CS 5263 Bioinformatics Lecture 6 Sequence Alignment Statistics

CS 5263 Bioinformatics Lecture 6: Sequence Alignment Statistics

Roadmap • Last lecture review – Affine gap penalty (more today) – Local sequence alignment – Statistics of substitution matrices • Statistics of alignment scores • Sequence alignment and FSA – Affine gap penalty – More complex models

Seq Alignment Algorithms • Global alignment – – – Basic: Needleman-Wunsch Variants (LCS, overlapping, …) Bounded DP (pruning search space) Linear space (divide-and-conquer) Affine gap penalty • Local Alignment – Basic: Smith-Waterman – All tricks in global alignment applicable • Bounded DP, linear space, affine gap

The local alignment problem Given two strings X = x 1……x. M, Y = y 1……y. N Find substrings x’, y’ whose similarity (optimal global alignment value) is maximum e. g. X = abcxdex Y = xxxcde x y X’ = cxde Y’ = c-de

The Smith-Waterman algorithm Initialization: F(0, j) = F(i, 0) = 0 0 F(i – 1, j) – d Iteration: F(i, j) = max F(i, j – 1) – d F(i – 1, j – 1) + (xi, yj)

The Smith-Waterman algorithm Termination: 1. If we want the best local alignment… FOPT = maxi, j F(i, j) 2. If we want all local alignments scoring > t For all i, j find F(i, j) > t, and trace back

Analysis • Time: – O(MN) for finding the best alignment – Depending on the number of sub-opt alignments • Memory: – O(MN) – O(M+N) possible

The statistics of alignment Where does (xi, yj) come from? Are two aligned sequences actually related?

Protein substitution matrix • : score to align amino acid s against t • ps, pt, frequency of s and t in database Scaling factor Log odds ratio • qst: the frequency that s is aligned to t in real homologous sequences

BLOSUM matrices • ps, pt, qst estimated from trusted alignments in the BLOCKS database • Eliminate near-identical sequences – BLOSUM-N: constructed from sequences where identity between any pair of sequences is less than N% – BLOSUM-62: good for most purposes 45 62 90 Weak homology Strong homology

DNA substitution matrix • Given the percent identity you would like to detect and some assumptions • You can get the substitution matrix by some calculation

Example • Assume p. A = p. C = p. T = p. G = 0. 25 • We want 88% identity • q. AA = q. CC = q. TT = q. GG = 0. 22 • The rest = 0. 12/12 = 0. 01 A A C 5 -7 -7 -7 C -7 5 G -7 -7 T G T -7 -7 5

Arbitrary substitution matrix • Even arbitrary substitution matrix has meaning • Better know what you are doing • Solve a polynomial function to obtain the scaling factor • Calculate target frequency qst • Calculate target percent identity
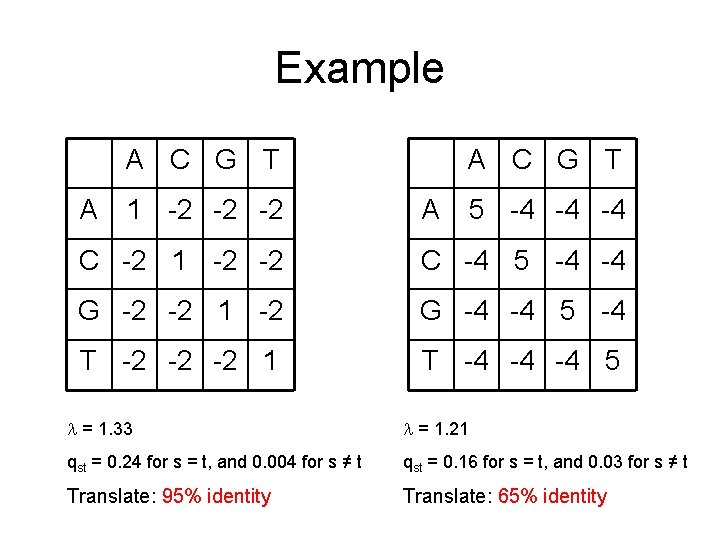
Example A C G T A 1 -2 -2 -2 A C G T A 5 -4 -4 -4 C -2 1 -2 -2 C -4 5 -4 -4 G -2 -2 1 -2 G -4 -4 5 -4 T -2 -2 -2 1 T -4 -4 -4 5 = 1. 33 = 1. 21 qst = 0. 24 for s = t, and 0. 004 for s ≠ t qst = 0. 16 for s = t, and 0. 03 for s ≠ t Translate: 95% identity Translate: 65% identity

Today • Significance of alignment score • Sequence alignment and FSA

Statistics of Alignment Scores • Q: How do we assess whether an alignment provides good evidence for homology? – Is a score 82 good? What about 180? • A: determine how likely it is that such an alignment score would result from chance

• Most of theory applies to local alignment • For global alignment, your best bet is to do Monte-Carlo simulation – Randomly shuffle your sequences before alignment – What’s the chance you can get a score as high as the real alignment?

• Procedure to estimate the significance of a global alignment – Given sequence X, Y – Global alignment score = S – Randomly shuffle sequence X (or Y) N times, obtain X 1, X 2, …, XN – Align each Xi with Y, let the score be Si – Plot the distribution of Si, and see where the real S locates

Mouse HEXA Human HEXA Score = 732 …………………………
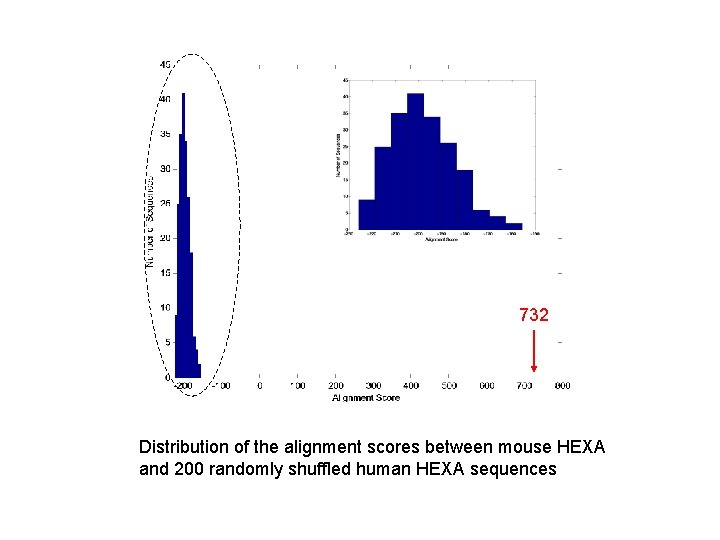
732 Distribution of the alignment scores between mouse HEXA and 200 randomly shuffled human HEXA sequences

Human HEXA Fly HEXO 1 Score = -74
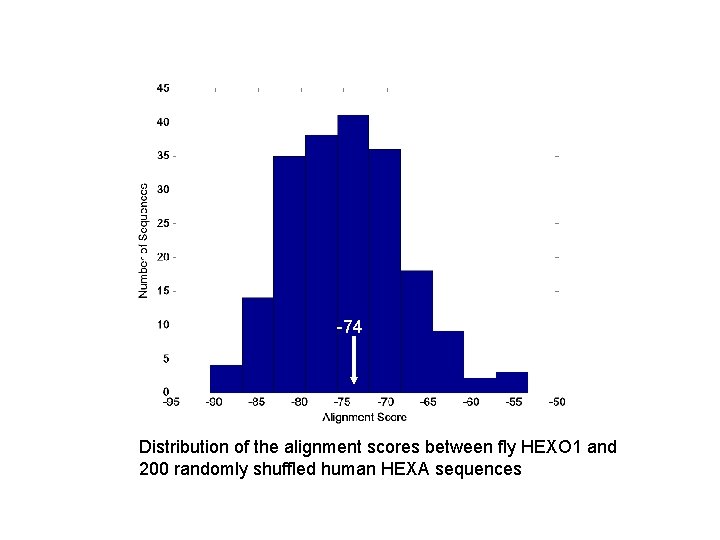
-74 Distribution of the alignment scores between fly HEXO 1 and 200 randomly shuffled human HEXA sequences

P-value of alignment • p-value – The probability that the alignment score can be obtained from aligning random sequences – Small p-value means the score is unlikely to happen by chance • A p-value 0. 05 means you are 95% sure that the result is significant.

What p-value is significant? • The most common thresholds are 0. 01 and 0. 05. • Is 95% enough? It depends on the cost associated with making a mistake. • Examples of costs: – Doing expensive wet lab validation. – Making clinical treatment decisions. – Misleading the scientific community. • Most sequence analysis uses more stringent thresholds because the p-values are not very accurate.
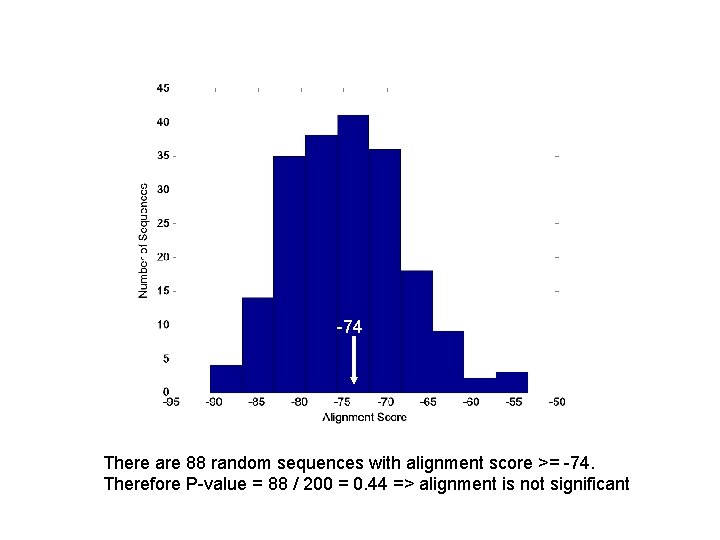
-74 There are 88 random sequences with alignment score >= -74. Therefore P-value = 88 / 200 = 0. 44 => alignment is not significant
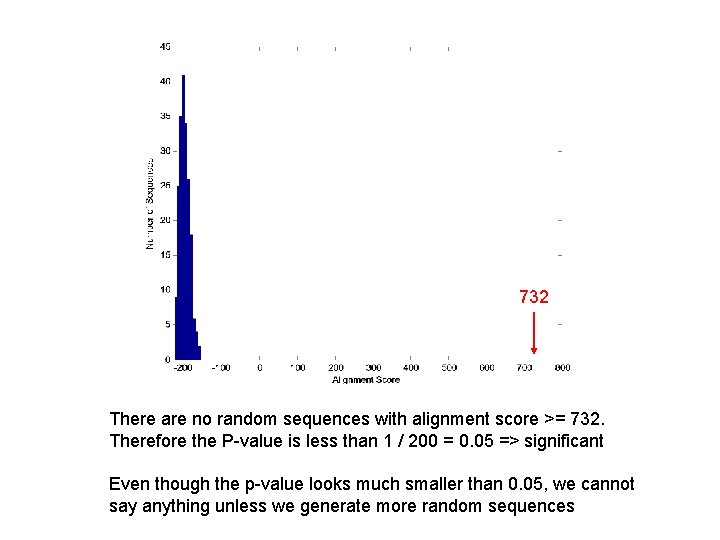
732 There are no random sequences with alignment score >= 732. Therefore the P-value is less than 1 / 200 = 0. 05 => significant Even though the p-value looks much smaller than 0. 05, we cannot say anything unless we generate more random sequences

Drawbacks • Monte-Carlo may take long time • Cannot accurately estimate p-value if p is small • To get 10 -5 p-value, have to align 105 random sequences – Unless we can fit a distribution • Such distribution may not be generalizable • No theory exists for global alignment score distribution

