Gapped BLAST and PSIBLAST a new generation of

Gapped BLAST and PSI-BLAST: a new generation of protein database search programs Stephen F. Altschul, Thomas L. Madden, Alejandro A. Schäffer, Jinghui Zhang, Zheng Zhang, Webb Miller and David J. Lipman Nucleic Acids Research, 1997, Vol. 25 No. 17 Oxford University Press

Index § § § Introduction BLAST 1. 0 BLAST 2. 0 Future works Conclusions 1

Index § § § Introduction BLAST 1. 0 BLAST 2. 0 Future works Conclusions 2

Introduction § The most used tool for searching protein and DNA similarities. § Variants • • BLASTP – TBLASP BLASTN – TBLASN BLASTX – TBLASTX … 3

Index § § § Introduction BLAST 1. 0 BLAST 2. 0 Future works Conclusions 4

BLAST 1. 0 § Features • One hit seeding • Ungapped extension § Pros • More specific searching § Cons • Computational and memory consumption 5

Index § § § Introduction BLAST 1. 0 BLAST 2. 0 Future works Conclusions 6

BLAST 2. 0 § Features • Two hit seeding • Gapped extension § Pros • Reduced computational and memory consumption • More relevant results § Cons • Less specific searching 7
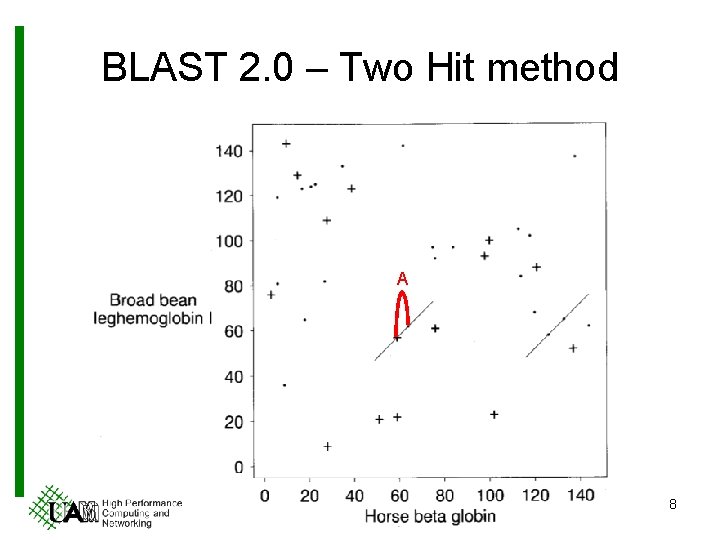
BLAST 2. 0 – Two Hit method A 8
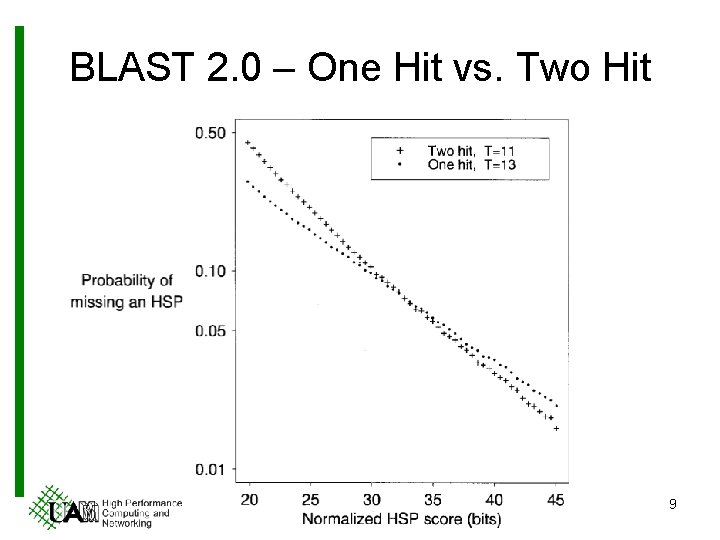
BLAST 2. 0 – One Hit vs. Two Hit 9

BLAST 2. 0 – Gapped Extension 1. Run an ungapped extension (as BLAST 1. 0) 2. If the HSP > Sg 3. Run a gapped extension 1. Apply Smith-Waterman algorithm 10

