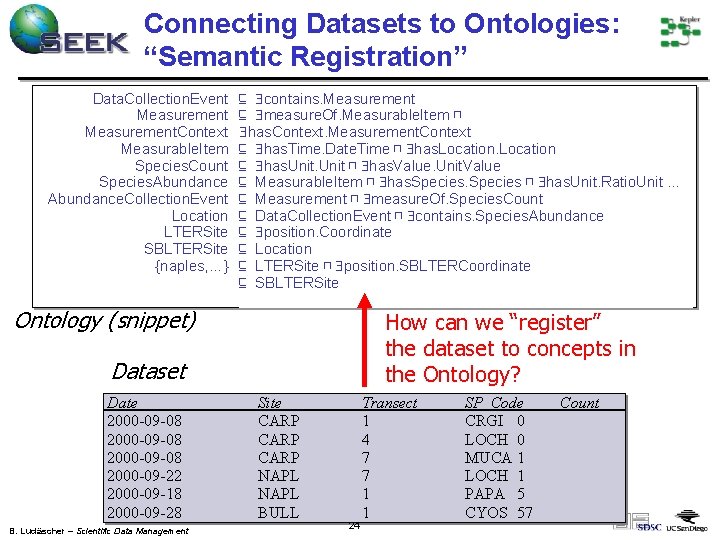
Connecting Datasets to Ontologies: “Semantic Registration” Data. Collection. Event Measurement. Context Measurable. Item Species. Count Species. Abundance. Collection. Event Location LTERSite SBLTERSite {naples, …} ⊑ contains. Measurement ⊑ measure. Of. Measurable. Item ⊓ has. Context. Measurement. Context ⊑ has. Time. Date. Time ⊓ has. Location ⊑ has. Unit ⊓ has. Value. Unit. Value ⊑ Measurable. Item ⊓ has. Species ⊓ has. Unit. Ratio. Unit … ⊑ Measurement ⊓ measure. Of. Species. Count ⊑ Data. Collection. Event ⊓ contains. Species. Abundance ⊑ position. Coordinate ⊑ Location ⊑ LTERSite ⊓ position. SBLTERCoordinate ⊑ SBLTERSite Ontology (snippet) How can we “register” the dataset to concepts in the Ontology? Dataset Date 2000 -09 -08 2000 -09 -22 2000 -09 -18 2000 -09 -28 B. Ludäscher – Scientific Data Management Site CARP NAPL BULL 24 Transect 1 4 7 7 1 1 SP_Code CRGI 0 LOCH 0 MUCA 1 LOCH 1 PAPA 5 CYOS 57 Count

Purpose of Semantic Registration Expose “hidden” information: – What do attributes represent? – What do specific values represent? – What conceptual “objects” are in the dataset? Capture connections between the dataset and ontology to: – Find existing datasets (or parts of datasets) via ontological concepts (discovery) – Enable integration of datasets (mediation) – Generate metadata for new data products (in a pipeline) B. Ludäscher – Scientific Data Management 25

Semantic Registration Framework Step 1: Data provider selects relevant ontological concepts (for the dataset) Step 2: The semantic registration system creates a structural representation based on chosen concepts (data provide refines if needed) Step 3: The data provider maps the dataset information to the generated structural representation B. Ludäscher – Scientific Data Management 26
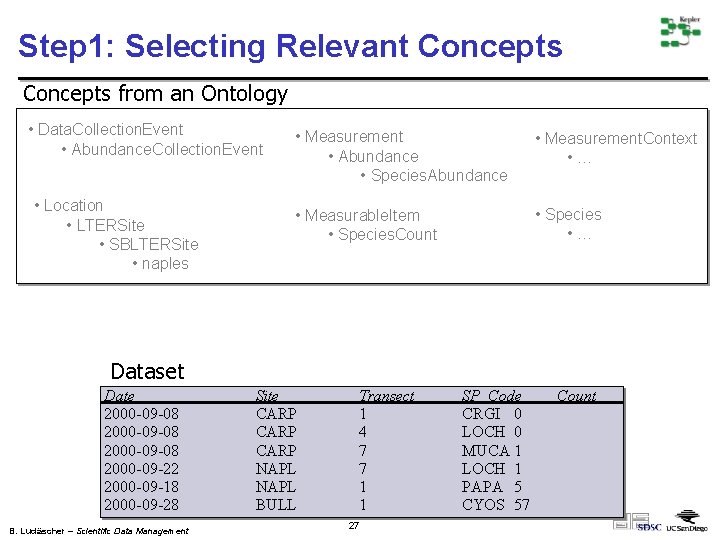
Step 1: Selecting Relevant Concepts from an Ontology • Data. Collection. Event • Abundance. Collection. Event • Location • LTERSite • SBLTERSite • naples • Measurement • Abundance • Species. Abundance • Measurement. Context • … • Measurable. Item • Species. Count • Species • … Dataset Date 2000 -09 -08 2000 -09 -22 2000 -09 -18 2000 -09 -28 B. Ludäscher – Scientific Data Management Site CARP NAPL BULL Transect 1 4 7 7 1 1 27 SP_Code CRGI 0 LOCH 0 MUCA 1 LOCH 1 PAPA 5 CYOS 57 Count
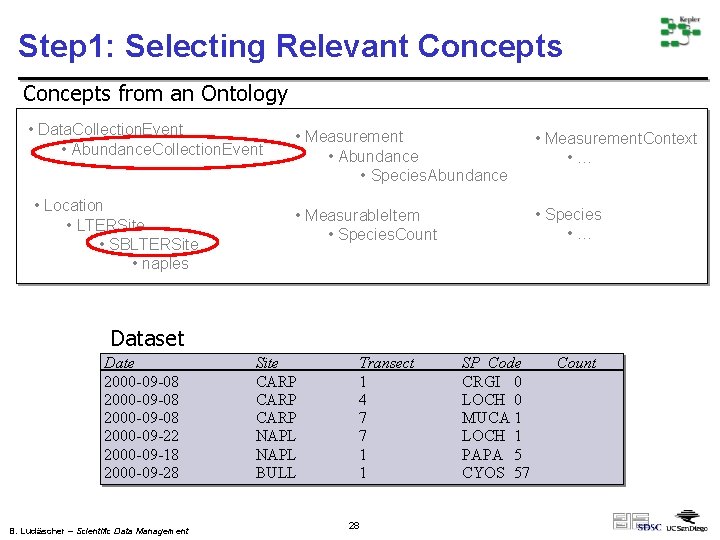
Step 1: Selecting Relevant Concepts from an Ontology • Data. Collection. Event • Abundance. Collection. Event • Location • LTERSite • SBLTERSite • naples • Measurement • Abundance • Species. Abundance • Measurement. Context • … • Measurable. Item • Species. Count • Species • … Dataset Date 2000 -09 -08 2000 -09 -22 2000 -09 -18 2000 -09 -28 B. Ludäscher – Scientific Data Management Site CARP NAPL BULL Transect 1 4 7 7 1 1 28 SP_Code CRGI 0 LOCH 0 MUCA 1 LOCH 1 PAPA 5 CYOS 57 Count
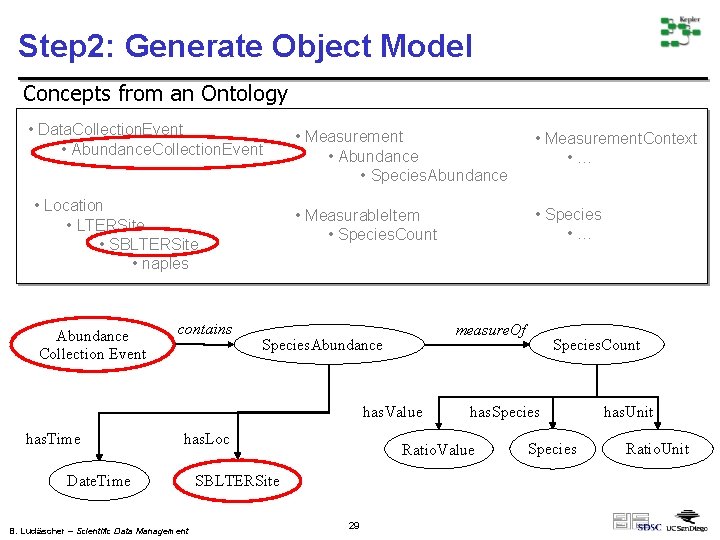
Step 2: Generate Object Model Concepts from an Ontology • Data. Collection. Event • Abundance. Collection. Event • Location • LTERSite • SBLTERSite • naples Abundance Collection Event contains • Measurement • Abundance • Species. Abundance • Measurement. Context • … • Measurable. Item • Species. Count • Species • … measure. Of Species. Abundance has. Value has. Time has. Loc Date. Time B. Ludäscher – Scientific Data Management has. Species Ratio. Value SBLTERSite 29 Species. Count Species has. Unit Ratio. Unit
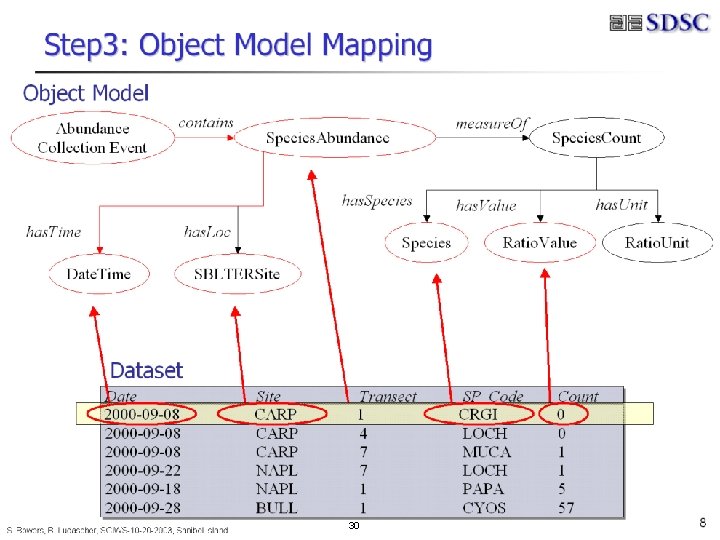
B. Ludäscher – Scientific Data Management 30
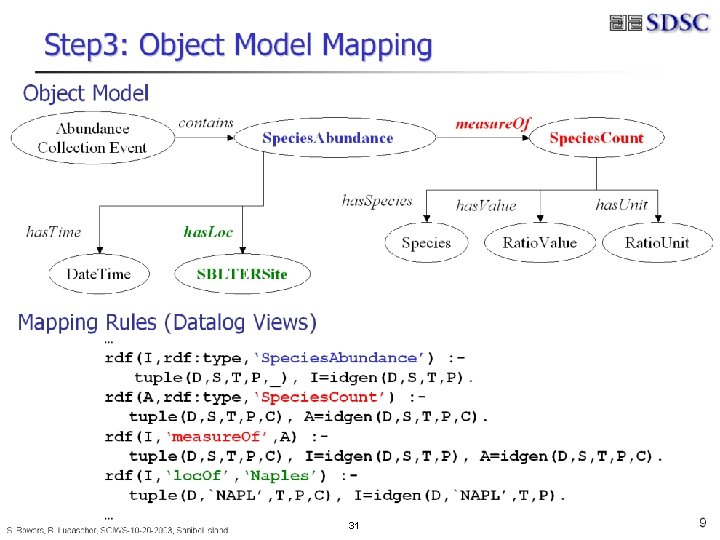
B. Ludäscher – Scientific Data Management 31
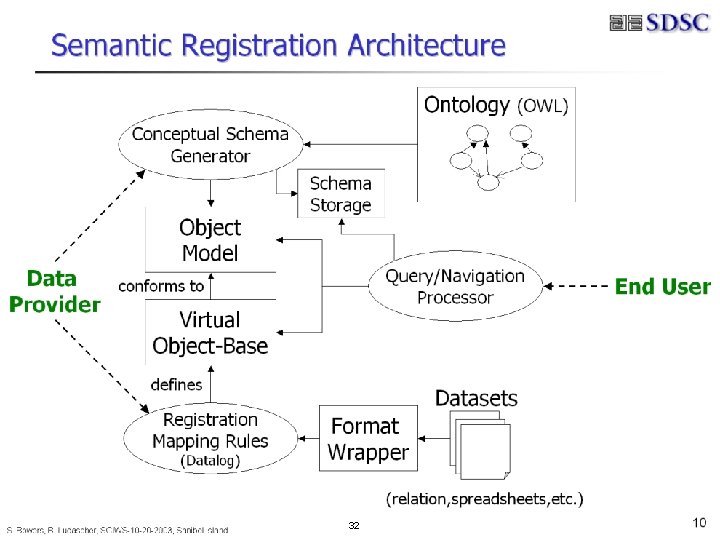
B. Ludäscher – Scientific Data Management 32

