Mass Spectrometry Dr S R Mane M Sc


























- Slides: 75

Mass Spectrometry Dr. S. R. Mane M. Sc. B. Ed. Ph. D. Associate Prof. & Head, Dept. of Chemistry, Smt. K. R. P. Kanya Mahavidyalaya, Islampur

Chemical Identification of Compound Comparison of Physical Properties Boiling Point Melting Point Elemental Analysis Burn the compound and measure the amounts of Density CO 2, H 2 O and other Optical rotation components that are Appearance produced to determine the Odor empirical formula Molecular formula

Introduction Modern techniques for structure determination of organic compounds include: o Mass spectrometry v Size and formula of the compound o Infrared spectroscopy v Functional groups present in the compound o Ultraviolet spectroscopy v Conjugated p electron system present in the compound o Nuclear magnetic resonance spectroscopy v Carbon-hydrogen framework of the compound

Introduction v MS is different than optical spectroscopy(UV-Vis, IR and NMR) as it does not measures the interaction of molecule with light or electromagnetic radiation v But since the record obtained resembles very much like optical lines in the spectrum so the name mass spectroscopy is given

Theorotical Background q In Mass spectrometer the neutral organic molecules(M) in vapor phase are bombarded with high energy electrons. q Due to this, energy of molecule increases and this excitation energy spread through out the molecular structure. q This energy is sufficient to knock out an electron of lowest ionization potential from the molecule and the molecular ion (M • +) is formed

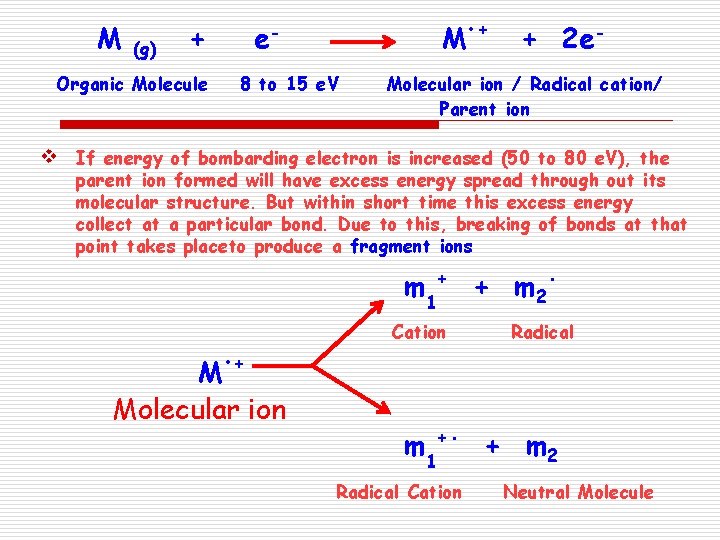
M (g) + Organic Molecule v e- M 8 to 15 e. V • + + 2 e- Molecular ion / Radical cation/ Parent ion If energy of bombarding electron is increased (50 to 80 e. V), the parent ion formed will have excess energy spread through out its molecular structure. But within short time this excess energy collect at a particular bond. Due to this, breaking of bonds at that point takes placeto produce a fragment ions m 1 + Cation M • + Molecular ion m 1 +. Radical Cation + m 2 . Radical + m 2 Neutral Molecule

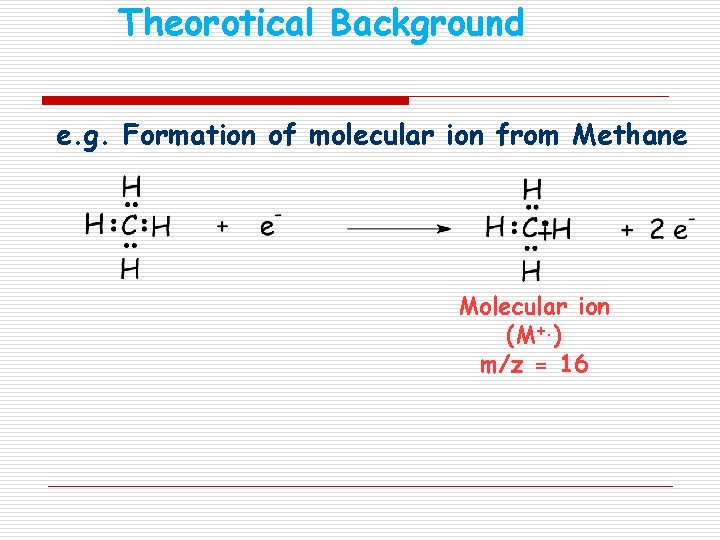
Theorotical Background e. g. Formation of molecular ion from Methane Molecular ion (M+. ) m/z = 16

Formation of Molecular ion from Ethane m/z = 15

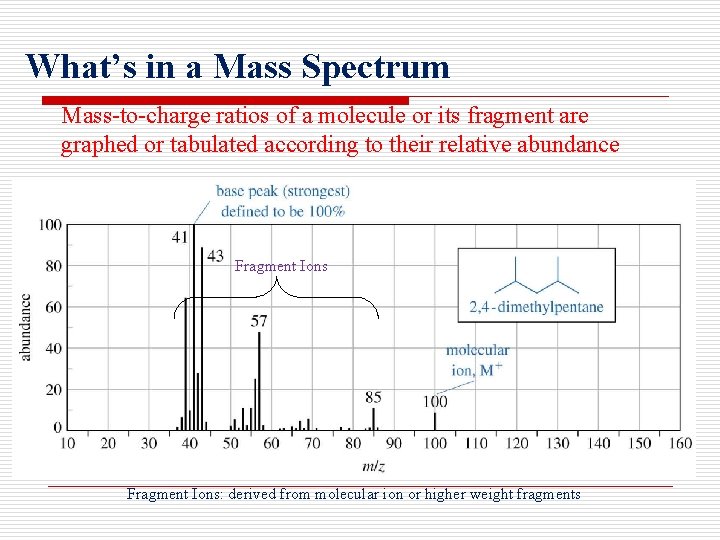
What’s in a Mass Spectrum Mass-to-charge ratios of a molecule or its fragment are graphed or tabulated according to their relative abundance Fragment Ions: derived from molecular ion or higher weight fragments

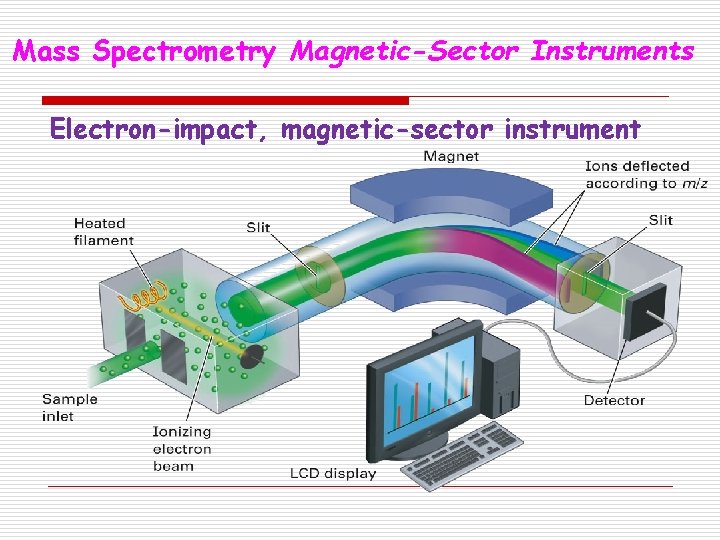
Mass Spectrometry Magnetic-Sector Instruments Electron-impact, magnetic-sector instrument

Background o Molecular ion (parent ion): n The radical cation corresponding to the mass of the original molecule o The molecular ion is usually the highest mass in the spectrum n Some exceptions w/specific isotopes n Some molecular ion peaks are absent.

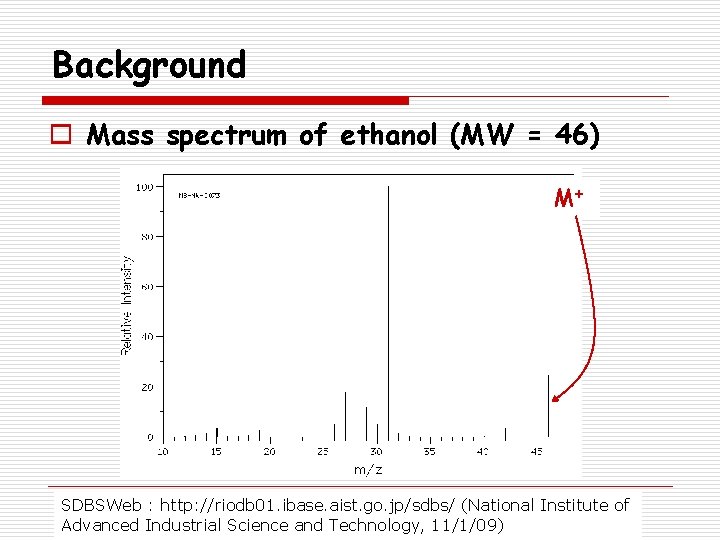
Background o Mass spectrum of ethanol (MW = 46) M+ SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/1/09)

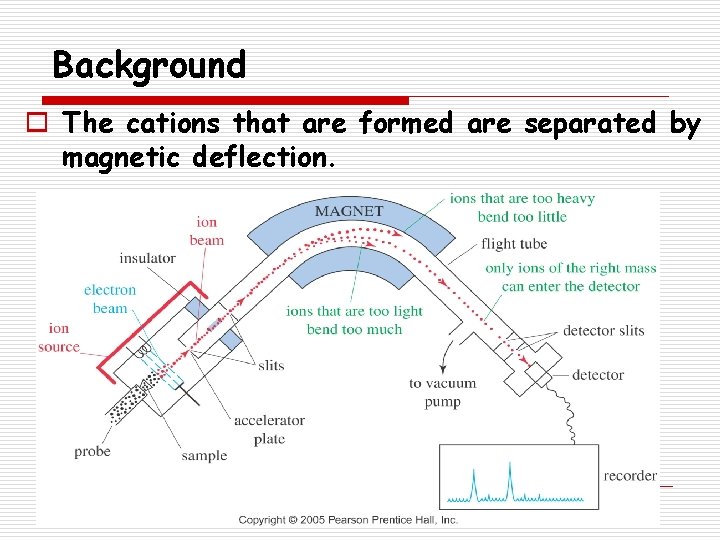
Background o The cations that are formed are separated by magnetic deflection.

