Canadian Bioinformatics Workshops www bioinformatics ca Module Title

Canadian Bioinformatics Workshops www. bioinformatics. ca
Module #: Title of Module 2

Module 5 Metabolomic Data Analysis Using Metabo. Analyst David Wishart Informatics and Statistics for Metabolomics May 26 -27, 2016

Learning Objectives • To become familiar with the standard metabolomics data analysis workflow • To become aware of key elements such as: data integrity checking, outlier detection, quality control, normalization, scaling, etc. • To learn how to use Metabo. Analyst to facilitate data analysis Module 5 bioinformatics. ca
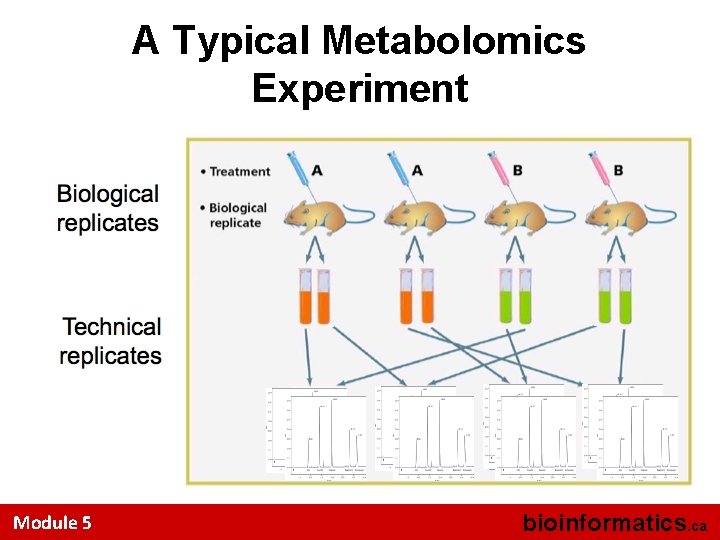
A Typical Metabolomics Experiment Module 5 bioinformatics. ca
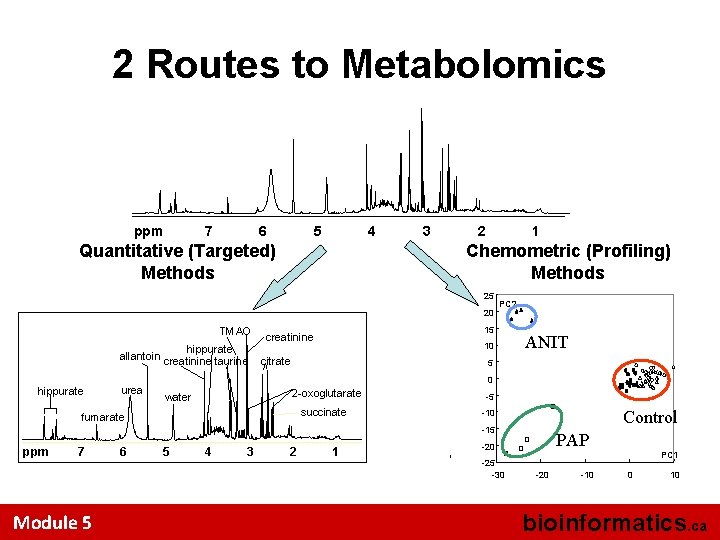
2 Routes to Metabolomics ppm 7 6 5 4 Quantitative (Targeted) Methods 3 2 1 Chemometric (Profiling) Methods 25 20 TMAO hippurate allantoin creatinine taurine hippurate urea PC 2 15 creatinine 10 citrate ANIT 5 0 2 -oxoglutarate water succinate fumarate -5 Control -10 -15 ppm 7 Module 5 6 5 4 3 2 1 PAP -20 -25 -30 -20 -10 PC 1 0 10 bioinformatics. ca

Metabolomics Data Workflow Chemometric Methods • Data Integrity Check • Spectral alignment or binning • Data normalization • Data QC/outlier removal • Data reduction & analysis • Compound ID Module 5 Targeted Methods • Data Integrity Check • Compound ID and quantification • Data normalization • Data QC/outlier removal • Data reduction & analysis bioinformatics. ca
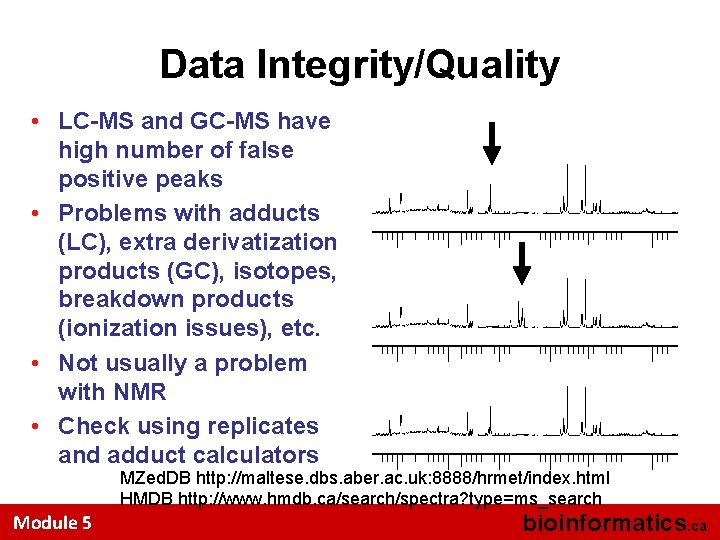
Data Integrity/Quality • LC-MS and GC-MS have high number of false positive peaks • Problems with adducts (LC), extra derivatization products (GC), isotopes, breakdown products (ionization issues), etc. • Not usually a problem with NMR • Check using replicates and adduct calculators MZed. DB http: //maltese. dbs. aber. ac. uk: 8888/hrmet/index. html HMDB http: //www. hmdb. ca/search/spectra? type=ms_search Module 5 bioinformatics. ca
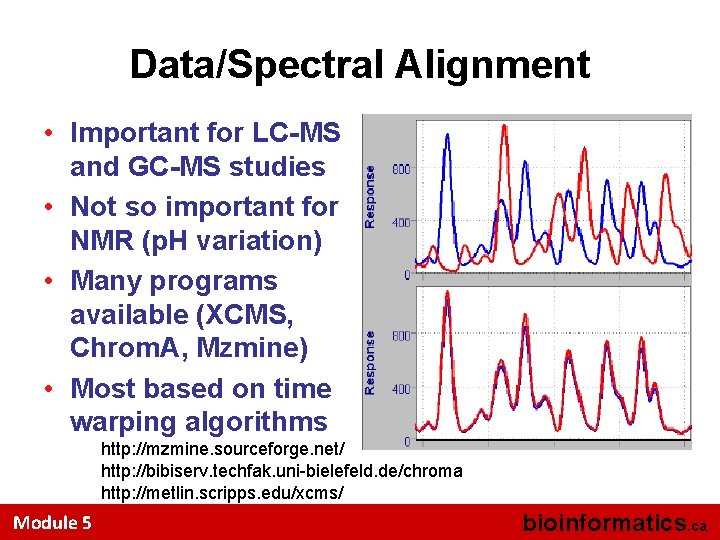
Data/Spectral Alignment • Important for LC-MS and GC-MS studies • Not so important for NMR (p. H variation) • Many programs available (XCMS, Chrom. A, Mzmine) • Most based on time warping algorithms http: //mzmine. sourceforge. net/ http: //bibiserv. techfak. uni-bielefeld. de/chroma http: //metlin. scripps. edu/xcms/ Module 5 bioinformatics. ca