Statistics for local alignment • Theory much more elegant • Score for ungapped local alignment follows extreme value distribution (Gumbel distribution) This distribution is characterized by a larger tail on the right.
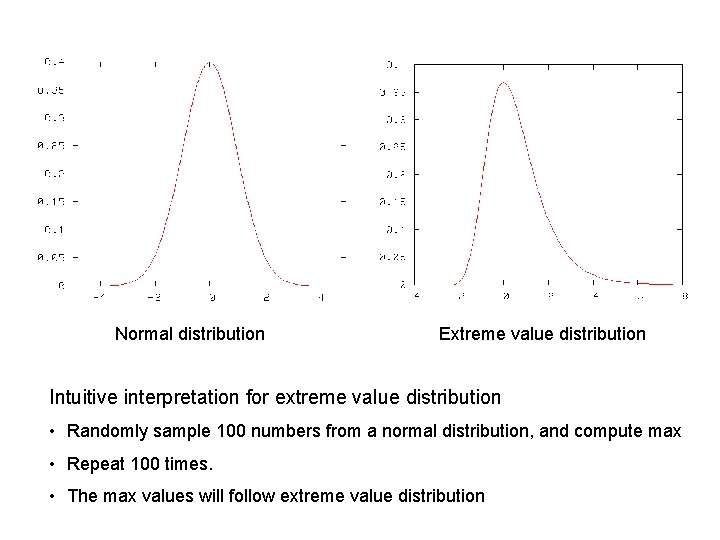
Normal distribution Extreme value distribution Intuitive interpretation for extreme value distribution • Randomly sample 100 numbers from a normal distribution, and compute max • Repeat 100 times. • The max values will follow extreme value distribution
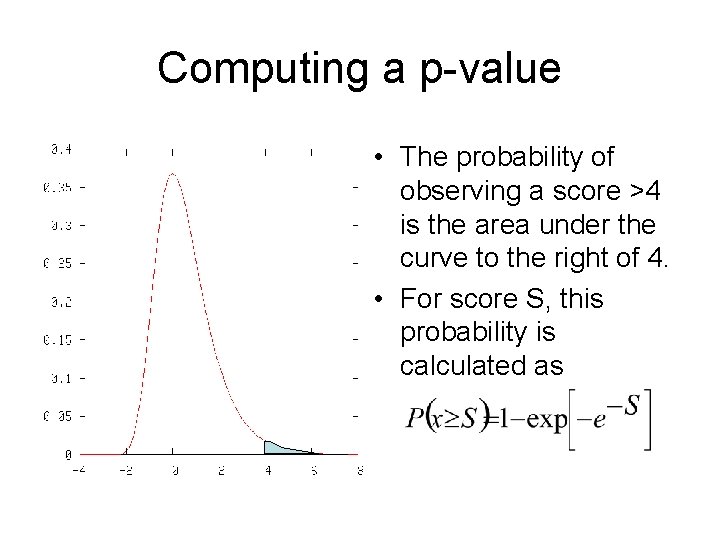
Computing a p-value • The probability of observing a score >4 is the area under the curve to the right of 4. • For score S, this probability is calculated as

Statistics for local alignment • How does this apply to sequence alignment? • Given two unrelated sequences of lengths M, N • Expected number of local alignments with score >= S can be calculated by – – E(S) = KMN exp[- S] Known as E-value : scaling factor as computed in last lecture K: empirical parameter ~ 0. 1 • Depend on sequence composition and substitution matrix

P-value for alignment score • P-value for a local alignment score S when P is small.

Example • You are aligning two sequences, each has 1000 bases • m = 1, s = -1, d = -inf (ungapped alignment) • You obtain a score 20 • Is this score significant?

• • • = ln 3 = 1. 1 E(S) = K MN exp{- S} E(20) = 0. 1 * 1000 * 3 -20 = 3 x 10 -5 P-value = 3 x 10 -5 << 0. 05 The alignment is significant
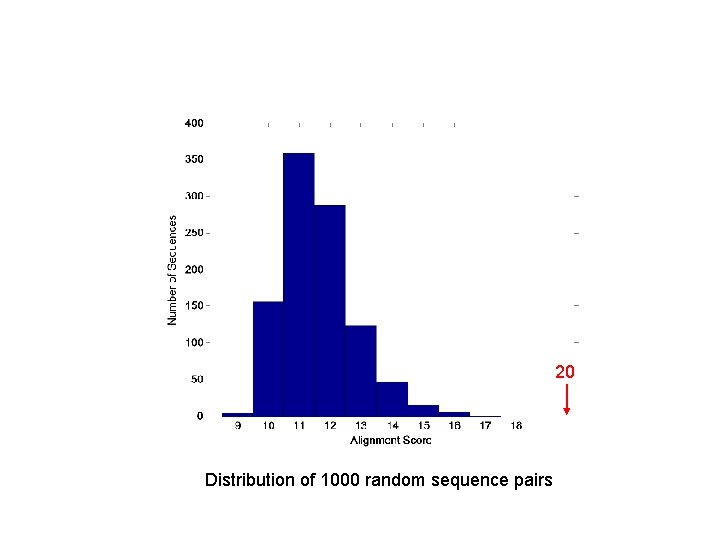
20 Distribution of 1000 random sequence pairs

Multiple-testing problem • What if you are searching a 1000 -base sequence against a database of 106 sequences (average length 1000 bases)? • How significant is a score 20 now? • You are essentially comparing 1000 bases with 1000 x 106 = 109 bases (ignore edge effect) • E(20) = 0. 1 * 1000 * 109 * 3 -20 = 30 • By chance we would expect to see 30 matches • P-value = 1 – e-30 = 0. 99999 • Not significant at all

Statistics for gapped local alignment • Theory not well developed • Extreme value distribution works well empirically • Need to estimate K and empirically – Given the database and substitution matrix, generate some random sequence pairs – Do local alignment – Fit an extreme value distribution to obtain K and

More on sequence alignment and FSA
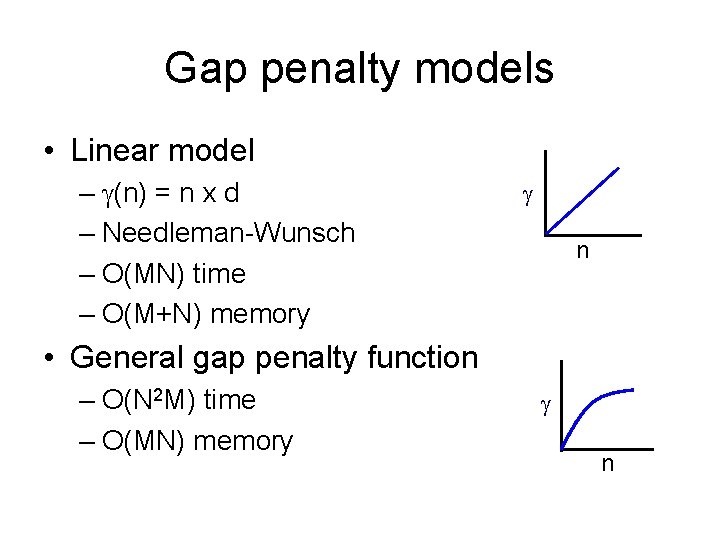
Gap penalty models • Linear model – (n) = n x d – Needleman-Wunsch – O(MN) time – O(M+N) memory n • General gap penalty function – O(N 2 M) time – O(MN) memory n
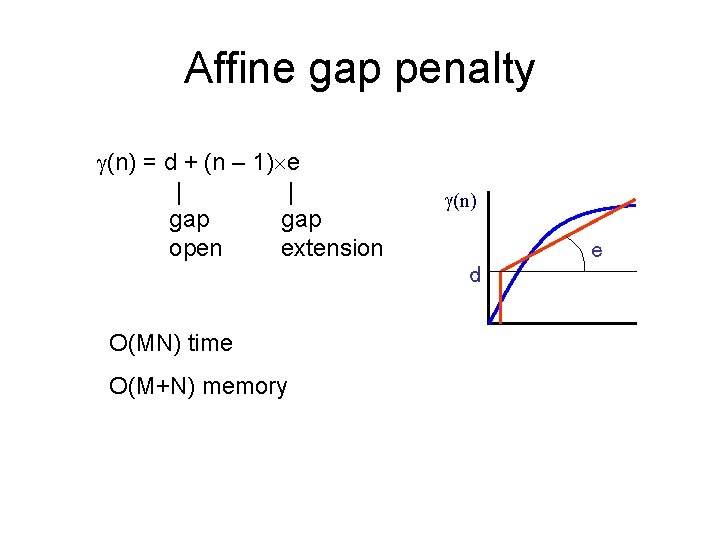
Affine gap penalty (n) = d + (n – 1) e | | gap open extension (n) e d O(MN) time O(M+N) memory
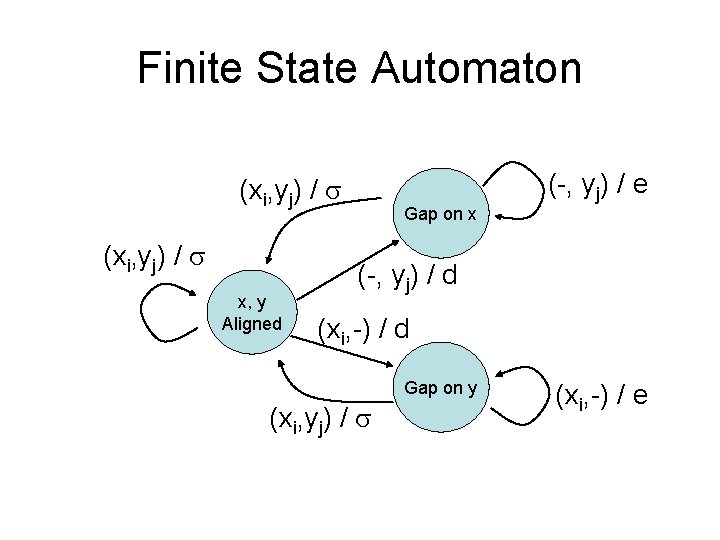
Finite State Automaton (xi, yj) / x, y Aligned Gap on x (-, yj) / e (-, yj) / d (xi, -) / d Gap on y (xi, yj) / (xi, -) / e
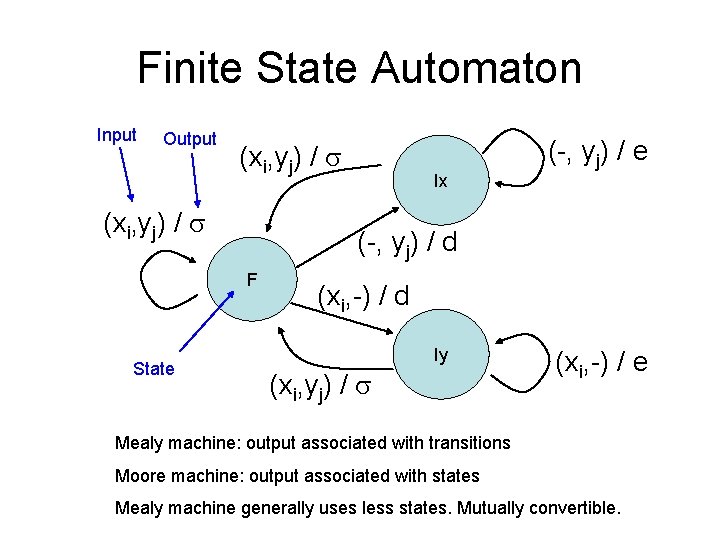
Finite State Automaton Input Output (xi, yj) / (-, yj) / d F State Ix (-, yj) / e (xi, -) / d Iy (xi, yj) / (xi, -) / e Mealy machine: output associated with transitions Moore machine: output associated with states Mealy machine generally uses less states. Mutually convertible.