Background o Only cations are detected. n Radicals are “invisible” in MS. o The amount of deflection observed depends on the mass to charge ratio (m/z). n Most cations formed have a charge of +1 so the amount of deflection observed is usually dependent on the mass of the ion.

Background o The resulting mass spectrum is a graph of the mass of each cation vs. its relative abundance. o The peaks are assigned an abundance as a percentage of the base peak. n the most intense peak in the spectrum o The base peak is not necessarily the same as the parent ion peak.

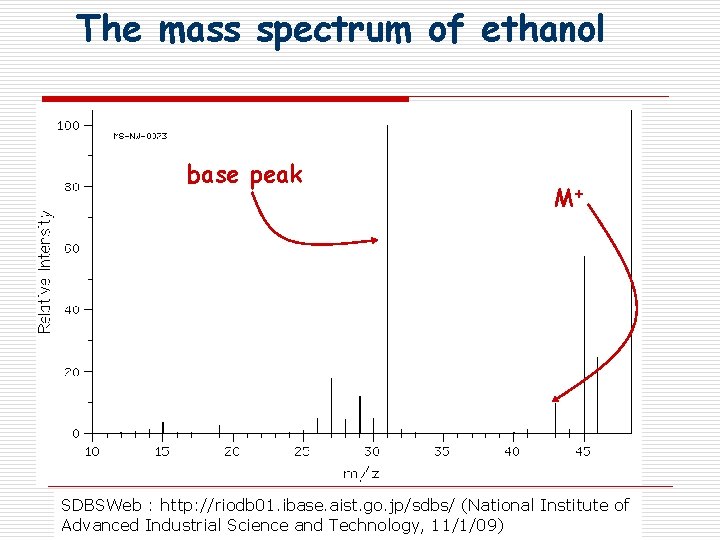
The mass spectrum of ethanol base peak M+ SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/1/09)

Background o Most elements occur naturally as a mixture of isotopes. n The presence of significant amounts of heavier isotopes leads to small peaks that have masses that are higher than the parent ion peak. o M+1 = a peak that is one mass unit higher than M+ o M+2 = a peak that is two mass units higher than M+

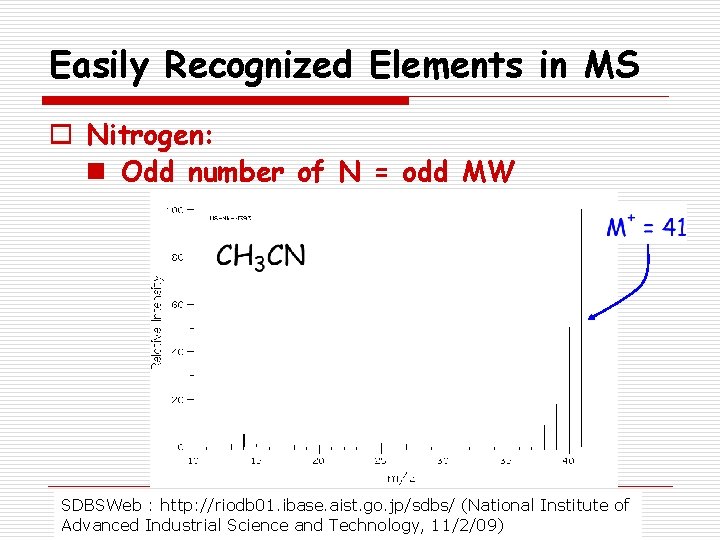
Easily Recognized Elements in MS o Nitrogen: n Odd number of N = odd MW SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/2/09)

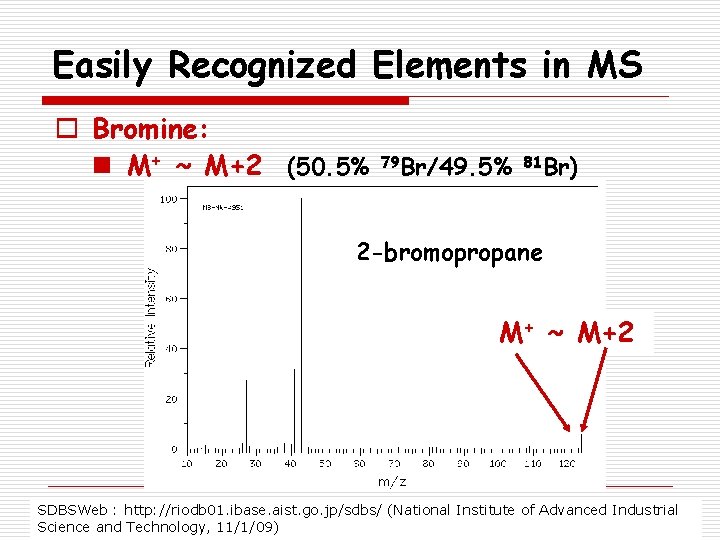
Easily Recognized Elements in MS o Bromine: n M+ ~ M+2 (50. 5% 79 Br/49. 5% 81 Br) 2 -bromopropane M+ ~ M+2 SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/1/09)

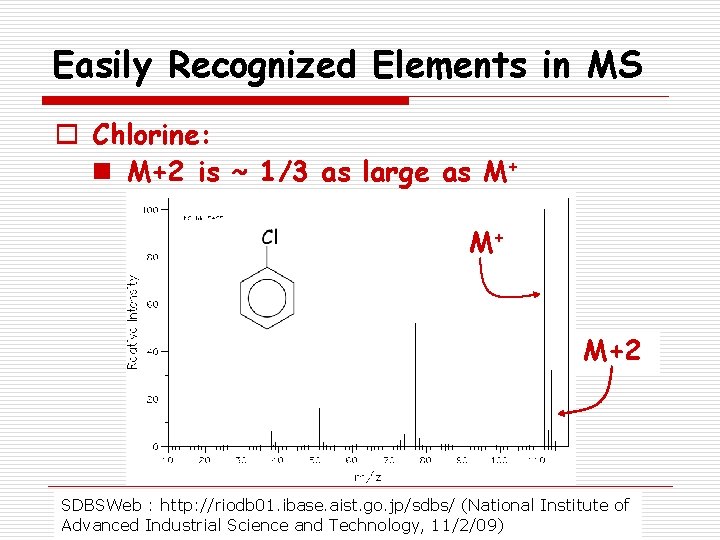
Easily Recognized Elements in MS o Chlorine: n M+2 is ~ 1/3 as large as M+ M+ M+2 SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/2/09)

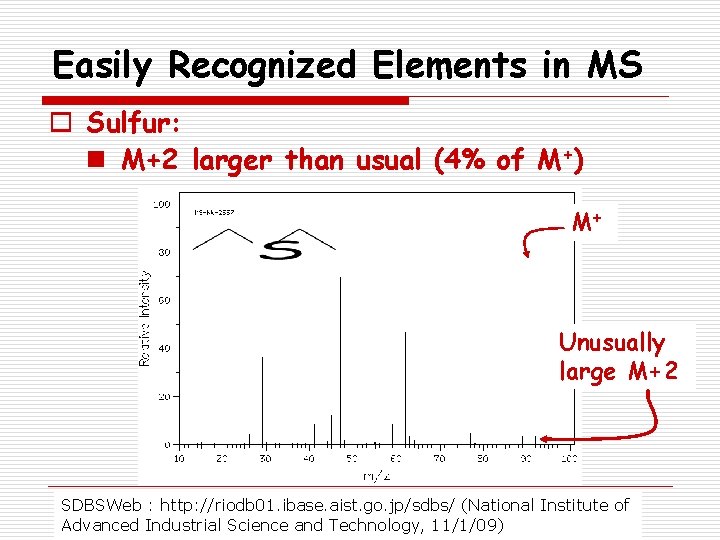
Easily Recognized Elements in MS o Sulfur: n M+2 larger than usual (4% of M+) M+ Unusually large M+2 SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/1/09)