Binning (3000 pts to 14 bins) xi, yi x = 232. 1 (AOC) y = 10 (bin #) bin 1 bin 2 bin 3 bin 4 bin 5 bin 6 bin 7 bin 8. . . Module 5 bioinformatics. ca
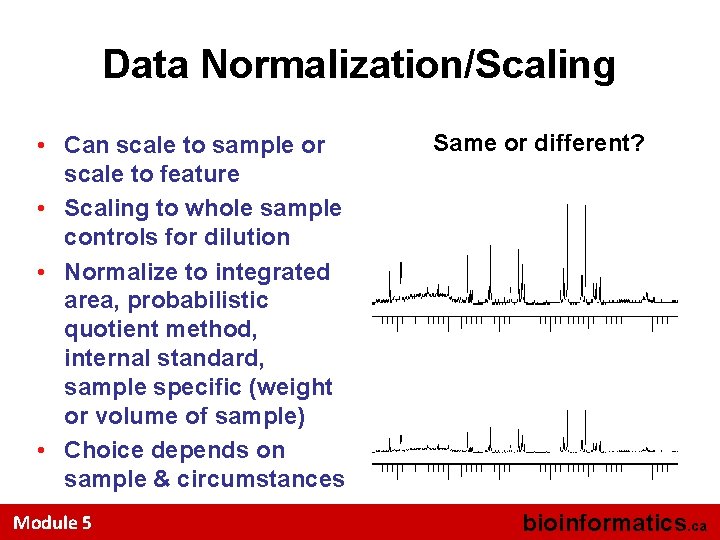
Data Normalization/Scaling • Can scale to sample or scale to feature • Scaling to whole sample controls for dilution • Normalize to integrated area, probabilistic quotient method, internal standard, sample specific (weight or volume of sample) • Choice depends on sample & circumstances Module 5 Same or different? bioinformatics. ca
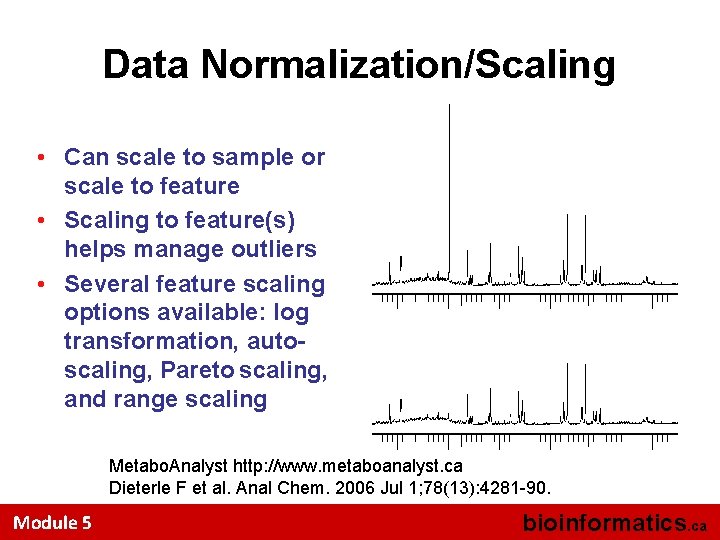
Data Normalization/Scaling • Can scale to sample or scale to feature • Scaling to feature(s) helps manage outliers • Several feature scaling options available: log transformation, autoscaling, Pareto scaling, and range scaling Metabo. Analyst http: //www. metaboanalyst. ca Dieterle F et al. Anal Chem. 2006 Jul 1; 78(13): 4281 -90. Module 5 bioinformatics. ca

Data QC, Outlier Removal & Data Reduction • Data filtering (remove solvent peaks, noise filtering, false positives, outlier removal -- needs justification) • Dimensional reduction or feature selection to reduce number of features or factors to consider (PCA or PLS-DA) • Clustering to find similarity Module 5 bioinformatics. ca

Metabo. Analyst http: //www. metaboanalyst. ca A comprehensive web server designed to process & analyze LC-MS, GC-MS or NMR-based metabolomic data Module 5 bioinformatics. ca

Metabo. Analyst History • 2009 v 1. 0 - Supports both univariate and multivariate data processing, including ttests, ANOVA, PCA, PLS-DA, colorful plots, with detailed explanations & summaries • 2012 v 2. 0 - Identifies significantly altered functions & pathways • 2015 v 3. 0 – Better performance, better graphical interactivity, biomarker analysis, power analysis, integration with gene expression data … Module 5 bioinformatics. ca

Metabo. Analyst Overview • • • Raw data processing Data reduction & statistical analysis Functional enrichment analysis Metabolic pathway analysis Power analysis and sample size estimation • Biomarker analysis • Integrative analysis Module 5 bioinformatics. ca

Metabo. Analyst Modules Data preprocessing Data normalization Data analysis Data interpretatio n 17 Module 5 bioinformatics. ca
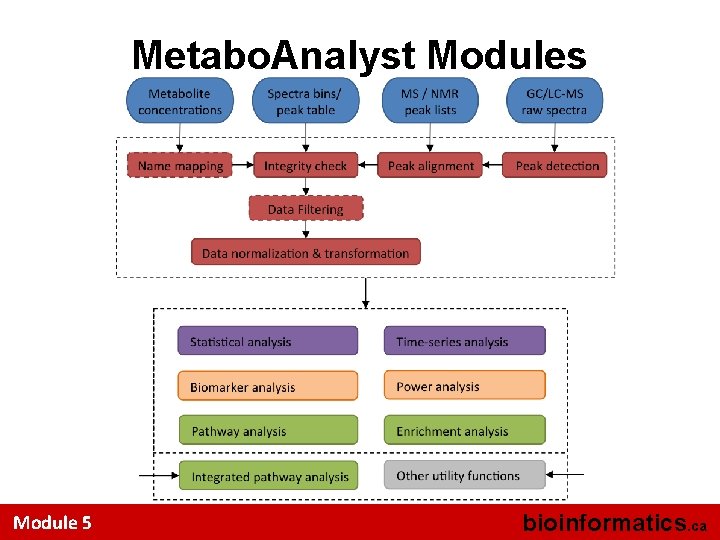
Metabo. Analyst Modules Module 5 bioinformatics. ca

Example Datasets Module 5 bioinformatics. ca

Example Datasets Module 5 bioinformatics. ca

Metabolomic Data Processing Module 5 bioinformatics. ca

Common Tasks • Purpose: to convert various raw data forms into data matrices suitable for statistical analysis • Supported data formats – Concentration tables (Targeted Analysis) – Peak lists (Untargeted) – Spectral bins (Untargeted) – Raw spectra (Untargeted) Module 5 bioinformatics. ca

Select a Module (Statistical Analysis) Module 5 bioinformatics. ca

Data Upload Module 5 bioinformatics. ca

Alternatively … Module 5 bioinformatics. ca

Data Set Selected • Here we have selected a data set from dairy cattle fed different proportions of cereal grains (0%, 15%, 30%, 45%) • The rumen was analyzed using NMR spectroscopy using quantitative metabolomic techniques • High grain diets are thought to be stressful on cows Module 5 bioinformatics. ca

Data Integrity Check Module 5 bioinformatics. ca

Data Normalization Samples = rows Compounds = columns Module 5 bioinformatics. ca

Data Normalization • At this point, the data has been transformed to a matrix with the samples in rows and the variables (compounds/peaks/bins) in columns • Metabo. Analyst offers three types of normalization, sample normalization, data transformation, and data scaling • Sample normalization aims to make each sample (row) comparable to each other (i. e. urine samples with different dilution effects) Module 5 bioinformatics. ca

