Using BLAST to Recherche dans des bases de

Using BLAST to Recherche dans des bases de Search Sequence données de séquences biologiques Databases Cédric Notredame

Outline -Evolution and Sequence Similarity -The inside of BLAST -Using BLAST -Adapting BLAST to your needs -Searching Protein Domains with BLAST -Digging Genomes

Two Minutes of the Evolutionnary Clock…
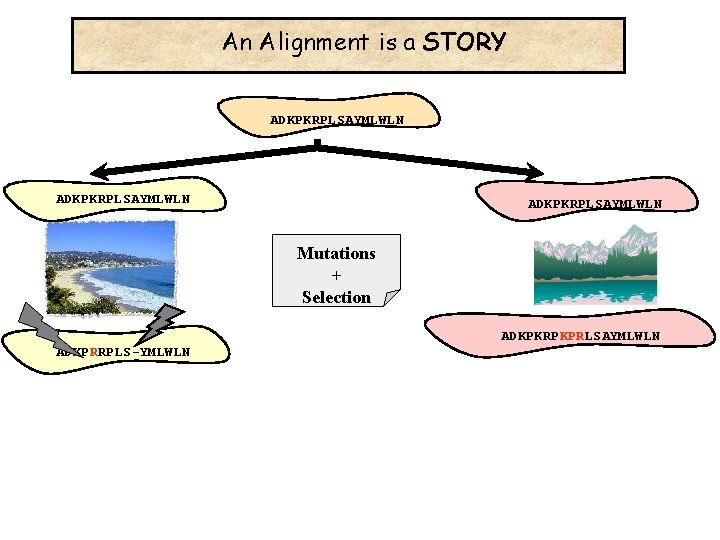
An Alignment is a STORY ADKPKRPLSAYMLWLN Mutations + Selection ADKPKRPKPRLSAYMLWLN ADKPRRPLS-YMLWLN
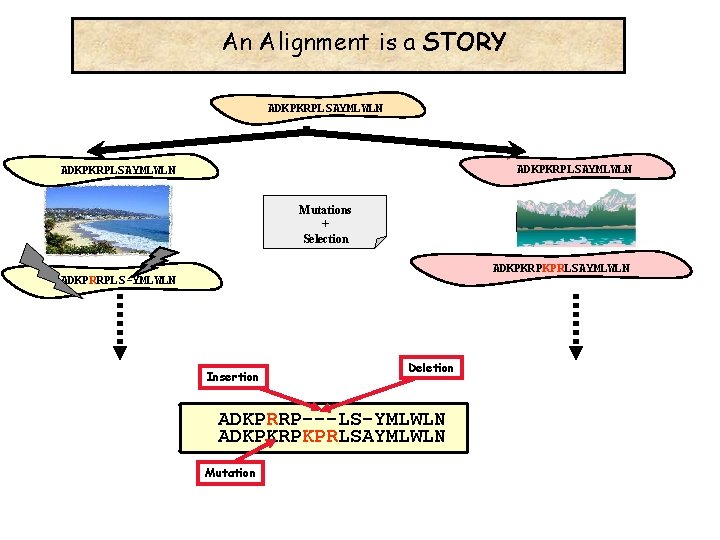
An Alignment is a STORY ADKPKRPLSAYMLWLN Mutations + Selection ADKPKRPKPRLSAYMLWLN ADKPRRPLS-YMLWLN Insertion Deletion ADKPRRP---LS-YMLWLN ADKPKRPKPRLSAYMLWLN Mutation
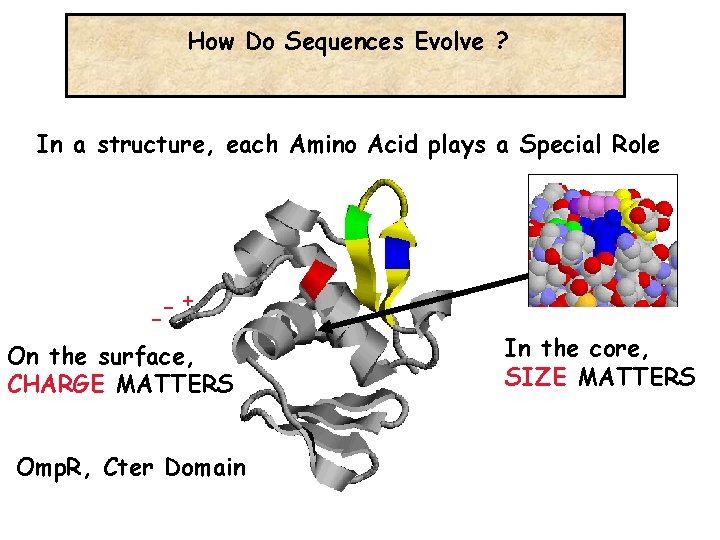
How Do Sequences Evolve ? In a structure, each Amino Acid plays a Special Role -+ On the surface, CHARGE MATTERS Omp. R, Cter Domain In the core, SIZE MATTERS
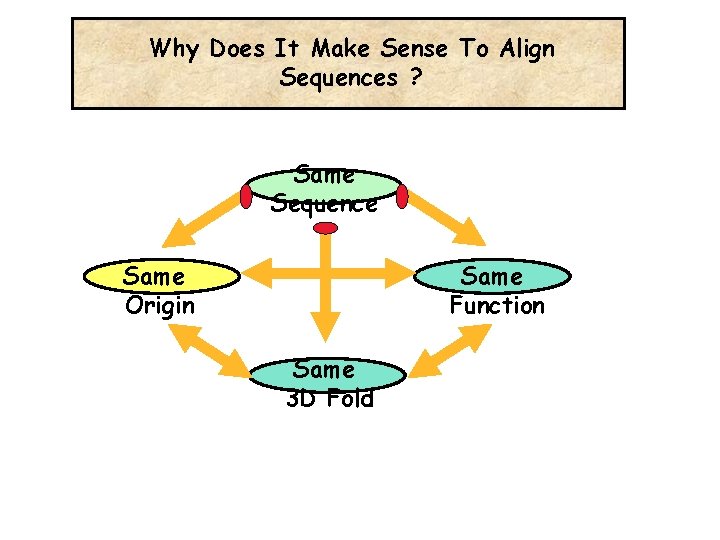
Why Does It Make Sense To Align Sequences ? Same Sequence Same Origin Same Function Same 3 D Fold
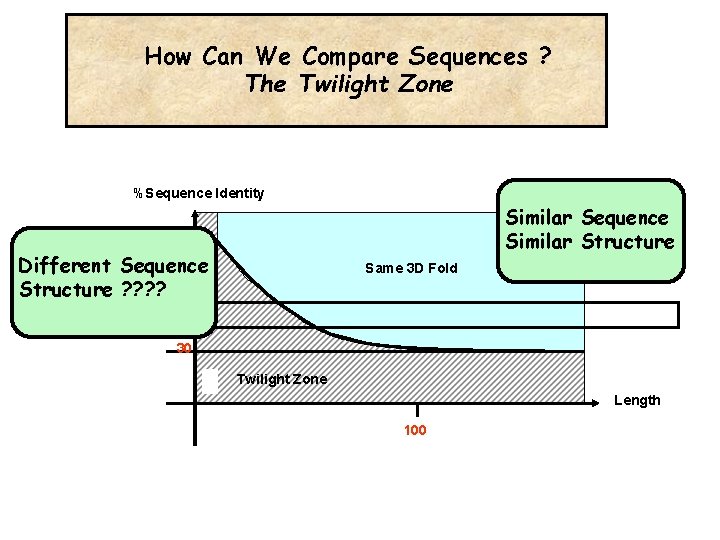
How Can We Compare Sequences ? The Twilight Zone %Sequence Identity Similar Sequence Similar Structure Different Sequence Structure ? ? Same 3 D Fold 30% 30 Twilight Zone Length 100
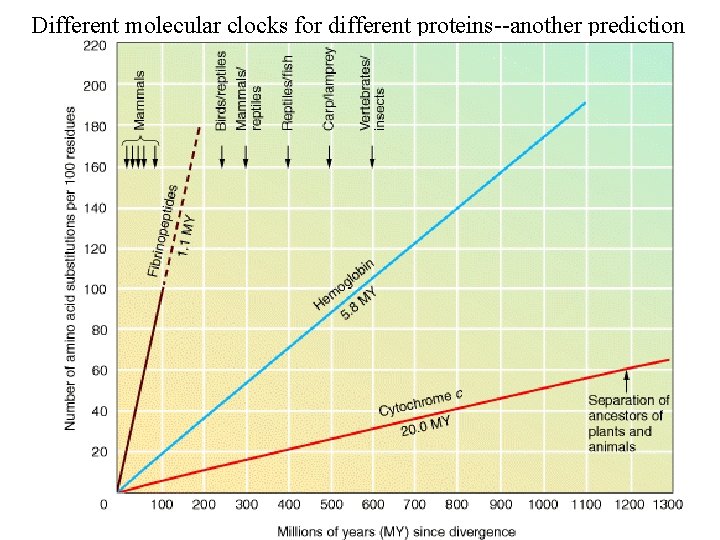
Different molecular clocks for different proteins--another prediction

A few Basic Definitions

A few Definitions Query : Your sequence Subject: The database against which you search Heuristic: Algorithm that does not guaranty the optimal solution

Other Important Definitions Identity Proportion of IDENTICAL residues between two sequences. Depends on the Alignment. Unit: the % id Similarity Proportion of SIMILAR residues Two residues are similar if their substitution cost is higher than 0. Depends on the matrix Unit: the %similarity Homology Sequences SIMILAR enough are sometimes HOMOLOGOUS HOMOLOGY COMMON ANCESTOR Unit: Yes or No! DIFFERENT sequences can also be Homologous

More Important Definitions Hit A sequence that matches your sequence and reported by BLAST. E-Value Expectation value How many times would you expect to find a hit by chance only? Depends on the alignment. Depends on the matrix Depends on the database Sensitive to Low complexity regions Unit: must be lower than 0. 0001 to mean something

A Good Hit Is Something You Would Not Expect by Chance

What is BLAST ?