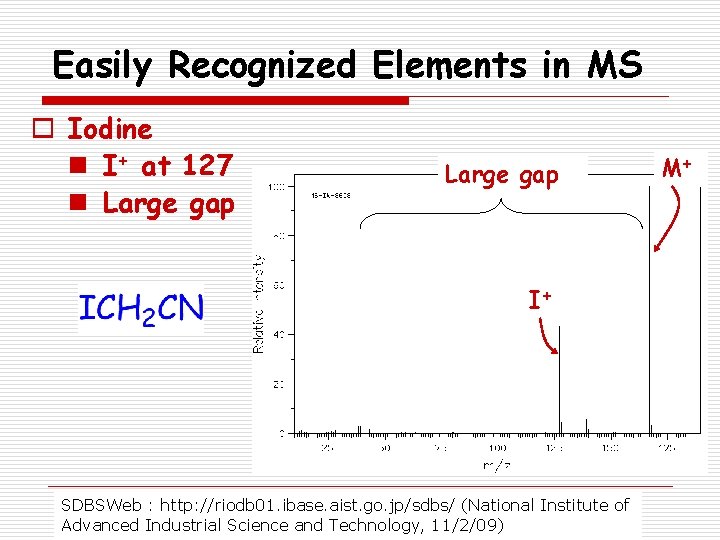
Easily Recognized Elements in MS o Iodine n I+ at 127 n Large gap I+ SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/2/09) M+

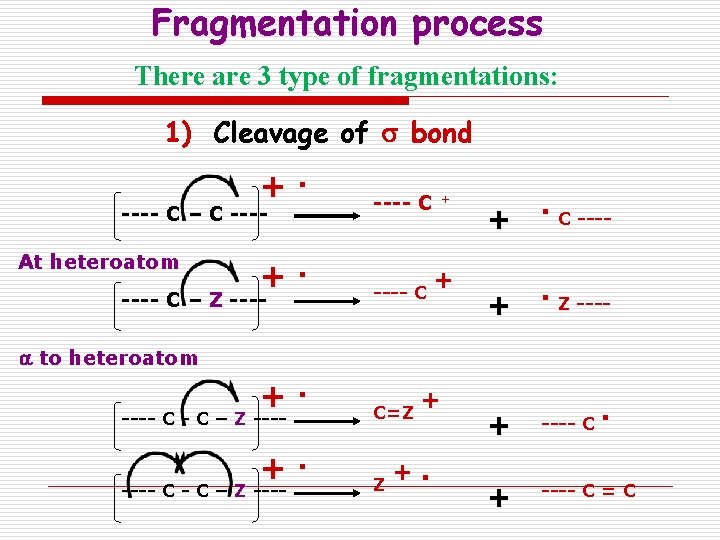
Fragmentation process There are 3 type of fragmentations: 1) Cleavage of s bond +. ---- C – C ---At heteroatom +. ---- C – Z ---- C + + + . C ---- + . Z ---- + ---- C = C a to heteroatom +. ---- C - C – Z ---- C=Z +. ---- C – Z ---- C - C – Z ---- + +. .

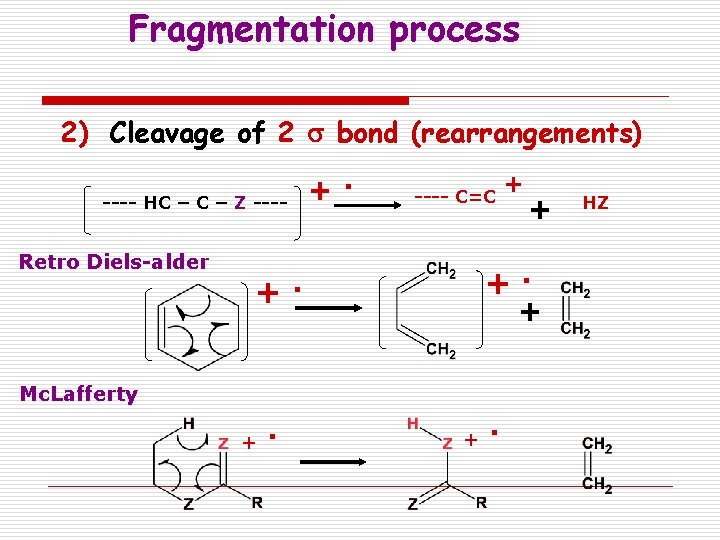
Fragmentation process 2) Cleavage of 2 s bond (rearrangements) +. ---- HC – Z ---- Retro Diels-alder + ---- C=C + + +. . + Mc. Lafferty . + + . HZ

Fragmentation process There are 3 type of fragmentations: 3) Cleavage of Complex rearrangements

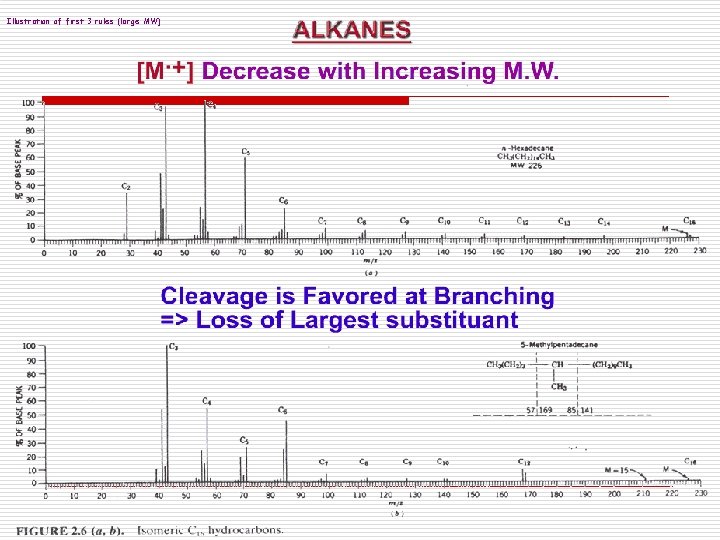
Fragmentation rules in MS 1. Intensity of M. + is Larger for linear chain than for branched compound 2. Intensity of M. + decrease with Increasing M. W. (fatty acid is an exception) 3. Cleavage is favored at branching reflecting the Increased stability of the. R R CH+ < R C+ ion R + + R Stability order: CH 3 < R-CH 2 < R’ R CH R” Loss of Largest Subst. Is most favored

Illustration of first 3 rules (large MW)

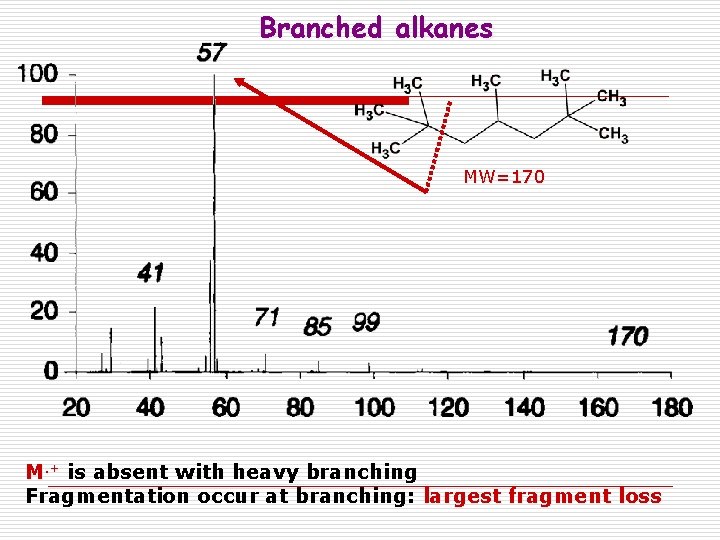
Branched alkanes MW=170 M. + is absent with heavy branching Fragmentation occur at branching: largest fragment loss

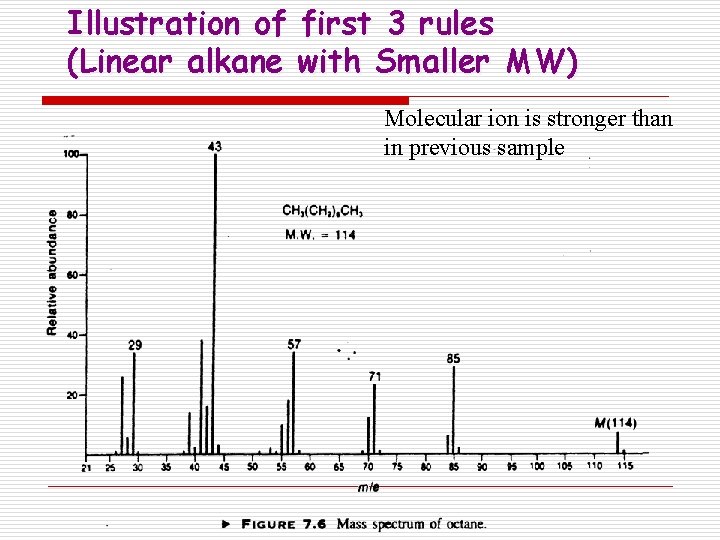
Illustration of first 3 rules (Linear alkane with Smaller MW) Molecular ion is stronger than in previous sample

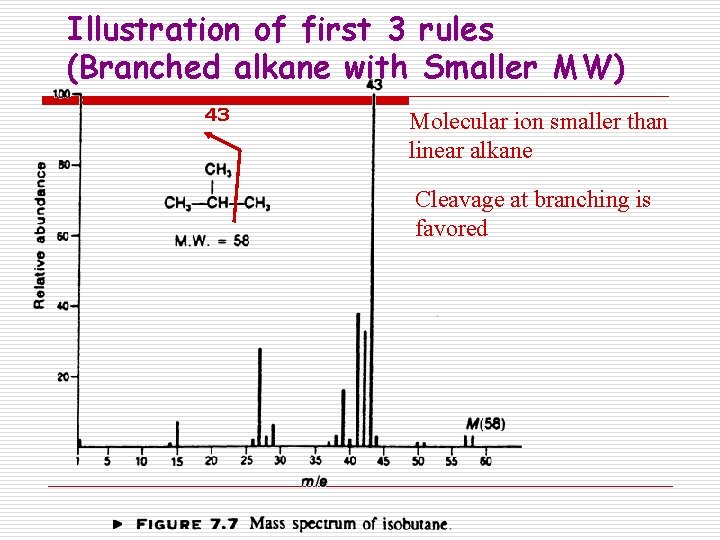
Illustration of first 3 rules (Branched alkane with Smaller MW) 43 Molecular ion smaller than linear alkane Cleavage at branching is favored

