I 519 Introduction to Bioinformatics Fall 2012 Sequence

I 519 Introduction to Bioinformatics, Fall 2012 Sequence Database Searching (Basic Tools and Advanced Methods)

What’s database searching § Goal: find similar (homologous) sequences of a query sequence in a sequence of database § Input: query sequence & database § Output: hits (pairwise alignments)

Basics of database searching § Core: pairwise alignment algorithm § Speed (fast sequence comparison) § Relevance of the search results (statistical tests) § Recovering all information of interest – The results depend of the search parameters like gap penalty, scoring matrix. – Sometimes searches with more than one matrix should be preformed § Specificity and sensitivity
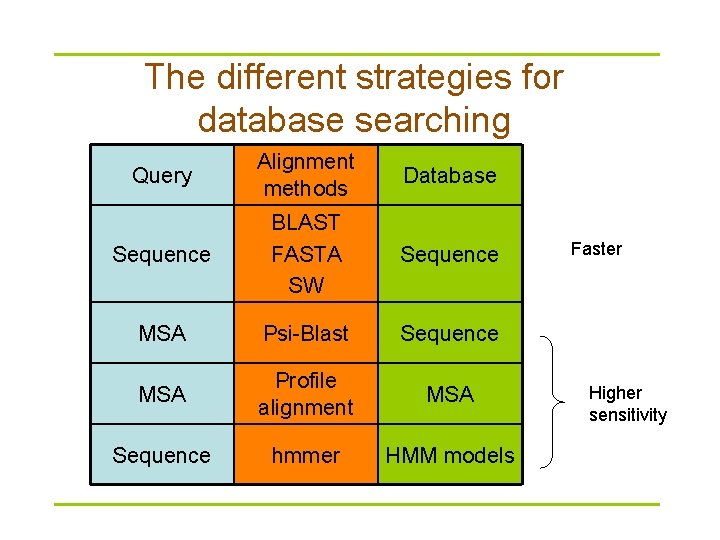
The different strategies for database searching Query Alignment methods Database Sequence BLAST FASTA SW Sequence MSA Psi-Blast Sequence MSA Profile alignment MSA Sequence hmmer HMM models Faster Higher sensitivity

What program to use for searching? § BLAST is fastest and easily accessed on the Web – A suite; BLASTP, BLASTN, BLASTX – lag behind the development of sequencing techniques § FASTA – more sensitive for DNA-DNA comparisons – FASTX and TFASTX can find similarities in sequences with frameshifts § Smith-Waterman is slower, but more sensitive – known as a “rigorous” or “exhaustive” search – SSEARCH in GCG and standalone FASTA

FASTA § Derived from logic of the dot plot – compute best diagonals from all frames of alignment § Word method looks for exact matches between words in query and test sequence – – hash tables (fast computer technique) DNA words are usually 6 bases protein words are 1 or 2 amino acids only searches for diagonals in region of word matches = faster searching
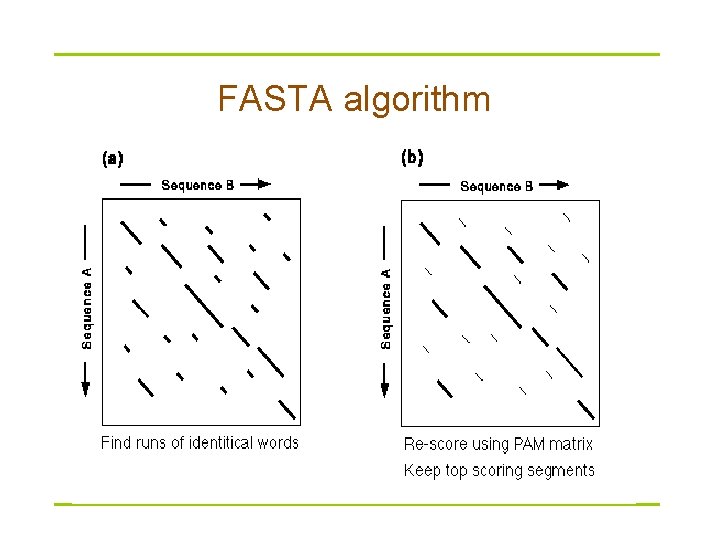
FASTA algorithm