BLAST Basic Local Alignment Search Tool BLAST is a Program Designed for RAPIDLY Comparing Your Sequence With every Sequence in a database and REPORT the most SIMILAR sequences

Database Search 1 -Query 2 -Comparison Engine LOCAL Alignment 3 -Database 4 -Statistical Evaluation (E-Value) PROBLEM: LOCAL ALIGNMENT (SW)TOO SLOW
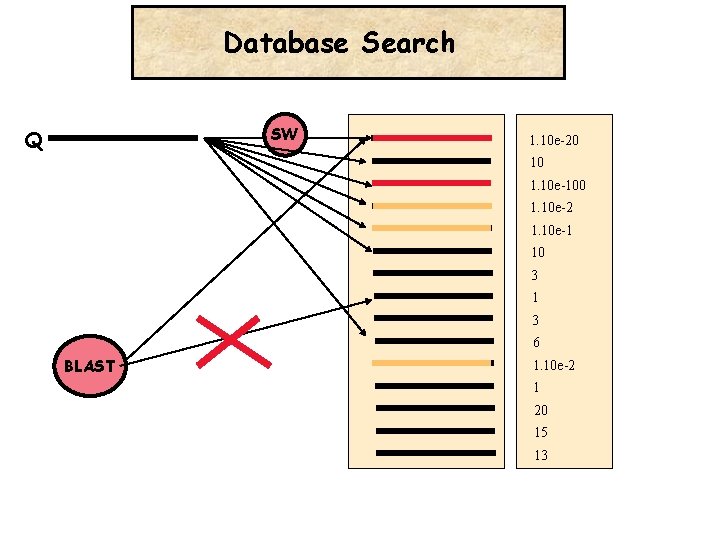
Database Search SW Q 1. 10 e-20 10 1. 10 e-100 1. 10 e-2 1. 10 e-1 10 3 1 3 6 BLAST 1. 10 e-2 1 20 15 13

BLAST Basic Local Alignment Search Tool BLAST is a Heuristic Smith and Waterman BLAST = 3 STEPS 1 -Decide who will be compared This is where Blast SAVES TIME This is where it LOSES HITS Most BLAST parameters refer to this step

BLAST Basic Local Alignment Search Tool BLAST is a Heuristic Smith and Waterman BLAST = 3 STEPS 1 -Decide who will be compared 2 -Check the most promising Hits 3 -Compute the E-value of the most interesting Hits

Heuristic. BLAST Algorithms A Bit of History Smith and Waterman • Exact Local Dynamic Programming, 1981 FASTA • Lipman and Pearson, 1985 • Looks for similar words (k-tup) on the same diagonal. • Comparison on the sequences one by one… BLAST • Altschul et al. , 1990 • The most widely cited tool in Biology • www. ncbi. nlm. nih. gov/Education/BLASTinfo/tut 1. html

The Inside of BLAST
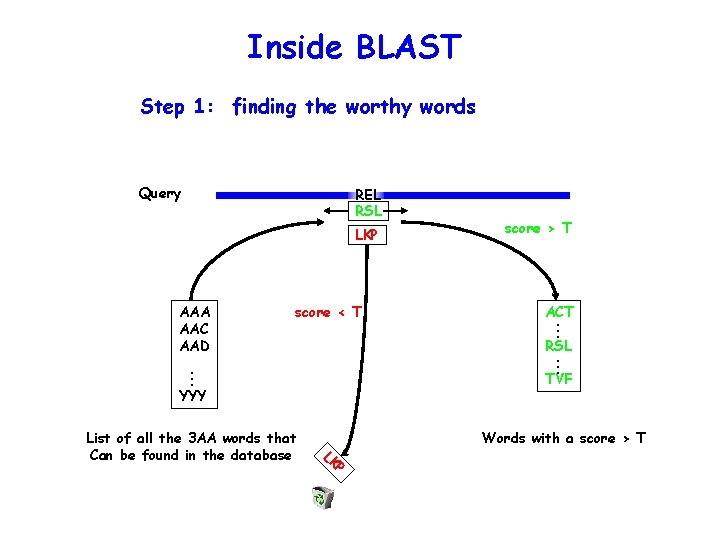
Inside BLAST Step 1: finding the worthy words Query REL RSL LKP score < T RSL . . . TVF YYY Words with a score > T P LK List of all the 3 AA words that Can be found in the database ACT. . . AAA AAC AAD score > T
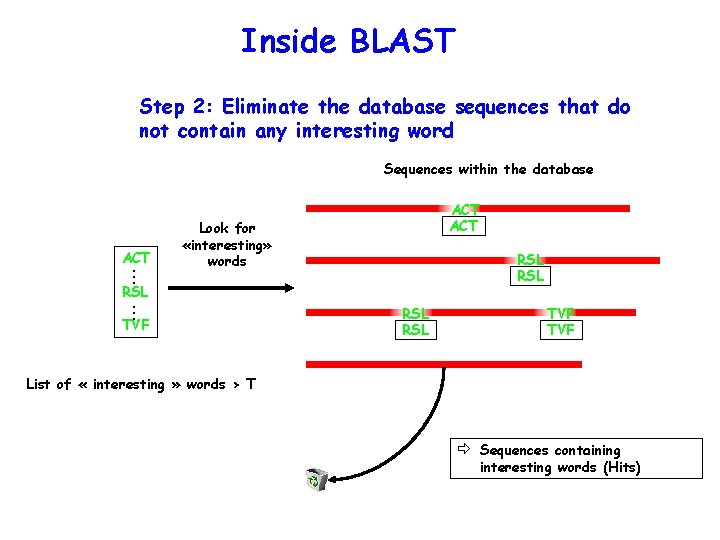
Inside BLAST Step 2: Eliminate the database sequences that do not contain any interesting word Sequences within the database ACT ACT . . . Look for «interesting» words RSL RSL . . . TVF RSL TVF List of « interesting » words > T ð Sequences containing interesting words (Hits)
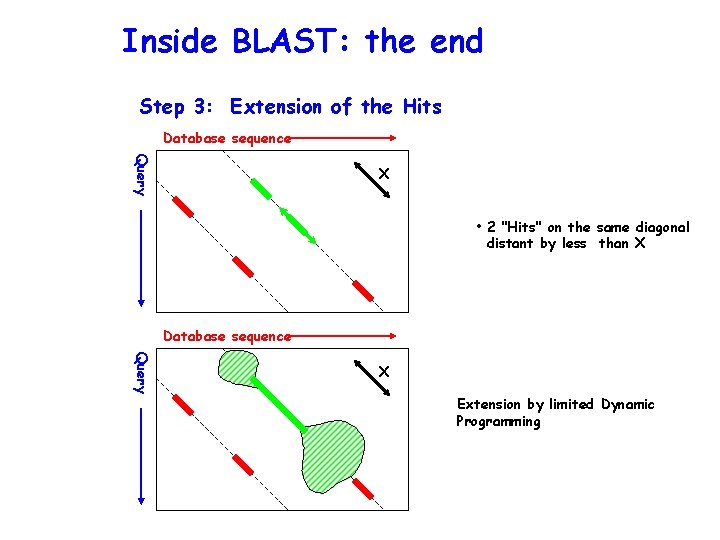
Inside BLAST: the end Step 3: Extension of the Hits Database sequence Query X • 2 "Hits" on the same diagonal distant by less than X Database sequence Query X Extension by limited Dynamic Programming

The Statistics in BLAST

BLAST Statistics: Raw Score Evaluation of the score • Raw Score ðSum of the substitutions and gap penalties. ðNot very informative