Data Normalization • Data transformation & data scaling aims to make each variable (column) comparable in scale to each other, thereby generating a “normal” distribution • This procedure is useful when variables are of very different orders of magnitude • Transformation operates on each data point itself – Log and cube root transformation • Scaling operates on each variable column – Autoscaling, Pareto scaling and range scaling Module 5 bioinformatics. ca
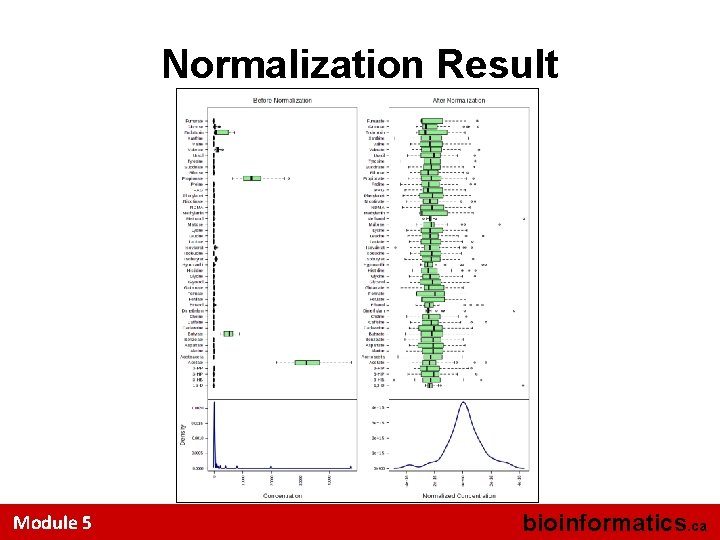
Normalization Result Module 5 bioinformatics. ca

Data Normalization • You cannot know a priori what the best normalization protocol will be • Metabo. Analyst allows you to interactively explore different normalization protocols and to visually inspect the degree of “normality” or Gaussian behavior • This example is nicely normalized Module 5 bioinformatics. ca

Next Steps • After normalization has been completed it is a good idea to look at your data a little further to identify outliers or noise that could/should be removed Module 5 bioinformatics. ca

Quality Control • Dealing with outliers – Detected mainly by visual inspection – May be corrected by normalization – May be excluded • Noise reduction – More of a concern for spectral bins/ peak lists – Usually improves downstream results Module 5 bioinformatics. ca
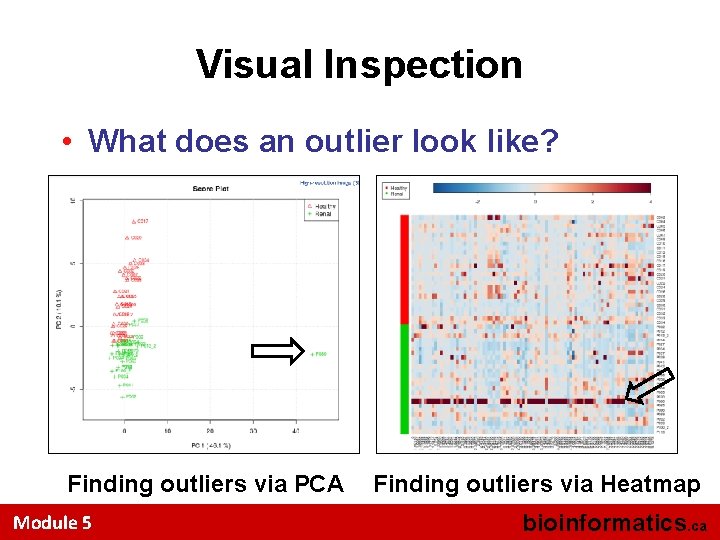
Visual Inspection • What does an outlier look like? Finding outliers via PCA Module 5 Finding outliers via Heatmap bioinformatics. ca
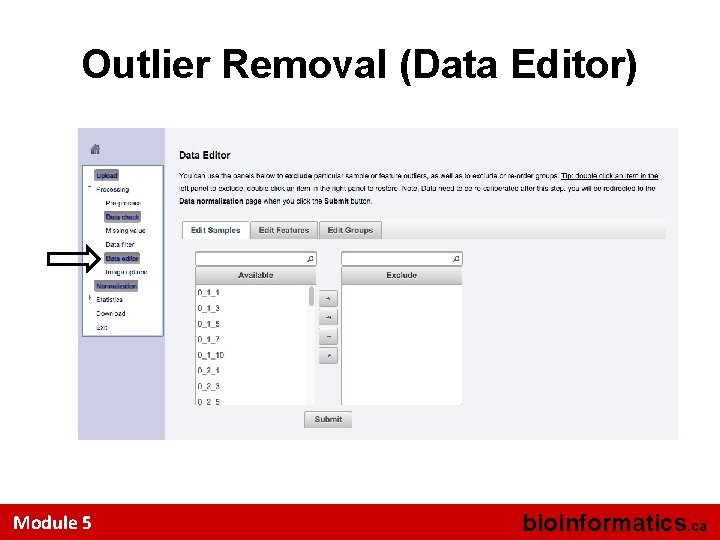
Outlier Removal (Data Editor) Module 5 bioinformatics. ca

Noise Reduction (Data Filtering) Module 5 bioinformatics. ca

Noise Reduction (cont. ) • Characteristics of noise & uninformative features – Low variances (default) – Low intensities Module 5 bioinformatics. ca

Data Reduction and Statistical Analysis Module 5 bioinformatics. ca

Common Tasks • To identify important features • To detect interesting patterns • To assess difference between the phenotypes • To facilitate classification or prediction • We will look at ANOVA, Multivariate Analysis (PCA, PLS-DA) and Clustering Module 5 bioinformatics. ca

Module 5 bioinformatics. ca

ANOVA • Looking at 4 different dairy cow populations – 0% grain in diet – 15% grain in diet – 30% grain in diet – 45% grain in diet • Try to identify those metabolites that are different between all groups or just between 0% and everything else Module 5 bioinformatics. ca
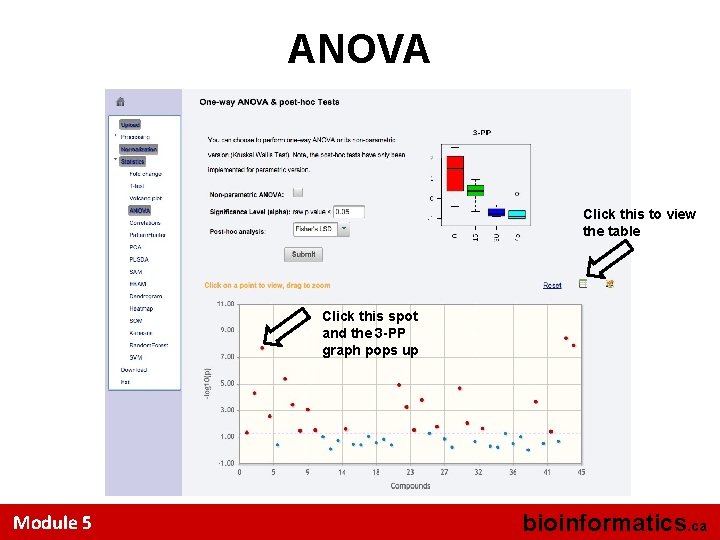
ANOVA Click this to view the table Click this spot and the 3 -PP graph pops up Module 5 bioinformatics. ca

View Individual Compounds Click this to see the uracil graphs Module 5 bioinformatics. ca