Makes longest diagonal § after all diagonals found, tries to join diagonals by adding gaps § computes alignments in regions of best diagonals
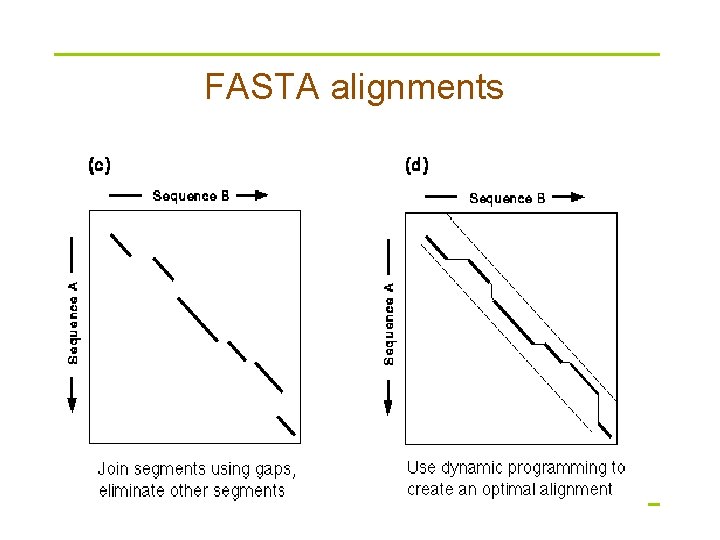
FASTA alignments

FASTA results: histogram !!SEQUENCE_LIST 1. 0 (Nucleotide) FASTA of: b 2. seq from: 1 to: 693 December 9, 2002 14: 02 TO: /u/browns 02/Victor/Search-set/*. seq Sequences: 2, 050 Symbols: 913, 285 Word Size: 6 Searching with both strands of the query. Scoring matrix: Gen. Run. Data: fastadna. cmp Constant pamfactor used Gap creation penalty: 16 Gap extension penalty: 4 Histogram Key: Each histogram symbol represents 4 search set sequences Each inset symbol represents 1 search set sequences z-scores computed from opt scores z-score obs exp (=) (*) < 20 0 0: 22 0 0: 24 3 0: = 26 2 0: = 28 5 0: == 30 11 3: *== 32 19 11: ==*== 34 38 30: =======*== 36 58 61: ========* 38 79 100: ========== * 40 134 140: =================* 42 167 171: =====================* 44 205 189: ========================*==== 46 209 192: ========================*===== 48 177 184: =======================*

FASTA results: list The best scores are: init 1 initn SW: PPI 1_HUMAN Begin: 1 End: 269 ! Q 00169 homo sapiens (human). phosph. . . 1854 SW: PPI 1_RABIT Begin: 1 End: 269 ! P 48738 oryctolagus cuniculus (rabbi. . . 1840 SW: PPI 1_RAT Begin: 1 End: 270 ! P 16446 rattus norvegicus (rat). pho. . . 1543 SW: PPI 1_MOUSE Begin: 1 End: 270 ! P 53810 musculus (mouse). phosph. . . 1542 SW: PPI 2_HUMAN Begin: 1 End: 270 ! P 48739 homo sapiens (human). phosph. . . 1533 SPTREMBL_NEW: BAC 25830 Begin: 1 End: 270 ! Bac 25830 musculus (mouse). 10, . . . 1488 SP_TREMBL: Q 8 N 5 W 1 Begin: 1 End: 268 ! Q 8 n 5 w 1 homo sapiens (human). simila. . . 1477 SW: PPI 2_RAT Begin: 1 End: 269 ! P 53812 rattus norvegicus (rat). pho. . . 1482 opt z-sc E(1018780). . 1854 2249. 3 1. 8 e-117 1840 2232. 4 1. 6 e-116 1543 1837 2228. 7 2. 5 e-116 1542 1836 2227. 5 2. 9 e-116 1533 1861. 0 7. 7 e-96 1488 1522 1847. 6 4. 2 e-95 1477 1522 1847. 6 4. 3 e-95 1482 1516 1840. 4 1. 1 e-94