BLAST Statistics: P Values Derived Statistics • p-value ðProbability of finding an alignment with such a score, by chance. ðThe lower, the better
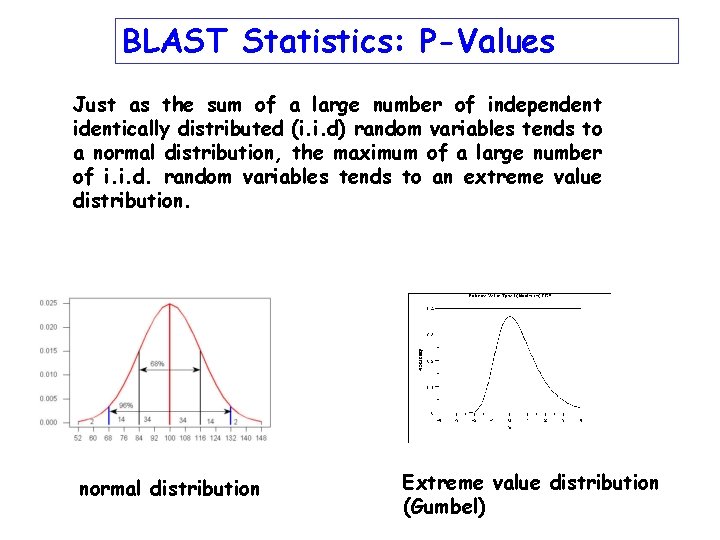
BLAST Statistics: P-Values Just as the sum of a large number of independent identically distributed (i. i. d) random variables tends to a normal distribution, the maximum of a large number of i. i. d. random variables tends to an extreme value distribution. normal distribution Extreme value distribution (Gumbel)

BLAST Statistics: P-Values P-Value: Probability that a random alignments obtains a score superior or Equal to X K must be calibrated with the database composition Lambda is calibrated with the matrix being used

BLAST Statistics: E-Values Derived Statistics • E-value ðNumber of alignments expected by chance ðThe lower, the better: <0. 00001 For Values Lower than 0. 0001, E-Value ~ P-Value The E-Values are easier to compare than P-Values

BLAST Statistics: Bit-Score • Bit Score ðEvaluates the amount of information in the alignment ðMakes it possible to compare alignments

BLAST Statistics: Booby Trap! The E-Value depends on N, the Database size. If N increases, some Hits can be lost

P 31383 Vs YEAST P 31383 Vs Uni. Prot

The Many Flavors of BLAST
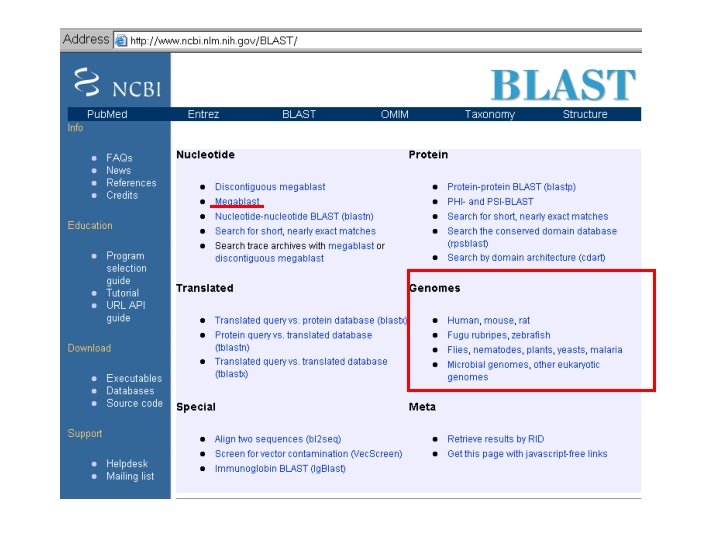
http: //blast. ncbi. nlm. nih. gov/Blast. cgi
http: //blast. ncbi. nlm. nih. gov/Blast. cgi
http: //blast. ncbi. nlm. nih. gov/Blast. cgi
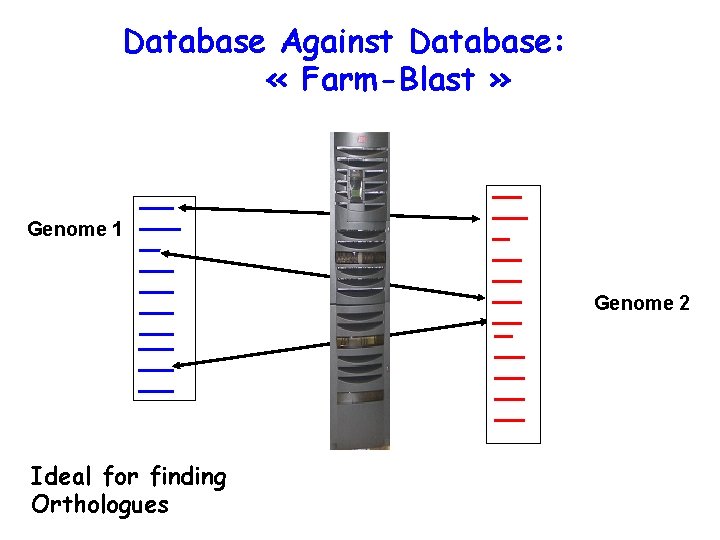
Database Against Database: « Farm-Blast » Genome 1 Genome 2 Ideal for finding Orthologues
The Classics 1 Sequence Vs A sequence Db
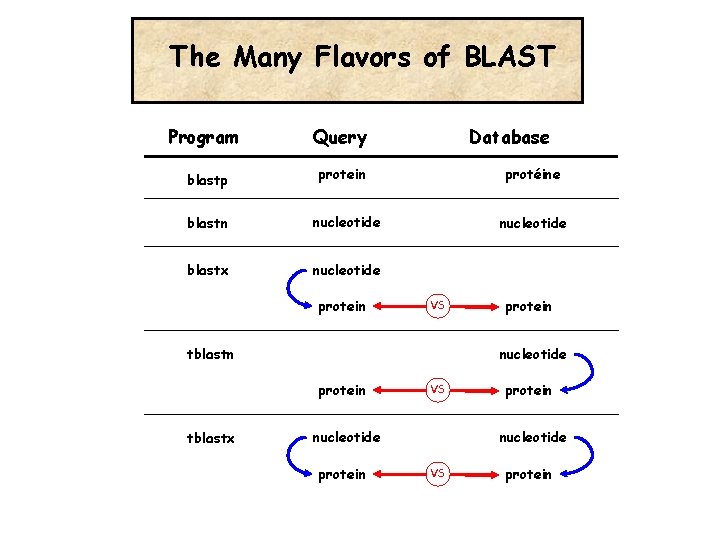
The Many Flavors of BLAST Program Query blastp protein blastn nucleotide blastx nucleotide protein Database protéine nucleotide VS tblastn nucleotide protein tblastx protein VS nucleotide protein nucleotide VS protein

The Many Flavors of BLAST Program Psi-blast Query protein Database protein RPS-blast protein Domain DART-blast protein mega-blast DNA Large DNA
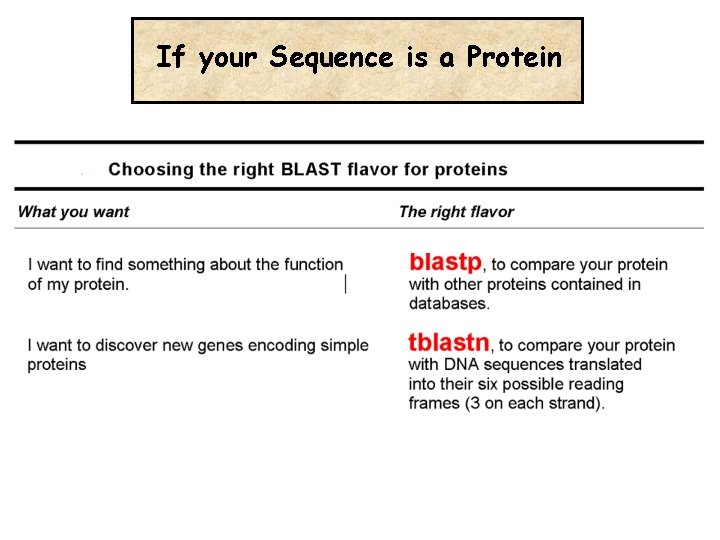
If your Sequence is a Protein
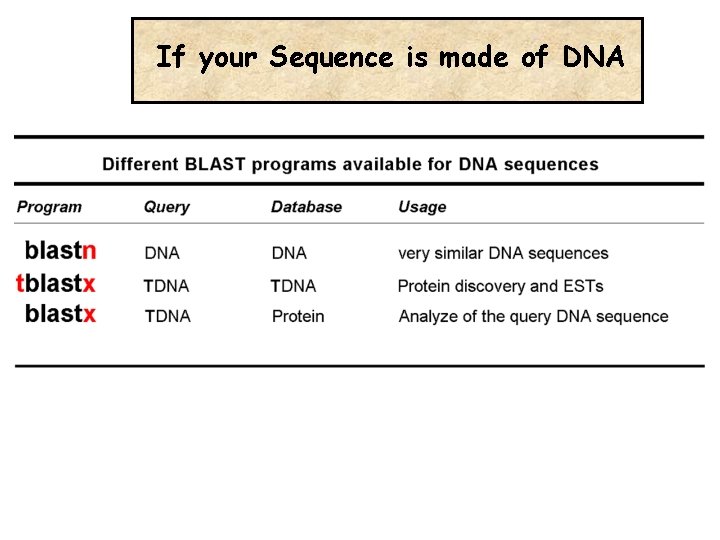
If your Sequence is made of DNA

BLASTing with DNA: Asking the right question.

Keeping an Eye on the Public Servers.