Scientific Workflows

Promoter Identification Workflow (PIW) B. Ludäscher – Scientific Data Management 34 Source: Matt Coleman (LLNL)
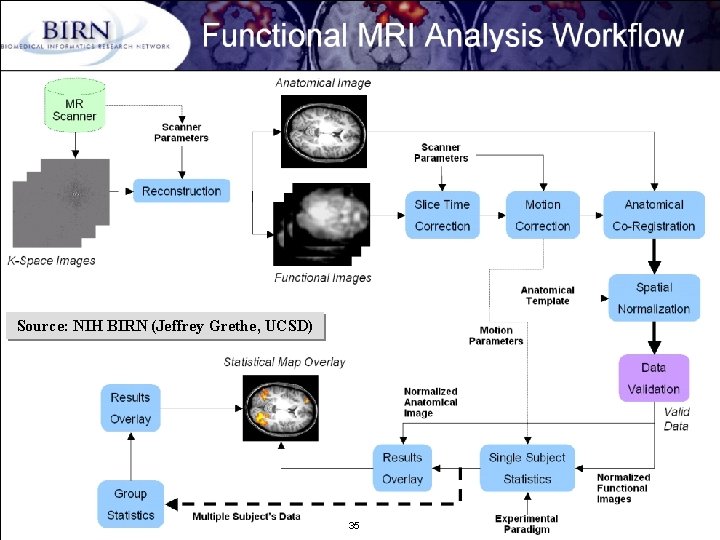
Source: NIH BIRN (Jeffrey Grethe, UCSD) B. Ludäscher – Scientific Data Management 35
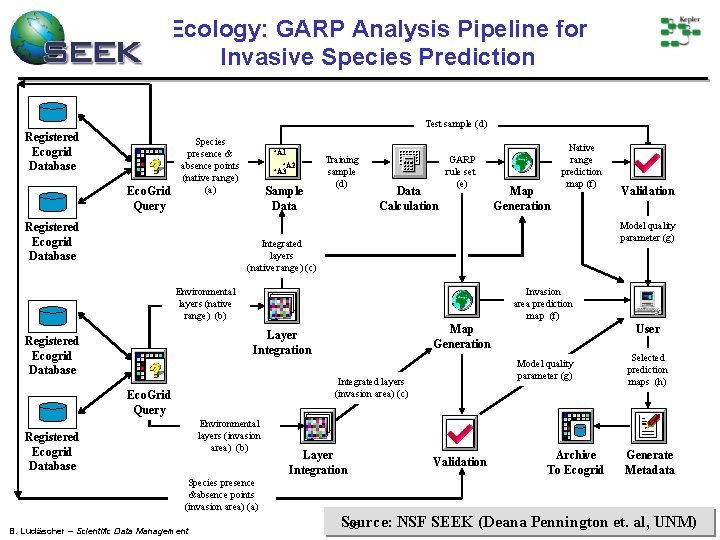
Ecology: GARP Analysis Pipeline for Invasive Species Prediction Test sample (d) Registered Ecogrid Database Eco. Grid Query Species presence & absence points (native range) (a) Registered Ecogrid Database +A 1 +A 2 +A 3 Sample Data Training sample (d) Data Calculation GARP rule set (e) Map Generation Native range prediction map (f) Model quality parameter (g) Integrated layers (native range) (c) Environmental layers (native range) (b) Invasion area prediction map (f) Map Generation Layer Integration Registered Ecogrid Database Environmental layers (invasion area) (b) Layer Integration User Model quality parameter (g) Integrated layers (invasion area) (c) Eco. Grid Query Registered Ecogrid Database Validation Archive To Ecogrid Selected prediction maps (h) Generate Metadata Species presence &absence points (invasion area) (a) B. Ludäscher – Scientific Data Management Source: NSF SEEK (Deana Pennington et. al, UNM) 36