FASTA results: alignment SCORES Init 1: 1515 Initn: 1565 Opt: 1687 z-score: 1158. 1 E(): 2. 3 e-58 >>GB_IN 3: DMU 09374 (2038 nt) initn: 1565 init 1: 1515 opt: 1687 Z-score: 1158. 1 expect(): 2. 3 e-58 66. 2% identity in 875 nt overlap (83 -957: 151 -1022) 60 70 80 90 100 110 u 39412. gb_pr CCCTTTGTGGCCGCCATGGACAATTCCGGGAAGCGGAGGCGATGGCGCTGTTGGCC || | ||||| DMU 09374 AGGCGGACATAAATCCTCGACATGGGTGACAACGAACAGAAGGCGCTCCAACTGATGGCC 130 140 150 160 170 180 120 130 140 150 160 170 u 39412. gb_pr GAGGCGGAGCGCAAAGTGAAGAACTCGCAGTCCTTCTTCTCTGGCCTCTTTGGAGGCTCA ||||| || ||| || ||||| || DMU 09374 GAGGCGGAGAAGAAGTTGACCCAGCAGAAGGGCTTTCTGGGATCGCTGTTCGGAGGGTCC 190 200 210 220 230 240 180 190 200 210 220 230 u 39412. gb_pr TCCAAAATAGAGGAAGCATGCGAAATCTACGCCAGAGCAGCAAACATGTTCAAAATGGCC ||| | ||||| || |||| || || DMU 09374 AACAAGGTGGAGGACGCCATCGAGTGCTACCAGCGGGCAACATGTTTAAGATGTCC 250 260 270 280 290 300 240 250 260 270 280 290 u 39412. gb_pr AAAAACTGGAGTGCTGCTGGAAACGCGTTCTGCCAGGCTGCACAGCTGCACCTGCAGCTC ||||| | |||||| ||| || | DMU 09374 AAAAACTGGACAAAGGCTGGGGAGTGCTTCTGCGAGGCGGCAACTCTACACGCGCGGGCT 310 320 330 340 350 360

FASTA on the web § Many websites offer FASTA searches – Various databases and various other services – Be sure to use FASTA 3 § Each server has its limits § Be aware that you are depending on the kindness of strangers.
![BLAST § [BLAST= Basic Local Alignment Search Tool] § The central idea of the BLAST § [BLAST= Basic Local Alignment Search Tool] § The central idea of the](http://slidetodoc.com/presentation_image_h/e8b29335a41eb07f6b7dece57cc81851/image-14.jpg)
BLAST § [BLAST= Basic Local Alignment Search Tool] § The central idea of the BLAST algorithm is that a statistically significant alignment is likely to contain a high-scoring pair of aligned words. § Uses word matching like FASTA – Similarity matching of words (3 aa’s, 11 bases) – Does not require identical words. § If no words are similar, then no alignment – won’t find matches for very short sequences § Original BLAST does not handle gaps well; the “gapped” blast is better