Using BLAST: The Basic Way
Database Search Result=Prediction Protein X IS or IS NOT homologous to the QUERRY.
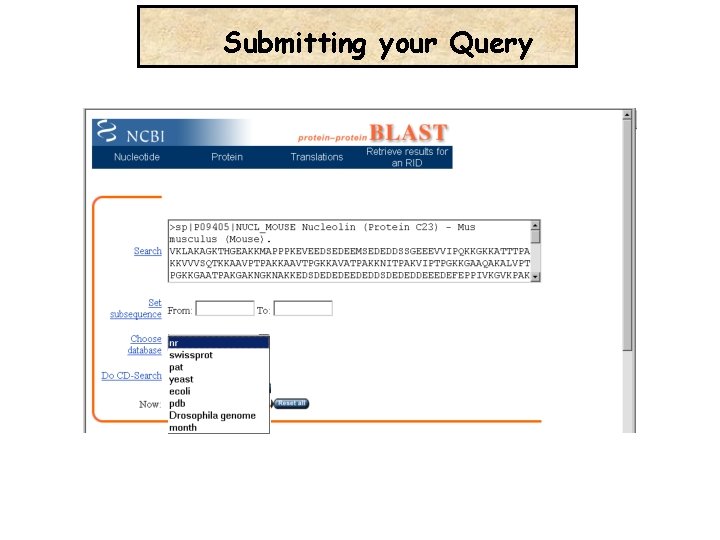
Submitting your Query

Understanding the BLAST Output Graphic Display Hit List Alignments

Understanding the Graphic Display
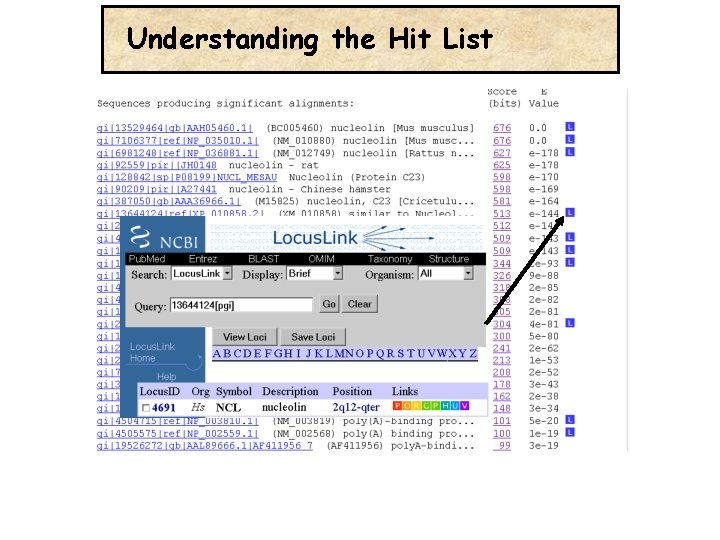
Understanding the Hit List

Understanding the Alignments Low Complexity

Low Complexity Regions ð Regions with a single residue repeated many times (like the AFGP) can produce meaningless alignments. ð The statistics expect ALL the regions to look the same « on average » . ð By default, BLAST replaces these regions with Xs

Reproducing The Experiment Everything you need to know to reproduce your search is at the bottom. BLAST searches are notoriously difficult to reproduce

Database Searches: A few Guidelines

Data. Base Search According to Pearson

Data. Base Search According to Pearson

Data. Base Search According to Pearson
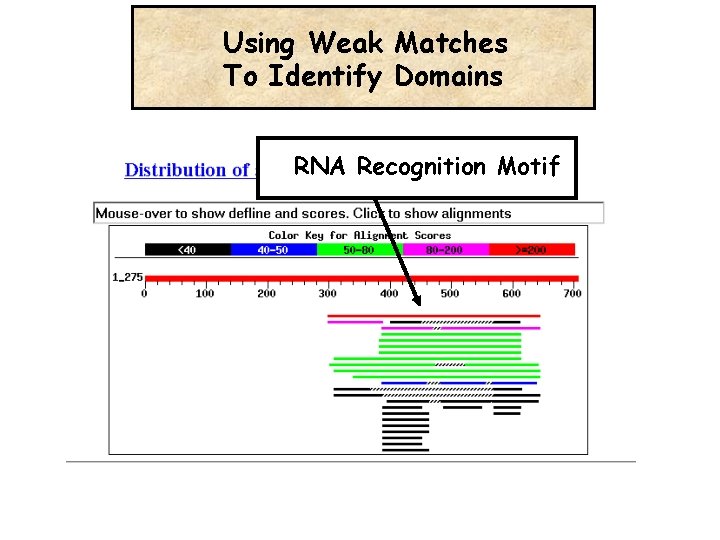
Using Weak Matches To Identify Domains RNA Recognition Motif

Three Short-Sighted Witnesses are more Informative than a single eagle-eye witness

Using BLAST: Trouble Shooting

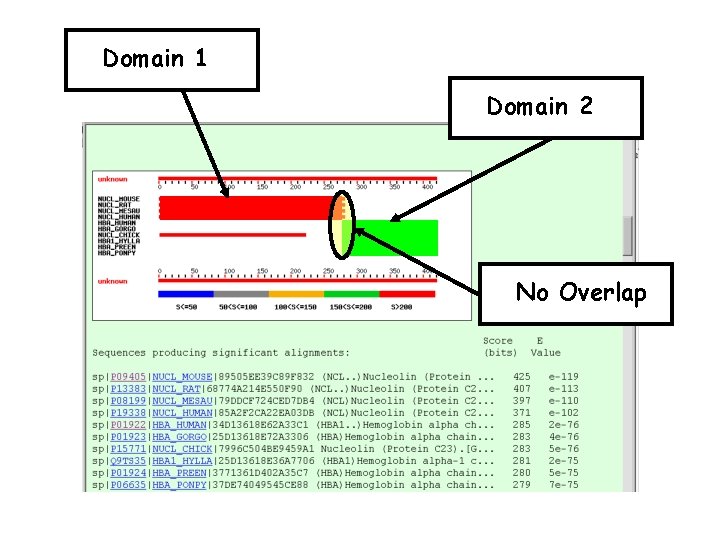
Domain 1 Domain 2 No Overlap
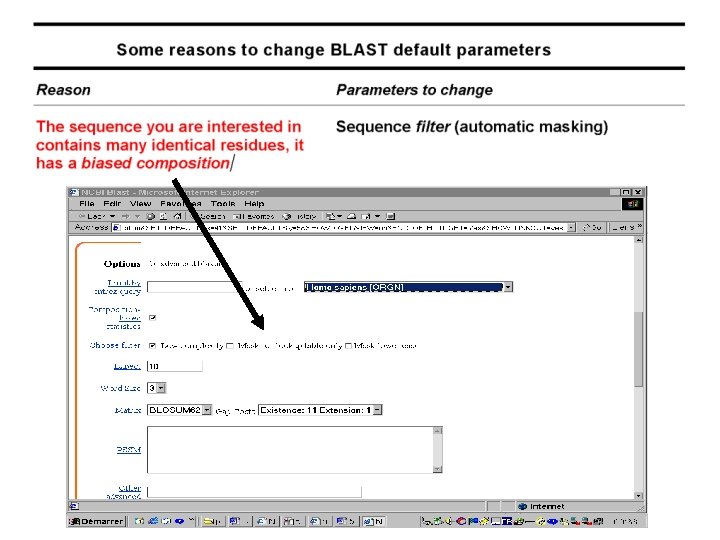

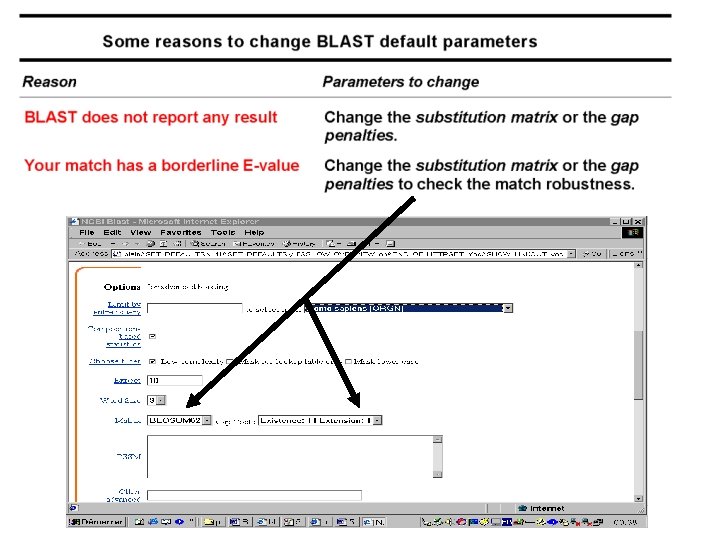


Advanced Blast on the EMBnet www. ch. embnet. org/software/a. BLAST. html • More choice on the databases • Change all the parameters