What’s Next? • Click and compare different compounds to see which ones are most different or most similar between the 4 groups • Click on the Correlation link (under the ANOVA link) to generate a heat map that displays the pairwise compound correlations and compound clusters Module 5 bioinformatics. ca
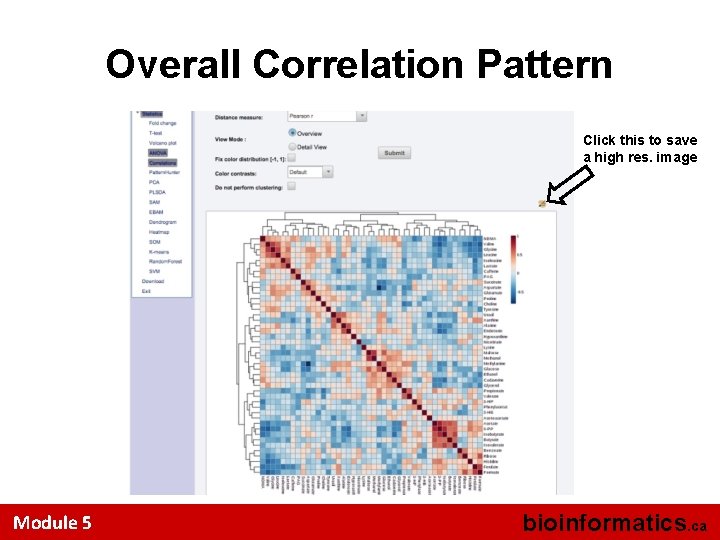
Overall Correlation Pattern Click this to save a high res. image Module 5 bioinformatics. ca
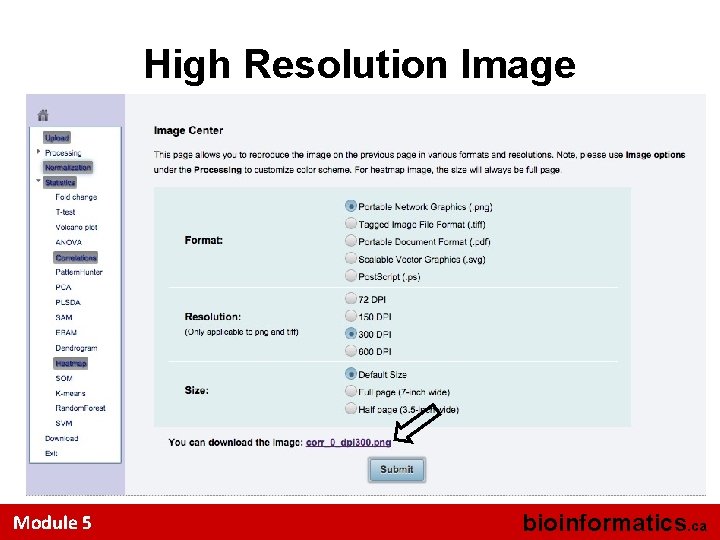
High Resolution Image Module 5 bioinformatics. ca
What’s Next? • When looking at >2 groups it is often useful to look for patterns or trends within particular metabolites • Use Pattern Hunter to find these trends Module 5 bioinformatics. ca

Pattern Matching • Looking for compounds showing interesting patterns of change • Essentially a method to look for linear trends or periodic trends in the data • Best for data that has 3 or more groups Module 5 bioinformatics. ca
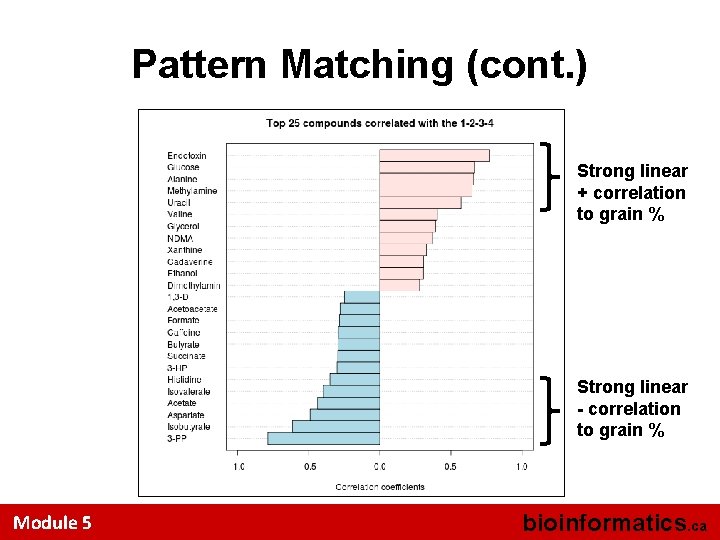
Pattern Matching (cont. ) Strong linear + correlation to grain % Strong linear - correlation to grain % Module 5 bioinformatics. ca
Module 5 bioinformatics. ca

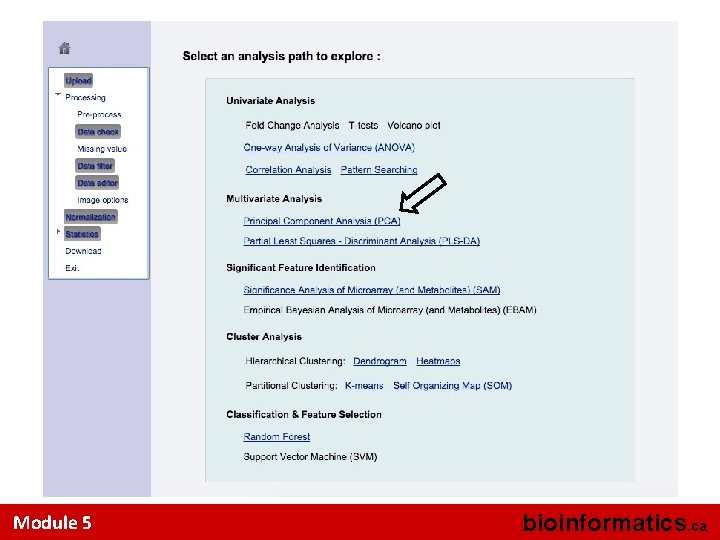
Multivariate Analysis • Use PCA option to view the separation (if any) in the 4 groups • Look at the 2 D PCA Score Plot – 2 most significant principal components • Look at the 2 D PCA Loading Plot • Look at the PCA Plot in 3 D – 3 most significant principal components • Options for viewing are located in the top tabs Module 5 bioinformatics. ca
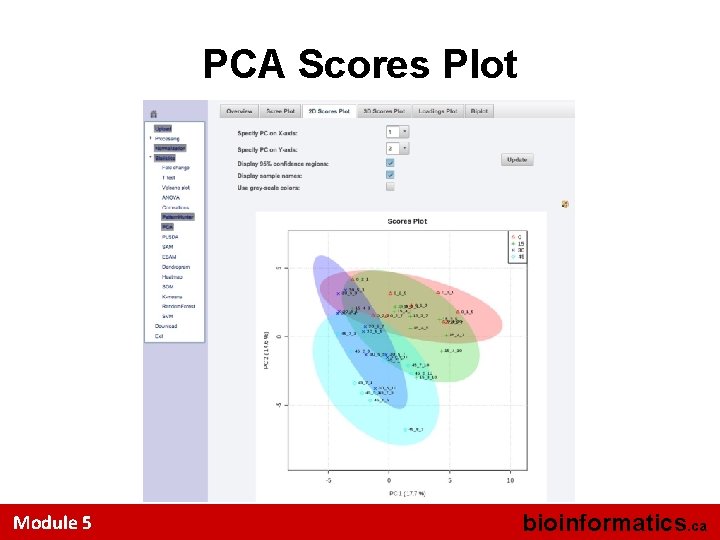
PCA Scores Plot Module 5 bioinformatics. ca
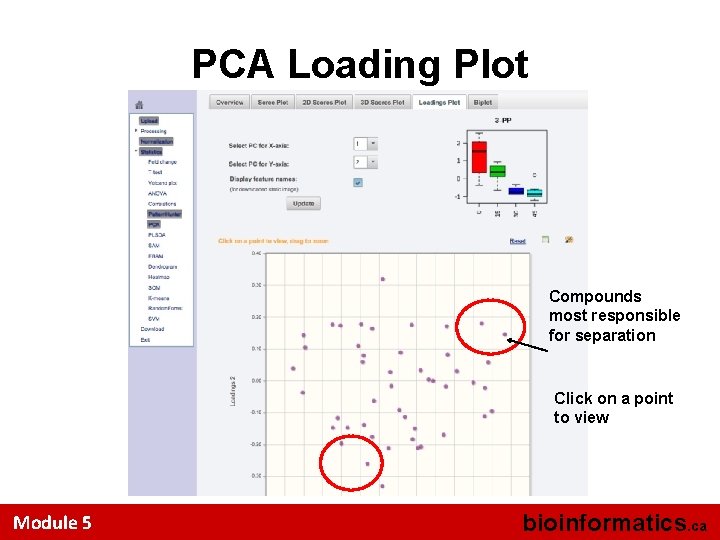
PCA Loading Plot Compounds most responsible for separation Click on a point to view Module 5 bioinformatics. ca
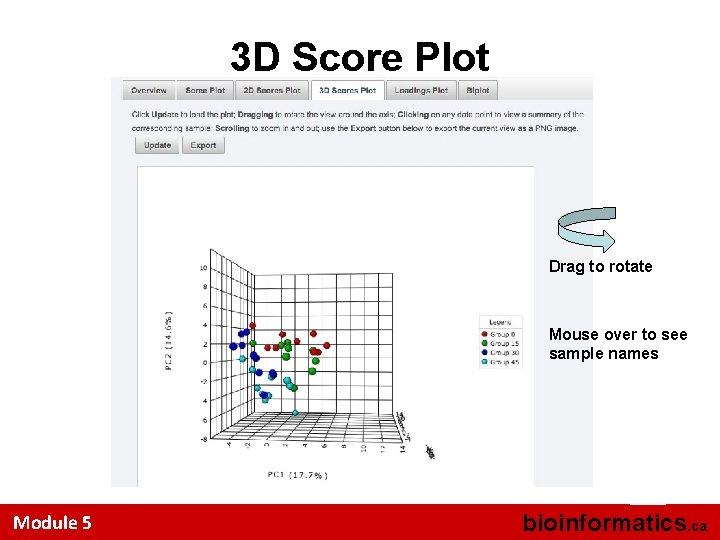
3 D Score Plot Drag to rotate Mouse over to see sample names 55 Module 5 bioinformatics. ca