Mealy machine • A Mealy machine is a 6 -tuple, (S, S 0, Σ, Λ, T, G), consisting of the following: – a finite set of states (S) – a start state (also called initial state) S 0 which is an element of (S) – a finite set called the input alphabet (Σ) – a finite set called the output alphabet (Λ) – a transition function (T : S × Σ → S) – an output function (G : S × Σ → Λ)
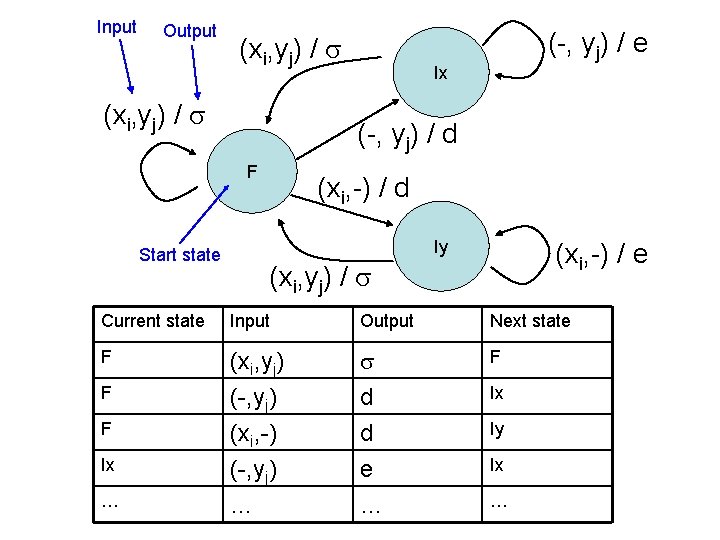
Input Output (xi, yj) / Ix (-, yj) / d F Start state (-, yj) / e (xi, -) / d Iy (xi, -) / e (xi, yj) / Current state Input Output Next state F (xi, yj) (-, yj) (xi, -) d d F (-, yj) … e … Ix F F Ix … Ix Iy …
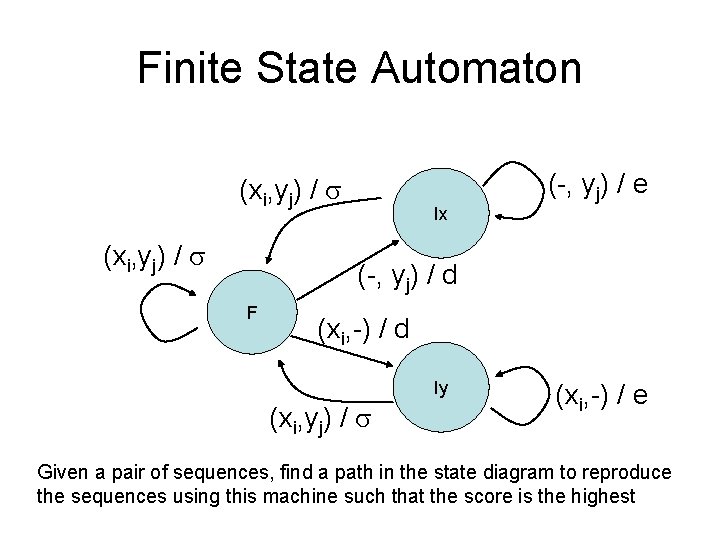
Finite State Automaton (xi, yj) / Ix (-, yj) / e (-, yj) / d F (xi, -) / d Iy (xi, yj) / (xi, -) / e Given a pair of sequences, find a path in the state diagram to reproduce the sequences using this machine such that the score is the highest
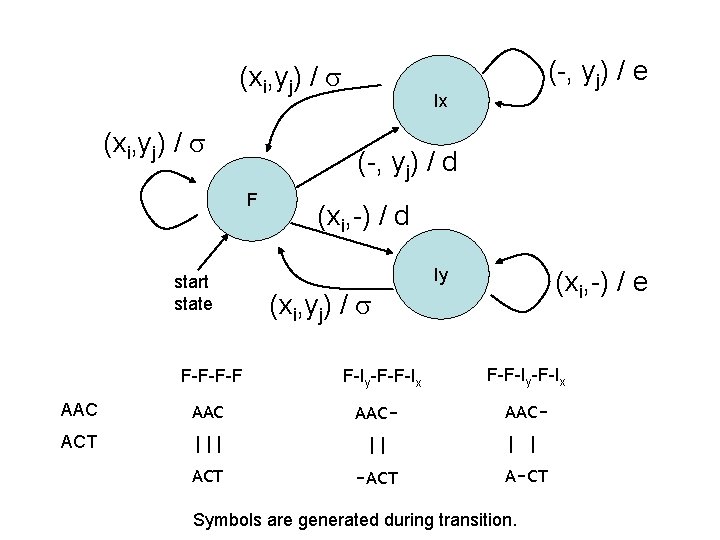
(xi, yj) / F-F-F-F Ix (-, yj) / d F start state (-, yj) / e (xi, -) / d Iy (xi, -) / e (xi, yj) / F-Iy-F-F-Ix AAC AAC- ACT ||| || ACT -ACT F-F-Iy-F-Ix AAC| | A-CT Symbols are generated during transition.
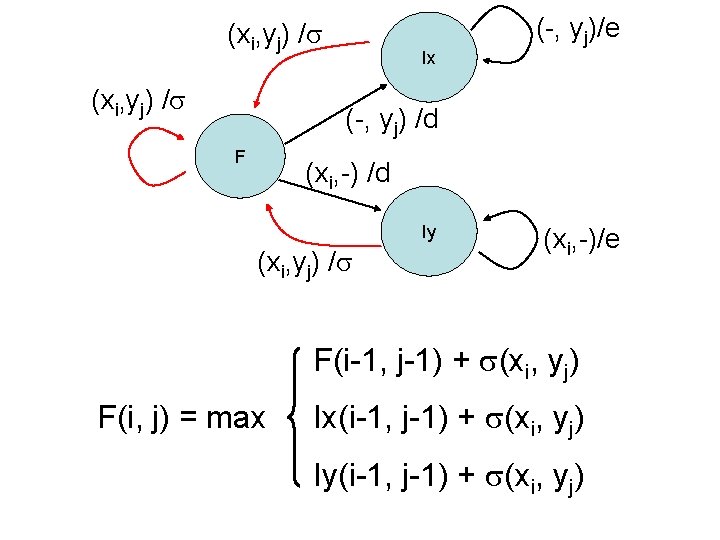
(xi, yj) / Ix (-, yj)/e (-, yj) /d F (xi, -) /d Iy (xi, yj) / (xi, -)/e F(i-1, j-1) + (xi, yj) F(i, j) = max Ix(i-1, j-1) + (xi, yj) Iy(i-1, j-1) + (xi, yj)
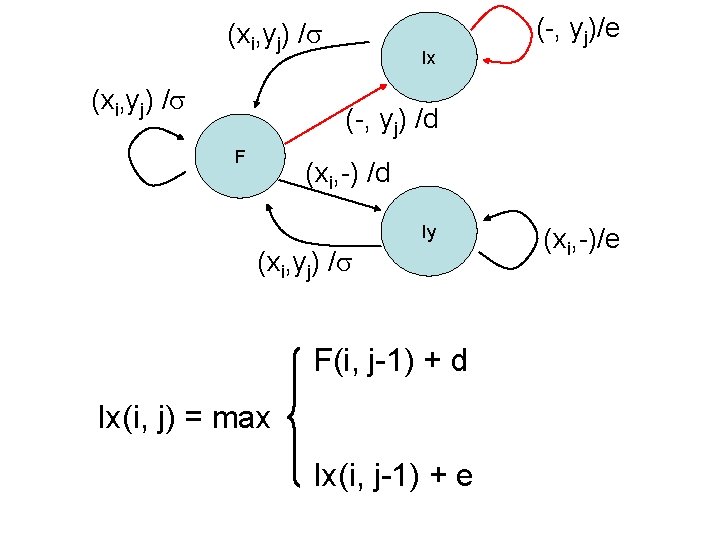
(xi, yj) / Ix (-, yj)/e (-, yj) /d F (xi, -) /d Iy (xi, yj) / F(i, j-1) + d Ix(i, j) = max Ix(i, j-1) + e (xi, -)/e
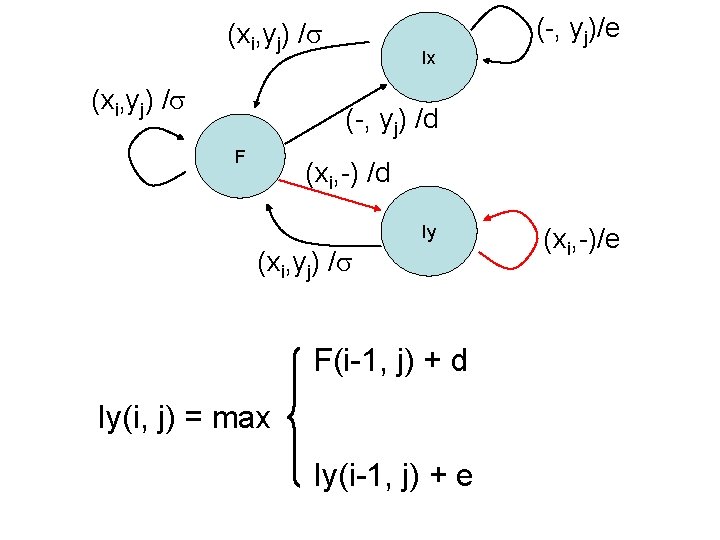
(xi, yj) / Ix (-, yj)/e (-, yj) /d F (xi, -) /d Iy (xi, yj) / F(i-1, j) + d Iy(i, j) = max Iy(i-1, j) + e (xi, -)/e
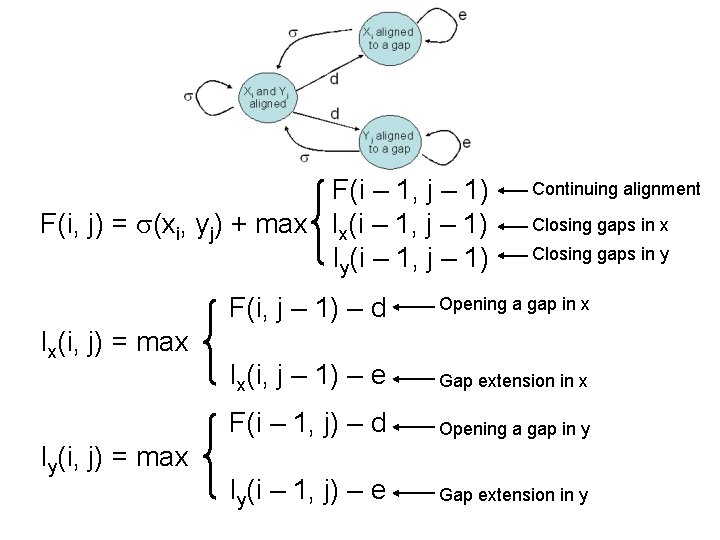
F(i – 1, j – 1) F(i, j) = (xi, yj) + max Ix(i – 1, j – 1) Iy(i – 1, j – 1) Ix(i, j) = max Iy(i, j) = max Continuing alignment Closing gaps in x Closing gaps in y F(i, j – 1) – d Opening a gap in x Ix(i, j – 1) – e Gap extension in x F(i – 1, j) – d Opening a gap in y Iy(i – 1, j) – e Gap extension in y
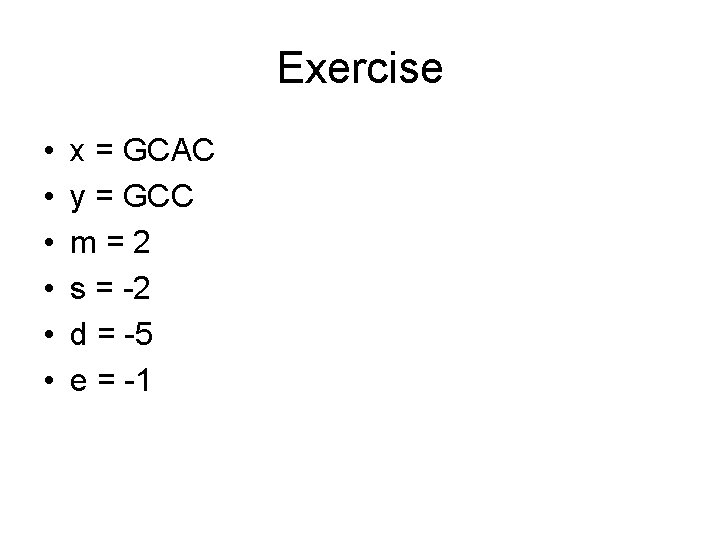
Exercise • • • x = GCAC y = GCC m = 2 s = -2 d = -5 e = -1
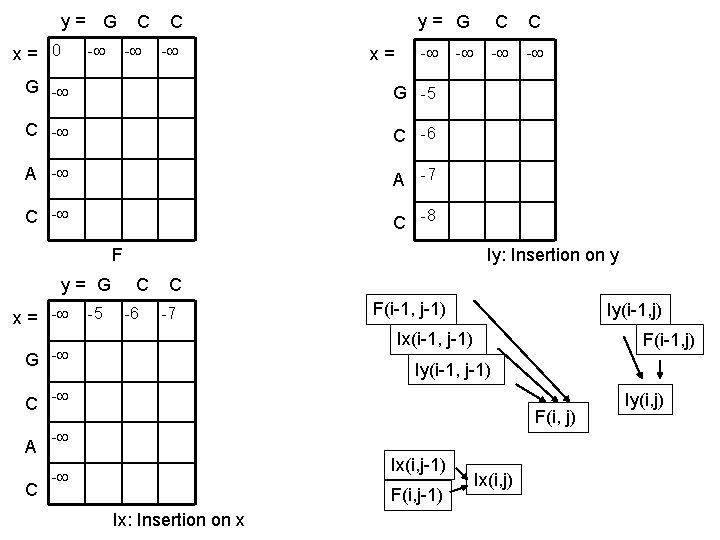
y = G C C x = 0 - - - y = G C C x = - G -5 C -6 A -7 C -8 - - F - Iy: Insertion on y y = G C C x = - -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - Iy(i-1, j-1) C - A C F(i-1, j) F(i, j) - Ix(i, j-1) - F(i, j-1) Ix: Insertion on x Ix(i, j) Iy(i, j)
