CS 5263 Bioinformatics Multiple Sequence Alignment Multiple Sequence

CS 5263 Bioinformatics Multiple Sequence Alignment

Multiple Sequence Alignment • Motivation: – A faint similarity between two sequences becomes very significant if present in many sequences • Definition – Given N sequences x 1, x 2, …, x. N: Insert gaps (-) in each sequence xi, such that • All sequences have the same length L • Score of the alignment is maximum • Two issues – How to score an alignment? – How to find a (nearly) optimal alignment?

Scoring function - first assumption • Columns are independent – Similar in pair-wise alignment • Therefore, the score of an alignment is the sum of all columns • Need to decide how to score a single column

Scoring function (cont’d) • Ideally: – An n-dimensional matrix, where n is the number of sequences – E. g. (A, C, C, G, -) for aligning 5 sequences – Total number of parameters: (k+1)n, where k is the alphabet size • Direct estimation of such scores is difficult – Too many parameters to estimate – Even more difficult if need to consider phylogenetic relationships Phylogenetic tree or evolution tree x ? y z w v

Scoring Function (cont’d) • Compromises: – Compute from pair-wise scores • Option 1: Based on sum of all pair-wise scores • Option 2: Based on scores with a consensus sequence – Other options • Consider tree topology explicitly • Information-theory based score • Difficult to optimize

Scoring Function: Sum Of Pairs Definition: Induced pairwise alignment A pairwise alignment induced by the multiple alignment Example: Induces: - x: AC-GCGG-C y: AC-GC-GAG z: GCCGC-GAG x: ACGCGG-C; y: ACGC-GAC; - x: AC-GCGG-C; z: GCCGC-GAG; - y: AC-GCGAG z: GCCGCGAG -

Sum Of Pairs (cont’d) • The sum-of-pairs (SP) score of an alignment is the sum of the scores of all induced pairwise alignments S(m) = k<l s(mk, ml): score of induced alignment (k, l)
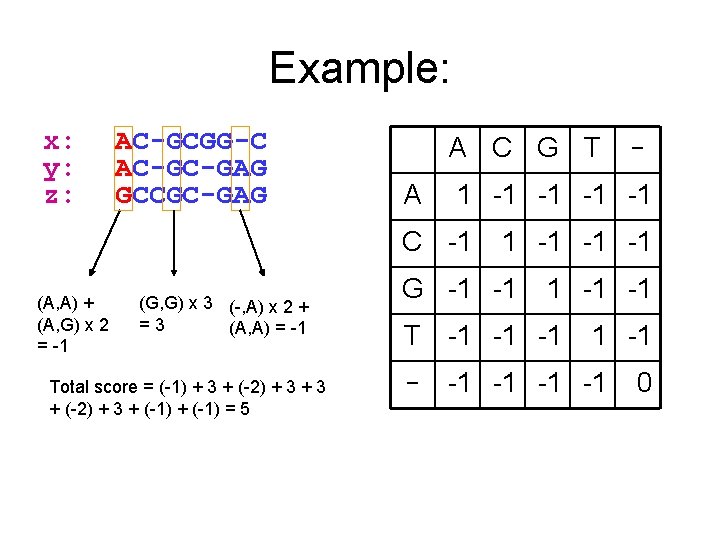
Example: x: y: z: AC-GCGG-C AC-GC-GAG GCCGC-GAG A C G T A 1 -1 -1 C -1 (A, A) + (A, G) x 2 = -1 (G, G) x 3 (-, A) x 2 + =3 (A, A) = -1 Total score = (-1) + 3 + (-2) + 3 + (-1) = 5 - 1 -1 -1 -1 G -1 -1 -1 T -1 -1 -1 -1 0

Sum Of Pairs (cont’d) • Drawback: no evolutionary characterization – Every sequence derived from all others • Heuristic way to incorporate evolution tree – Weighted Sum of Pairs: S(m) = k<l wkl s(mk, ml) wkl: weight decreasing with distance Human Mouse Duck Chicken