Module 5 bioinformatics. ca

Multivariate Analysis • Use PLS-DA option to view the separation of the 4 (labeled) groups • PLS-DA “rotates” the PCA axes to maximize separation • Look at the 2 D PLS Scores Plot • Look at the Q 2 and R 2 Values (Cross Validation) • Use the VIP plot to ID important metabolites (VIP > 1. 2) Module 5 bioinformatics. ca
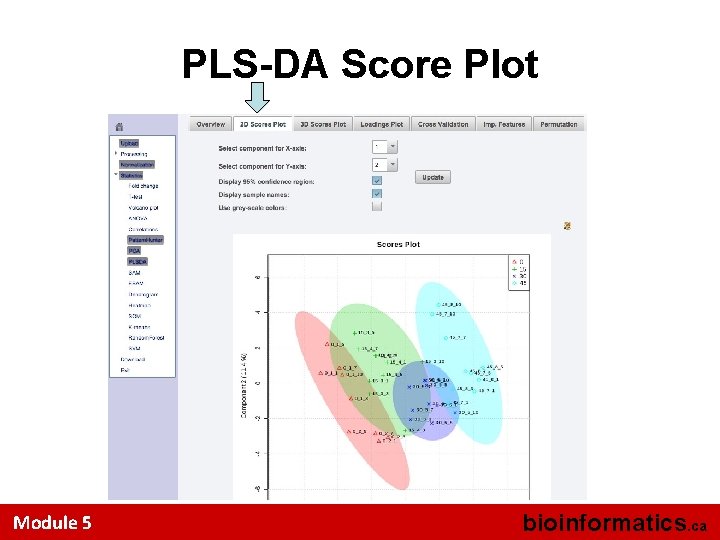
PLS-DA Score Plot Module 5 bioinformatics. ca
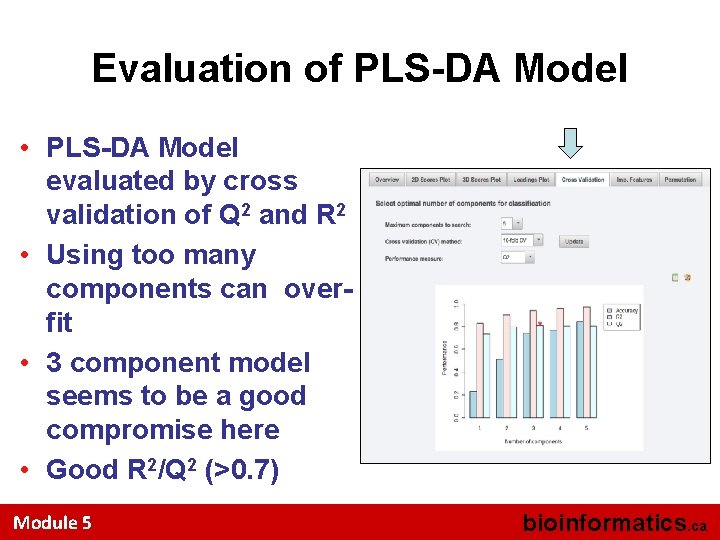
Evaluation of PLS-DA Model • PLS-DA Model evaluated by cross validation of Q 2 and R 2 • Using too many components can overfit • 3 component model seems to be a good compromise here • Good R 2/Q 2 (>0. 7) Module 5 bioinformatics. ca
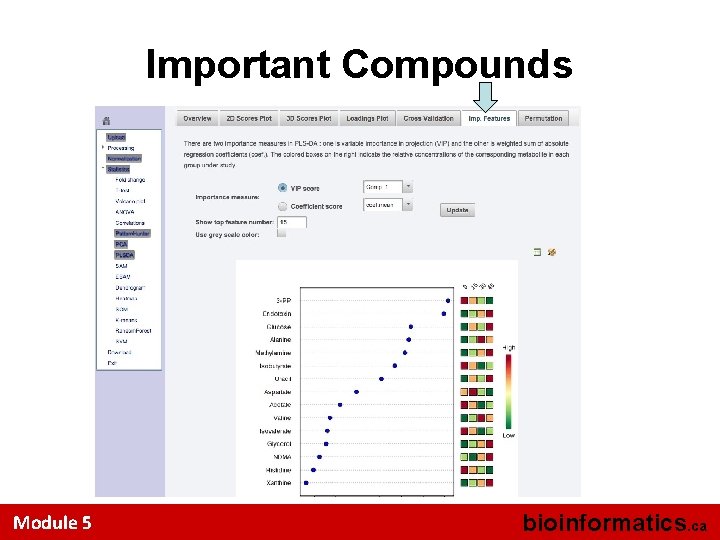
Important Compounds Module 5 bioinformatics. ca

