What is Blast WhatWhy Standalone Blast LocatingDownloading Blast
What is Blast What/Why Standalone Blast Locating/Downloading Blast Using Blast You need: Your sequence to Blast and the database to search against

National Center for Biotechnology Information (NCBI) – 1988 BLAST – Basic Local Alignment Search Tool Blast finds regions of similarity between biological sequences. The program compares nucleotide or protein sequences to sequence databases NCBI maintains several large annotated databases that are freely available to the public for download. YOU MAY WANT TO USE YOUR OWN DATABASE
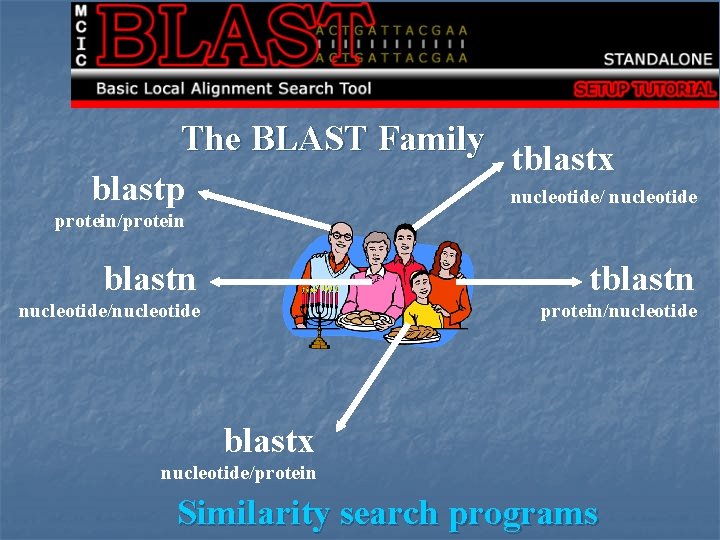
The BLAST Family tblastx blastp nucleotide/ nucleotide protein/protein blastn tblastn nucleotide/nucleotide protein/nucleotide blastx nucleotide/protein Similarity search programs
n Why Standalone Blast The main advantage of Standalone Blast is to be able to create and Blast your own databases n Create a folder called Blast on your C: / drive to store the downloaded file n Get Blast Here n Scroll n Click http: //www. ncbi. nlm. nih. gov/ Down To Here FTP site
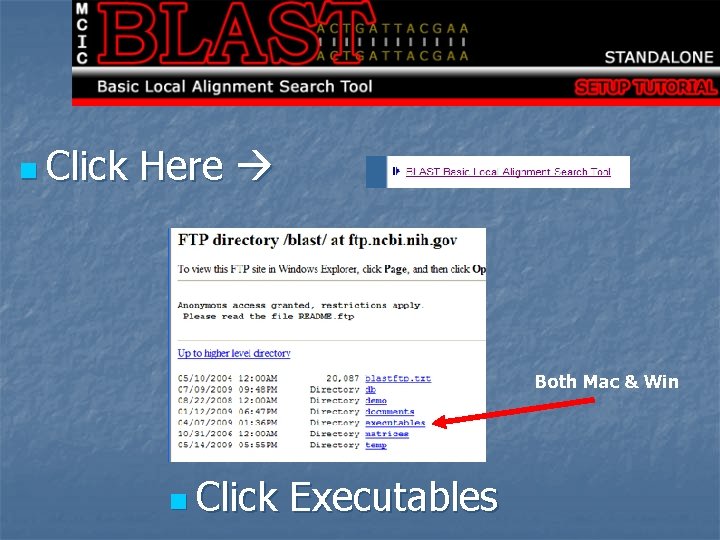
n Click Here Both Mac & Win n Click Executables
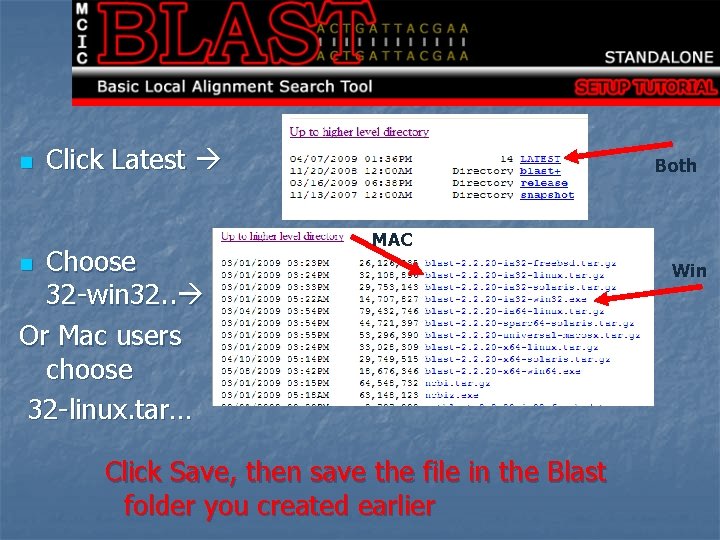
n Click Latest Choose 32 -win 32. . Or Mac users choose 32 -linux. tar… Both MAC n Click Save, then save the file in the Blast folder you created earlier Win
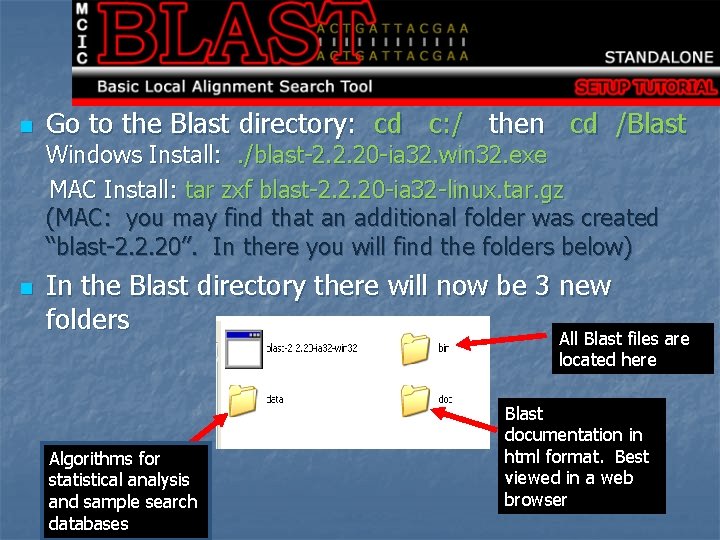
n Go to the Blast directory: cd c: / then cd /Blast Windows Install: . /blast-2. 2. 20 -ia 32. win 32. exe MAC Install: tar zxf blast-2. 2. 20 -ia 32 -linux. tar. gz (MAC: you may find that an additional folder was created “blast-2. 2. 20”. In there you will find the folders below) n In the Blast directory there will now be 3 new folders All Blast files are located here Algorithms for statistical analysis and sample search databases Blast documentation in html format. Best viewed in a web browser
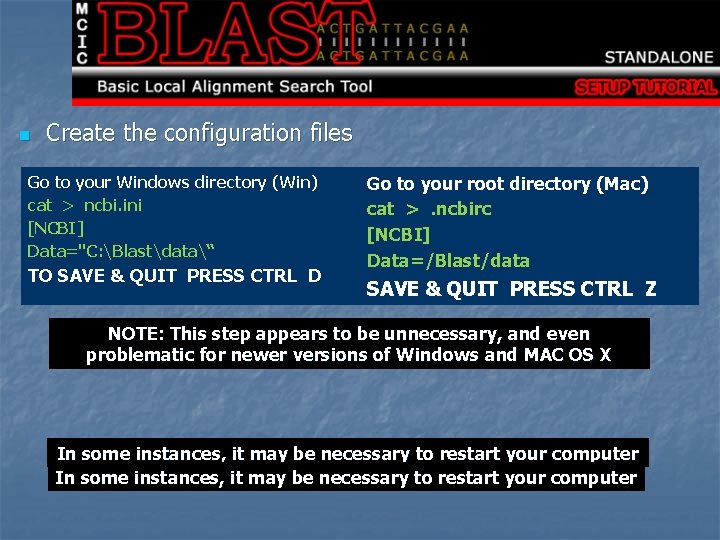
n Create the configuration files Go to your Windows directory (Win) cat > ncbi. ini [NCBI] Data="C: Blastdata“ TO SAVE & QUIT PRESS CTRL D Go to your root directory (Mac) cat >. ncbirc [NCBI] Data=/Blast/data SAVE & QUIT PRESS CTRL Z NOTE: This step appears to be unnecessary, and even problematic for newer versions of Windows and MAC OS X In some instances, it may be necessary to restart your computer