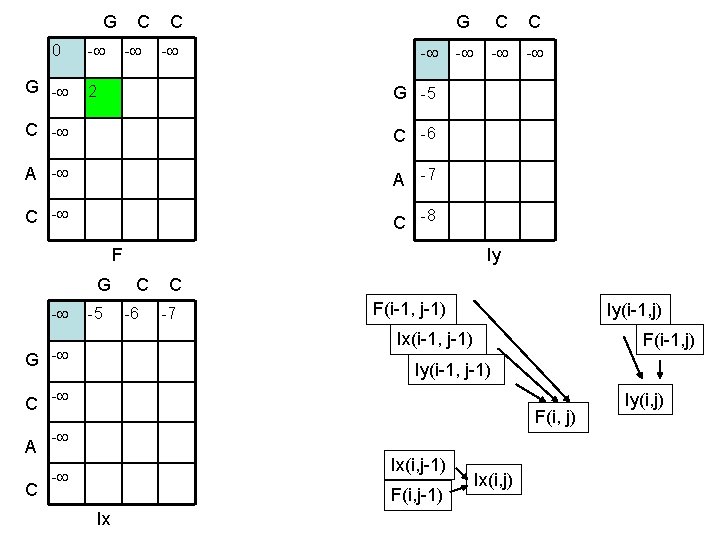
G C C 0 G - - 2 G C C - - G -5 C -6 A -7 C -8 F Iy G C C - -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - Iy(i-1, j-1) C - A C F(i-1, j) F(i, j) - Ix(i, j-1) - F(i, j-1) Ix Ix(i, j) Iy(i, j)
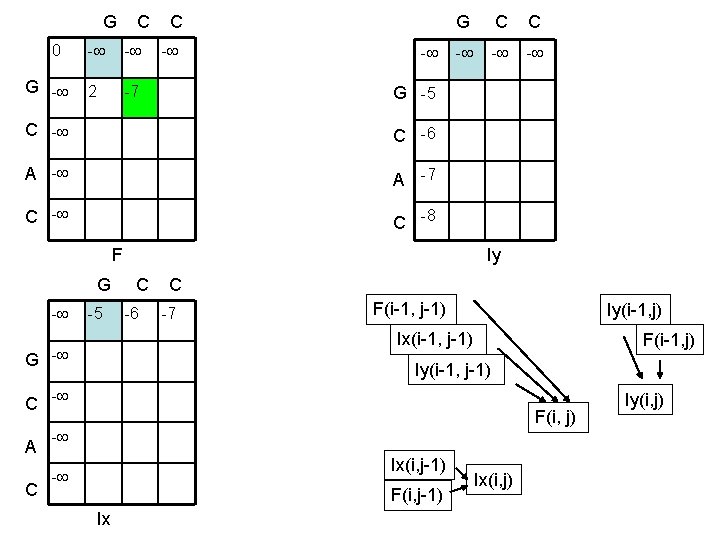
G C C 0 G - - - 2 -7 - G C C - - G -5 C -6 A -7 C -8 F Iy G C C - -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - Iy(i-1, j-1) C - A C F(i-1, j) F(i, j) - Ix(i, j-1) - F(i, j-1) Ix Ix(i, j) Iy(i, j)
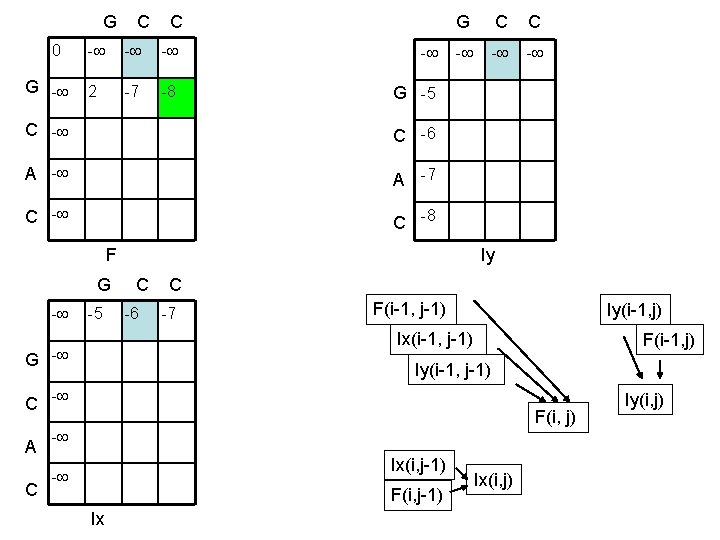
G C C 0 G - G C C - - 2 -7 -8 G -5 C -6 A -7 C -8 - - F - Iy G C C - -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - Iy(i-1, j-1) C - A C F(i-1, j) F(i, j) - Ix(i, j-1) - F(i, j-1) Ix Ix(i, j) Iy(i, j)
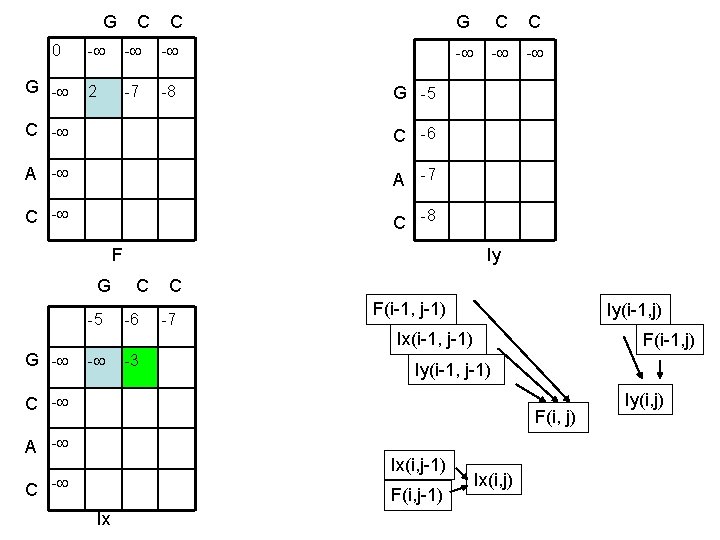
G C C 0 G - - 2 -7 -8 G C C - - - G -5 C -6 A -7 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 F(i-1, j) Iy(i-1, j-1) C - F(i, j) A - Ix(i, j-1) C - F(i, j-1) Ix Ix(i, j) Iy(i, j)
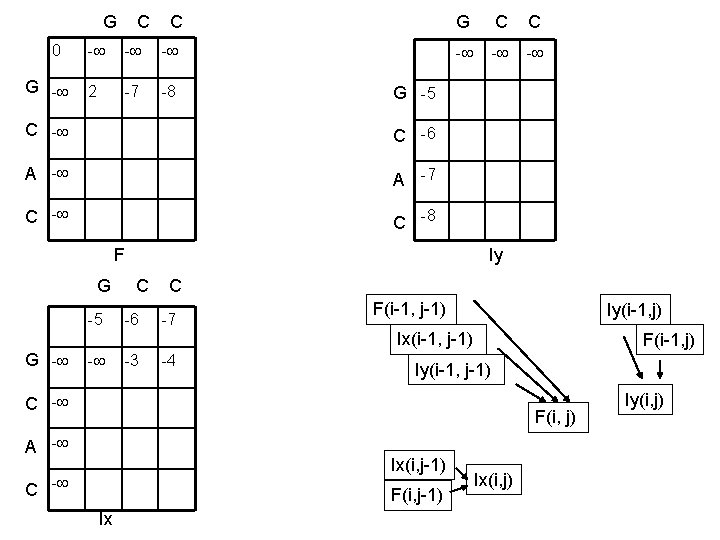
G C C 0 G - - 2 -7 -8 G C C - - - G -5 C -6 A -7 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 -4 F(i-1, j) Iy(i-1, j-1) C - F(i, j) A - Ix(i, j-1) C - F(i, j-1) Ix Ix(i, j) Iy(i, j)
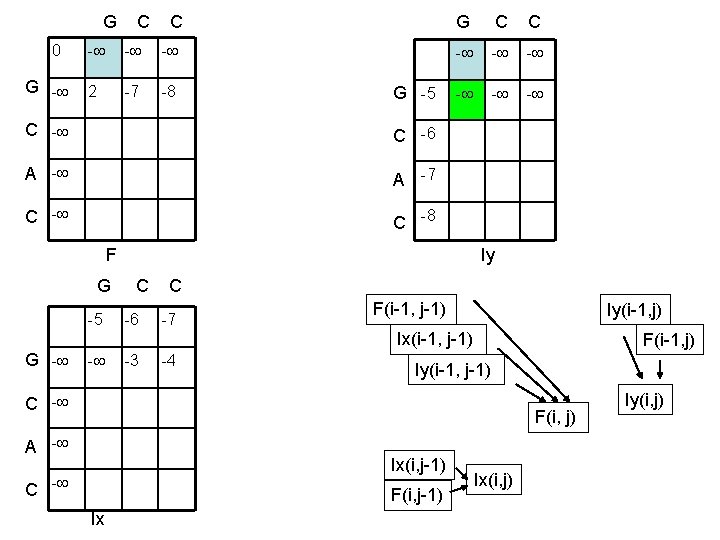
G C C 0 G - - 2 -7 -8 G C C G -5 C -6 A -7 C -8 - - - F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 -4 F(i-1, j) Iy(i-1, j-1) C - F(i, j) A - Ix(i, j-1) C - F(i, j-1) Ix Ix(i, j) Iy(i, j)
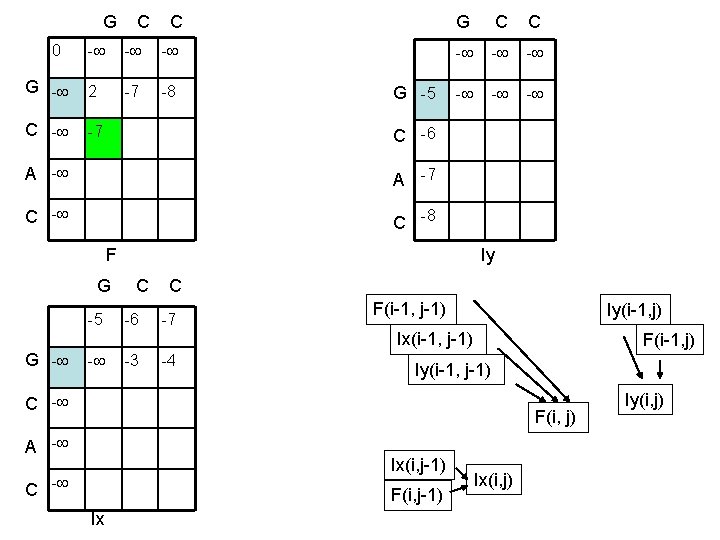
G C C 0 - - - G - 2 -7 -8 C - -7 G C C G -5 - - - C -6 A -7 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 -4 F(i-1, j) Iy(i-1, j-1) C - F(i, j) A - Ix(i, j-1) C - F(i, j-1) Ix Ix(i, j) Iy(i, j)
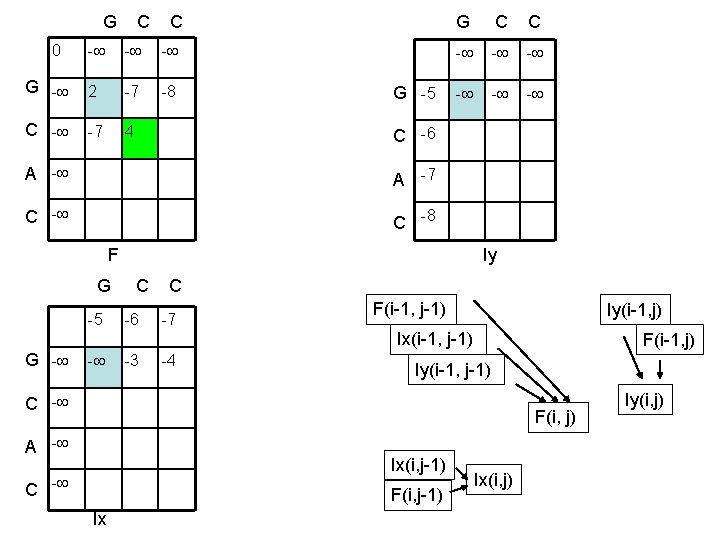
G C C 0 - - - G - 2 -7 -8 C - -7 4 G C C G -5 - - - C -6 A -7 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 -4 F(i-1, j) Iy(i-1, j-1) C - F(i, j) A - Ix(i, j-1) C - F(i, j-1) Ix Ix(i, j) Iy(i, j)
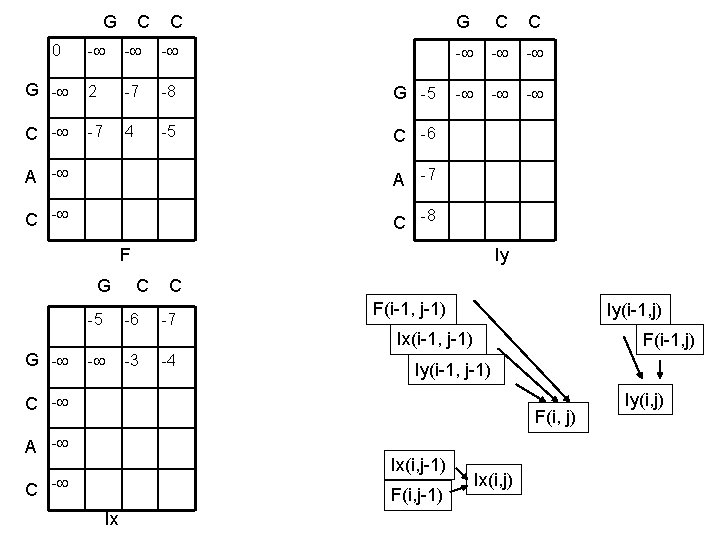
G C C 0 G C C - - - G - 2 -7 -8 G -5 C - -7 4 -5 C -6 A -7 C -8 - - - F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 -4 F(i-1, j) Iy(i-1, j-1) C - F(i, j) A - Ix(i, j-1) C - F(i, j-1) Ix Ix(i, j) Iy(i, j)
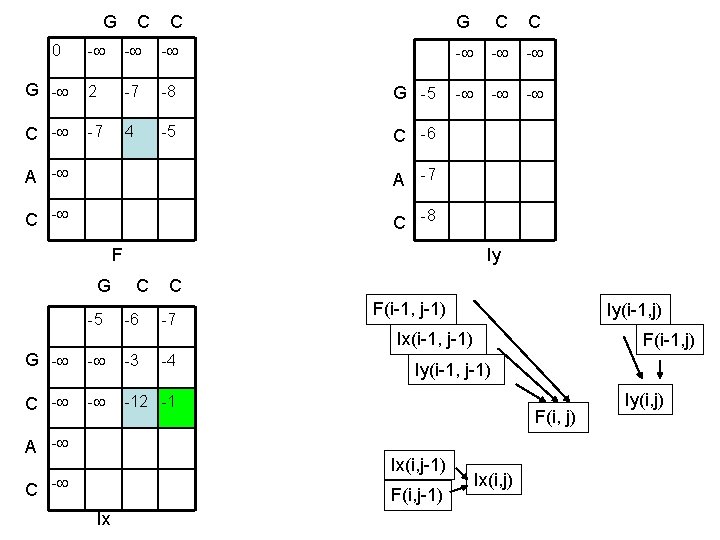
G C C 0 G C C - - - G - 2 -7 -8 G -5 C - -7 4 -5 C -6 A -7 C -8 - - - F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - -4 Iy(i-1, j-1) F(i, j) Ix(i, j-1) C - F(i, j-1) Ix F(i-1, j) Ix(i, j) Iy(i, j)