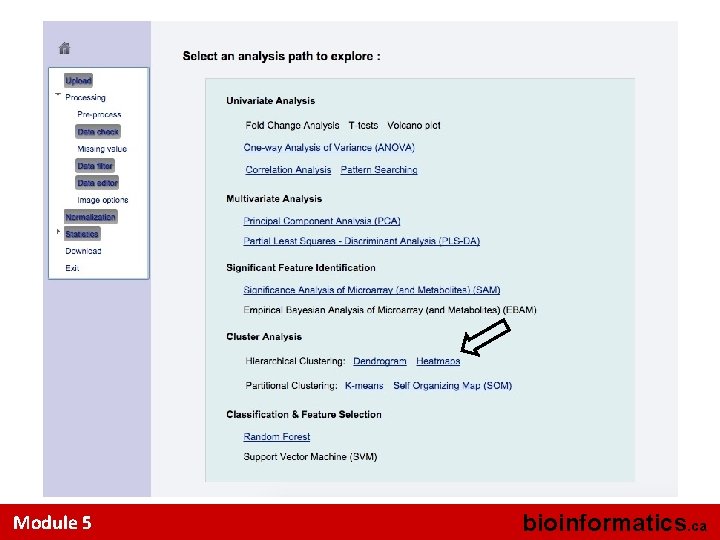
Model Validation Note, permutation is computationally intensive. It is not performed by default. Users need to set the permutation number and press the submit button Module 5 bioinformatics. ca
Module 5 bioinformatics. ca

Hierarchical Clustering (Heat Maps) • An alternative way of viewing or clustering multivariate data • Allows one to look at the behavior of individual metabolites • Can ask questions such as: which compounds have a low concentration in group 0, 15 but increase in the group 35 and 45? or which compound is the only one significantly increased in group 45? Module 5 bioinformatics. ca

Heatmap Visualization Note that the Heatmap is not being clustered on Rows. It is ordered by the class labels Module 5 bioinformatics. ca
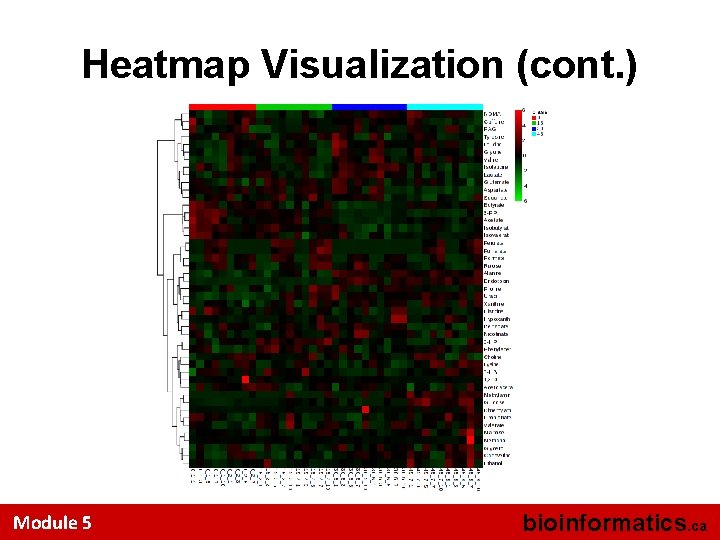
Heatmap Visualization (cont. ) Module 5 bioinformatics. ca

What’s Next? • Most of the multivariate analysis is now done • Metabo. Analyst has been keeping track of the plots or graphs you have generated • Now its time to generate a printed report that summarizes what you’ve done and what you’ve found Module 5 bioinformatics. ca
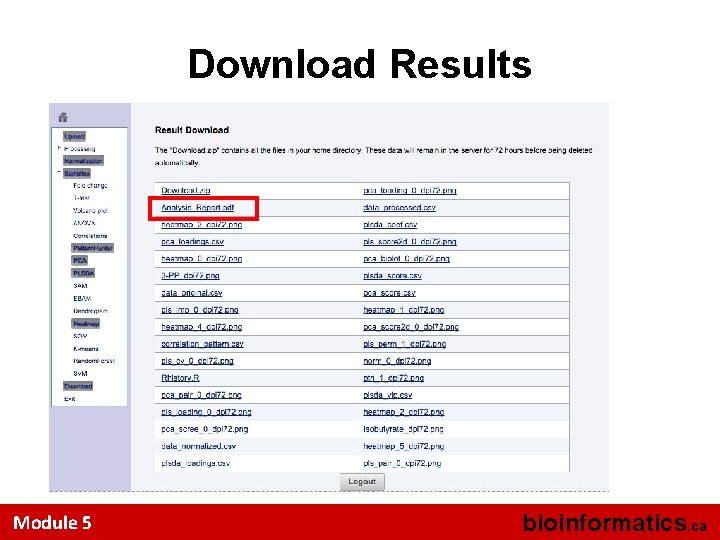
Download Results Module 5 bioinformatics. ca

Analysis Report Module 5 bioinformatics. ca

Select a Module (Enrichment Analysis) Module 5 bioinformatics. ca

Metabolite Set Enrichment Analysis (MSEA) http: //www. msea. ca Now part of Metaboanalyst Module 5 • Designed to handle lists of metabolites (with or without concentration data) • Modeled after Gene Set Enrichment Analysis (GSEA) • Supports over representation analysis (ORA), single sample profiling (SSP) and quantitative enrichment analysis (QEA) • Contains a library of 6300 pre-defined metabolite sets including 85 pathway sets & 850 disease sets bioinformatics. ca

Enrichment Analysis • Purpose: To test if there are biologically meaningful groups of metabolites that are significantly enriched in your data • Biological meaningful in terms of: – – Pathways Diseases Genetic variations Localization • Currently, MSEA only supports human metabolomic data Module 5 bioinformatics. ca

MSEA • Accepts 3 kinds of input files – list of metabolite names only (ORA – over representation analysis) – list of metabolite names + concentration data from a single sample (SSP – single sample profiling) – a concentration table with a list of metabolite names + concentrations for multiple samples/patients (QEA – quantitative enrichment analysis) Module 5 bioinformatics. ca
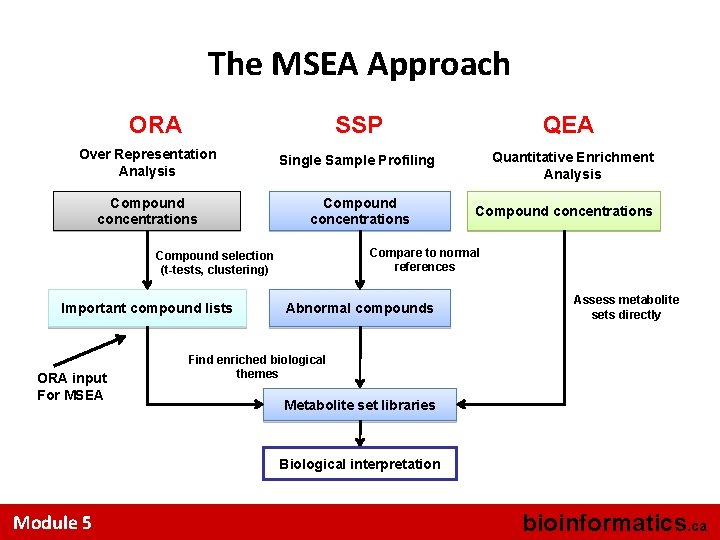
The MSEA Approach ORA SSP QEA Over Representation Analysis Single Sample Profiling Quantitative Enrichment Analysis Compound concentrations Compare to normal references Compound selection (t-tests, clustering) Important compound lists ORA input For MSEA Compound concentrations Abnormal compounds Assess metabolite sets directly Find enriched biological themes Metabolite set libraries Biological interpretation Module 5 bioinformatics. ca

Data Set Selected • Here we are using a collection of metabolites identified by NMR (compound list + concentrations) from the urine from 77 lung and colon cancer patients, some of whom were suffering from cachexia (muscle wasting) Module 5 bioinformatics. ca