n NOW YOUR BLAST PROGRAM HAS BEEN INSTALLED Test Blast using the single sequence and database provided ** Change to the Bio. Download directory where your database and query sequence is stored. n Formatdb Prepare the database before Blasting To execute formatdb -i my. Dbase -p F you need the correct path /cygdrive/c/Blast/bin/formatdb -i. /TA 496 Seq 1. txt –p F -i the file to create into a searchable database -p T = protein database and F = nucleotide database
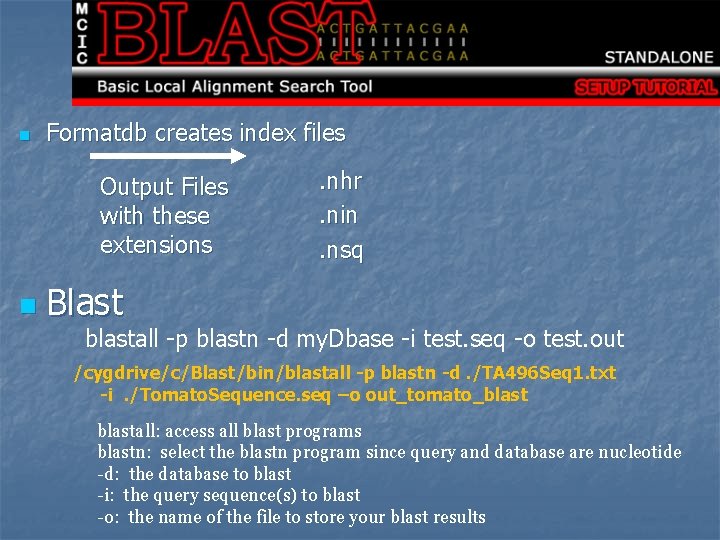
n Formatdb creates index files Output Files with these extensions n . nhr. nin. nsq Blast blastall -p blastn -d my. Dbase -i test. seq -o test. out /cygdrive/c/Blast/bin/blastall -p blastn -d. /TA 496 Seq 1. txt -i. /Tomato. Sequence. seq –o out_tomato_blastall: access all blast programs blastn: select the blastn program since query and database are nucleotide -d: the database to blast -i: the query sequence(s) to blast -o: the name of the file to store your blast results

n Additional Blast Parameters e-value e n Output of Blast Files & Parsing n Additional Blast Documentation
- Slides: 11