Adapting BLAST To your Problem
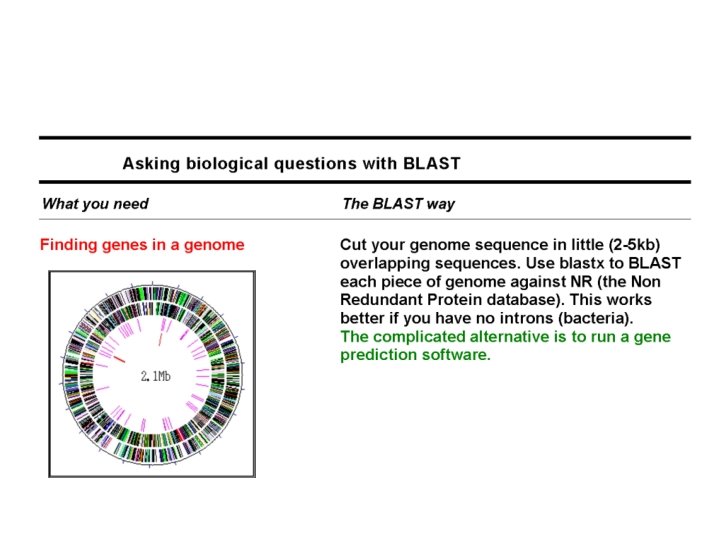

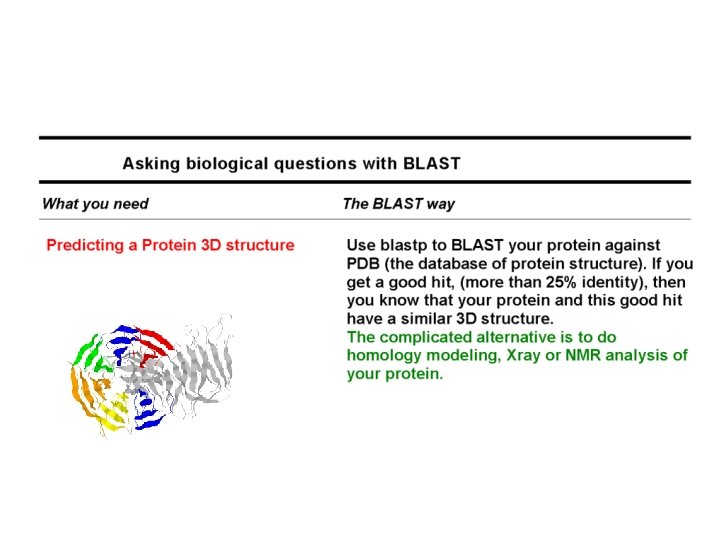
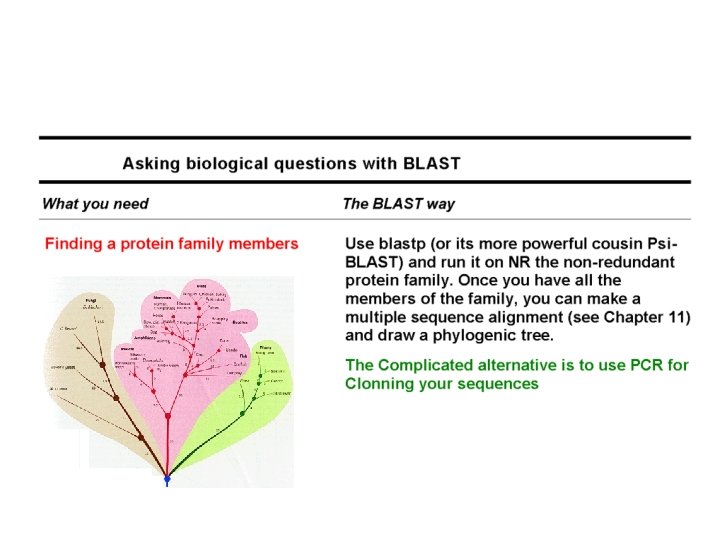

Domain-Flavored BLAST
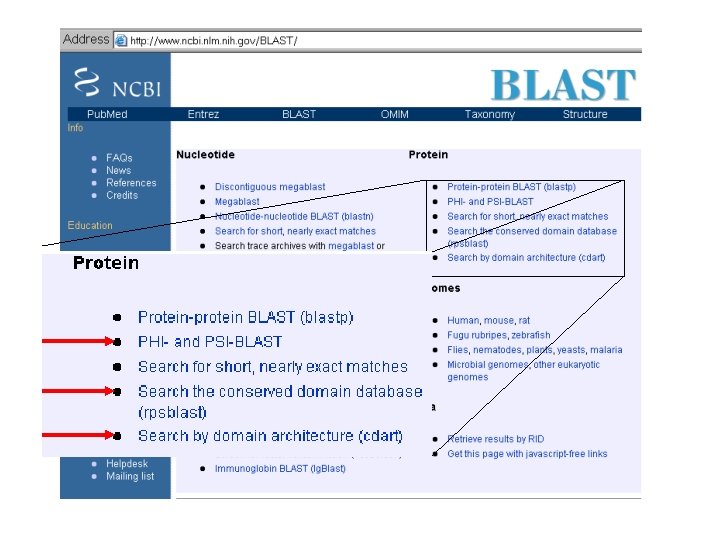

Psi-BLAST

BLAST latest Flavor PSI-BLAST -Position Specific Iterated Version of BLAST. -Uses Profiles. -More Sensitive.
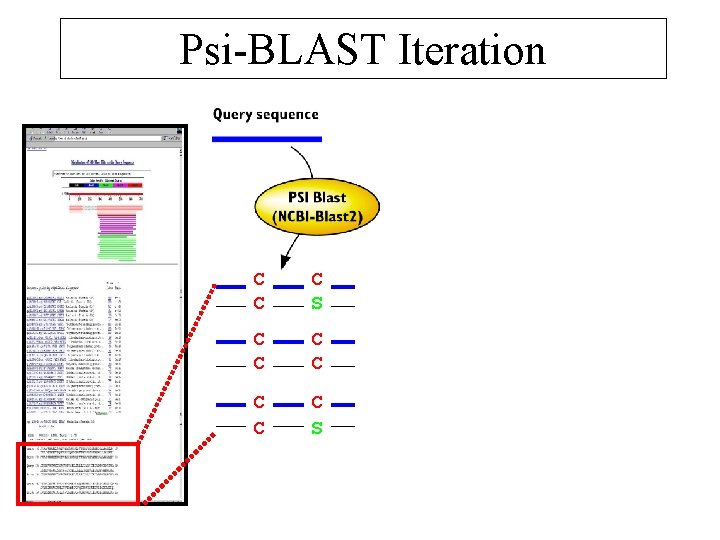
Psi-BLAST Iteration C C CC S C CC C C S
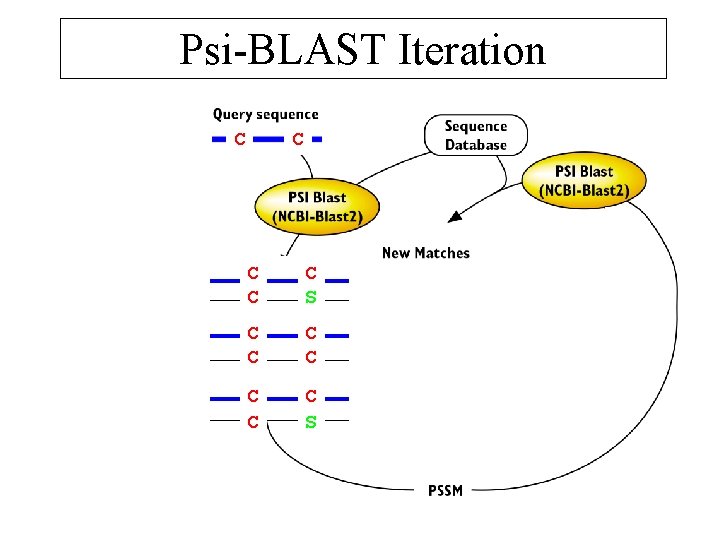
Psi-BLAST Iteration C C CC S C CC C C S
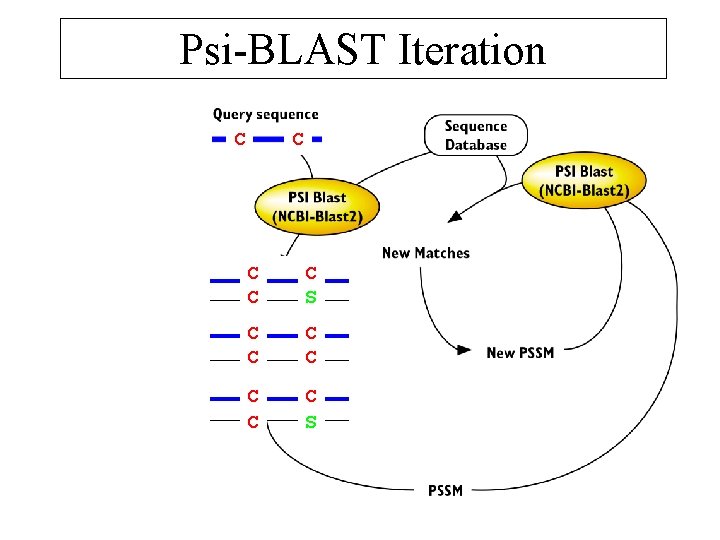
Psi-BLAST Iteration C C CC S C CC C C S

BLAST PSSM or weight matrix M Y C E Q A 0 2 -1 0 0 S -1 -1 -1 0 -1 C -1 -1 10 1 -1. . Y -1 6 -1 -1 -1 V -1 1 -1 -1 -1 U 0 0 0 E N 0 -1 0 0 0 5 C E S 0 -1 3 5 -1 -1 5 4 -1 0 -1 -1 -1 . .