Start with a Compound List for ORA Module 5 bioinformatics. ca

Upload Compound List Normally GSEA would require a list of all known genes for the given platform. Here we just use the list of metabolites found in KEGG. ORA is a “weak” analysis in MSEA Module 5 bioinformatics. ca

Perform Compound Name Standardization Module 5 bioinformatics. ca
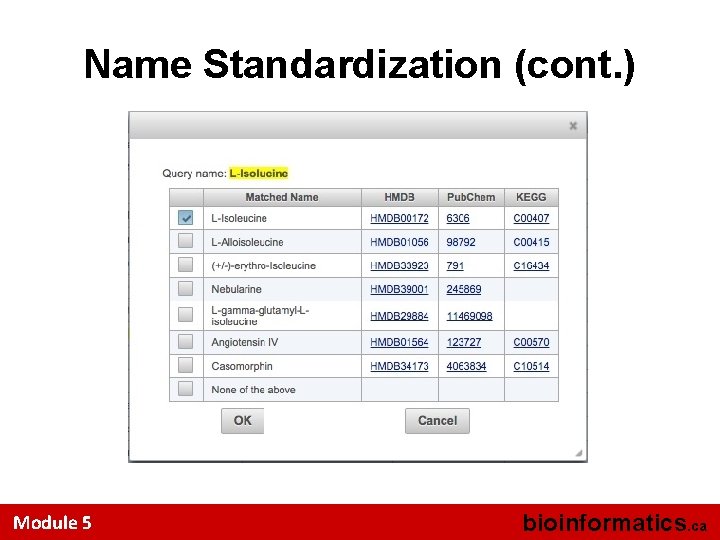
Name Standardization (cont. ) Module 5 bioinformatics. ca

Select a Metabolite Set Library Module 5 bioinformatics. ca
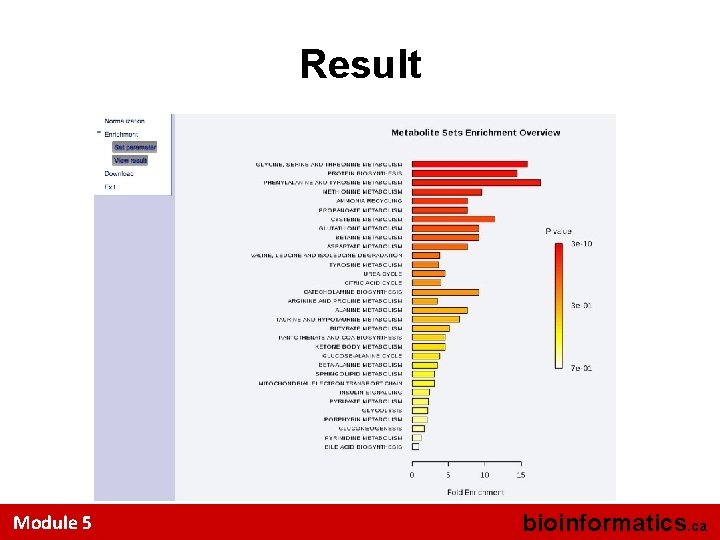
Result Module 5 bioinformatics. ca
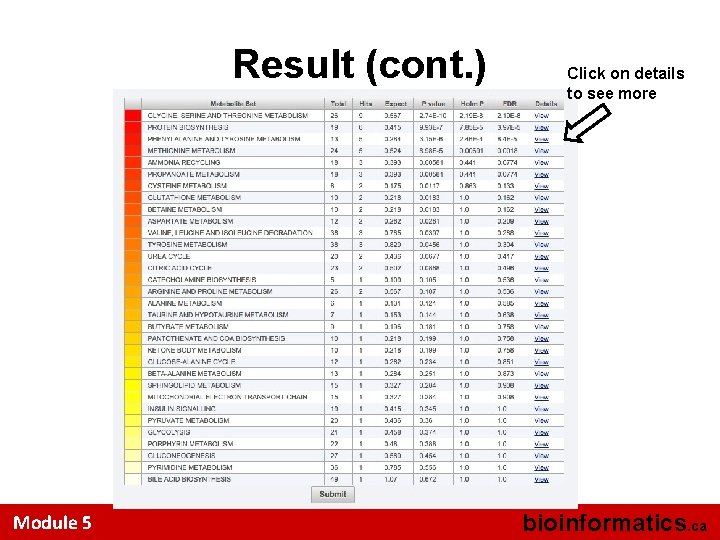
Result (cont. ) Module 5 Click on details to see more bioinformatics. ca

The Matched Metabolite Set Click on SMPDB to see more information Module 5 bioinformatics. ca

Phenylalanine and Tyrosine Metabolism in SMPDB Module 5 bioinformatics. ca
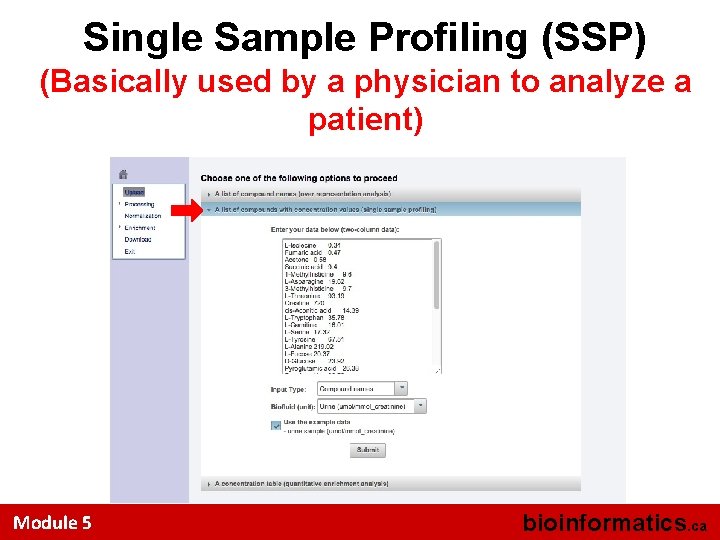
Single Sample Profiling (SSP) (Basically used by a physician to analyze a patient) Module 5 bioinformatics. ca
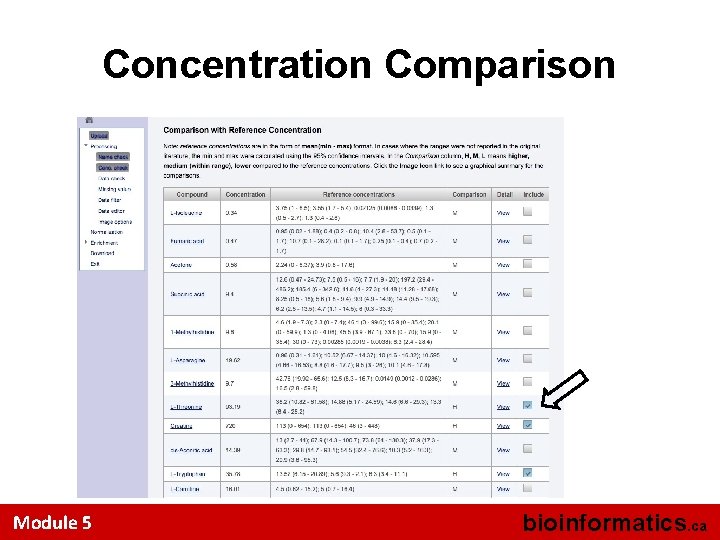
Concentration Comparison Module 5 bioinformatics. ca
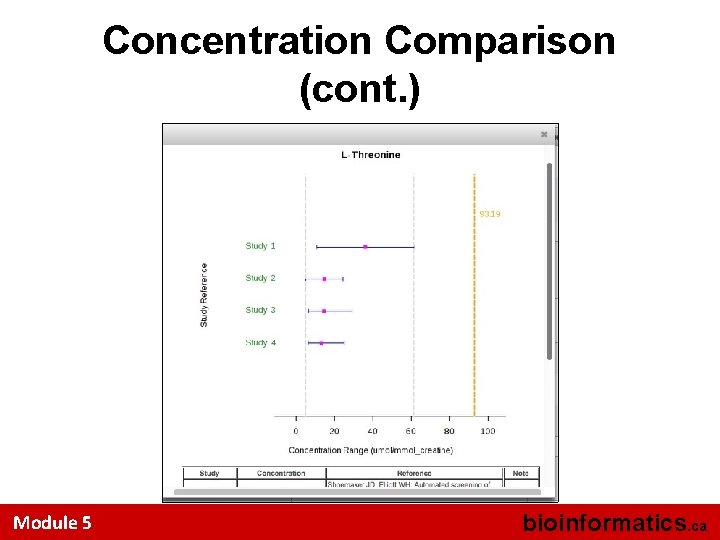
Concentration Comparison (cont. ) Module 5 bioinformatics. ca