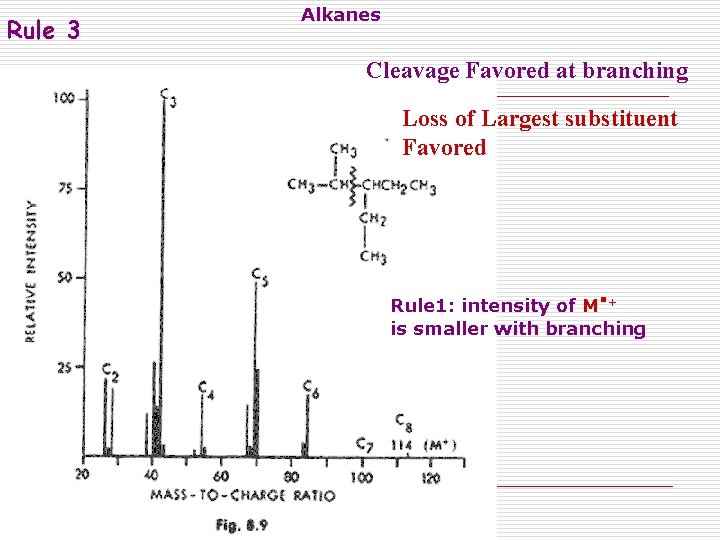
Rule 3 Alkanes Cleavage Favored at branching Loss of Largest substituent Favored . Rule 1: intensity of M + is smaller with branching

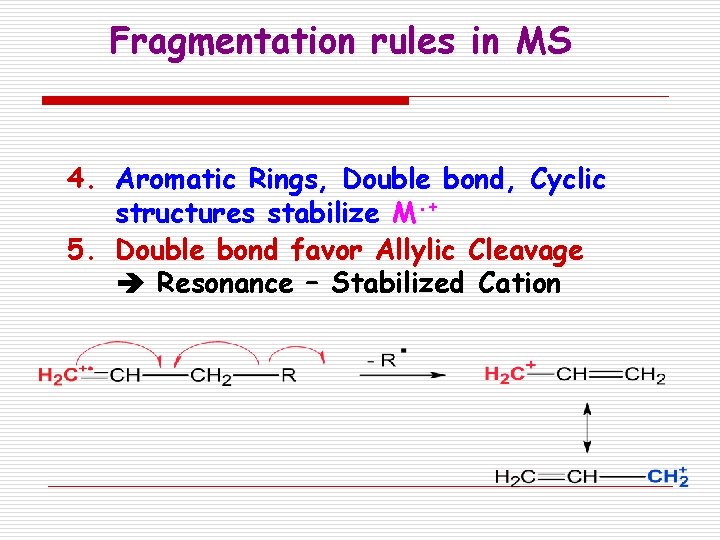
Fragmentation rules in MS 4. Aromatic Rings, Double bond, Cyclic structures stabilize M. + 5. Double bond favor Allylic Cleavage Resonance – Stabilized Cation

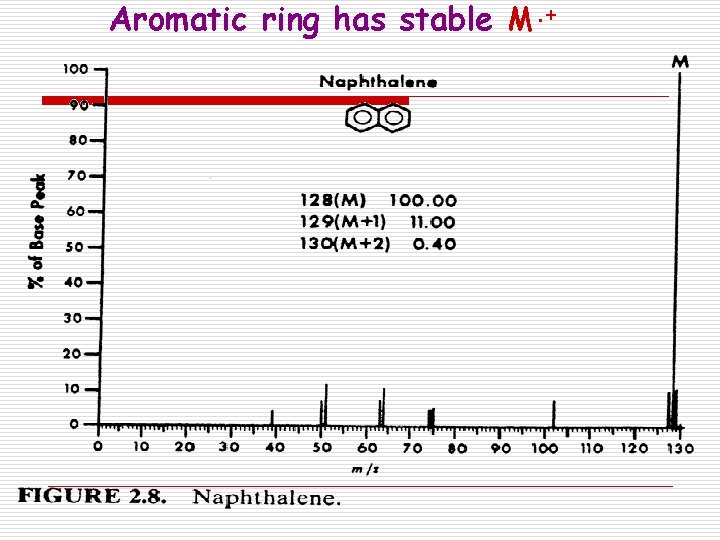
Aromatic ring has stable M. +

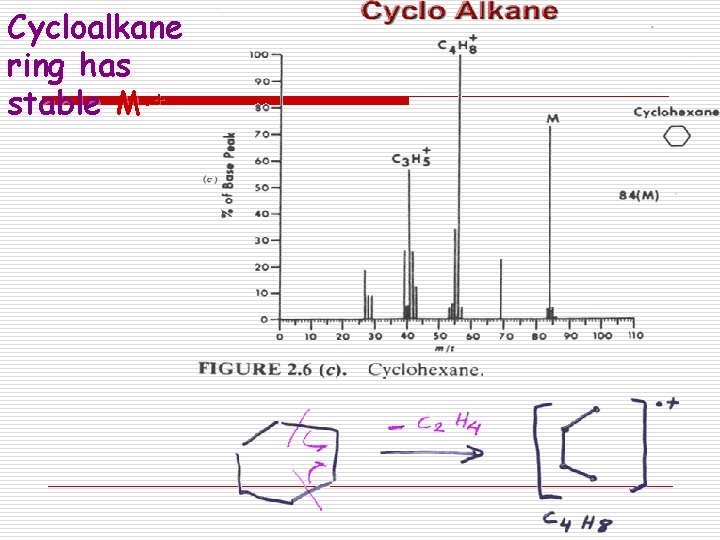
Cycloalkane ring has stable M. +

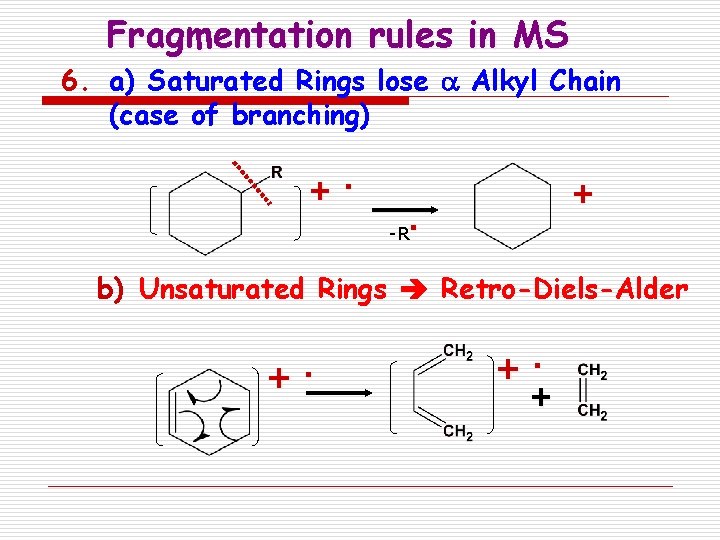
Fragmentation rules in MS 6. a) Saturated Rings lose a Alkyl Chain (case of branching) +. -R + . b) Unsaturated Rings Retro-Diels-Alder + . +

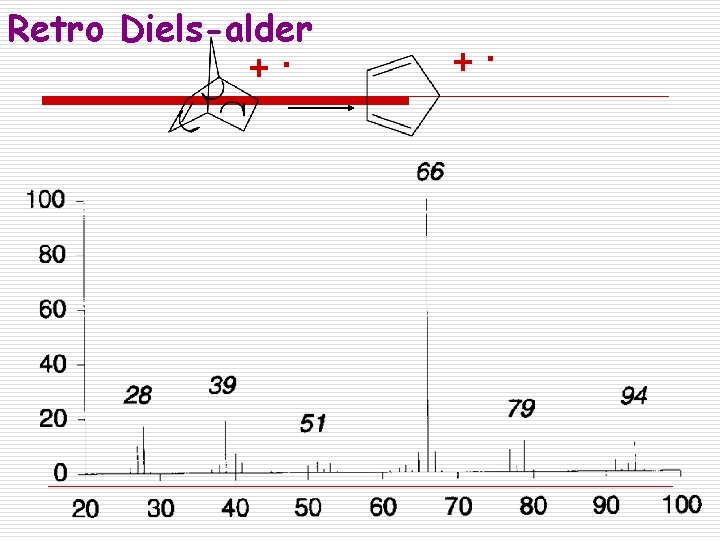
Retro Diels-alder +.