Consensus score -AGGCTATCACCTGACCTCCAGGCCGA--TGCCC--TAG-CTATCAC--GACCGC--GGTCGATTTGCCCGAC CAG-CTATCAC--GACCGC----TCGATTTGCTCGAC Consensus sequence: CAG-CTATCAC--GACCGC--GGTCGATTTGCCCGAC • Find optimal consensus string m* to maximize S(m) = i s(m*, mi) s(mk, ml): score of pairwise alignment (k, l)

Multiple Sequence Alignments Algorithms • Can also be global or local – We only talk about global for now • A simple method – Do pairwise alignment between all pairs – Combine the pairwise alignments into a single multiple alignment – Is this going to work?
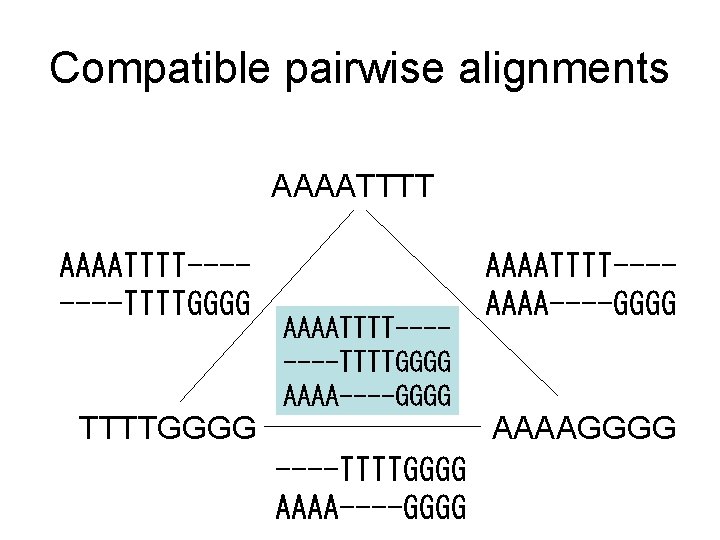
Compatible pairwise alignments AAAATTTT-------TTTTGGGG AAAA----GGGG TTTTGGGG AAAATTTT---AAAA----GGGG AAAAGGGG ----TTTTGGGG AAAA----GGGG
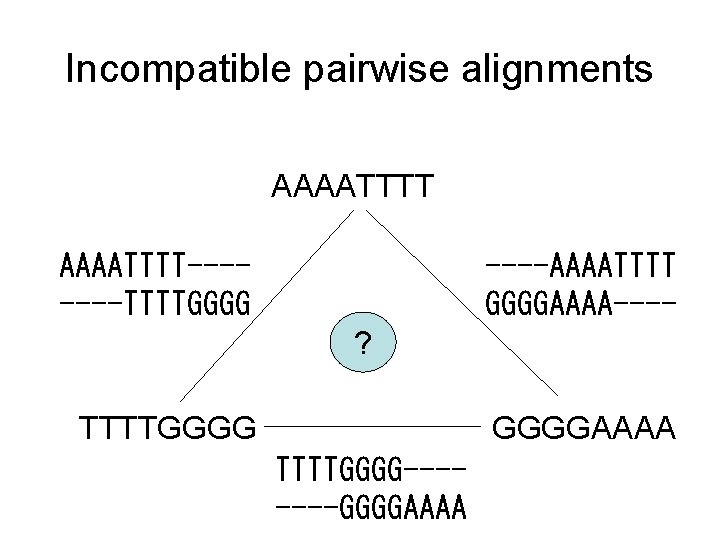
Incompatible pairwise alignments AAAATTTT-------TTTTGGGG ----AAAATTTT GGGGAAAA---? TTTTGGGGAAAA TTTTGGGG-------GGGGAAAA