Word match Extend hits
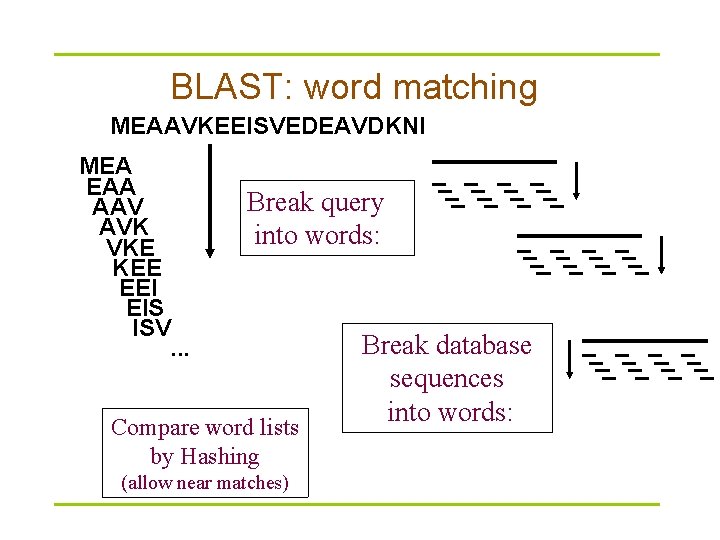
BLAST: word matching MEAAVKEEISVEDEAVDKNI MEA EAA AAV AVK VKE KEE EEI EIS ISV Break query into words: . . . Compare word lists by Hashing (allow near matches) Break database sequences into words:
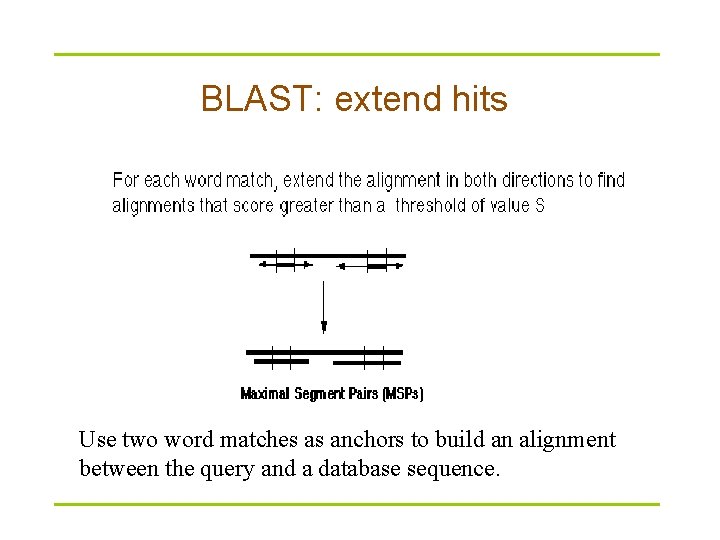
BLAST: extend hits Use two word matches as anchors to build an alignment between the query and a database sequence.

Gapped BLAST algorithm § The NCBI’s BLAST website now both use “gapped BLAST” § This algorithm is more complex than the original BLAST § It requires two word matches close to each other on a pair of sequences (i. e. with a gap) before it creates an alignment allow gaps (using Smith. Waterman algorithm constrained by a band width).
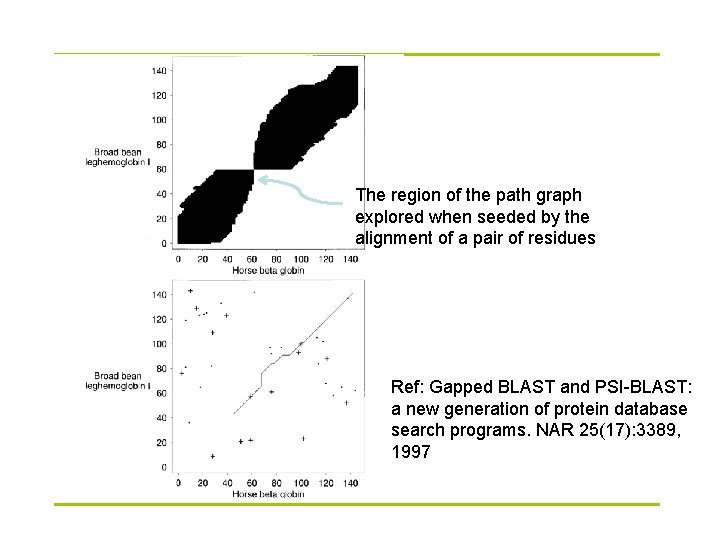
The region of the path graph explored when seeded by the alignment of a pair of residues Ref: Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. NAR 25(17): 3389, 1997

Statistical tests § Evaluate the probability of an event taking place by chance. § Very little is known about the random distribution of optimal global alignment scores § Statistics for the scores of local alignments, are well understood (particularly true for local alignments lacking gaps) § P-value – Randomized data – Distribution under the same setup

BLAST statistics § E-value is equivalent to standard P value (based on Karlin-Altschul theorem) – Karlin-Alschul equation is probably one of the most recognized equation in bioinformatics. – E-value represents the likelihood that the observed alignment is due to chance alone – E-value: the number of alignments expected by chance (E) during a sequence database search. E = 1 indicates that an alignment this good would happen by chance with any random sequence searched against the same database.