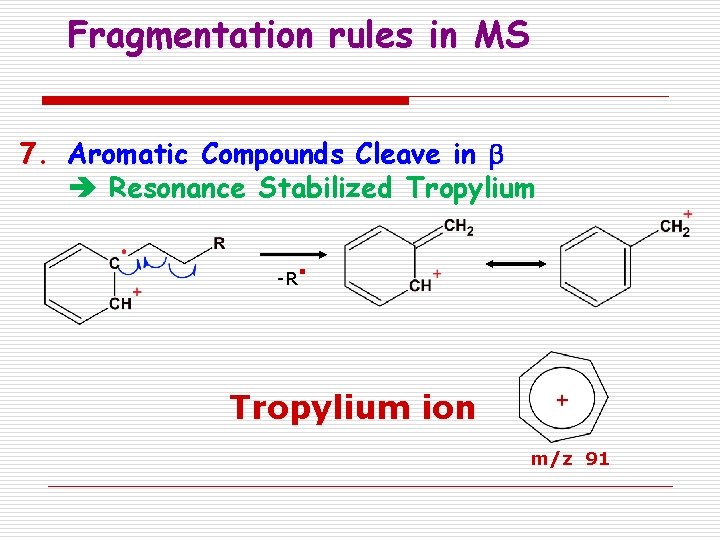
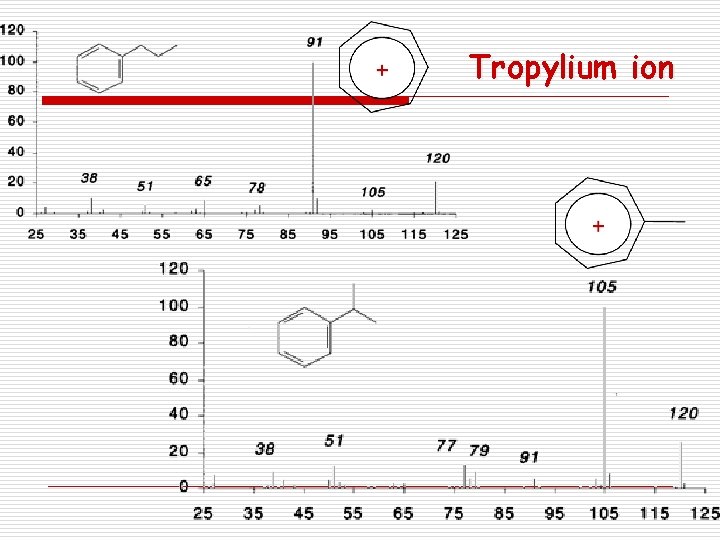
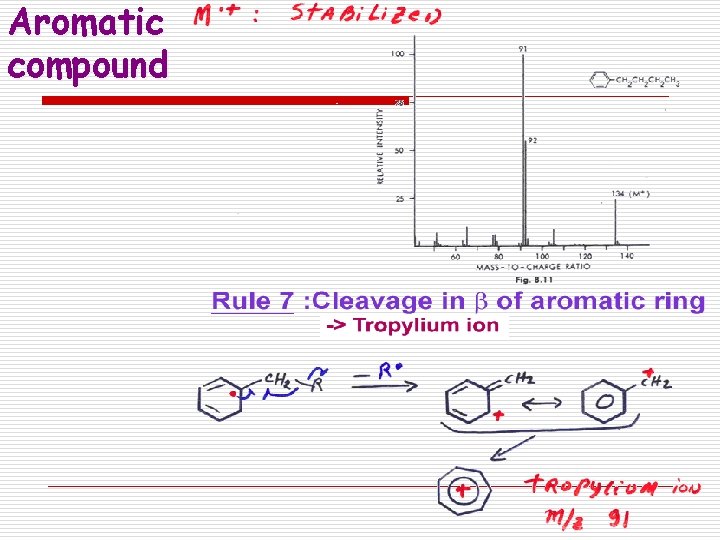
Fragmentation rules in MS 7. Aromatic Compounds Cleave in b Resonance Stabilized Tropylium -R . Tropylium ion m/z 91

Tropylium ion

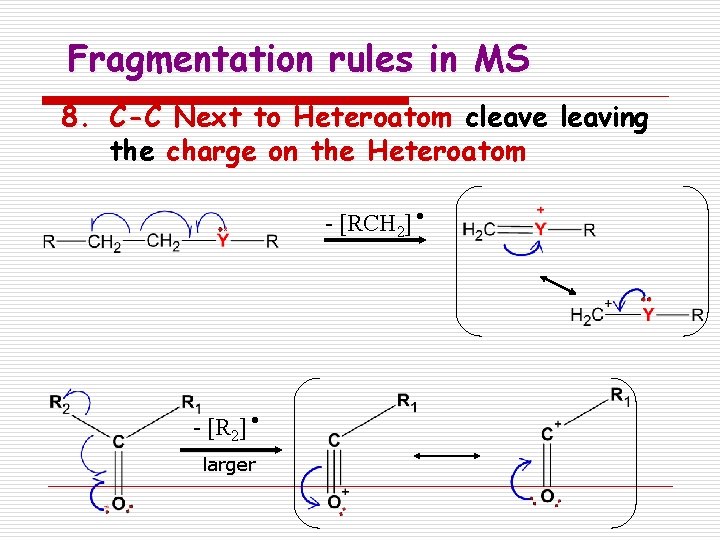
Fragmentation rules in MS 8. C-C Next to Heteroatom cleave leaving the charge on the Heteroatom - [RCH 2] - [R 2] larger

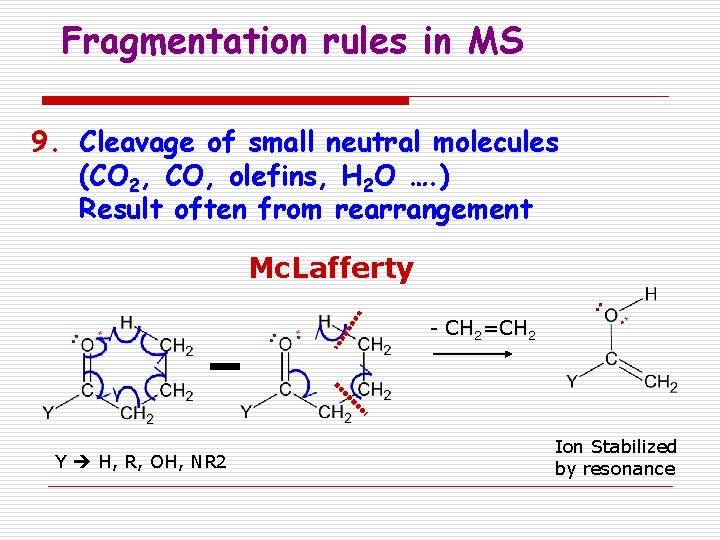
Fragmentation rules in MS 9. Cleavage of small neutral molecules (CO 2, CO, olefins, H 2 O …. ) Result often from rearrangement Mc. Lafferty - CH 2=CH 2 Y H, R, OH, NR 2 Ion Stabilized by resonance

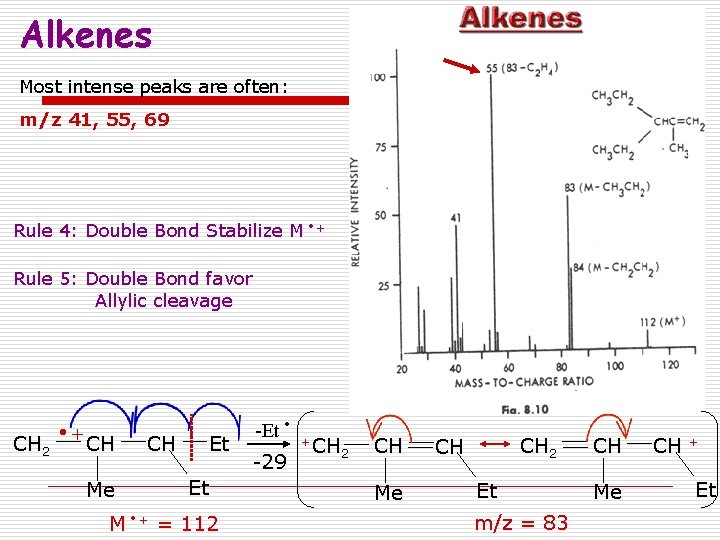
Alkenes Most intense peaks are often: m/z 41, 55, 69 Rule 4: Double Bond Stabilize M + Rule 5: Double Bond favor Allylic cleavage CH 2 + CH Me CH Et Et M + = 112 -Et -29 +CH 2 CH Me CH 2 CH Et m/z = 83 CH Me CH + Et