Asking a Question With Psi-BLAST

Asking a Question With Psi-BLAST Is the Leghemoglobin related to the Human Hemoglobin ?
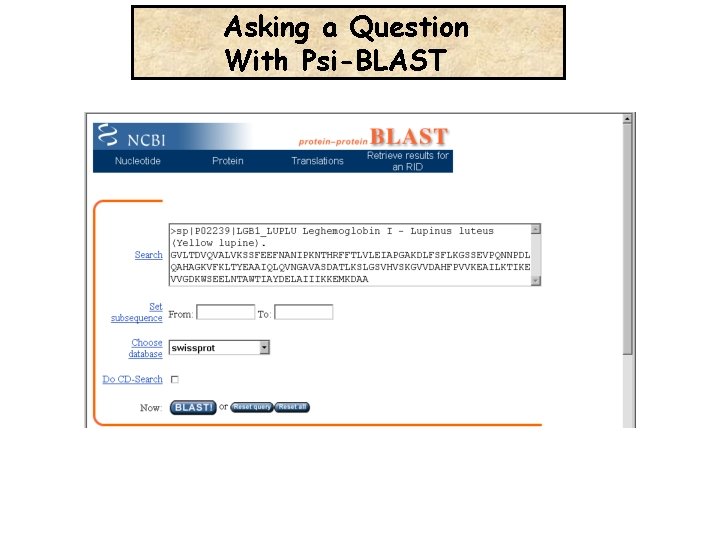
Asking a Question With Psi-BLAST
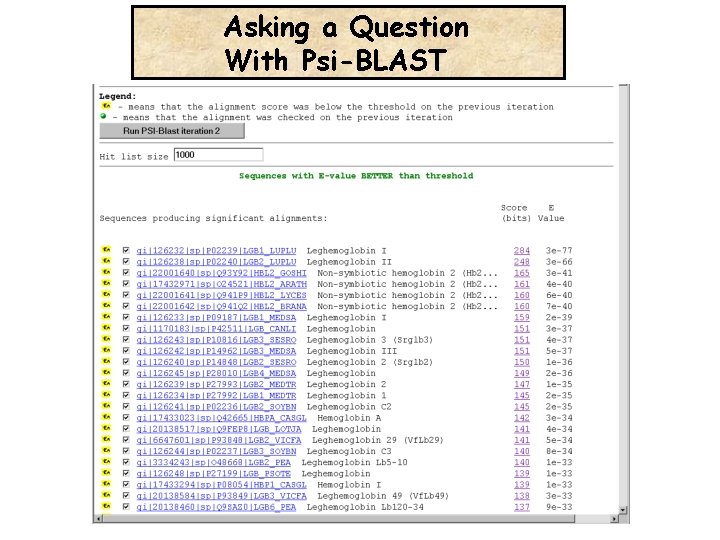
Asking a Question With Psi-BLAST
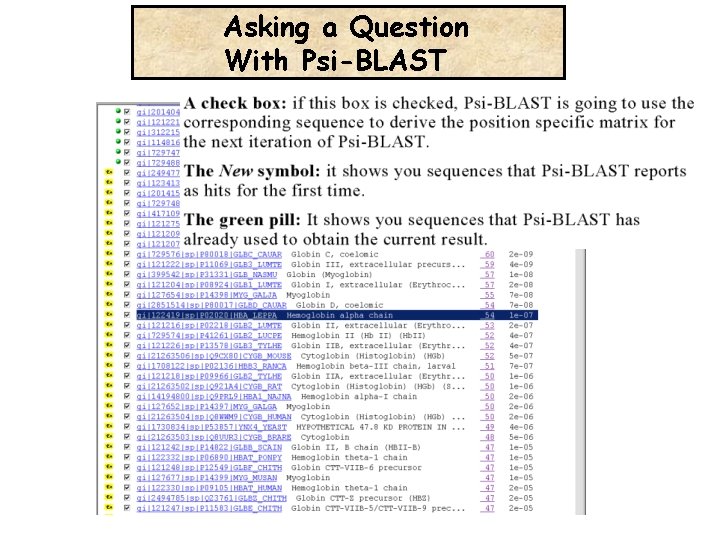
Asking a Question With Psi-BLAST


Which Domain Organisation For Your Protein: (Reverse PSI-BLAST)
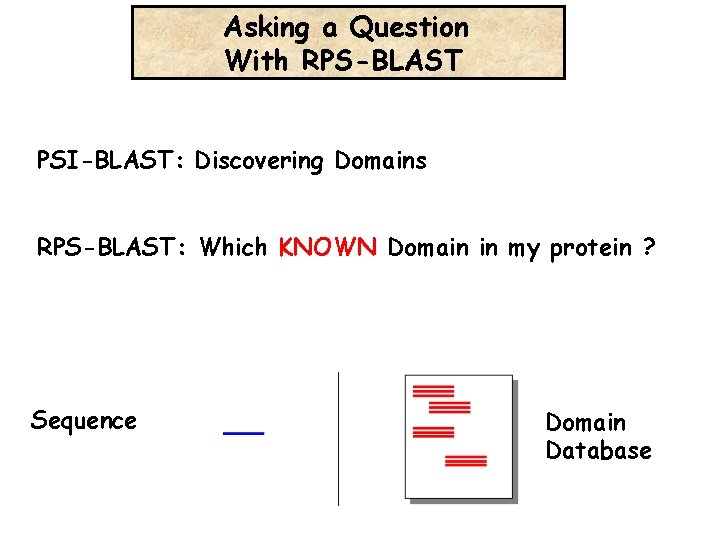
Asking a Question With RPS-BLAST PSI-BLAST: Discovering Domains RPS-BLAST: Which KNOWN Domain in my protein ? Sequence Domain Database
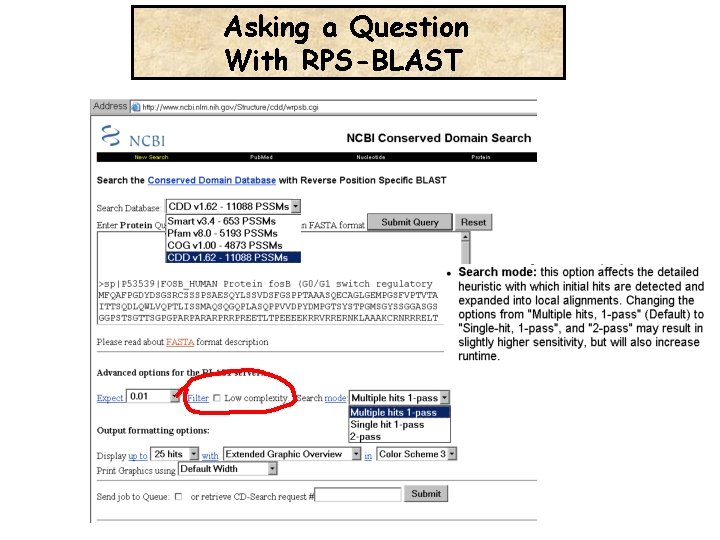
Asking a Question With RPS-BLAST

False Hits caused by the domain low complexity (see Evalues)
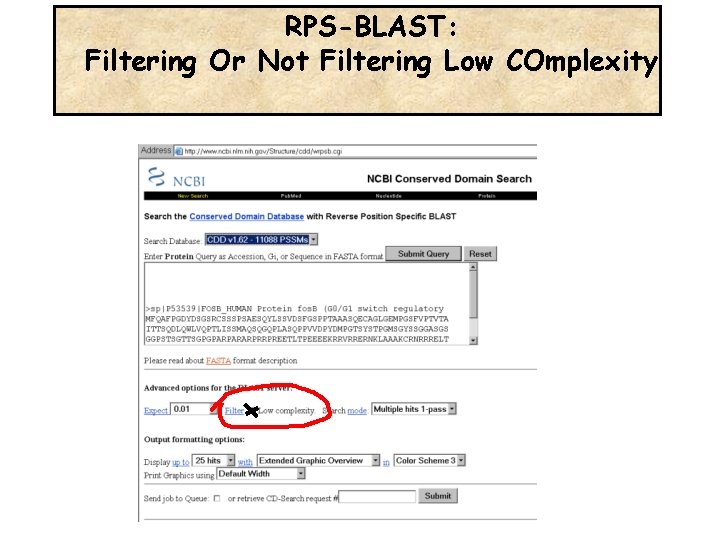
RPS-BLAST: Filtering Or Not Filtering Low COmplexity


How Many Proteins Have the same Domain Structure as Mine ? (CDART)

Asking a Question With CDART: Conserved Domain Architecture Retrieval Tool Finds the proteins that contain the same domains as your protein.