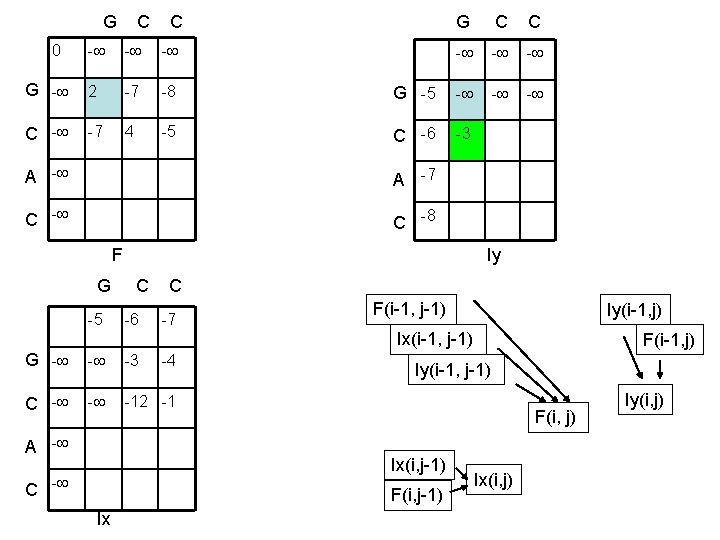
G C C 0 - - - G - 2 -7 -8 C - -7 4 -5 G C C - - - G -5 - - - C -6 -3 A -7 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - -4 Iy(i-1, j-1) F(i, j) Ix(i, j-1) C - F(i, j-1) Ix F(i-1, j) Ix(i, j) Iy(i, j)
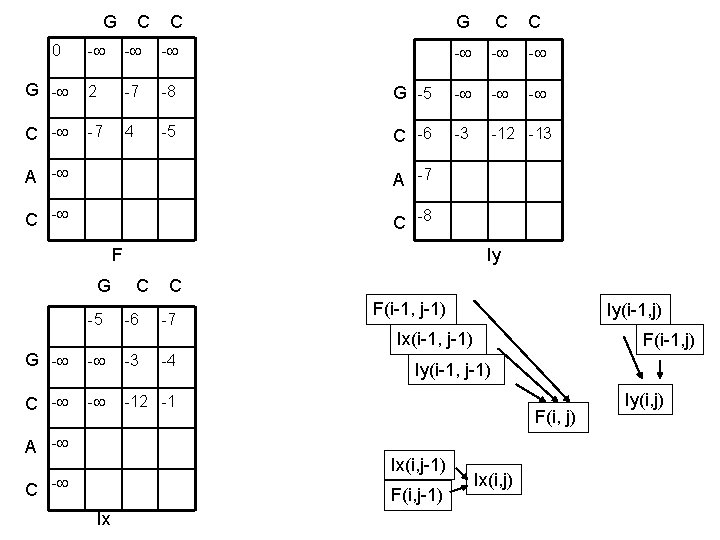
G C C 0 - - - G - 2 -7 -8 C - -7 4 -5 G C C - - - G -5 - - - C -6 -3 -12 -13 A -7 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - -4 Iy(i-1, j-1) F(i, j) Ix(i, j-1) C - F(i, j-1) Ix F(i-1, j) Ix(i, j) Iy(i, j)
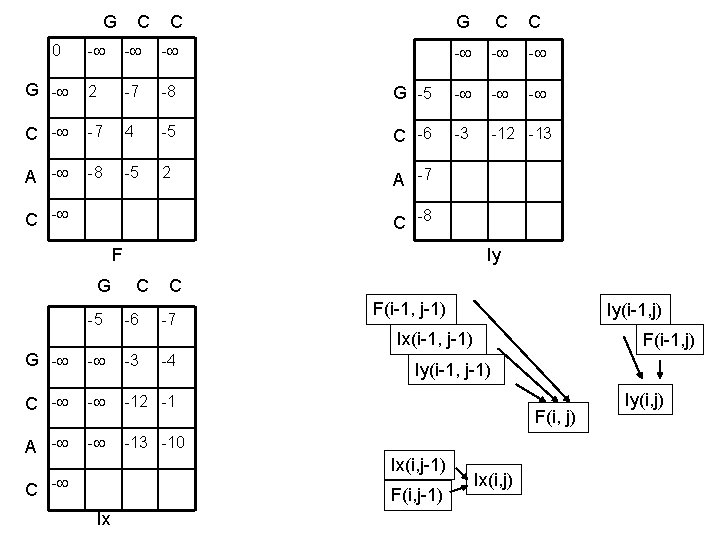
G C C 0 - - - G - 2 -7 -8 C - -7 4 A - -8 -5 G C C - - - G -5 - -5 C -6 -3 -12 -13 2 A -7 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - - -13 -10 -4 Iy(i-1, j-1) F(i, j) Ix(i, j-1) C - F(i, j-1) Ix F(i-1, j) Ix(i, j) Iy(i, j)
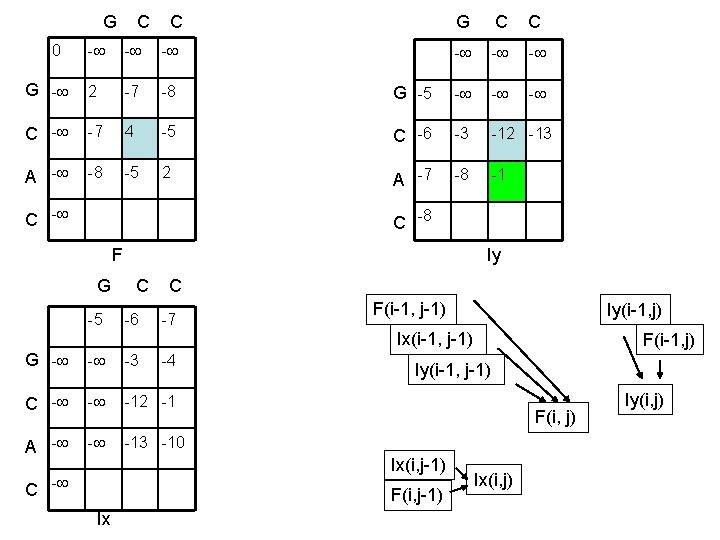
G C C 0 - - - G - 2 -7 -8 C - -7 4 A - -8 -5 G C C - - - G -5 - -5 C -6 -3 -12 -13 2 A -7 -8 -1 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - - -13 -10 -4 Iy(i-1, j-1) F(i, j) Ix(i, j-1) C - F(i, j-1) Ix F(i-1, j) Ix(i, j) Iy(i, j)
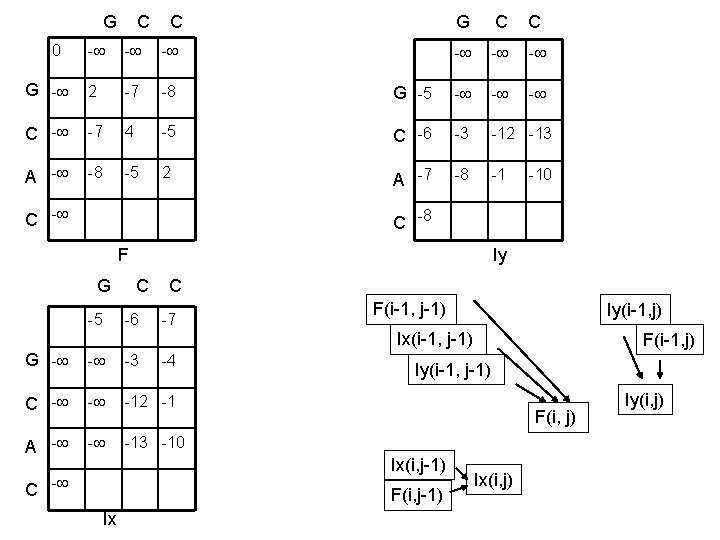
G C C 0 - - - G - 2 -7 -8 C - -7 4 A - -8 -5 G C C - - - G -5 - -5 C -6 -3 -12 -13 2 A -7 -8 -1 C - -10 C -8 F Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - - -13 -10 -4 Iy(i-1, j-1) F(i, j) Ix(i, j-1) C - F(i, j-1) Ix F(i-1, j) Ix(i, j) Iy(i, j)
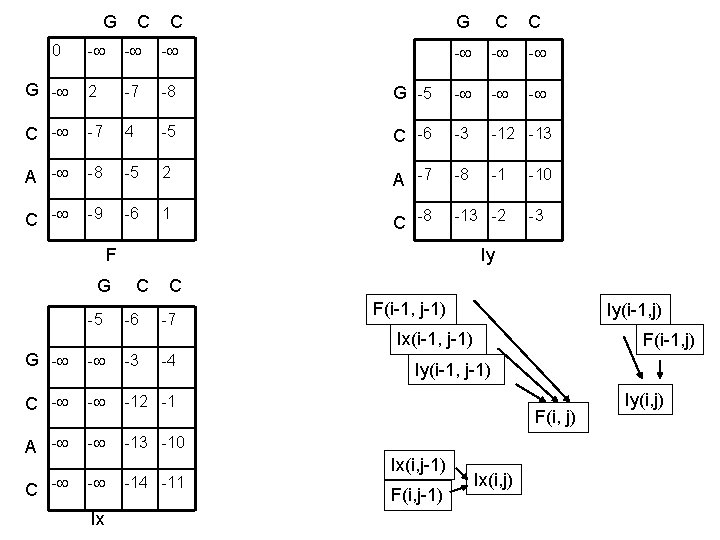
G C C 0 - - - G - 2 -7 -8 C - -7 4 A - -8 C - -9 G C C - - - G -5 - -5 C -6 -3 -12 -13 -5 2 A -7 -8 -1 -6 1 C -8 -13 -2 F -10 -3 Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - - -13 -10 C - - Ix -4 -11 F(i-1, j) Iy(i-1, j-1) F(i, j) Ix(i, j-1) F(i, j-1) Ix(i, j) Iy(i, j)
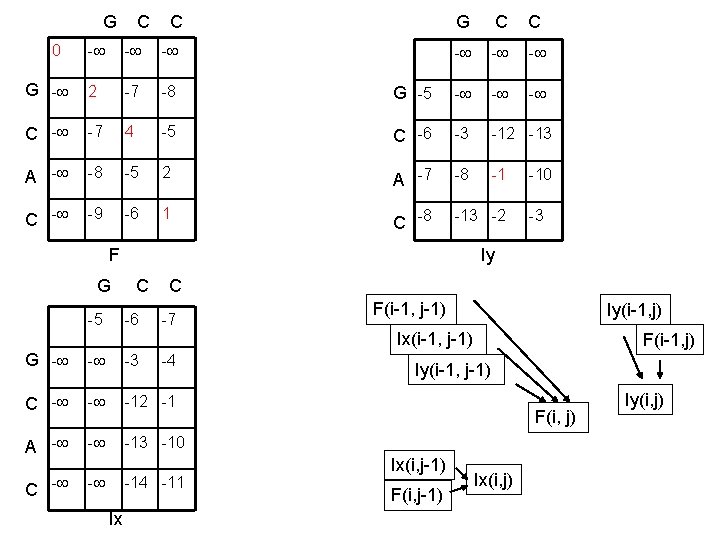
G C C 0 - - - G - 2 -7 -8 C - -7 4 A - -8 C - -9 G C C - - - G -5 - -5 C -6 -3 -12 -13 -5 2 A -7 -8 -1 -6 1 C -8 -13 -2 F -10 -3 Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - - -13 -10 C - - -4 -11 Ix F(i-1, j) Iy(i-1, j-1) F(i, j) Ix(i, j-1) F(i, j-1) Ix(i, j) Iy(i, j)
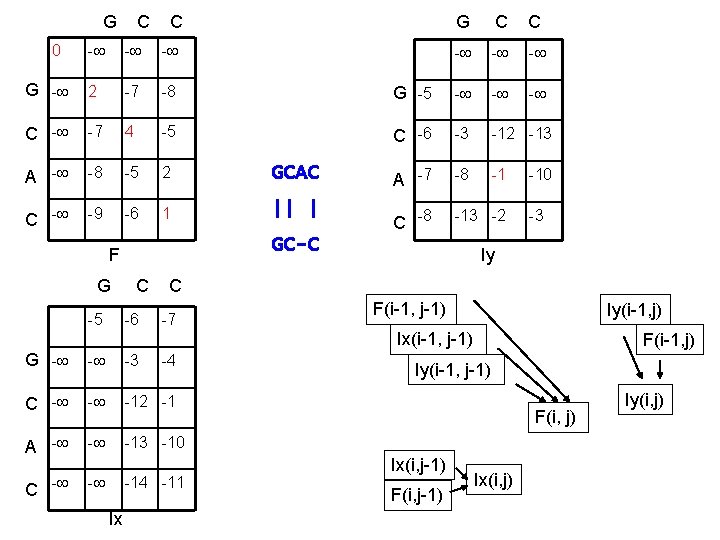
G C C 0 - - - G - 2 -7 -8 C - -7 4 -5 A - -8 -5 2 C - -9 -6 1 G C C - - - G -5 - - - C -6 -3 -12 -13 GCAC A -7 -8 -1 || | C -8 -13 -2 GC-C F -10 -3 Iy G C C -5 -6 -7 F(i-1, j-1) Iy(i-1, j) Ix(i-1, j-1) G - - -3 C - - -12 -1 A - - -13 -10 C - - -4 -11 Ix F(i-1, j) Iy(i-1, j-1) F(i, j) Ix(i, j-1) F(i, j-1) Ix(i, j) Iy(i, j)