Multidimensional Dynamic Programming (MDP) Generalization of Needleman-Wunsh: • Find the longest path in a high-dimensional cube – As opposed to a two-dimensional grid • Uses a N-dimensional matrix – As apposed to a two-dimensional array • Entry F(i 1, …, ik) represents score of optimal alignment for s 1[1. . i 1], … sk[1. . ik] F(i 1, i 2, …, i. N) = max(all neighbors of a cell) (F(nbr)+S(current))

Multidimensional Dynamic Programming (MDP) • Example: in 3 D (three sequences): (i-1, j-1, k-1) • 23 – 1 = 7 neighbors/cell F(i-1, j-1, k-1) + S(xi, xj, xk), F(i-1, j-1, k ) + S(xi, xj, -), F(i-1, j , k-1) + S(xi, -, xk), F(i, j, k) = max F(i , j-1, k-1) + S(-, xj, xk), F(i-1, j , k ) + S(xi, -, -), F(i , j-1, k ) + S(-, xj, -), F(i , j , k-1) + S(-, -, xk) (i-1, j-1, k) (i, j-1, k-1) (i, j-1, k) (i-1, j, k-1) (i-1, j, k) (i, j, k-1) (i, j, k)

Multidimensional Dynamic Programming (MDP) Running Time: 1. Size of matrix: LN; Where L = length of each sequence N = number of sequences 2. Neighbors/cell: 2 N – 1 Therefore…………… O(2 N LN)

Faster MDP • Carrillo & Lipman, 1988 – Branch and bound – Other heuristics • Implemented in a tool called MSA • Practical for about 6 sequences of length about 200 -300.

Faster MDP • Basic idea: bounds of the optimal score of a multiple alignment can be pre-computed – Upper-bound: sum of optimal pair-wise alignment scores, i. e. S(m) = k<l s(mk, ml) k<l s(k, l) Optimal msa Score of the alignment between k and l induced by m Score of optimal alignment between k and l – lower-bounded: score computed by any approximate algorithm (such as the ones we’ll talk next) – For any partial path, if Scurrent + Sperspective < lowerbound, can give up that path – Guarantees optimality

Progressive Alignment • • Multiple Alignment is NP-hard Most used heuristic: Progressive Alignment Algorithm: 1. 2. 3. 4. Align two of the sequences xi, xj Fix that alignment Align a third sequence xk to the alignment xi, xj Repeat until all sequences are aligned Running Time: O(NL 2) Each alignment takes O(L 2) Repeat N times

Progressive Alignment x y z w • When evolutionary tree is known: – Align closest first, in the order of the tree Example: Order of alignments: 1. (x, y) 2. (z, w) 3. (xy, zw)

Progressive Alignment: CLUSTALW: most popular multiple protein alignment Algorithm: 1. Find all dij: alignment dist (xi, xj) • High alignment score => short distance 2. Construct a tree (Neighbor-joining hierarchical clustering. Will discuss in future) 3. Align nodes in order of decreasing similarity + a large number of heuristics

CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD
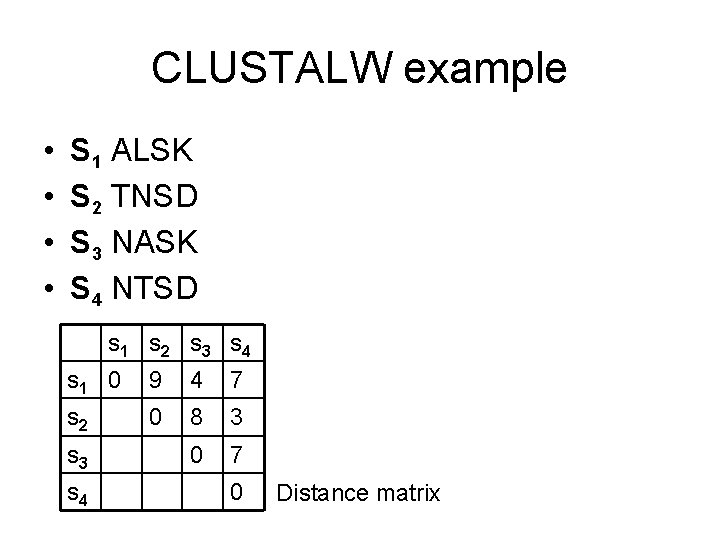
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 Distance matrix
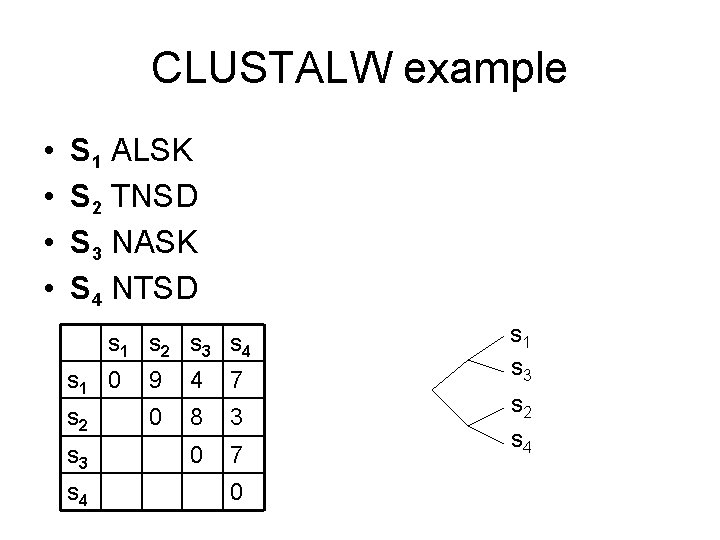
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 s 1 s 3 s 2 s 4
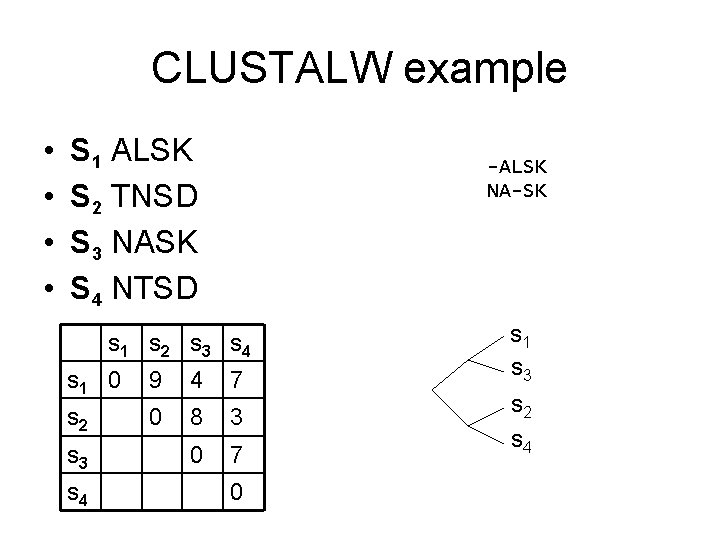
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD -ALSK NA-SK s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 s 1 s 3 s 2 s 4
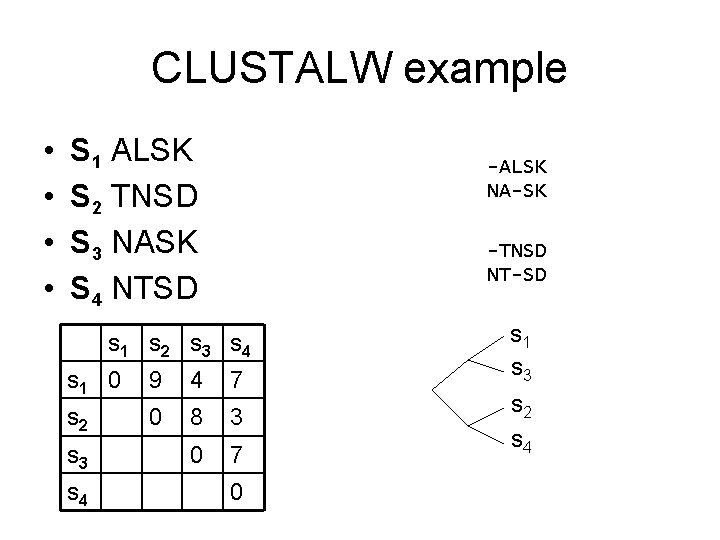
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD -ALSK NA-SK -TNSD NT-SD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 s 1 s 3 s 2 s 4
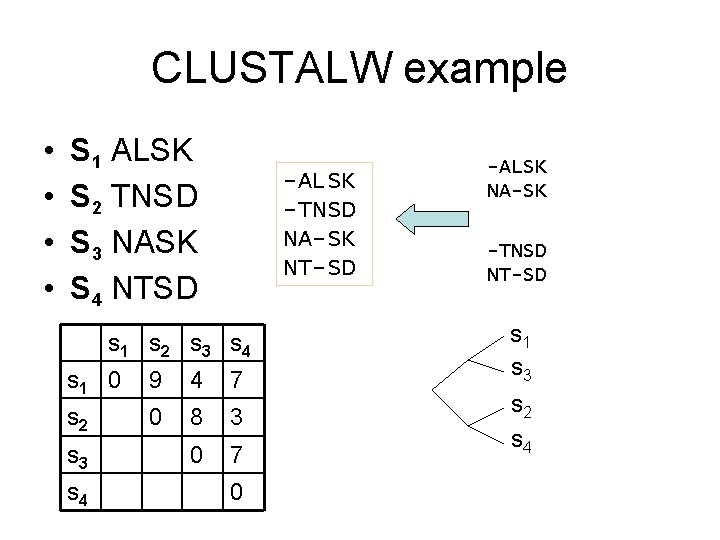
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD -ALSK -TNSD NA-SK NT-SD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 -ALSK NA-SK -TNSD NT-SD s 1 s 3 s 2 s 4