Asking a Question With CDART PSI-BLAST: Discovering Domains RPS-BLAST: Which known Domain in my protein ? CDART: -Which proteins have the SAME DOMAIN ORGANIZATION as my proteins ? Which domains are COMMONLY ASSOCIATED with the domain I am interested in ?
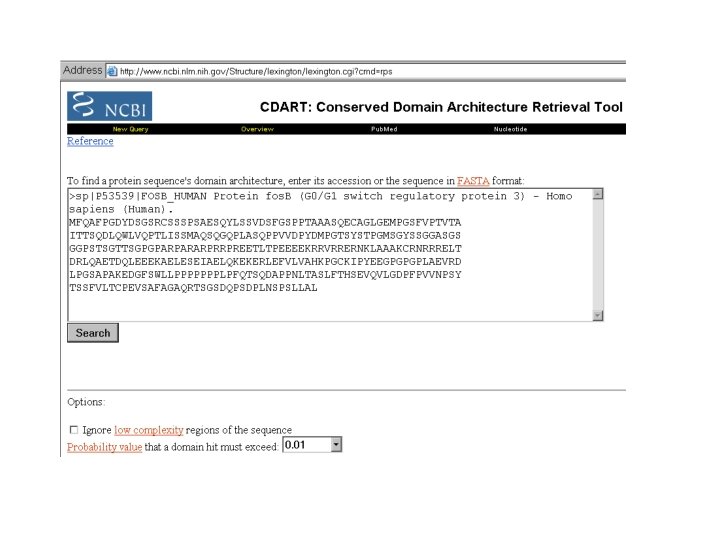
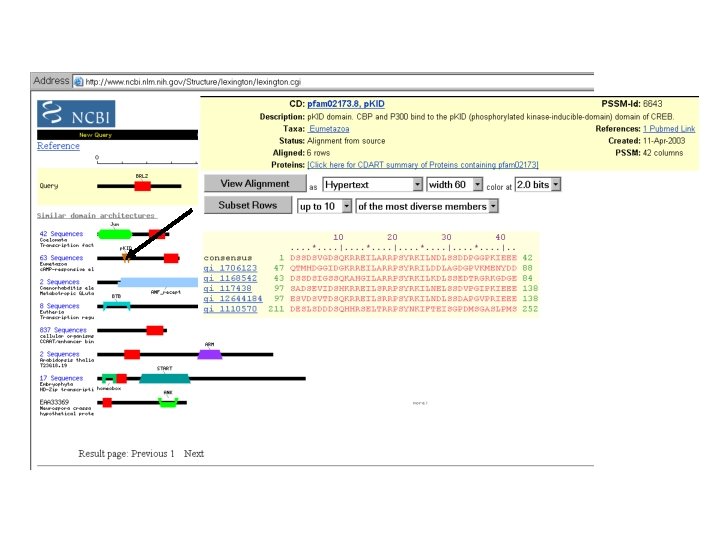

Filtering: -By Domain -By Species

-I want to Find all the Insect proteins containing a June/Fos organisation.
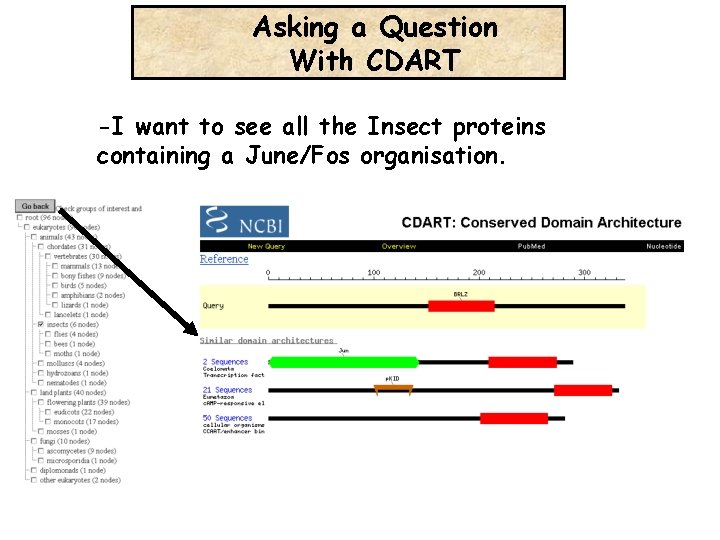
Asking a Question With CDART -I want to see all the Insect proteins containing a June/Fos organisation.

Asking a Question With CDART -I want to see all the Insect proteins containing a June/Fos organisation.
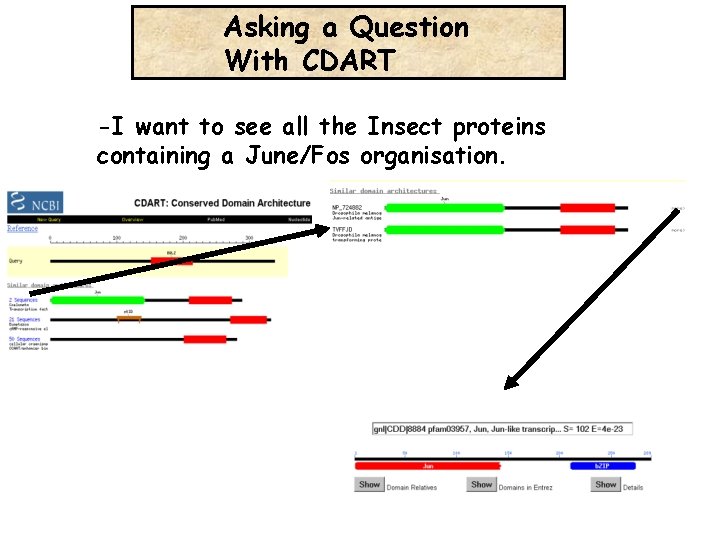
Asking a Question With CDART -I want to see all the Insect proteins containing a June/Fos organisation.
Genome Flavored BLAST
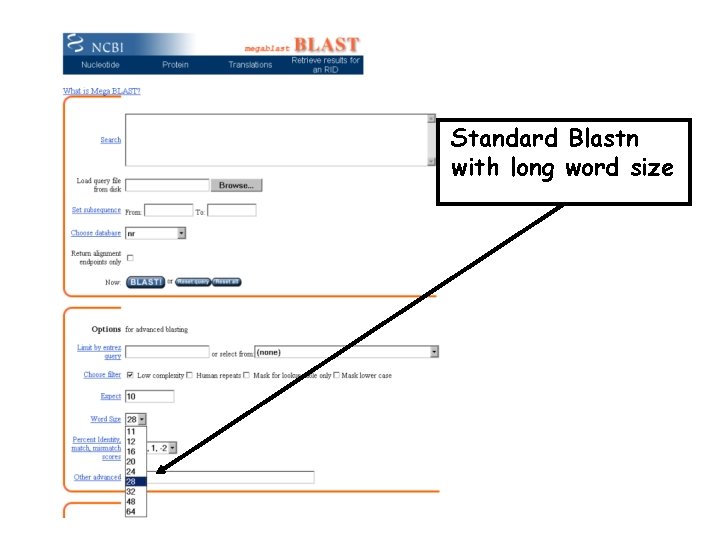
Standard Blastn with long word size
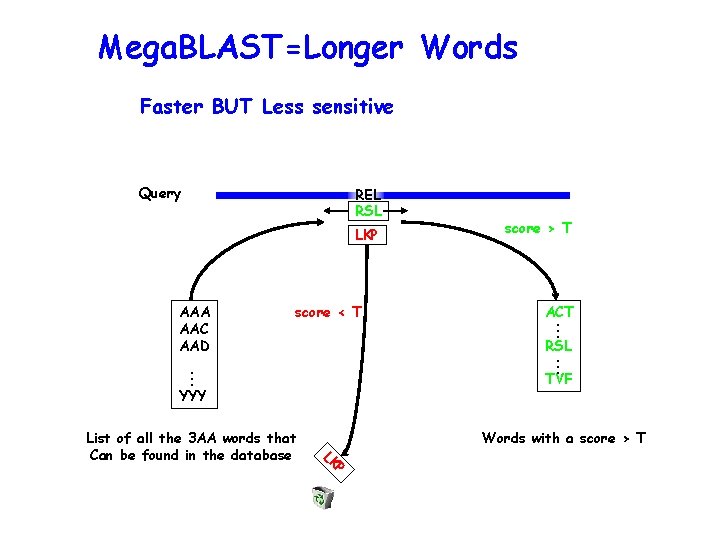
Mega. BLAST=Longer Words Faster BUT Less sensitive Query REL RSL LKP score < T RSL . . . TVF YYY Words with a score > T P LK List of all the 3 AA words that Can be found in the database ACT. . . AAA AAC AAD score > T
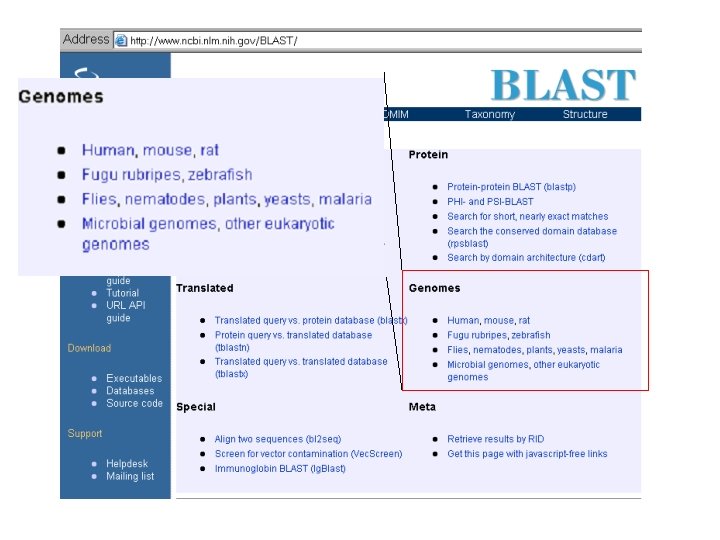