Scientific Workflows: Some Findings • More dataflow than (business) workflow • Need for “programming extension” – Iterations over lists (foreach); filtering; functional composition; generic & higher-order operations (zip, map(f), …) • Need for abstraction and nested workflows • Need for data transformations • Need for rich user interaction & workflow steering: – pause / revise / resume – select & branch; e. g. , web browser capability at specific steps as part of a coordinated SWF • Need for high-throughput transfers (“grid-enabling”, “streaming”) • Need for persistence of intermediate products data provenance (“virtual data” concept) B. Ludäscher – Scientific Data Management 37

Our Starting Point: Dataflow Process Networks and Ptolemy II read! see! try! Source: Edward Lee et al. http: //ptolemy. eecs. berkeley. edu/ptolemy. II/

Kepler Team, Projects, Sponsors • Ilkay Altintas SDM • Chad Berkley SEEK • Shawn Bowers SEEK • Jeffrey Grethe BIRN • Christopher H. Brooks Ptolemy II • Zhengang Cheng SDM • Efrat Jaeger GEON Matt Jones SEEK • • Edward A. Lee Ptolemy II • Kai Lin GEON • Ashraf Memon GEON • Bertram Ludaescher BIRN, GEON, SDM, SEEK • Steve Mock NMI • Steve Neuendorffer Ptolemy II • Mladen Vouk SDM • Yang Zhao Ptolemy II • … B. Ludäscher – Scientific Data Management 39 Ptolemy II
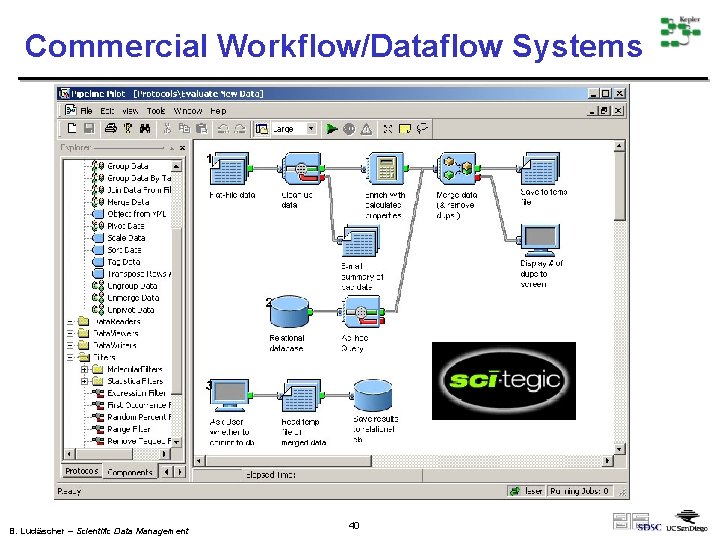
Commercial Workflow/Dataflow Systems B. Ludäscher – Scientific Data Management 40

SCIRun: Problem Solving Environments for Large-Scale Scientific Computing • • SCIRun: PSE for interactive construction, debugging, and steering of large-scale scientific computations Component model, based on generalized dataflow programming Steve Parker (cs. utah. edu) B. Ludäscher – Scientific Data Management 41

E-Science and Link-Up Buddies • … <UPDATE ME> … – Taverna, Scufl, Freefluo, . . – Discovery. Net – Triana – ICENI –… B. Ludäscher – Scientific Data Management 42
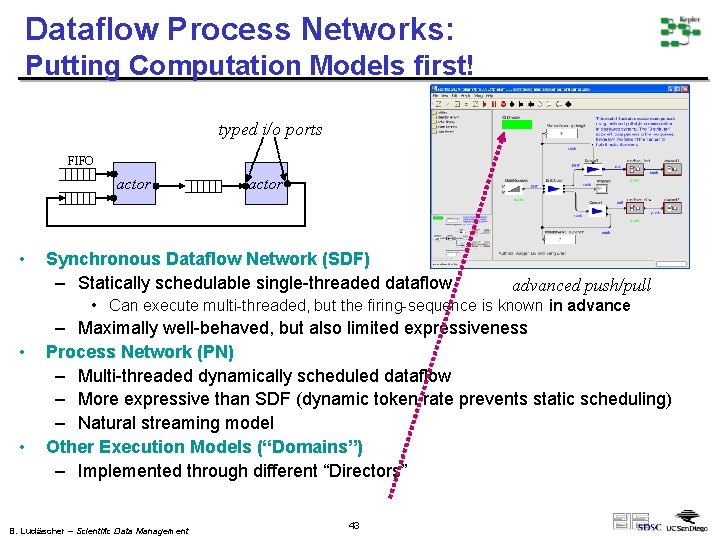
Dataflow Process Networks: Putting Computation Models first! typed i/o ports FIFO actor • actor Synchronous Dataflow Network (SDF) – Statically schedulable single-threaded dataflow advanced push/pull • Can execute multi-threaded, but the firing-sequence is known in advance • • – Maximally well-behaved, but also limited expressiveness Process Network (PN) – Multi-threaded dynamically scheduled dataflow – More expressive than SDF (dynamic token rate prevents static scheduling) – Natural streaming model Other Execution Models (“Domains”) – Implemented through different “Directors” B. Ludäscher – Scientific Data Management 43

Promoter Identification Workflow (PIW) B. Ludäscher – Scientific Data Management 44 Source: Matt Coleman (LLNL)
![Execution Semantics Promoter Identification Workflow in Ptolemy-II [SSDBM’ 03] B. Ludäscher – Scientific Data Execution Semantics Promoter Identification Workflow in Ptolemy-II [SSDBM’ 03] B. Ludäscher – Scientific Data](http://slidetodoc.com/presentation_image/bc647805a3f97ff46f2495b6a35d7a41/image-43.jpg)
Execution Semantics Promoter Identification Workflow in Ptolemy-II [SSDBM’ 03] B. Ludäscher – Scientific Data Management 45
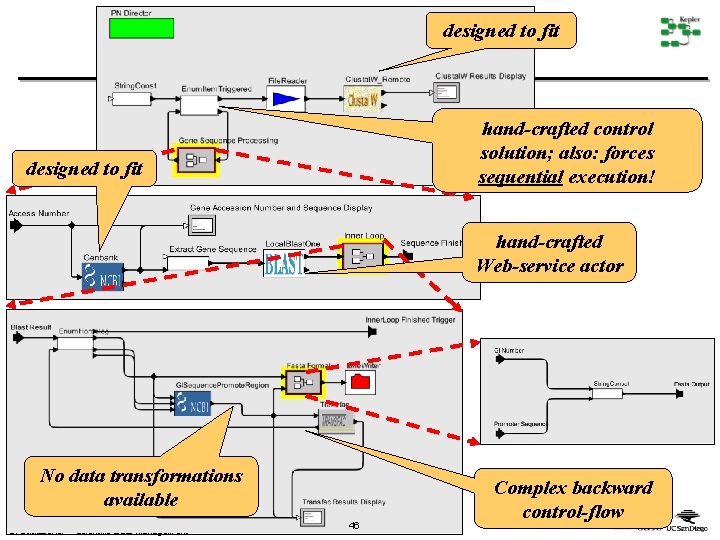
designed to fit hand-crafted control solution; also: forces sequential execution! designed to fit hand-crafted Web-service actor No data transformations available B. Ludäscher – Scientific Data Management 46 Complex backward control-flow
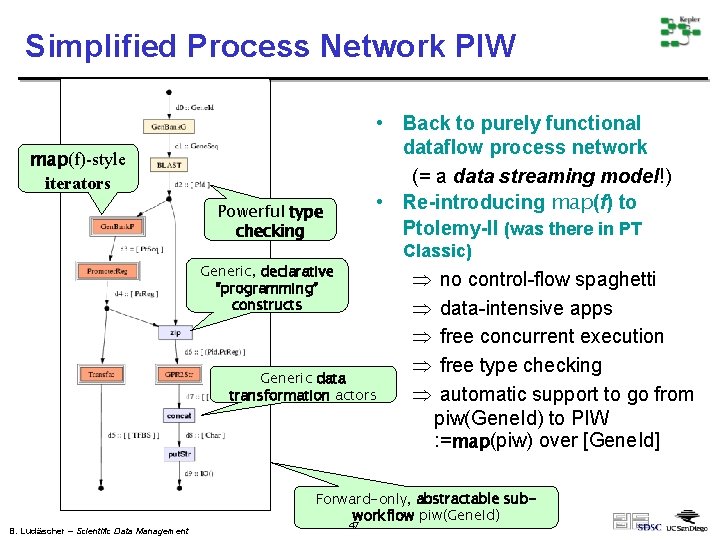
Simplified Process Network PIW • Back to purely functional dataflow process network (= a data streaming model!) • Re-introducing map(f) to Ptolemy-II (was there in PT map(f)-style iterators Powerful type checking Classic) Generic, declarative “programming” constructs Generic data transformation actors B. Ludäscher – Scientific Data Management Þ Þ Þ no control-flow spaghetti data-intensive apps free concurrent execution free type checking automatic support to go from piw(Gene. Id) to PIW : =map(piw) over [Gene. Id] Forward-only, abstractable subworkflow piw(Gene. Id) 47
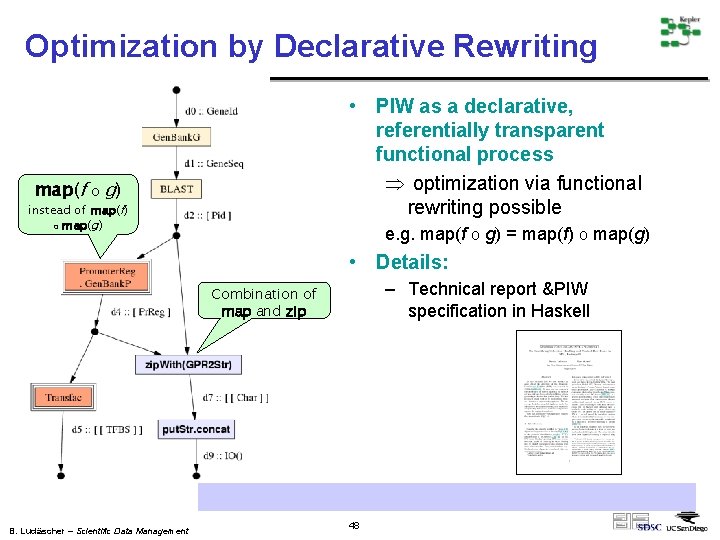
Optimization by Declarative Rewriting map(f o • PIW as a declarative, referentially transparent functional process Þ optimization via functional rewriting possible g) instead of map(f) o map(g) e. g. map(f o g) = map(f) o map(g) • Details: – Technical report &PIW specification in Haskell Combination of map and zip http: //kbi. sdsc. edu/Sci. DAC-SDM/scidac-tn-map-constructs. pdf B. Ludäscher – Scientific Data Management 48