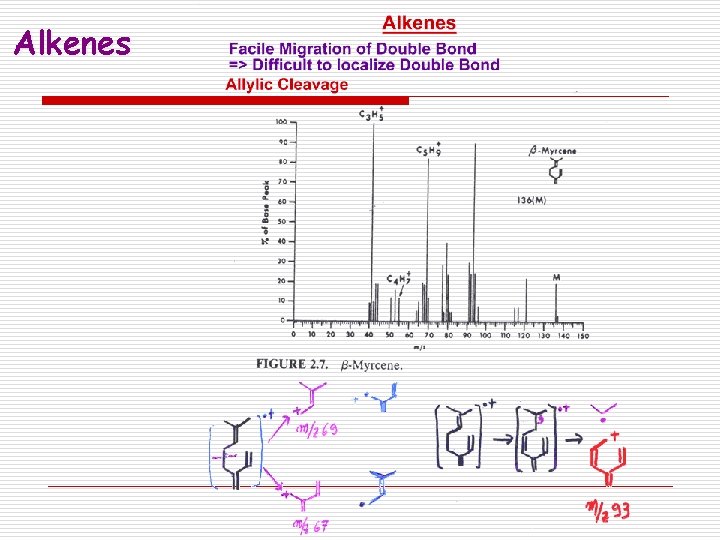
Alkenes

Aromatic compound

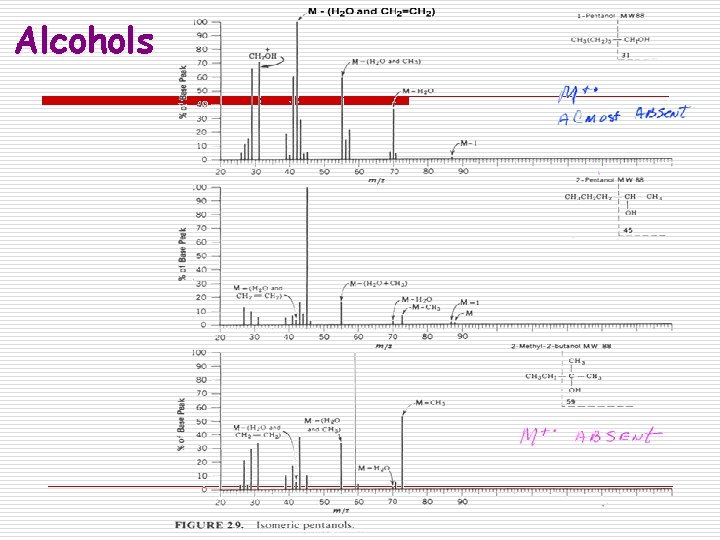
Alcohols

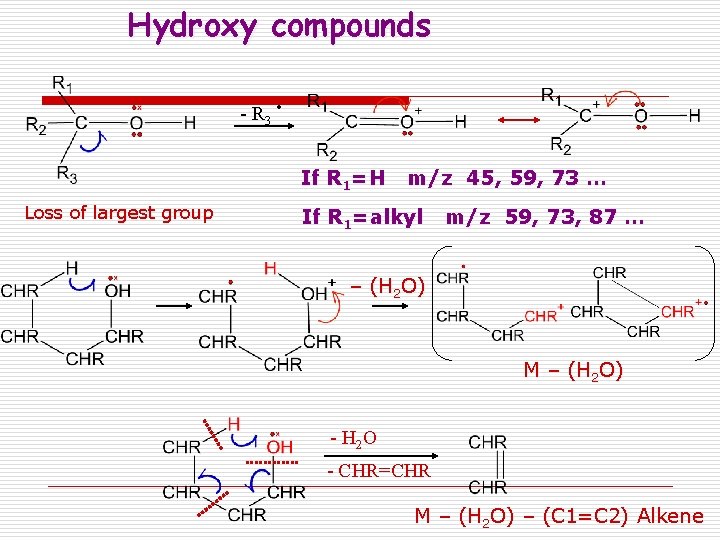
Hydroxy compounds - R 3 If R 1=H m/z 45, 59, 73 … Loss of largest group If R 1=alkyl m/z 59, 73, 87 … – (H 2 O) M – (H 2 O) - H 2 O - CHR=CHR M – (H 2 O) – (C 1=C 2) Alkene

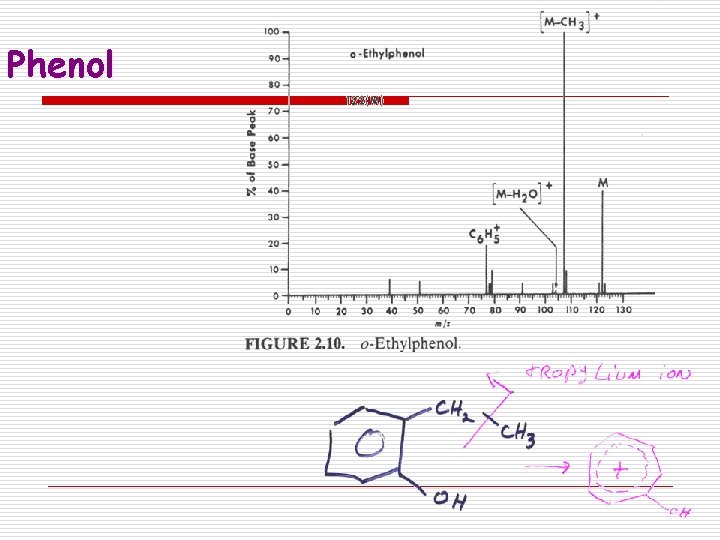
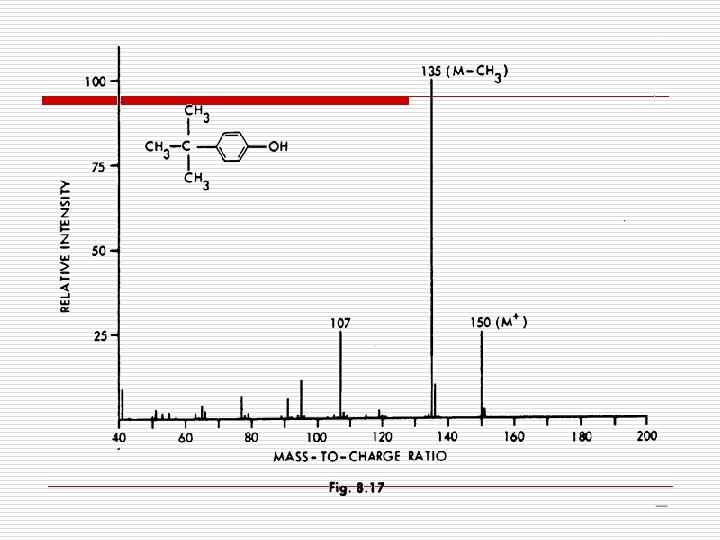
Phenol


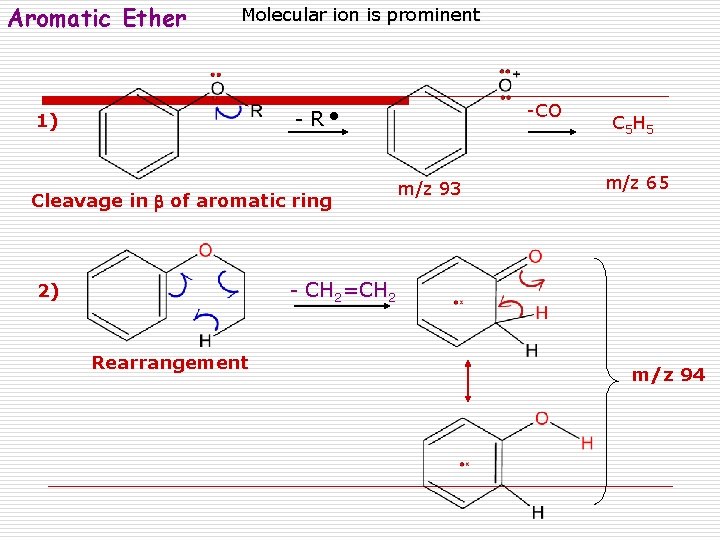
Aromatic Ether Molecular ion is prominent -CO - R 1) Cleavage in b of aromatic ring m/z 93 C 5 H 5 m/z 65 - CH 2=CH 2 2) Rearrangement m/z 94

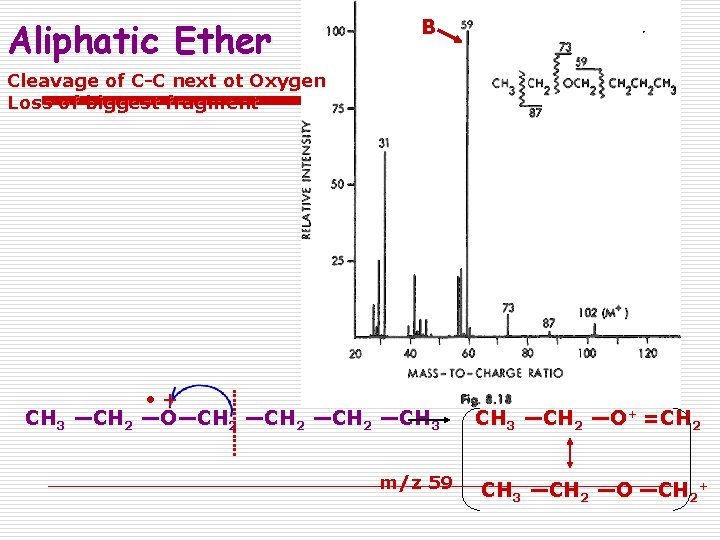
Aliphatic Ether B Cleavage of C-C next ot Oxygen Loss of biggest fragment • + CH 3 —CH 2 —O—CH 2 —CH 3 m/z 59 CH 3 —CH 2 —O+ =CH 2 CH 3 —CH 2 —O —CH 2+

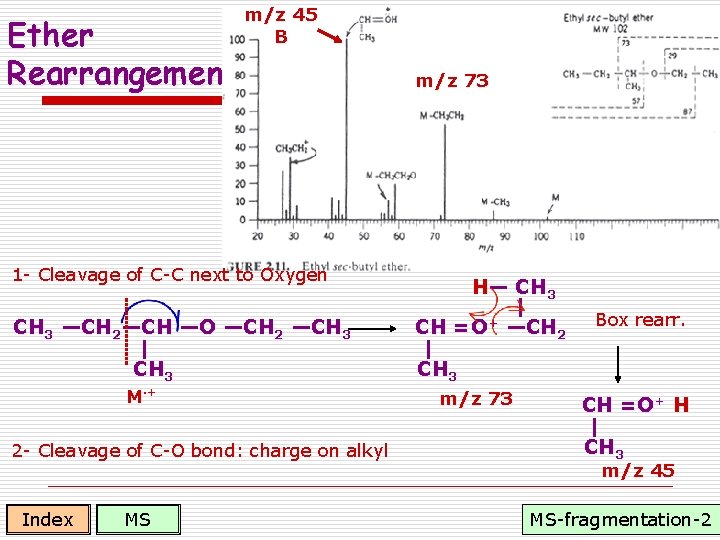
Ether Rearrangement m/z 45 B m/z 73 1 - Cleavage of C-C next to Oxygen CH 3 —CH 2—CH —O —CH 2 —CH 3 M·+ 2 - Cleavage of C-O bond: charge on alkyl Index MS H— CH 3 CH =O+ —CH 2 Box rearr. CH 3 m/z 73 CH =O+ H CH 3 m/z 45 MS-fragmentation-2

Fragmentation Patterns o The impact of the stream of high energy electrons often breaks the molecule into fragments, commonly a cation and a radical. n Bonds break to give the most stable cation. n Stability of the radical is less important.