Exercising FSA • How do you make an FSA for the Needleman-Wunsch algorithm?
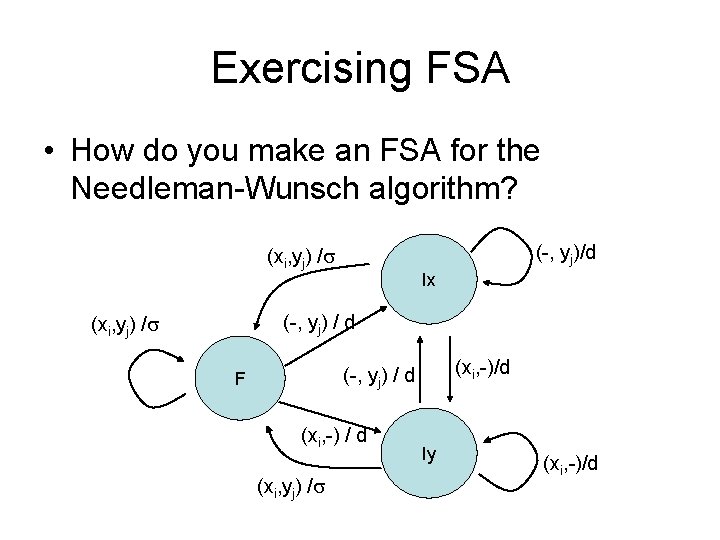
Exercising FSA • How do you make an FSA for the Needleman-Wunsch algorithm? (-, yj)/d (xi, yj) / Ix (-, yj) / d (xi, yj) / (xi, -)/d (-, yj) / d F (xi, -) / d (xi, yj) / Iy (xi, -)/d
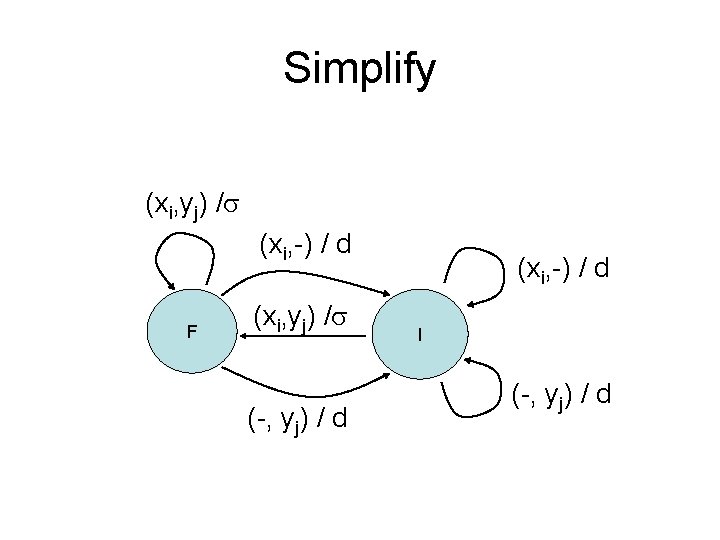
Simplify (xi, yj) / (xi, -) / d F (xi, yj) / (-, yj) / d (xi, -) / d I (-, yj) / d
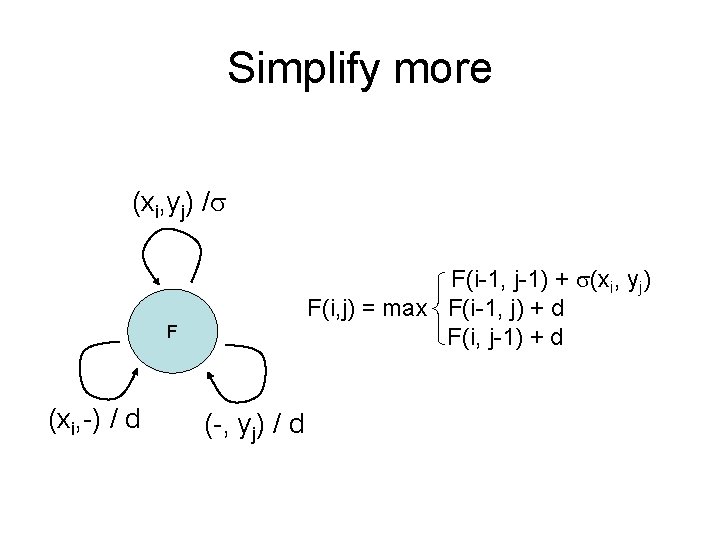
Simplify more (xi, yj) / F(i-1, j-1) + (xi, yj) F(i, j) = max F(i-1, j) + d F(i, j-1) + d F (xi, -) / d (-, yj) / d