Web Services & Scientific Workflows in Kepler • Web services = individual components (“actors”) • “Minute-Made” Application Integration: – Plugging-in and harvesting web service components is easy and fast • Rich SWF modeling semantics (“directors” and more): – Different and precise dataflow models of computation – Clear and composable component interaction semantics Web service composition and application integration tool • Coming soon: – Shrinked wrapped, pre-packaged “Kepler-to-Go” (v 0. 8) – SWFs with structural and semantic data types (better design support) – Grid-enabled web services (for big data, big computations, …) – Different deployment models (SWF WS, web site, applet, …) B. Ludäscher – Scientific Data Management 49

KEPLER Core Capabilities (1/2) • Designing scientific workflows – Composition of actors (tasks) to perform a scientific WF • Actor prototyping • Accessing heterogeneous data – Data access wizard to search and retrieve Grid-based resources – Relational DB access and query – Ability to link to EML data sources B. Ludäscher – Scientific Data Management 50

KEPLER Core Capabilities (2/2) • Data transformation actors to link heterogeneous data • Executing scientific workflows – Distributed and/or local computation – Various models for computational semantics and scheduling – SDF and PN: PN Most common for scientific workflows • External computing environments: – C++, Python, C (… Perl--planned. . . ) • Deploying scientific tasks and workflows as web services themselves (… planned …) B. Ludäscher – Scientific Data Management 51
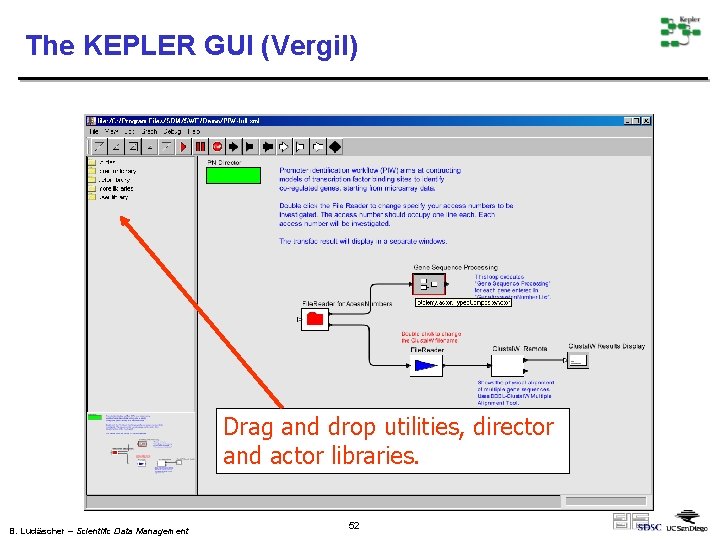
The KEPLER GUI (Vergil) Drag and drop utilities, director and actor libraries. B. Ludäscher – Scientific Data Management 52
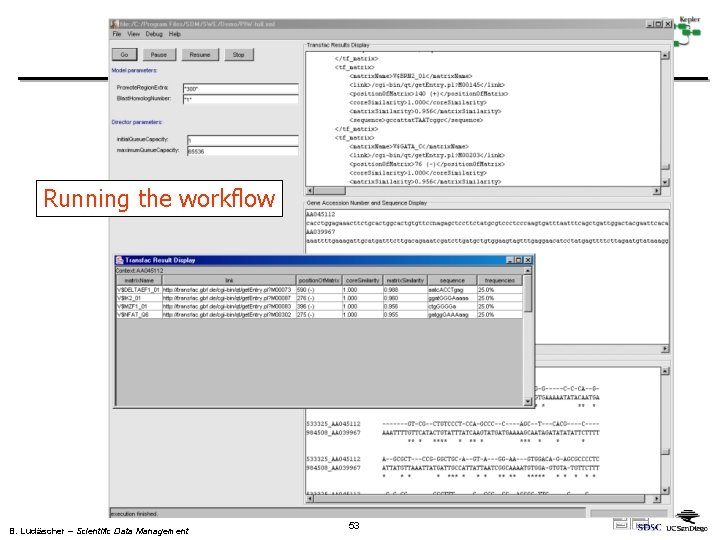
Running the workflow B. Ludäscher – Scientific Data Management 53