Iterative Refinement Problems with progressive alignment: • Depend on pair-wise alignments • If sequences are very distantly related, much higher likelihood of errors • Initial alignments are “frozen” even when new evidence comes Example: x: y: GAAGTT GAC-TT Frozen! z: w: GAACTG GTACTG Now clear: correct y should be GA-CTT

Iterative Refinement Algorithm (Barton-Stenberg): 1. 2. 3. 4. Align most similar xi, xj Align xk most similar to (xixj) Repeat 2 until (x 1…x. N) are aligned For j = 1 to N, Remove xj, and realign to x 1…xj-1 xj+1…x. N 5. Repeat 4 until convergence Progressive alignment
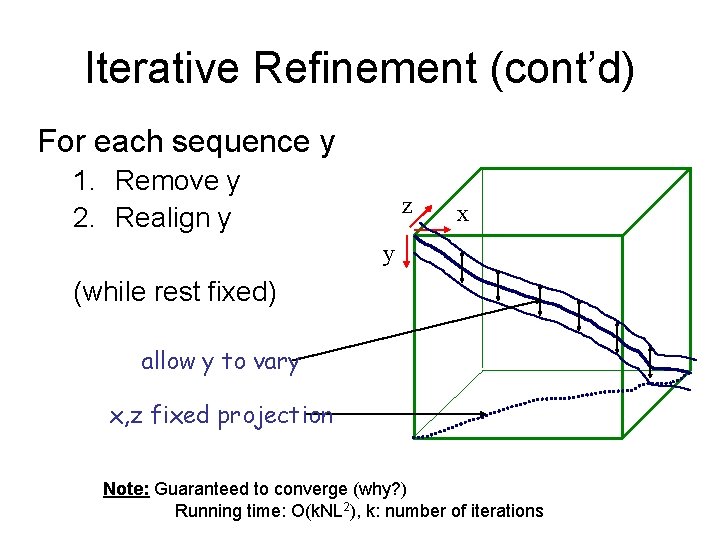
Iterative Refinement (cont’d) For each sequence y 1. Remove y 2. Realign y z x y (while rest fixed) allow y to vary x, z fixed projection Note: Guaranteed to converge (why? ) Running time: O(k. NL 2), k: number of iterations