How to compute BLAST E-value Raw score (S) Bit score (S’) m: the effective query length n: the effective database length E P (the probability of getting at least one HSP)
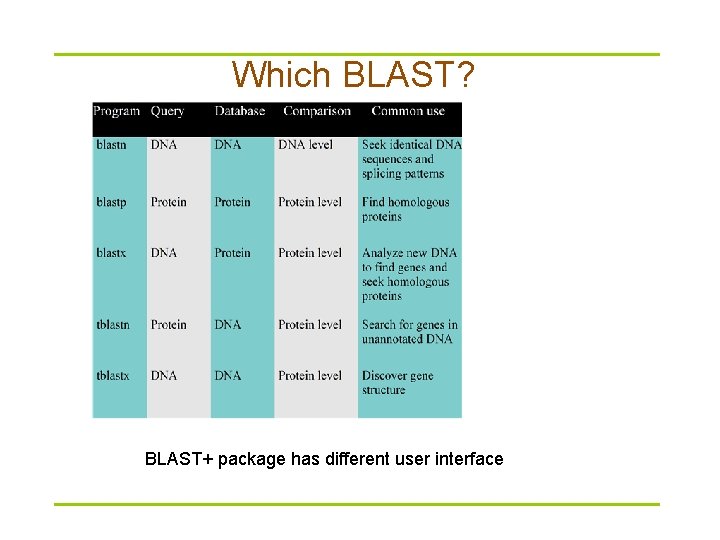
Which BLAST? BLAST+ package has different user interface

Which database? § For blastp (blastx) – – – Non-redundant protein sequences (nr) Reference proteins (refseq_protein) Swissprotein sequences (swissprot) Patented protein sequences (pat) Protein Data Bank proteins (pdb) Environmental samples (env_nr) § For blastn (tblastx) – – Human genomic plus transcript (not for tblastx) Mouse genomic plus transcript (not available for tblastx) Nucleotide collection (nr/nt) etc

BLAST is approximate § BLAST makes similarity searches very quickly because it takes shortcuts. – looks for short, nearly identical “words” (11 bases) § It also makes errors – misses some important similarities – makes many incorrect matches • easily fooled by repeats or skewed composition

Interpretation of BLAST hits § very low E values (e-100) are homologs § moderate E values are related genes § long list of gradually declining of E values indicates a large gene family § long regions of moderate similarity are more significant than short regions of high identity

Is the hit with smallest E-value the closest sequence to the query? § Not necessarily § Some people argue that more strict phylogeny analysis is needed for further conclusion – Why? Different evolutionary rates, etc

Biological relevance § It is up to you, the biologist to scrutinize these alignments and determine if they are significant. § Were you looking for a short region of nearly identical sequence or a larger region of general similarity? § Are the mismatches conservative ones? § Are the matching regions important structural components of the genes or just introns and flanking regions?

20 tips to improve your BLAST searches § “Don't Use the Default Parameters” (but what parameters? ) § “Treat BLAST Searches as Scientific Experiments § “View BLAST Reports Graphically” (depends!) § “Be Skeptical of Hypothetical Proteins” § “Look for Stop Codons and Frame-Shifts to find Pseudo-Genes” § “Parse BLAST Reports with Bioperl” (your own python, or perl, program) § “How to Lie with BLAST Statistics” (“Lies, dammed lies, and statistics”) § … Ref: BLAST by Ian Korf; Mark Yandell; Joseph Bedell, 2003.