Distributed SWFs in KEPLER • Web and Grid Service plug-ins – WSDL, and whatever comes after GWSDL – Proxy. Init, Globus. Grid. Job, Grid. FTP, Data. Access. Wizard • WS Harvester – Imports all the operations of a specific WS (or of all the WSs in a UDDI repository) as Kepler actors • WS-deployment interface (…ongoing work…) • XSLT and XQuery transformers to link non-fitting services together B. Ludäscher – Scientific Data Management 54
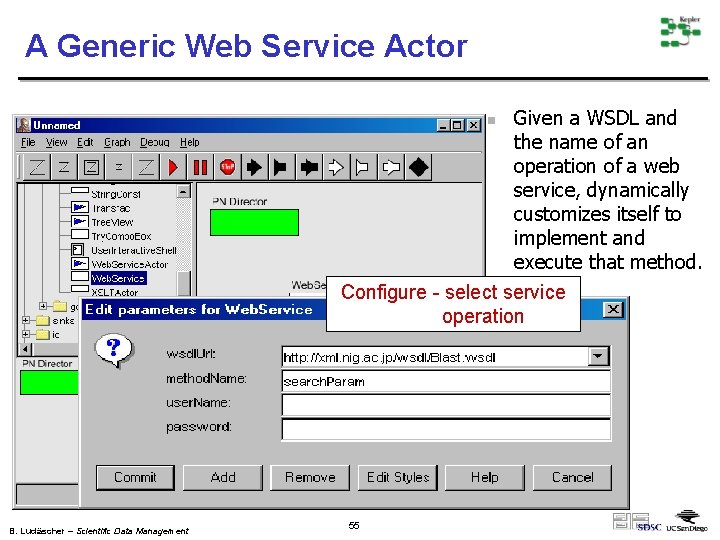
A Generic Web Service Actor Given a WSDL and the name of an operation of a web service, dynamically customizes itself to implement and execute that method. Configure - select service operation n B. Ludäscher – Scientific Data Management 55
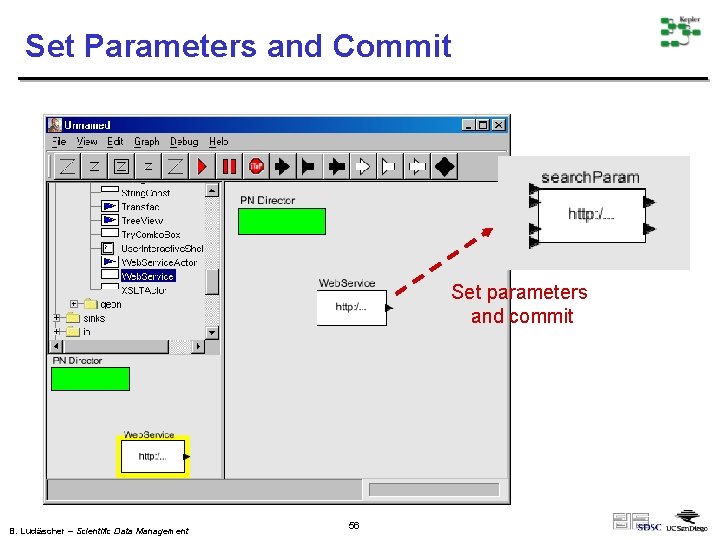
Set Parameters and Commit Set parameters and commit B. Ludäscher – Scientific Data Management 56
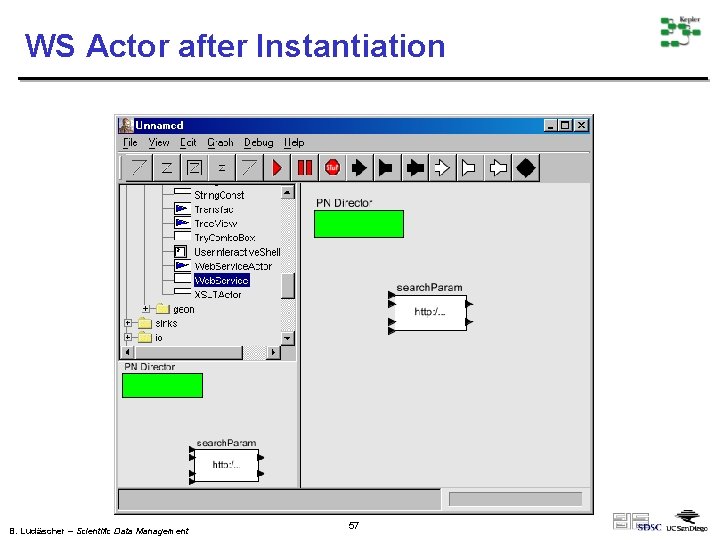
WS Actor after Instantiation B. Ludäscher – Scientific Data Management 57
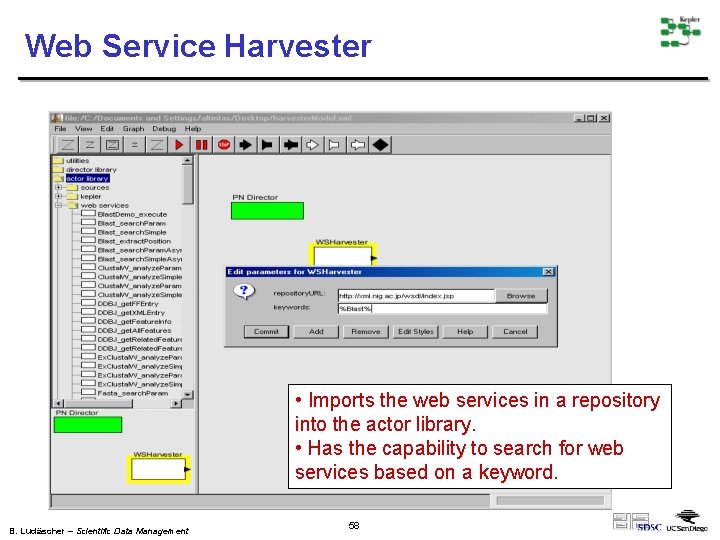
Web Service Harvester • Imports the web services in a repository into the actor library. • Has the capability to search for web services based on a keyword. B. Ludäscher – Scientific Data Management 58
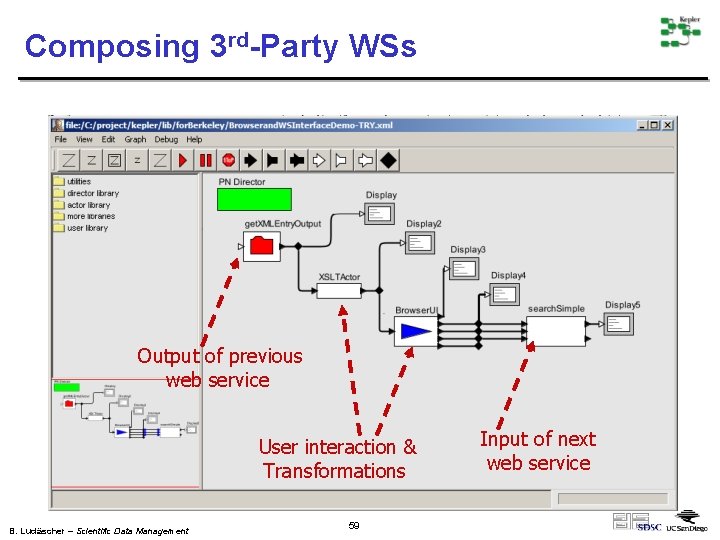
Composing 3 rd-Party WSs Output of previous web service User interaction & Transformations B. Ludäscher – Scientific Data Management 59 Input of next web service
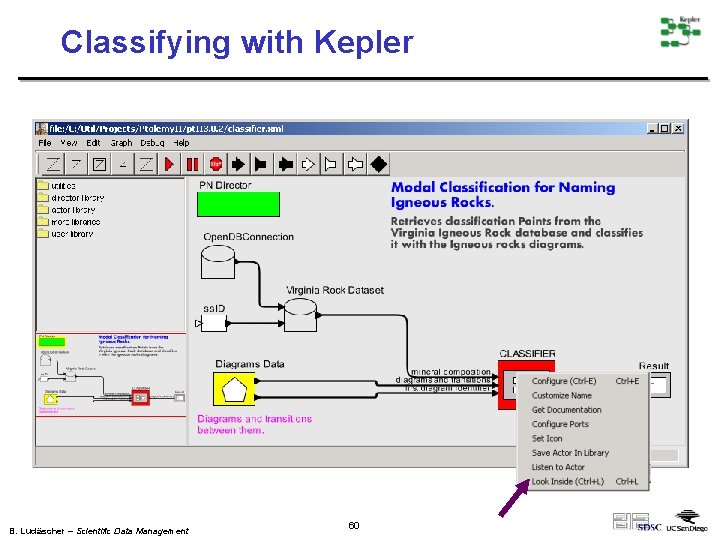
Classifying with Kepler B. Ludäscher – Scientific Data Management 60
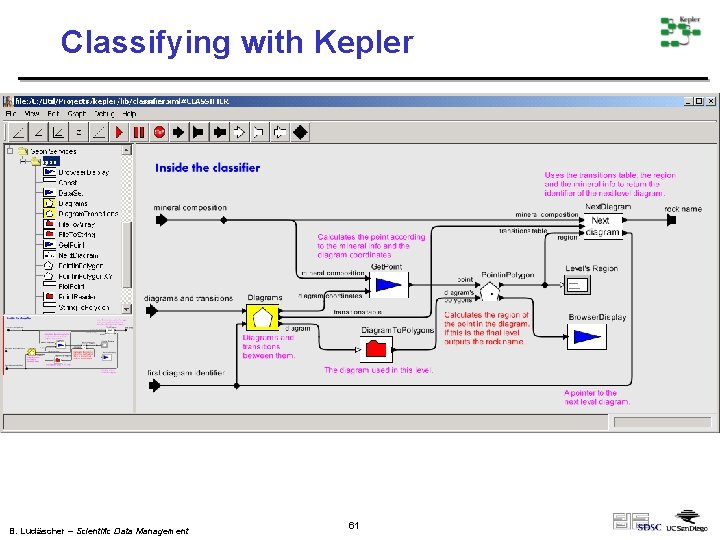
Classifying with Kepler B. Ludäscher – Scientific Data Management 61
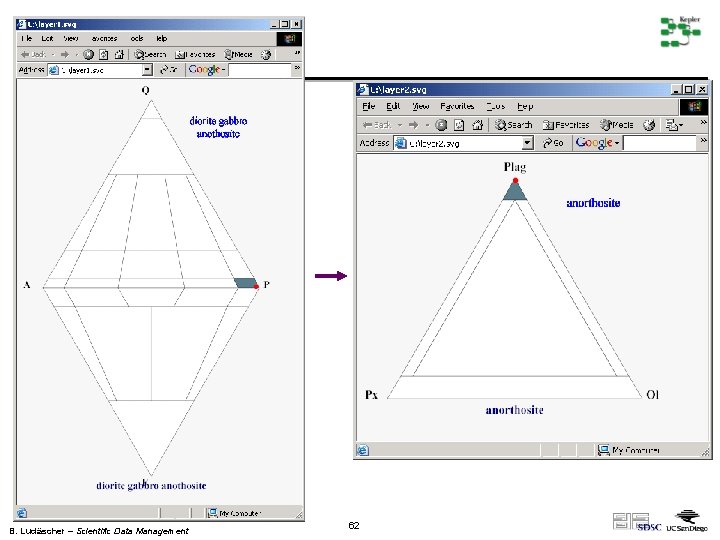
B. Ludäscher – Scientific Data Management 62
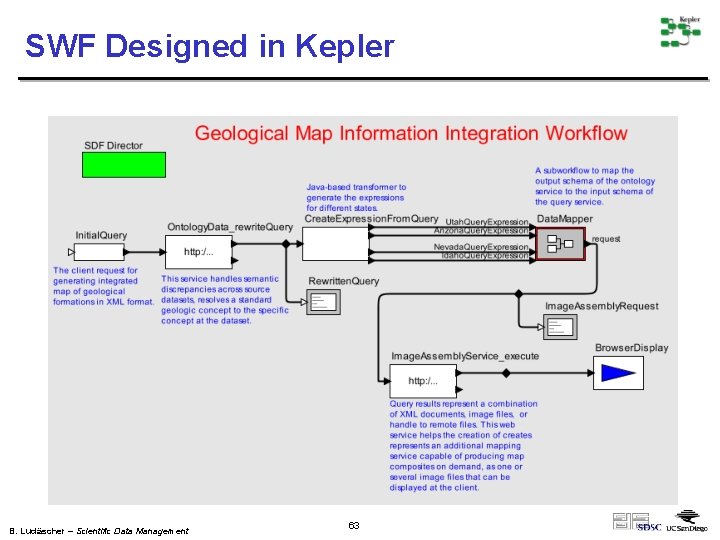
SWF Designed in Kepler B. Ludäscher – Scientific Data Management 63
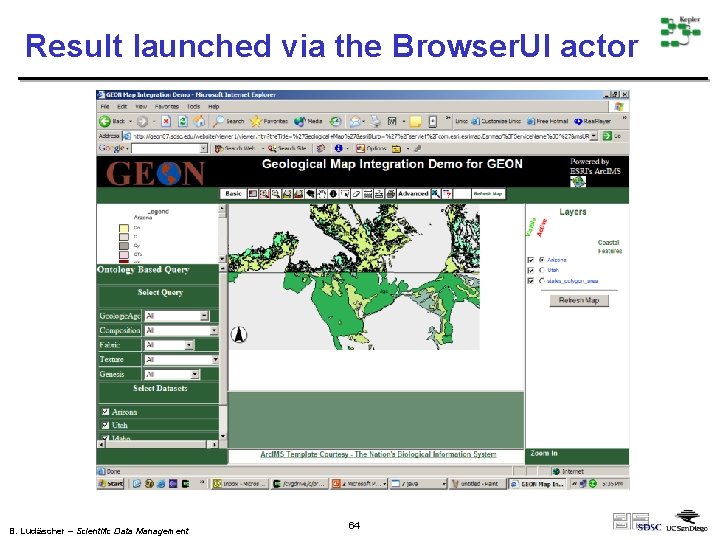
Result launched via the Browser. UI actor B. Ludäscher – Scientific Data Management 64
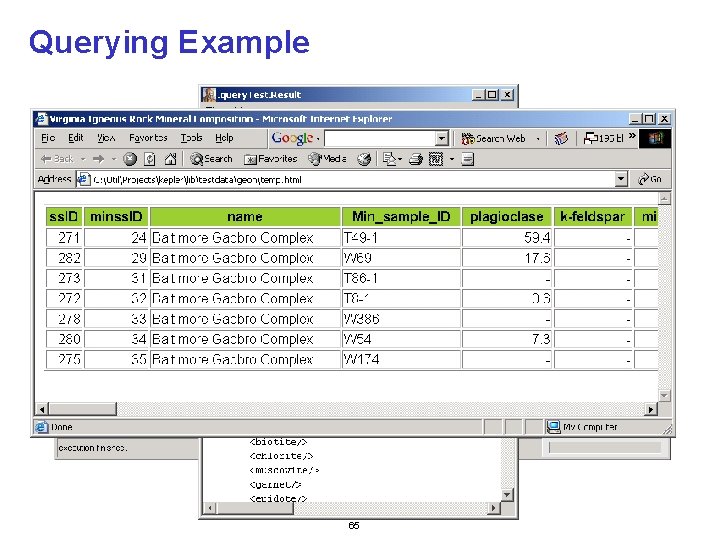
Querying Example 65