The Nc. Bi Bl. As. T GEno. Me Sec. Tion is Mes. Sy

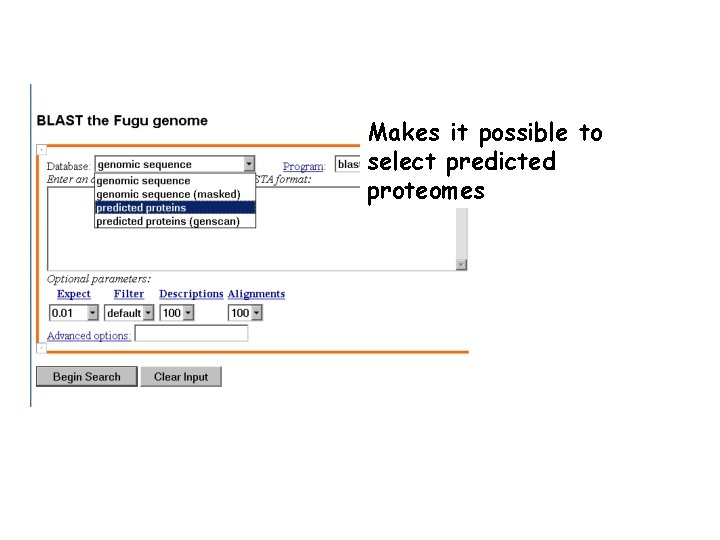
Makes it possible to select predicted proteomes
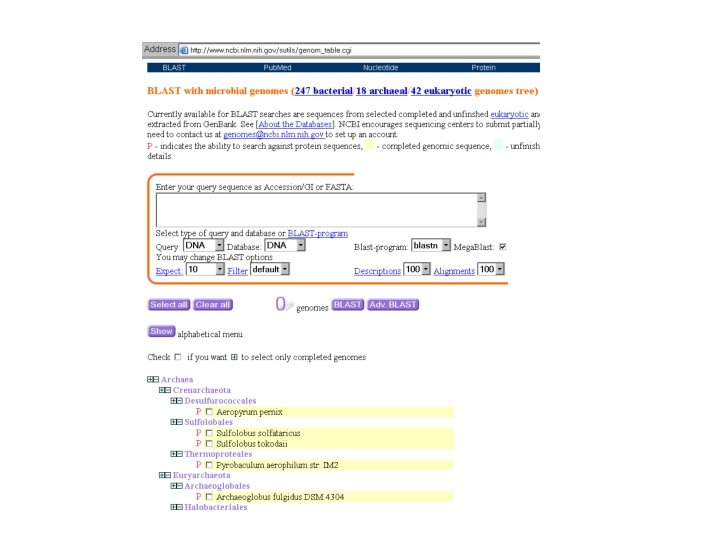

Venter-BLAST

When it comes to BLASTing Eukaryotic Genomes: WWW. ENSEMBL. ORG

Asking a Question With ENSEMBL-BLAST ENSEMBL: WHERE are located the genes coding for Homologues of my protein
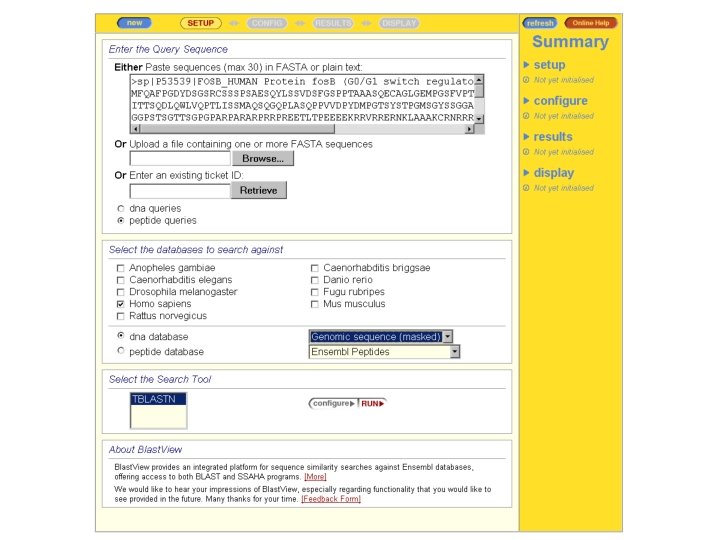
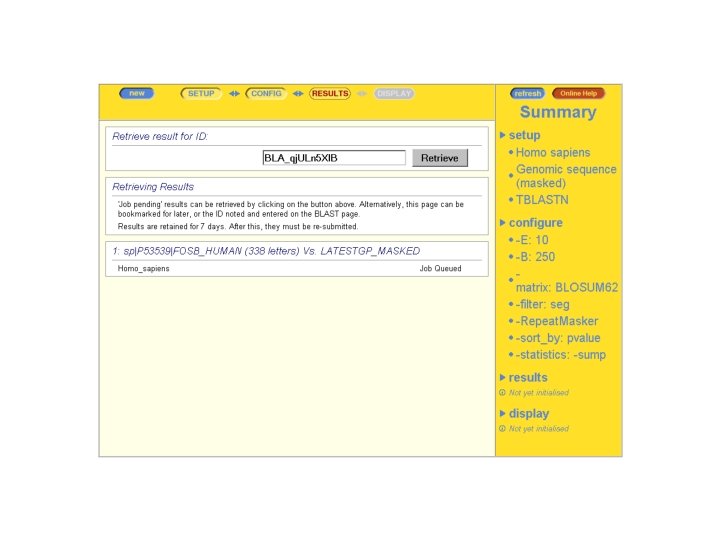
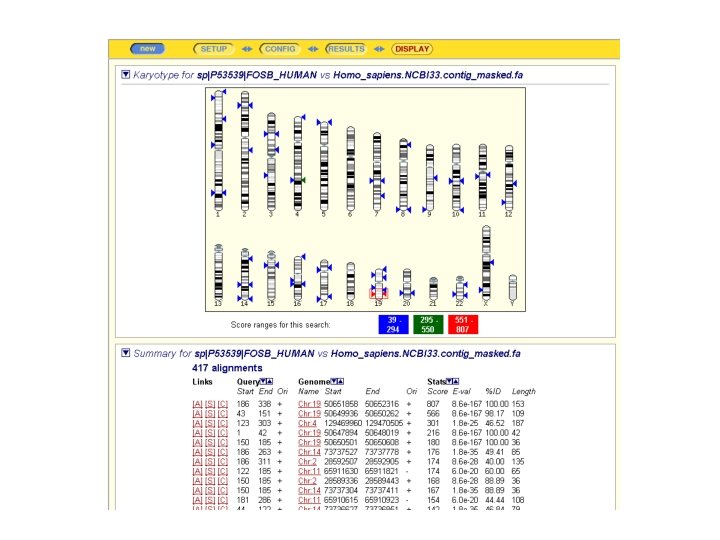

CONCLUSION

Searching Databases -BLAST is a fast approximation for the Full Local Dynamic Programming. It is convenient to scan Databases. -BLAST computes the Statistical Significance of the Alignments (E-Value, P-Value). -The main pitfall to avoid are low complexity regions

Searching Databases -USE blastp the best educated blast to discover the function of your protein -USE Psi-Blast to find remote homologues -USE RPS-Blast to find domains in your protein (Interpro for EBI) -USE ENSEMBL-Blast for the human Genome

A few Extra Ressources
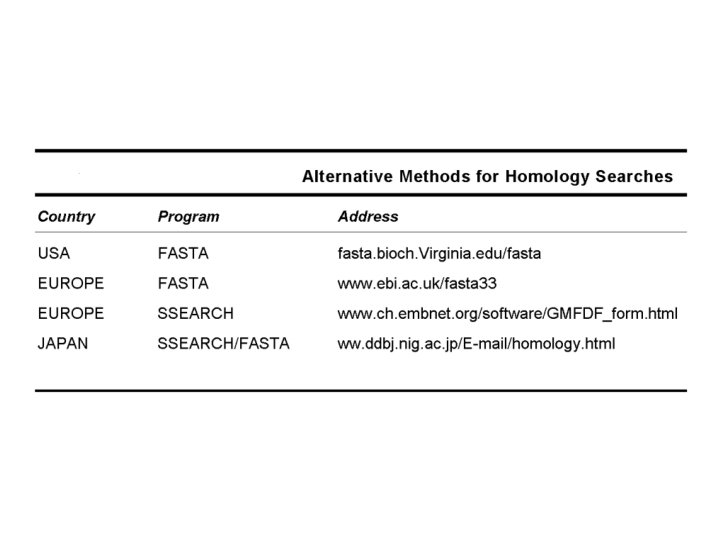



Tunning BLAST

BLAST Tunning

- Slides: 131