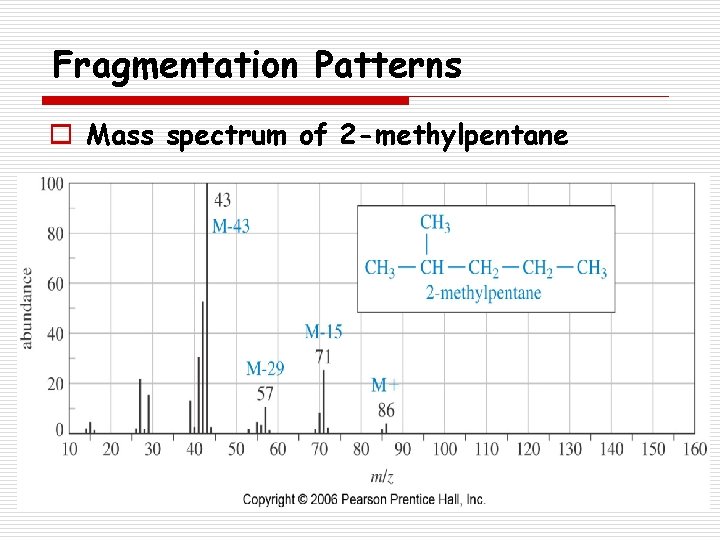
Fragmentation Patterns o Alkanes n Fragmentation often alkyl groups: o Loss of methyl o Loss of propyl o Loss of butyl splits off simple M+ M+ - 15 29 43 57 n Branched alkanes tend to fragment forming the most stable carbocations.

Fragmentation Patterns o Mass spectrum of 2 -methylpentane

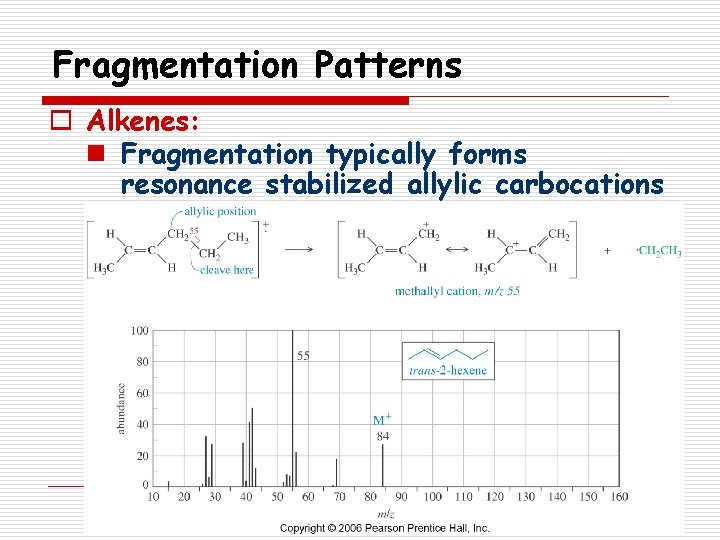
Fragmentation Patterns o Alkenes: n Fragmentation typically forms resonance stabilized allylic carbocations

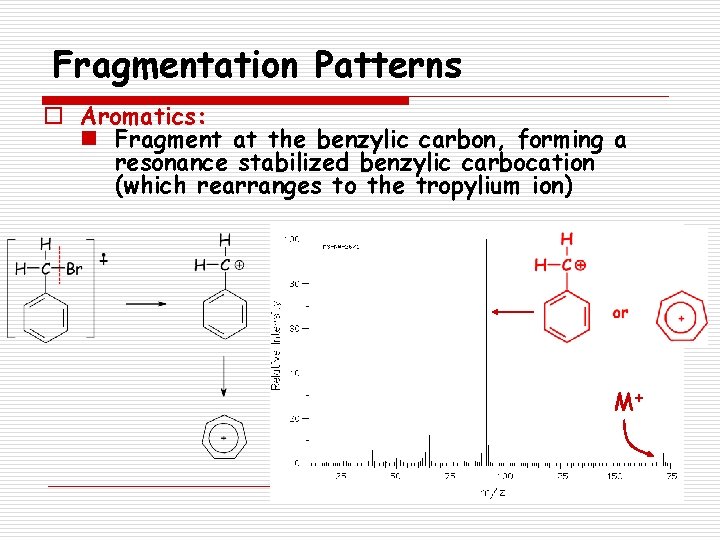
Fragmentation Patterns o Aromatics: n Fragment at the benzylic carbon, forming a resonance stabilized benzylic carbocation (which rearranges to the tropylium ion) M+

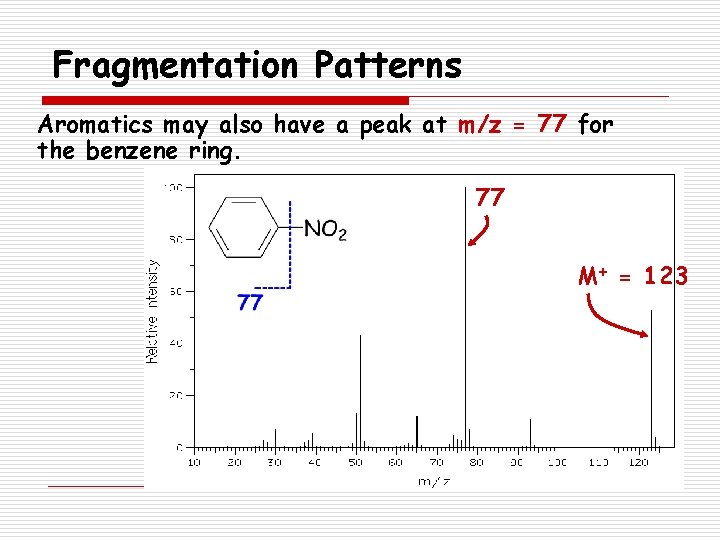
Fragmentation Patterns Aromatics may also have a peak at m/z = 77 for the benzene ring. 77 M+ = 123

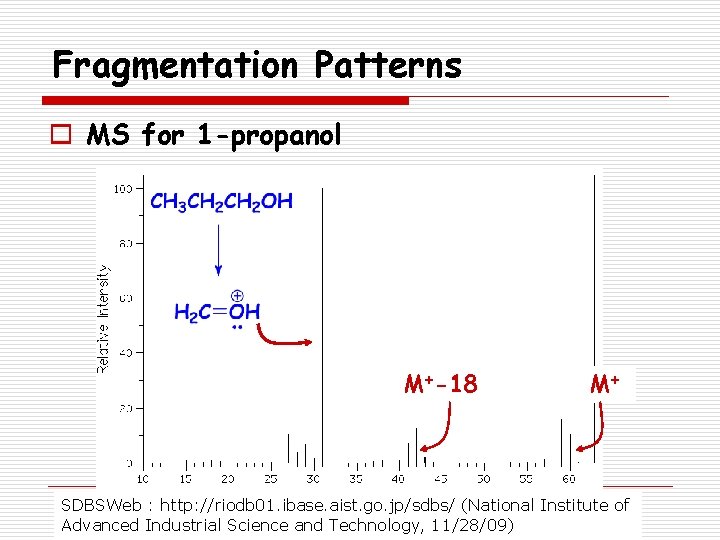
Fragmentation Patterns o Alcohols n Fragment easily resulting in very small or missing parent ion peak n May lose hydroxyl radical or water o M+ - 17 or M+ - 18 n Commonly lose an alkyl group attached to the carbinol carbon forming an oxonium ion. o 1 o alcohol usually has prominent peak at m/z = 31 corresponding to H 2 C=OH+

Fragmentation Patterns o MS for 1 -propanol M+-18 M+ SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/28/09)

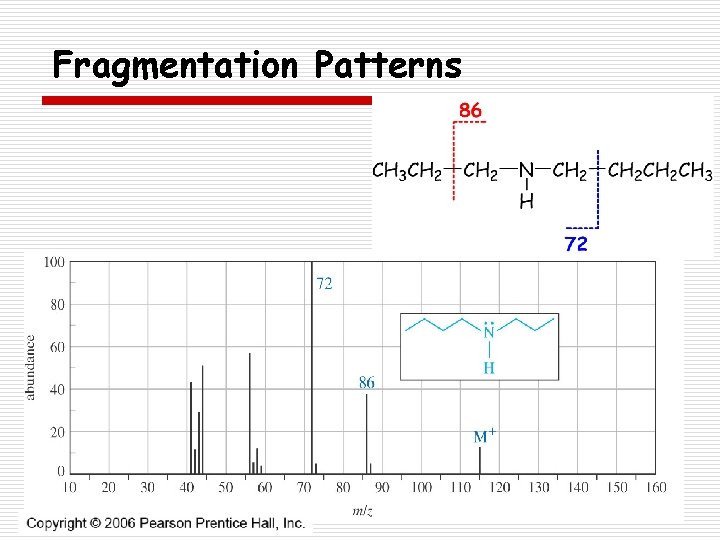
Fragmentation Patterns o Amines n Odd M+ (assuming an odd number of nitrogens are present) n a-cleavage dominates forming an iminium ion