KEPLER and YOU • Kepler … – is a community-based, cross-project, open source collaboration – uses web services as basic building blocks – has a joint CVS repository, mailing lists, web site, … – is gaining momentum thanks to contributors and contributions • BSD-style license allows commercial spin-offs – a pre-packaged, shrink-wrapped version (“Kepler-to-GO”) coming soon to a place near you… B. Ludäscher – Scientific Data Management 66

Now back to the “Semantics Stuff”
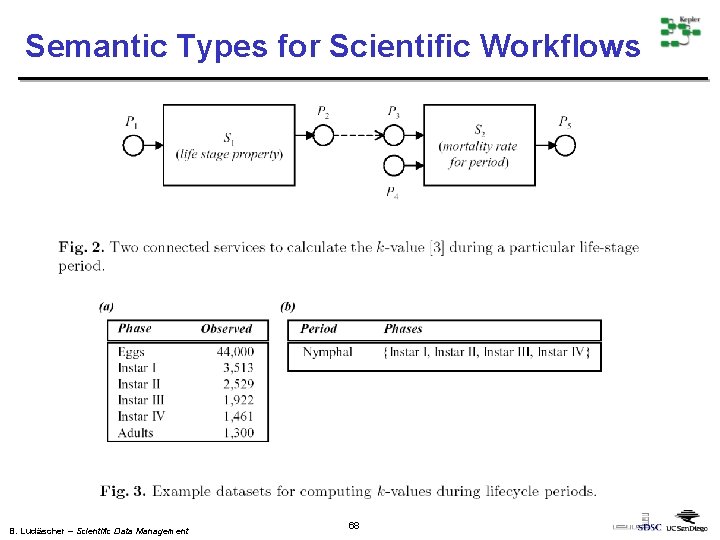
Semantic Types for Scientific Workflows B. Ludäscher – Scientific Data Management 68
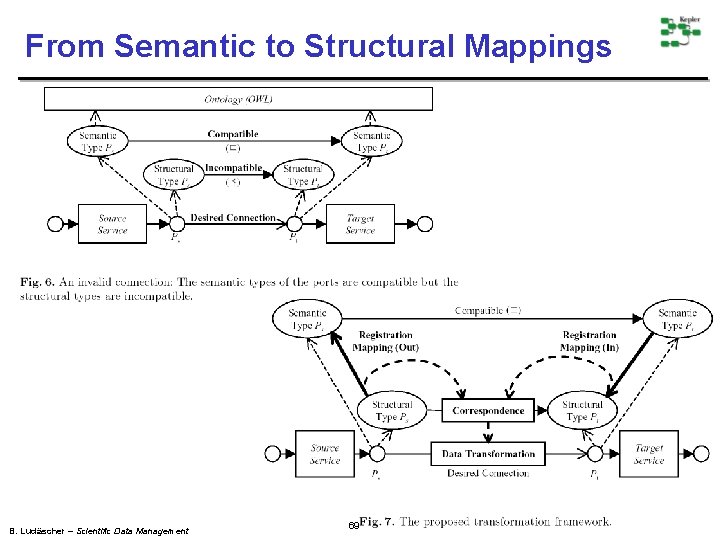
From Semantic to Structural Mappings B. Ludäscher – Scientific Data Management 69