A more difficult alignment problem • (A gene finder indeed!) • X is a genomic sequence (DNA) – X encodes a gene – May contain introns • Y is an ORF from another species – Contains only exons • We want to compare X against Y – Conservation is on the level of amino acids
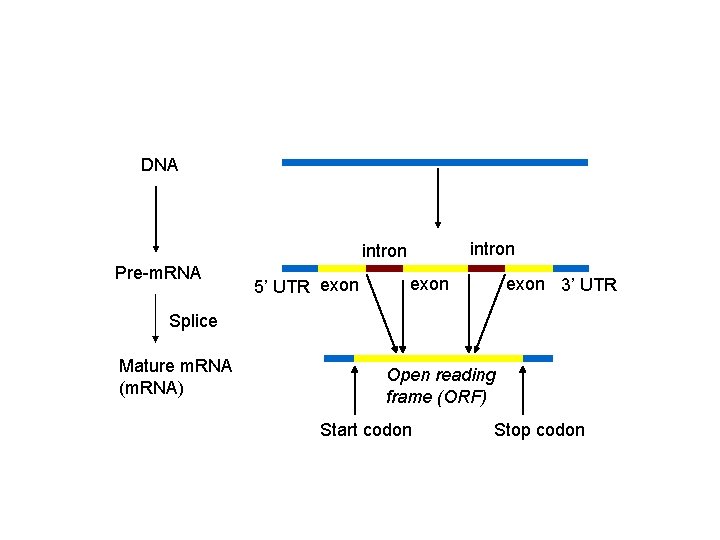
DNA intron Pre-m. RNA 5’ UTR exon 3’ UTR Splice Mature m. RNA (m. RNA) Open reading frame (ORF) Start codon Stop codon
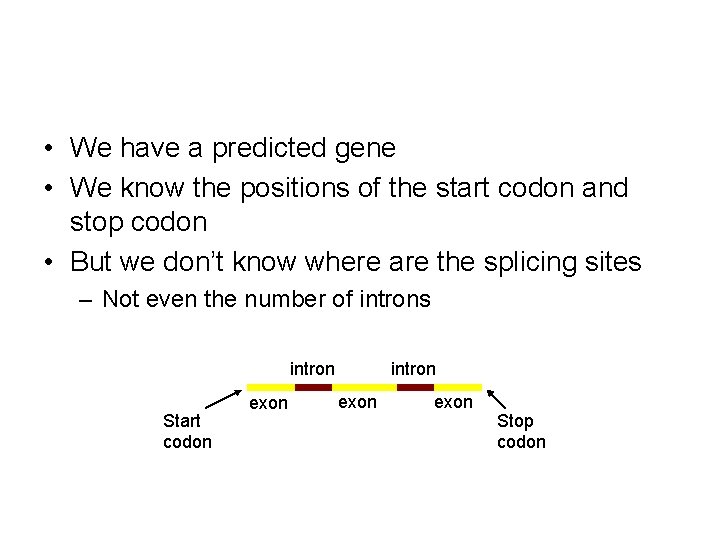
• We have a predicted gene • We know the positions of the start codon and stop codon • But we don’t know where are the splicing sites – Not even the number of introns intron Start codon exon intron exon Stop codon
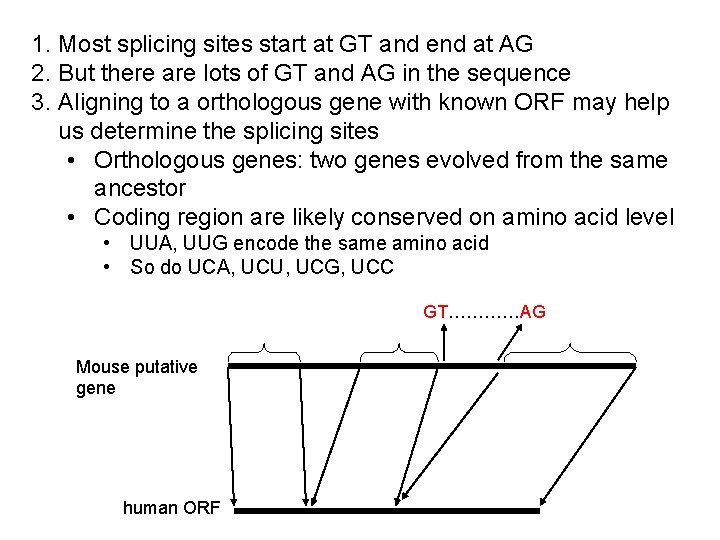
1. Most splicing sites start at GT and end at AG 2. But there are lots of GT and AG in the sequence 3. Aligning to a orthologous gene with known ORF may help us determine the splicing sites • Orthologous genes: two genes evolved from the same ancestor • Coding region are likely conserved on amino acid level • UUA, UUG encode the same amino acid • So do UCA, UCU, UCG, UCC GT…………AG Mouse putative gene human ORF
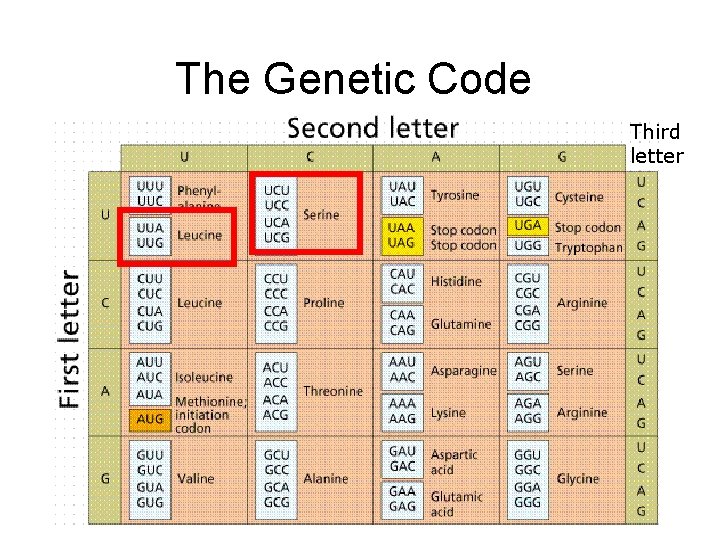
The Genetic Code Third letter
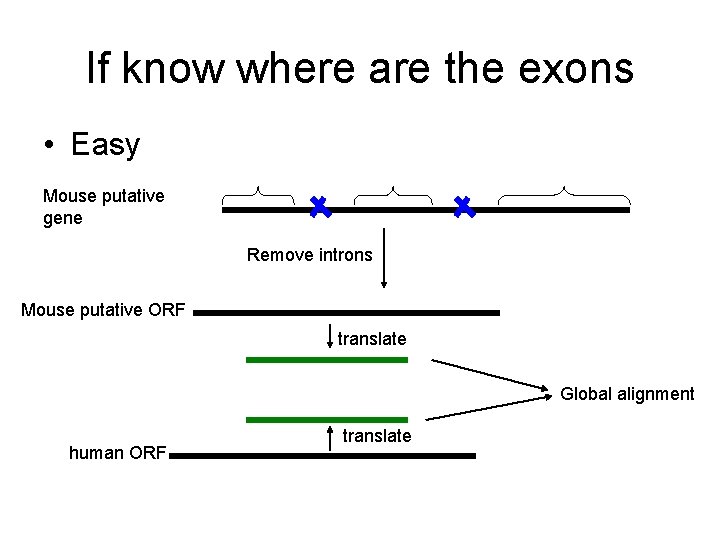
If know where are the exons • Easy Mouse putative gene Remove introns Mouse putative ORF translate Global alignment human ORF translate
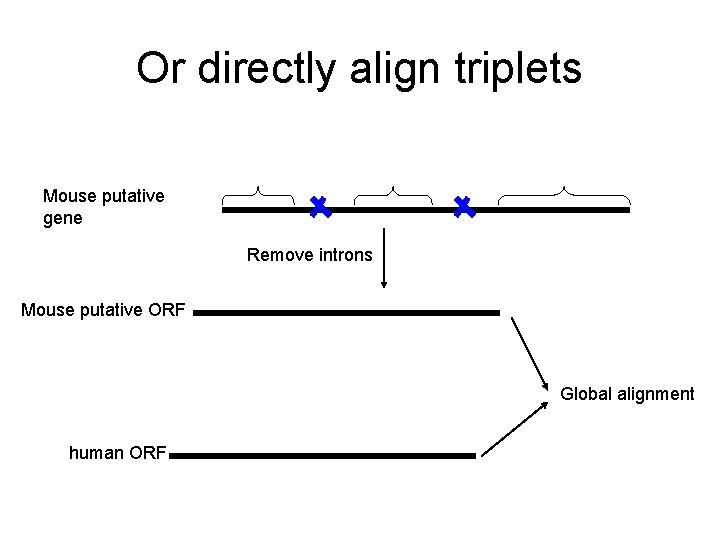
Or directly align triplets Mouse putative gene Remove introns Mouse putative ORF Global alignment human ORF
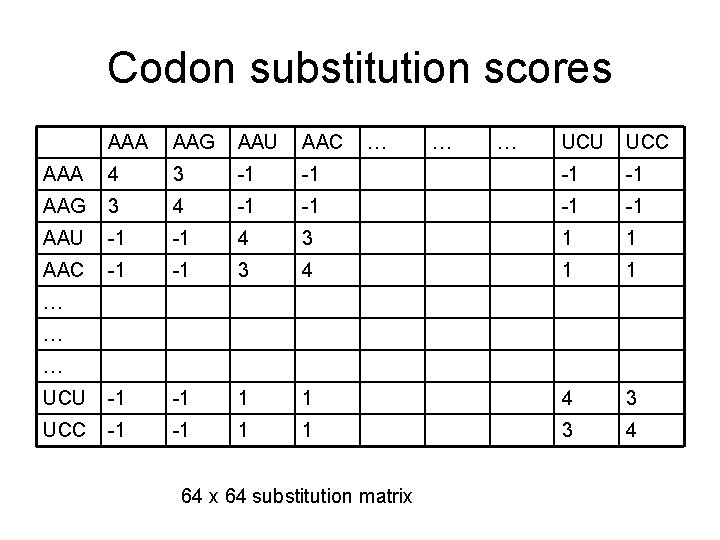
Codon substitution scores AAA AAG AAU AAC AAA 4 3 -1 AAG 3 4 AAU -1 AAC … … … UCU UCC -1 -1 4 3 1 1 -1 -1 3 4 1 1 UCU -1 -1 1 1 4 3 UCC -1 -1 1 1 3 4 … … … 64 x 64 substitution matrix
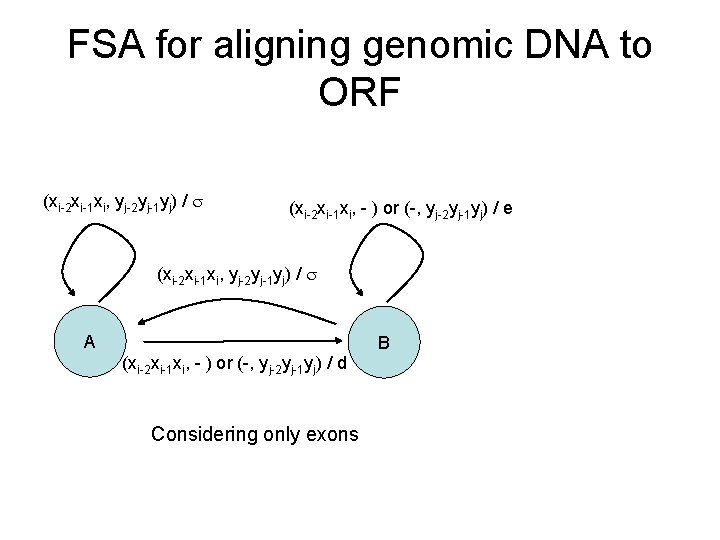
FSA for aligning genomic DNA to ORF (xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e (xi-2 xi-1 xi, yj-2 yj-1 yj) / A B (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d Considering only exons
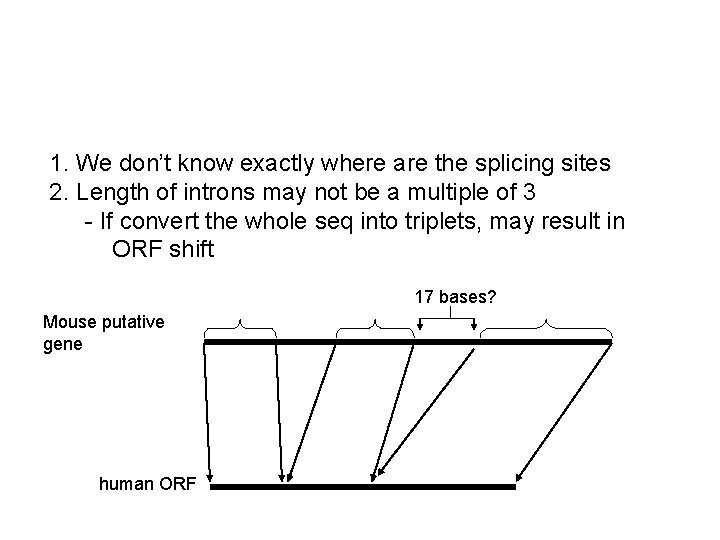
1. We don’t know exactly where are the splicing sites 2. Length of introns may not be a multiple of 3 - If convert the whole seq into triplets, may result in ORF shift 17 bases? Mouse putative gene human ORF
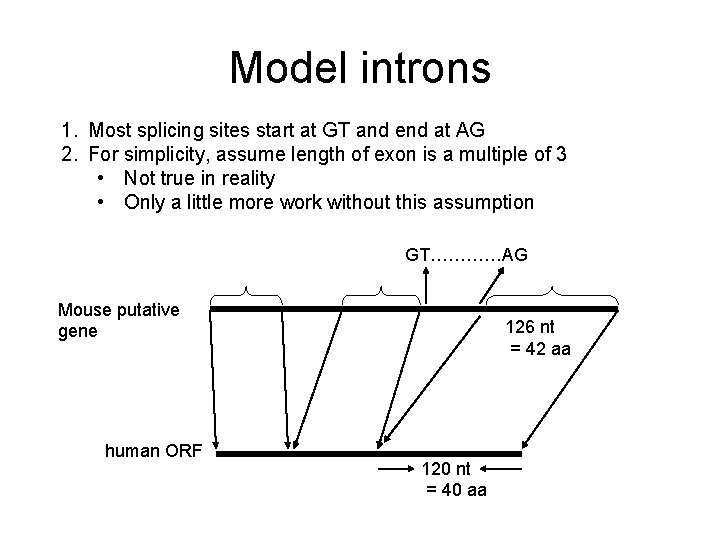
Model introns 1. Most splicing sites start at GT and end at AG 2. For simplicity, assume length of exon is a multiple of 3 • Not true in reality • Only a little more work without this assumption GT…………AG Mouse putative gene human ORF 126 nt = 42 aa 120 nt = 40 aa
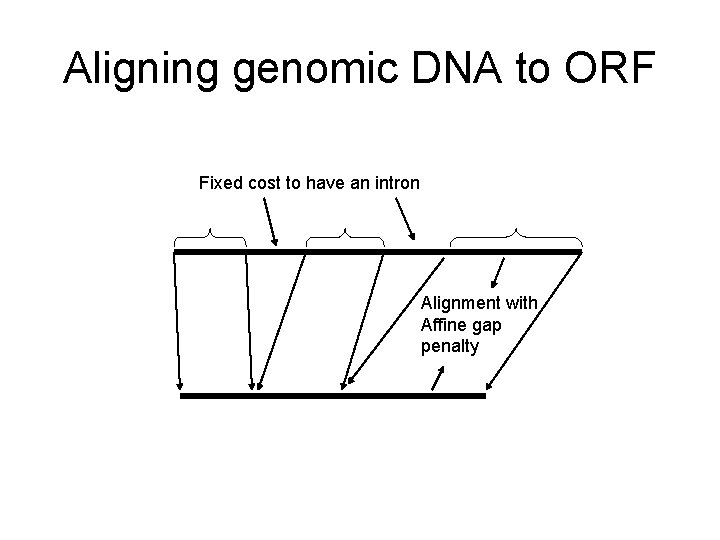