Borderline similarity § What to do with matches with E values that are not so impressive? (e. g. , E values > 0. 01, smaller numbers are more significant) § this is the “Twilight Zone” § retest these sequences and look for related hits (not just your original query sequence) § similarity is (is not) transitive: – if A~B and B~C, then A~C

Distant homology detection § Use more data (MSA) § Use advanced approaches (like HMM)

PSI-BLAST § Position Specific Iterated BLAST (PSIBLAST) § Basic idea – Use results from BLAST query to construct a profile matrix (or PSSM) – Search database with profile instead of query sequence § Iterate § Position-specific scoring matrices are an extension of substitution scoring matrices
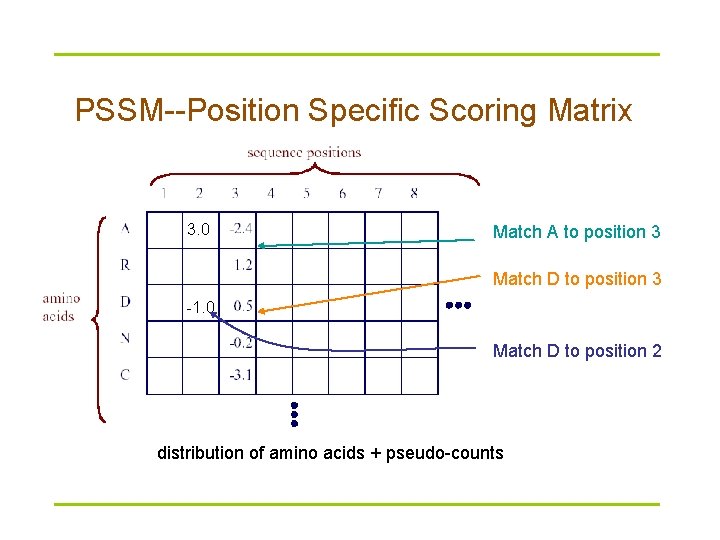
PSSM--Position Specific Scoring Matrix 3. 0 Match A to position 3 Match D to position 3 -1. 0 Match D to position 2 distribution of amino acids + pseudo-counts
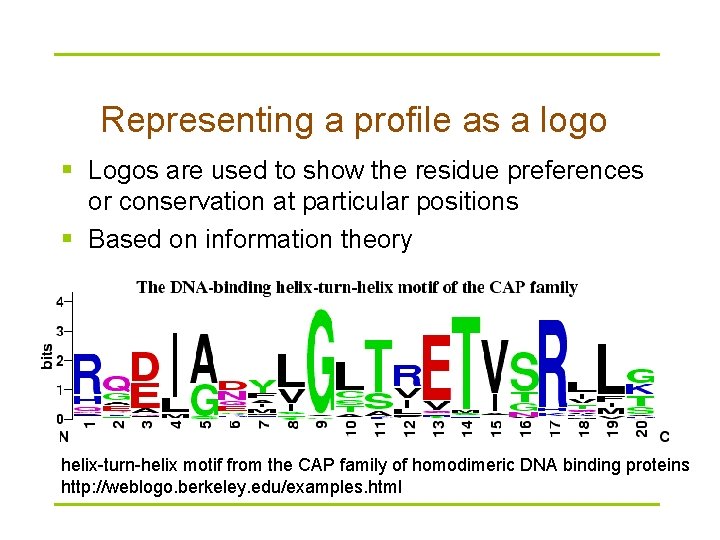
Representing a profile as a logo § Logos are used to show the residue preferences or conservation at particular positions § Based on information theory helix-turn-helix motif from the CAP family of homodimeric DNA binding proteins http: //weblogo. berkeley. edu/examples. html

Profile-profile alignment (alignment of alignments) § Inputs: two MSAs (profiles) § Alignment of two MSAs (similar to pairwise sequence alignment) § Many existing programs § Scoring functions vary – Dot product (FFAS)

FFAS scoring function Dot product of two amino acid “frequency” vectors

Sequence identity scoring zones § >25 -30%: homology zone § 15 -25%: twilight zone § <15%: midnight zone Weak sequence similarity detection is still not solved! And sequence similarity != structural similarity (for proteins) != functional similarity

Readings § Chapter 5 (Pairwise Sequence Alignment and Database Searching) § Box 5. 1 – saving space by throwing away the intermediate calculations § “Time can be saved with a loss of rigor by not calculating the whole matrix” (Figure 5. 18)
- Slides: 38