Fragmentation Patterns

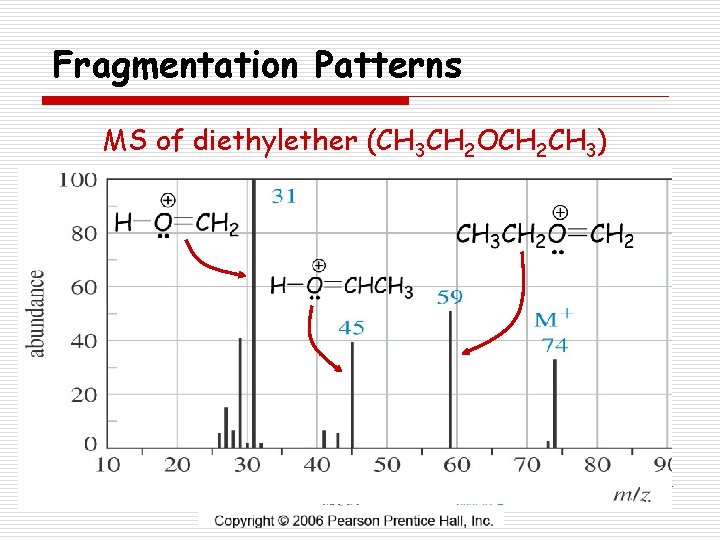
Fragmentation Patterns o Ethers n a-cleavage forming oxonium ion n Loss of alkyl group forming a carbocation

Fragmentation Patterns MS of diethylether (CH 3 CH 2 OCH 2 CH 3)

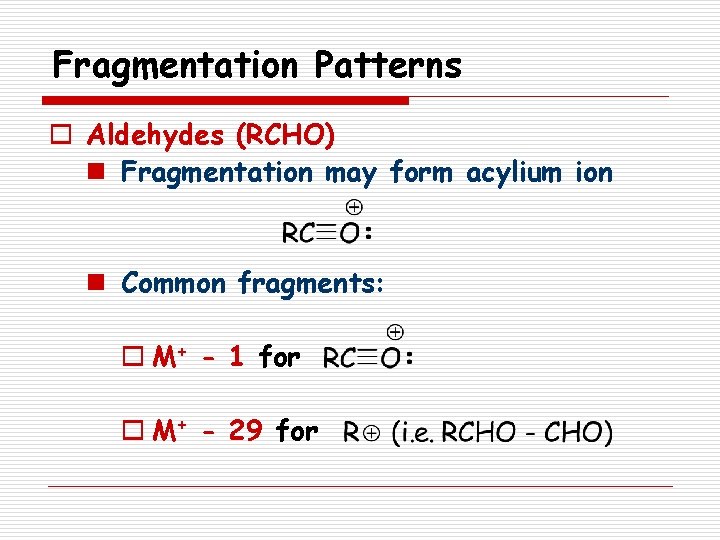
Fragmentation Patterns o Aldehydes (RCHO) n Fragmentation may form acylium ion n Common fragments: o M+ - 1 for o M+ - 29 for

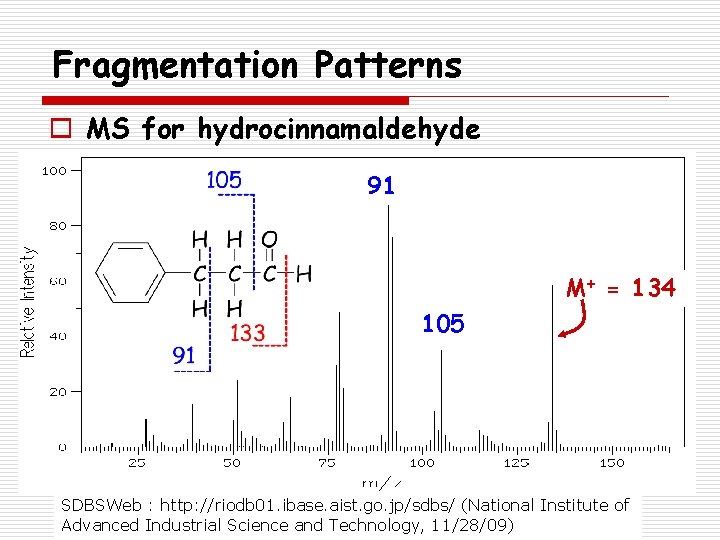
Fragmentation Patterns o MS for hydrocinnamaldehyde 91 M+ = 134 105 SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/28/09)

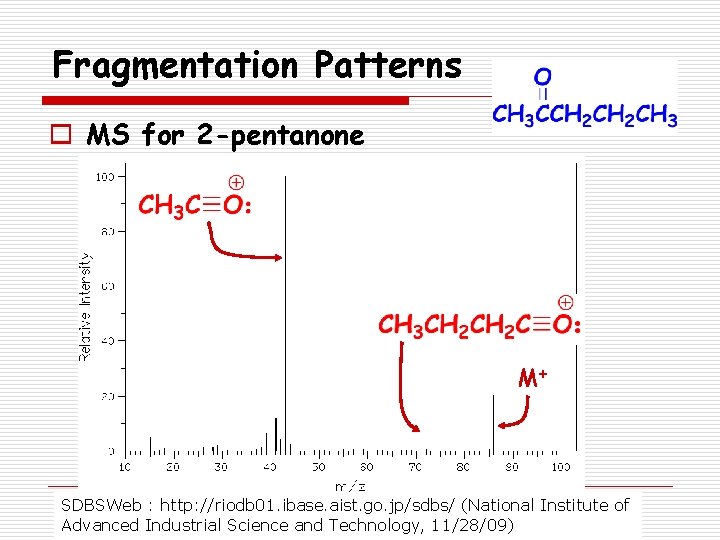
Fragmentation Patterns o Ketones n Fragmentation leads to formation of acylium ion: o Loss of R forming o Loss of R’ forming

Fragmentation Patterns o MS for 2 -pentanone M+ SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/28/09)

Fragmentation Patterns o Esters (RCO 2 R’) n Common fragmentation patterns include: o Loss of OR’ n peak at M+ - OR’ o Loss of R’ n peak at M+ - R’

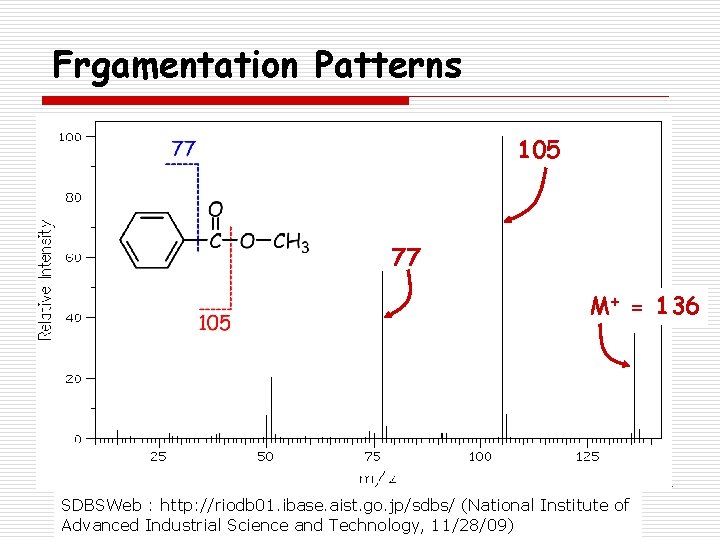
Frgamentation Patterns 105 77 M+ = 136 SDBSWeb : http: //riodb 01. ibase. aist. go. jp/sdbs/ (National Institute of Advanced Industrial Science and Technology, 11/28/09)

Rule of Thirteen o The “Rule of Thirteen” can be used to identify possible molecular formulas for an unknown hydrocarbon, Cn. Hm. n Step 1: n = M+/13 (integer only, use remainder in step 2) n Step 2: m = n + remainder from step 1

Rule of Thirteen o Example: The formula for a hydrocarbon with M+ =106 can be found: n Step 1: n = 106/13 = 8 (R = 2) n Step 2: m = 8 + 2 = 10 n Formula: C 8 H 10

Rule of Thirteen o If a heteroatom is present, n Subtract the mass of each heteroatom from the MW n Calculate the formula for the corresponding hydrocarbon n Add the heteroatoms to the formula

Rule of Thirteen Example: A compound with a molecular ion peak at m/z = 102 has a strong peak at 1739 cm-1 in its IR spectrum. Determine its molecular formula.

GC-Mass Spec: Experiment 23 o Mass Spec can be combined with gas chromatography to analyze mixtures of compounds. n GC separates the components of the mixture. n Each component is analyzed by the Mass Spectrometer.

GC-Mass Spec: Experiment 23 o Assignment: n Observe the GC-mass spec experiment o Record experimental conditions n Analyze the mass spectrum of each component of your mixture: o Parent ion peak? o Heteroatoms apparent from spectrum? o A minimum of 1 or two significant fragments and their structures

GC-Mass Spec: Experiment 23 o Assignment (cont. ): n Using the Mass Spec data, retention times, and boiling points, identify the components of your mixture. n Write three paragraphs (one per compound) summarizing and interpreting all data. See your data sheet for more details.