Aligning genomic DNA to ORF Fixed cost to have an intron Alignment with Affine gap penalty
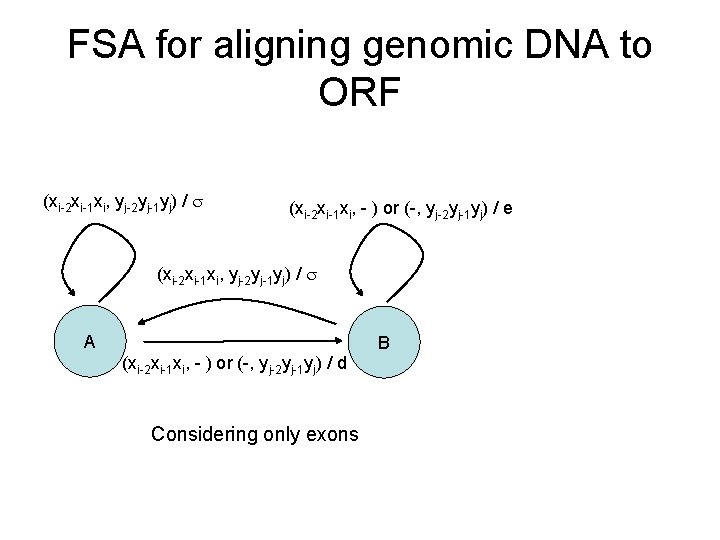
FSA for aligning genomic DNA to ORF (xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e (xi-2 xi-1 xi, yj-2 yj-1 yj) / A B (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d Considering only exons
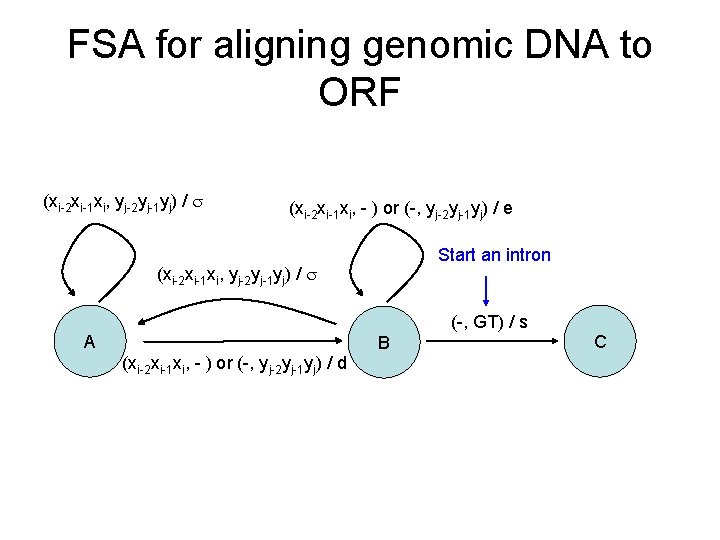
FSA for aligning genomic DNA to ORF (xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e Start an intron (xi-2 xi-1 xi, yj-2 yj-1 yj) / (-, GT) / s A B (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d C
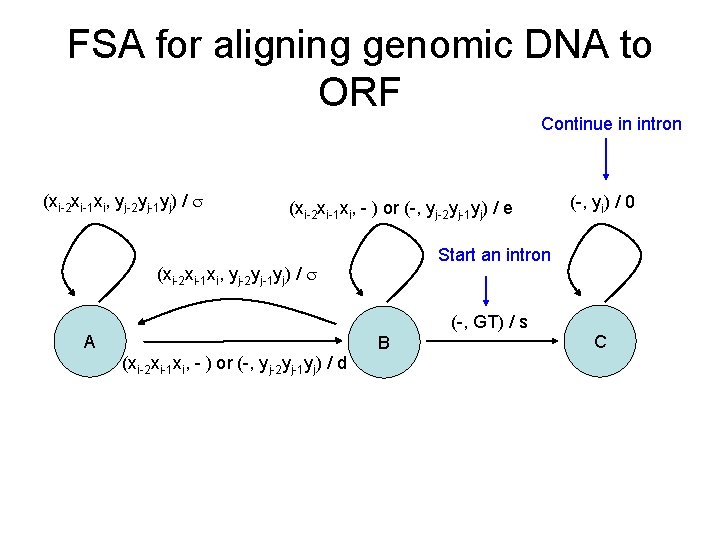
FSA for aligning genomic DNA to ORF Continue in intron (xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e Start an intron (xi-2 xi-1 xi, yj-2 yj-1 yj) / (-, GT) / s A B (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d (-, yi) / 0 C
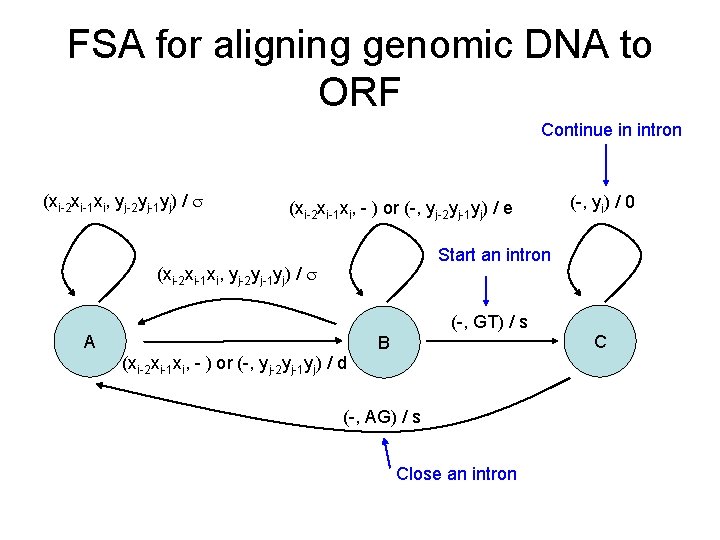
FSA for aligning genomic DNA to ORF Continue in intron (xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e (-, yi) / 0 Start an intron (xi-2 xi-1 xi, yj-2 yj-1 yj) / (-, GT) / s A B (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d (-, AG) / s Close an intron C
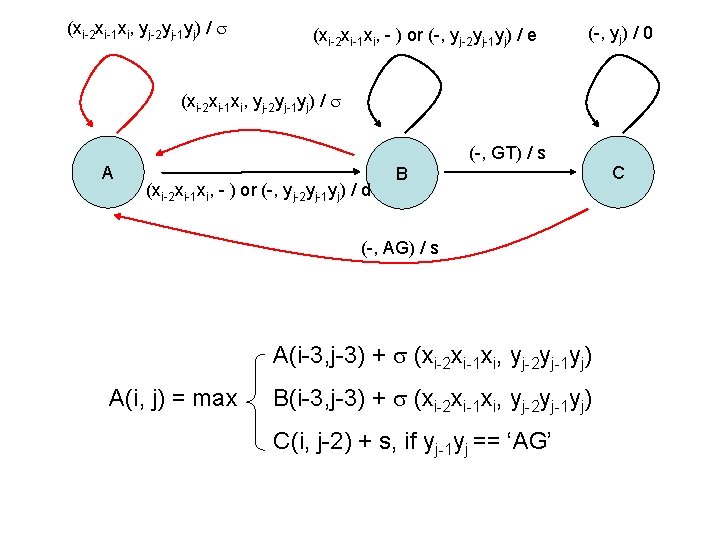
(xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e (-, yj) / 0 (xi-2 xi-1 xi, yj-2 yj-1 yj) / A (-, GT) / s (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d B (-, AG) / s A(i-3, j-3) + (xi-2 xi-1 xi, yj-2 yj-1 yj) A(i, j) = max B(i-3, j-3) + (xi-2 xi-1 xi, yj-2 yj-1 yj) C(i, j-2) + s, if yj-1 yj == ‘AG’ C
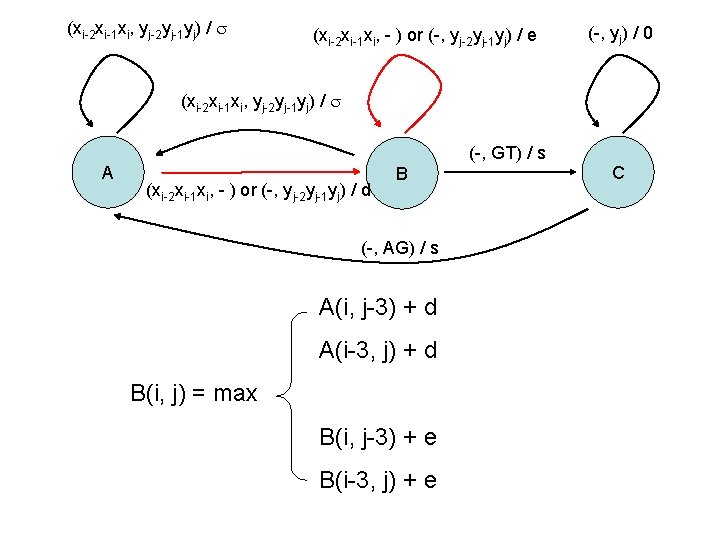
(xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e (-, yj) / 0 (xi-2 xi-1 xi, yj-2 yj-1 yj) / A (-, GT) / s (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d B (-, AG) / s A(i, j-3) + d A(i-3, j) + d B(i, j) = max B(i, j-3) + e B(i-3, j) + e C
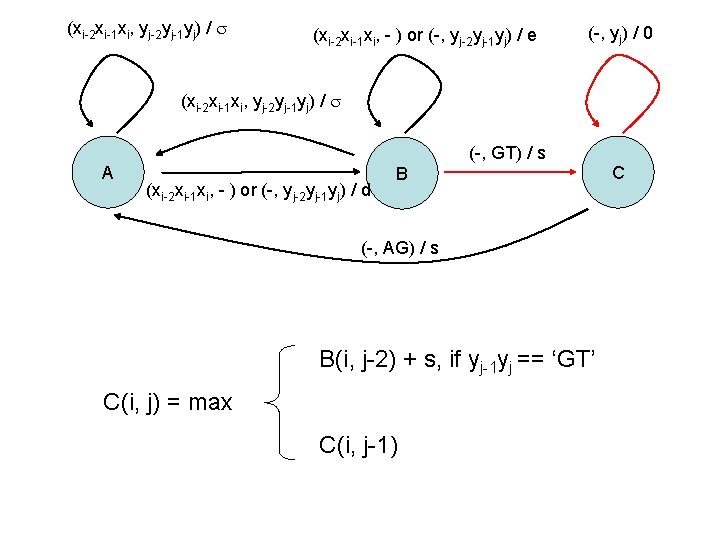
(xi-2 xi-1 xi, yj-2 yj-1 yj) / (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / e (-, yj) / 0 (xi-2 xi-1 xi, yj-2 yj-1 yj) / A (-, GT) / s (xi-2 xi-1 xi, - ) or (-, yj-2 yj-1 yj) / d B (-, AG) / s B(i, j-2) + s, if yj-1 yj == ‘GT’ C(i, j) = max C(i, j-1) C
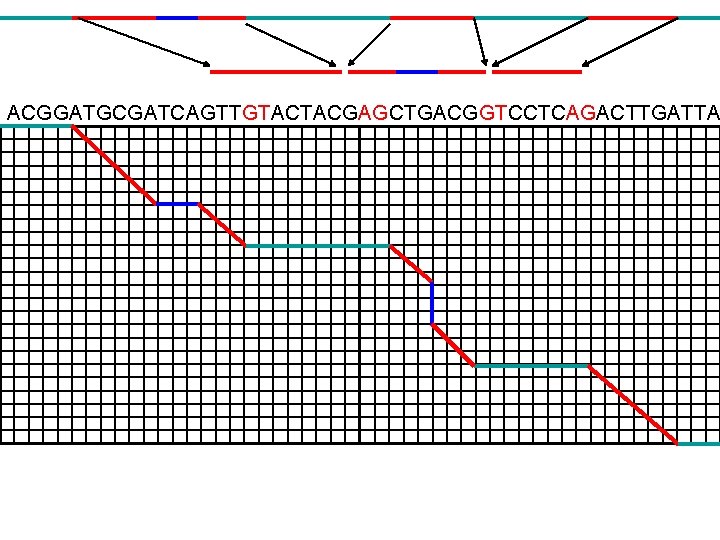
ACGGATGCGATCAGTTGTACTACGAGCTGACGGTCCTCAGACTTGATTA

• There is a close relationship between dynamic programming, FSA, regular expression, and regular grammar • Using FSA, you can design more complex alignment algorithms • If you can draw the state diagram for a problem, it can be easily formulated into a DP problem – In particular, Hidden Markov Models – Will discuss more in a few weeks
- Slides: 95