BLAST 2. 0 – What’s Sg? 11
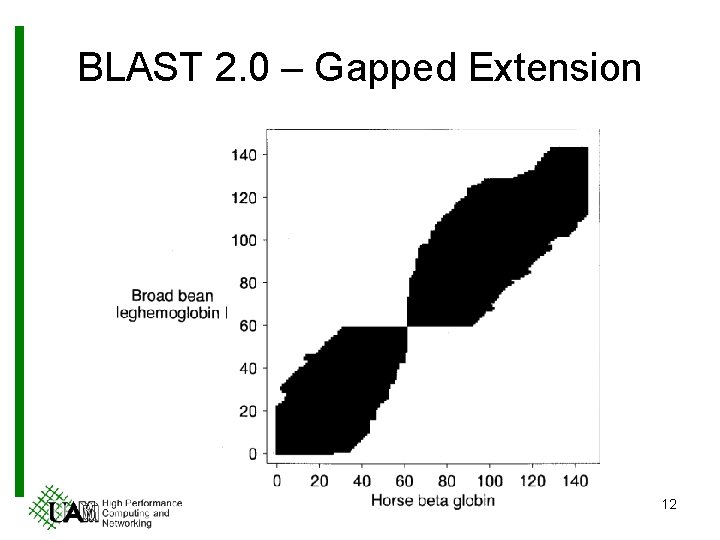
BLAST 2. 0 – Gapped Extension 12
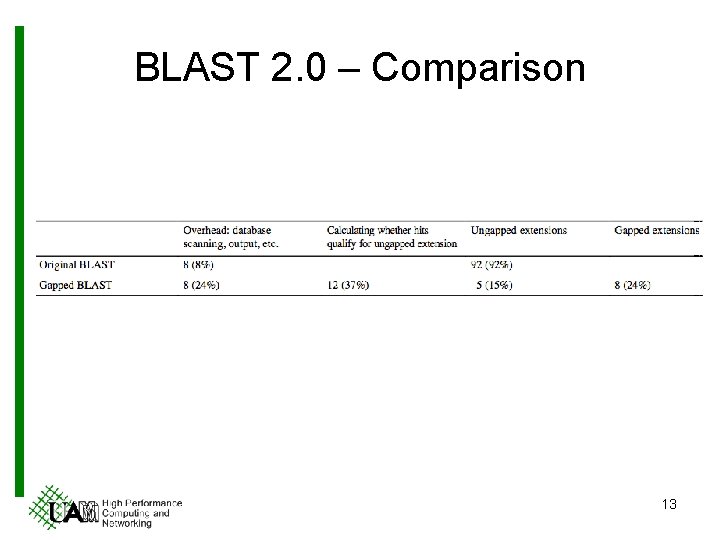
BLAST 2. 0 – Comparison 13

PSI-BLAST § Features • Use BLAST 2. 0 output for building position-specific scoring matrices (PSSM). • Run an iterated variant of BLAST 2. 0 to process PSSM. – Different scoring method. § Pros • More sensitivity. § Cons • More computational consumption. 14
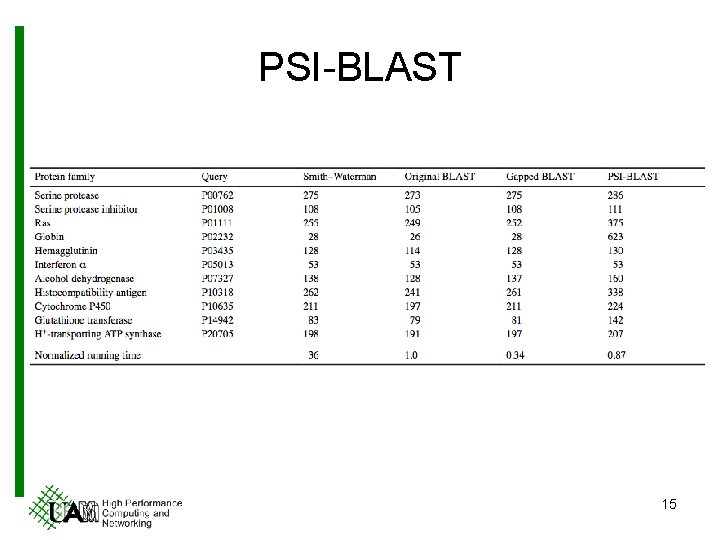
PSI-BLAST 15
Future Works § High-performance computing. § Medicine Specialized § DNA sequencers 16

Conclusions § BLAST 2. 0 is more optimized than BLAST 1. 0, by returning less results but more relevant. § High-performance approaches can optimized seeding and extension stages. § New heuristic models that improve previous techniques. 17
- Slides: 18