Structural and Semantic Mappings B. Ludäscher – Scientific Data Management 70
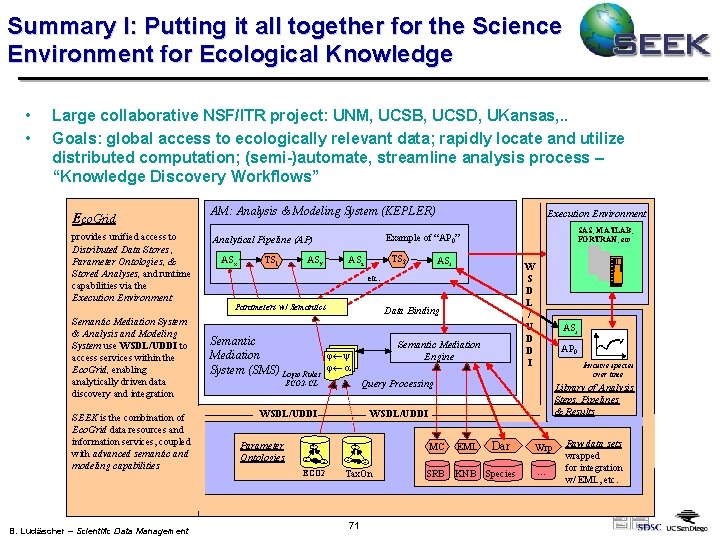
Summary I: Putting it all together for the Science Environment for Ecological Knowledge • • Large collaborative NSF/ITR project: UNM, UCSB, UCSD, UKansas, . . Goals: global access to ecologically relevant data; rapidly locate and utilize distributed computation; (semi-)automate, streamline analysis process – “Knowledge Discovery Workflows” Eco. Grid provides unified access to Distributed Data Stores , Parameter Ontologies, & Stored Analyses, and runtime capabilities via the Execution Environment Semantic Mediation System & Analysis and Modeling System use WSDL/UDDI to access services within the Eco. Grid, enabling analytically driven data discovery and integration SEEK is the combination of Eco. Grid data resources and information services, coupled with advanced semantic and modeling capabilities B. Ludäscher – Scientific Data Management AM: Analysis & Modeling System (KEPLER) TS 1 SAS, MATLAB, FORTRAN, etc Example of “AP 0” Analytical Pipeline (AP) ASx Execution Environment ASy TS 2 ASz ASr W S D L / U D D I etc. Parameters w/ Semantics Data Binding Semantic j¬y Mediation System (SMS) Logic Rules j¬ a Semantic Mediation Engine Invasive species over time Library of Analysis Steps, Pipelines & Results WSDL/UDDI ECO 2 C C AP 0 Query Processing ECO 2 -CL Parameter Ontologies ASr C ECO 2 C C EML SRB KNB Species C Tax. On 71 Dar MC Wrp . . . Raw data sets wrapped for integration w/ EML, etc.

Outline 1. Motivation: Traditional vs Scientific Data Integration 2. Semantic (a. k. a. Model-Based) Mediation 3. Scientific Workflows (a. k. a. Analysis Pipelines) 4. DB Theory Appetizer: Web Service Composition Through Declarative Queries B. Ludäscher – Scientific Data Management 72

Planning with Limited Access Patterns (back to GAV mediation …) • User query Q: answer(ISBN, Author, Title) book(ISBN, Author, Title), catalog(ISBN, Author), not library(ISBN). • Limited (web service) APIs (access patterns): – Src 1. books: in: ISBN out: Author, Title – Src 1. books: in: Author out: ISBN, Title – Src 2. catalog: in: {} out: ISBN, Author – Src 3. library: in: {} out: ISBN • Note: Q is not executable, but feasible (equivalent to executable Q’: catalog ; book ; not library) B. Ludäscher – Scientific Data Management 73
![Query Feasibility is as hard as Containment • Theorem [EDBT’ 04]: For UCQneg queries Query Feasibility is as hard as Containment • Theorem [EDBT’ 04]: For UCQneg queries](http://slidetodoc.com/presentation_image/bc647805a3f97ff46f2495b6a35d7a41/image-72.jpg)
Query Feasibility is as hard as Containment • Theorem [EDBT’ 04]: For UCQneg queries Q: Q is feasible iff ans(Q) Q • The answerable part ans(Q) can be computed in quadratic time. Idea: scan Q for answerable literals, rescan, repeat until ans(Q) is reached • Checking query containment Q 1 Q 2 is hard: – Already NP-complete for CQ (conjunctive queries) – Undecidable for FO (first-order logic queries) B. Ludäscher – Scientific Data Management 74

Conjunctive Query Containment • • Given: conjunctive queries Q 1, Q 2 (aka Select-Project-Join queries) Problem: Is answers(D, Q 1) answers(D, Q 2) for all databases D? If yes, we say that “Q 1 is contained in Q 2”; short: Q 1 Q 2 Examples: Q 1: answer(X) student(X, cs) Q 2: answer(X) student(X, Dept), advisor(X, Y), dept(Y, cs) Q 3: answer(X) student(X, Dept) • Quiz: – Q 1 Q 2 ? – No: not every student X necessarily has an adviser Y who is in the cs department! – Q 1 Q 3 ? – Yes: every cs student in some department (crux of the “proof”: Dept = cs) Homework: What about Q 1 Q 2 if we know that every student must have an advisor from the same department? B. Ludäscher – Scientific Data Management 75

The World’s Shortest Conjunctive Query Containment Checker (an NP-complete problem): 7 lines in Prolog … Quiz: 1. find the bug in the 7 lines of code 2. Fix the bug (hint: add one more line of code) Moral: Short programs can be buggy too B. Ludäscher – Scientific Data Management 76