Iterative Refinement Example: align (x, y), (z, w), (xy, zw): x: y: z: w: GAAGTTA GAC-TTA GAACTGA GTACTGA After realigning y: x: y: z: w: GAAGTTA G-ACTTA GAACTGA GTACTGA + 3 matches

Iterative Refinement • Example not handled well: x: y 1: y 2: y 3: GAAGTTA GAC-TTA z: w: GAACTGA GTACTGA Realigning any single yi changes nothing
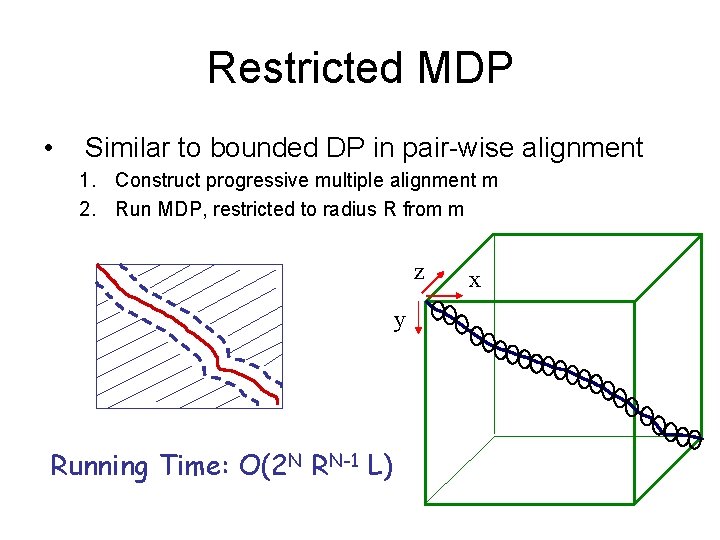
Restricted MDP • Similar to bounded DP in pair-wise alignment 1. Construct progressive multiple alignment m 2. Run MDP, restricted to radius R from m z y Running Time: O(2 N RN-1 L) x

Restricted MDP x: y 1: y 2: y 3: GAAGTTA GAC-TTA z: w: GAACTGA GTACTGA • Within radius 1 of the optimal Restricted MDP will fix it.

Other approaches • Statistical learning methods – Profile Hidden Markov Models – Will discuss in future lectures • Consistency-based methods – Still rely on pairwise alignment • But consider a third seq when aligning two seqs • If block A in seq x aligns to block B in seq y, and both aligns to block C in seq z, we have higher confidence to say that the alignment between A-B is reliable • Essentially: change scoring system according to consistency • Than applied DP as in other approaches – Pioneered by a tool called T-Coffee

Multiple alignment tools • Clustal W (Thompson, 1994) – Most popular • T-Coffee (Notredame, 2000) – Another popular tool – Consistency-based – Slower than clustal. W, but generally more accurate for more distantly related sequences • MUSCLE (Edgar, 2004) – Iterative refinement – More efficient than most others • DIALIGN (Morgenstern, 1998, 1999, 2005) – “local” • Align-m (Walle, 2004) – “local” • PROBCONS (Do, 2004) – Probabilistic consistency-based – Best accuracy on benchmarks • Pro. DA (Phuong, 2006) – Allow repeated and shuffled regions

In summary • Multiple alignment scoring functions – Sum of pairs – Other funcs exist, but less used • Multiple alignment algorithms: – MDP • Optimal • too slow • Branch & Bound doesn’t solve the problem entirely – – Progressive alignment: clustal. W Iterative refinement Restricted MDP Consistency-based Heuristic
- Slides: 37