Summary II: Got milk/eggs/meat/wool? Or: “Die eierlegende Wollmilchsau …” • Data Integration – query rewriting under GAV/LAV – w/ binding pattern constraints – distributed query processing • Semantic Mediation – semantic integrity constraints, reasoning w/ plans, automated deduction – deductive database/logic programming technology, AI “stuff”. . . – Semantic Web technology • Scientific Workflow Management – more procedural than database mediation (the scientist is the “query planner”) – deployment using web services B. Ludäscher – Scientific Data Management 77
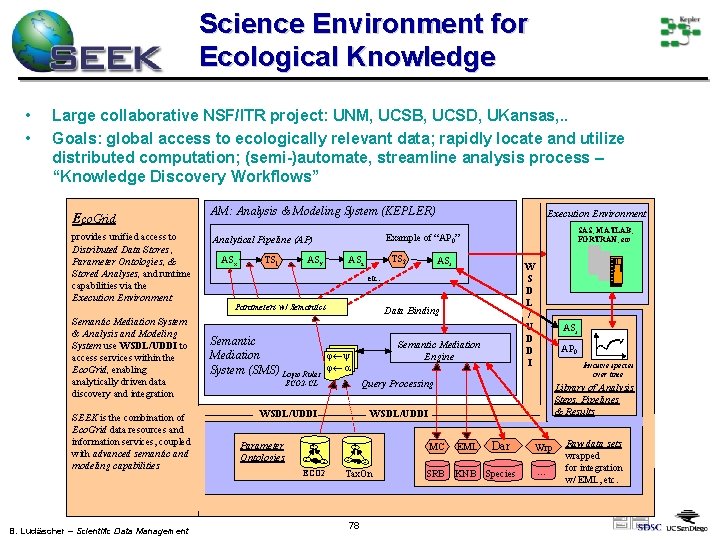
Science Environment for Ecological Knowledge • • Large collaborative NSF/ITR project: UNM, UCSB, UCSD, UKansas, . . Goals: global access to ecologically relevant data; rapidly locate and utilize distributed computation; (semi-)automate, streamline analysis process – “Knowledge Discovery Workflows” Eco. Grid provides unified access to Distributed Data Stores , Parameter Ontologies, & Stored Analyses, and runtime capabilities via the Execution Environment Semantic Mediation System & Analysis and Modeling System use WSDL/UDDI to access services within the Eco. Grid, enabling analytically driven data discovery and integration SEEK is the combination of Eco. Grid data resources and information services, coupled with advanced semantic and modeling capabilities B. Ludäscher – Scientific Data Management AM: Analysis & Modeling System (KEPLER) TS 1 SAS, MATLAB, FORTRAN, etc Example of “AP 0” Analytical Pipeline (AP) ASx Execution Environment ASy TS 2 ASz ASr W S D L / U D D I etc. Parameters w/ Semantics Data Binding Semantic j¬y Mediation System (SMS) Logic Rules j¬ a Semantic Mediation Engine Invasive species over time Library of Analysis Steps, Pipelines & Results WSDL/UDDI ECO 2 C C AP 0 Query Processing ECO 2 -CL Parameter Ontologies ASr C ECO 2 C C EML SRB KNB Species C Tax. On 78 Dar MC Wrp . . . Raw data sets wrapped for integration w/ EML, etc.
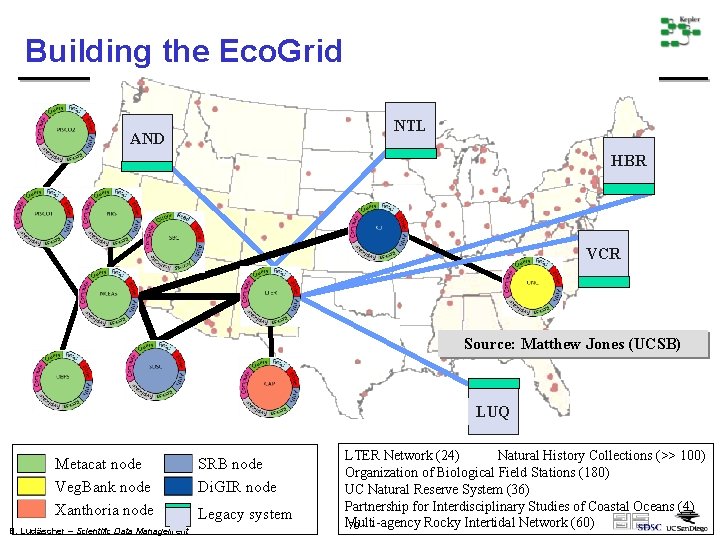
Building the Eco. Grid NTL AND HBR VCR Source: Matthew Jones (UCSB) LUQ Metacat node Veg. Bank node Xanthoria node B. Ludäscher – Scientific Data Management SRB node Di. GIR node Legacy system LTER Network (24) Natural History Collections (>> 100) Organization of Biological Field Stations (180) UC Natural Reserve System (36) Partnership for Interdisciplinary Studies of Coastal Oceans (4) Multi-agency Rocky Intertidal Network (60) 79
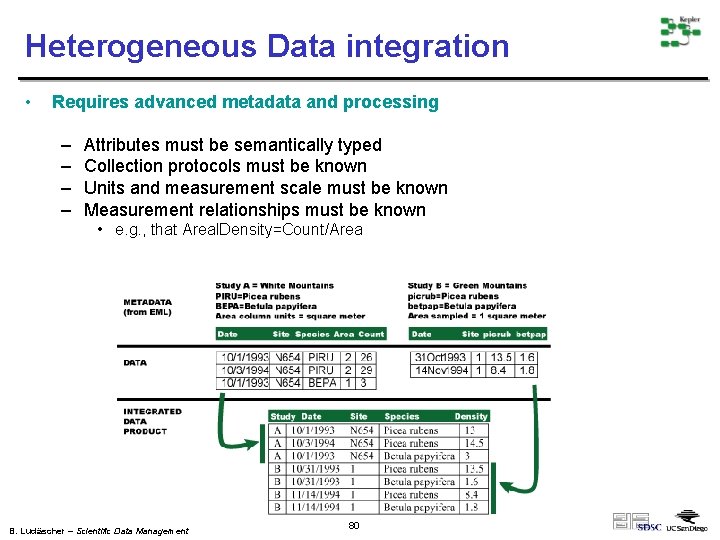
Heterogeneous Data integration • Requires advanced metadata and processing – – Attributes must be semantically typed Collection protocols must be known Units and measurement scale must be known Measurement relationships must be known • e. g. , that Areal. Density=Count/Area B. Ludäscher – Scientific Data Management 80
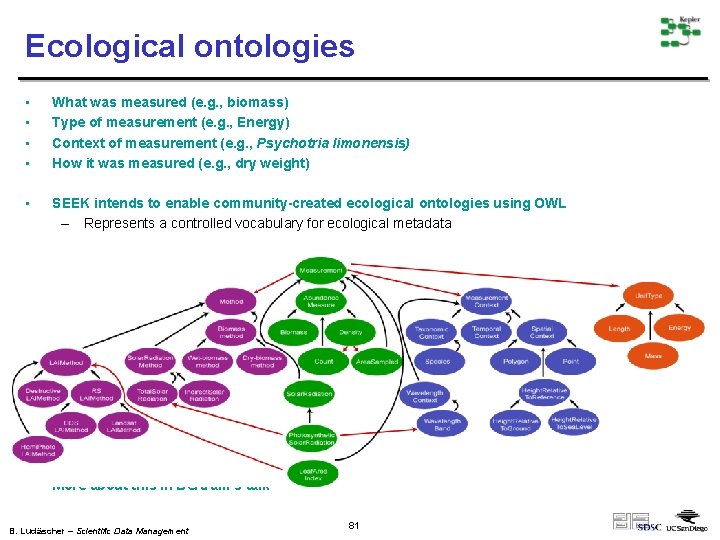
Ecological ontologies • • What was measured (e. g. , biomass) Type of measurement (e. g. , Energy) Context of measurement (e. g. , Psychotria limonensis) How it was measured (e. g. , dry weight) • SEEK intends to enable community-created ecological ontologies using OWL – Represents a controlled vocabulary for ecological metadata • More about this in Bertram’s talk B. Ludäscher – Scientific Data Management 81
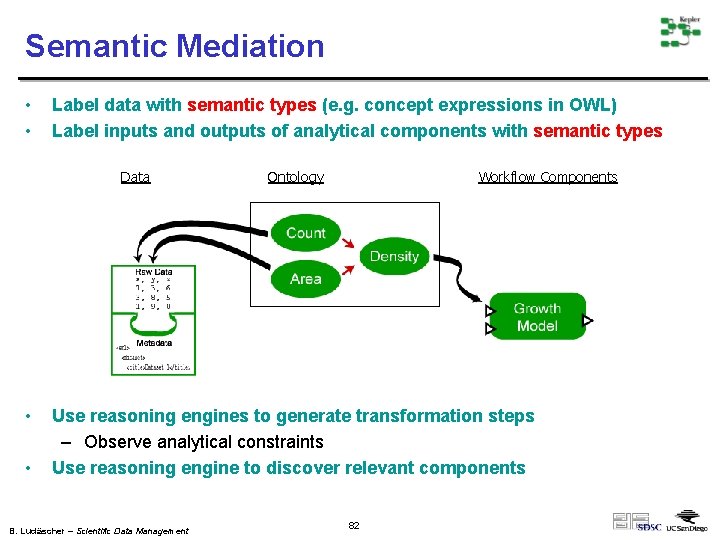
Semantic Mediation • • Label data with semantic types (e. g. concept expressions in OWL) Label inputs and outputs of analytical components with semantic types Data • • Ontology Workflow Components Use reasoning engines to generate transformation steps – Observe analytical constraints Use reasoning engine to discover relevant components B. Ludäscher – Scientific Data Management 82
- Slides: 80