Quantitative Enrichment Analysis (QEA) Module 5 bioinformatics. ca
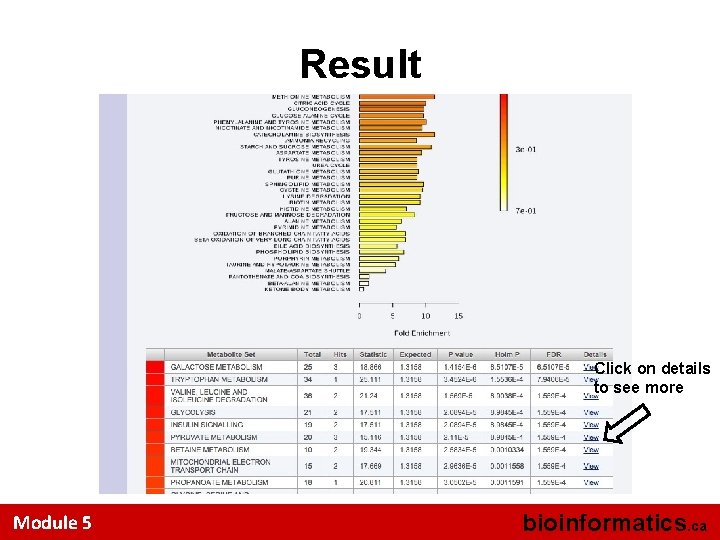
Result Click on details to see more Module 5 bioinformatics. ca
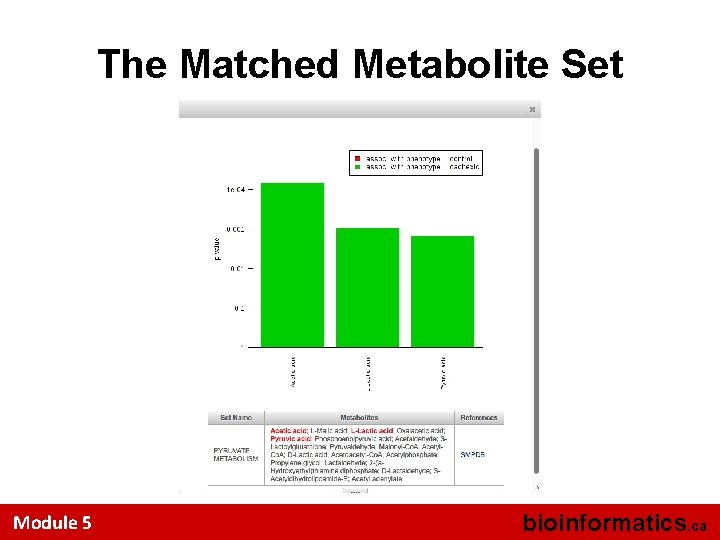
The Matched Metabolite Set Module 5 bioinformatics. ca

Select a Module (Pathway Analysis) Module 5 bioinformatics. ca

Pathway Analysis • Purpose: to extend and enhance metabolite set enrichment analysis for pathways by – Considering pathway structures – Supporting pathway visualization • Currently supports analysis for 21 diverse (model) organisms such as humans, mouse, drosophila, arabadopsis, E. coli, yeast, etc. (KEGG pathways only) Module 5 bioinformatics. ca

Data Set Selected • Here we are using a collection of metabolites identified by NMR (compound list + concentrations) from the urine from 77 lung and colon cancer patients, some of whom were suffering from cachexia (muscle wasting) Module 5 bioinformatics. ca

Pathway Analysis Module 5 bioinformatics. ca
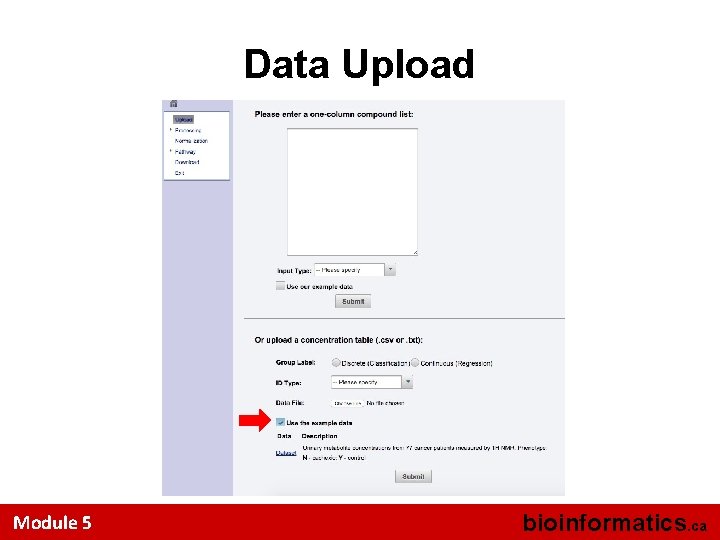
Data Upload Module 5 bioinformatics. ca

Perform Data Normalization Module 5 bioinformatics. ca

Select Pathway Libraries Module 5 bioinformatics. ca

Perform Network Topology Analysis Module 5 bioinformatics. ca
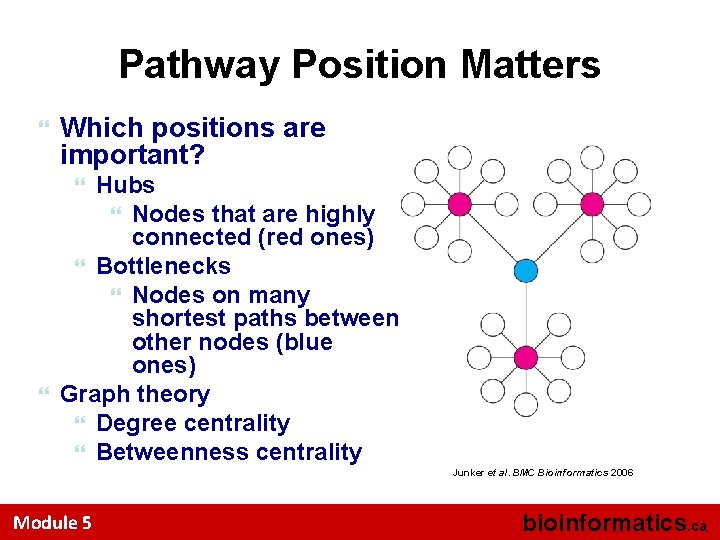
Pathway Position Matters Which positions are important? Hubs Nodes that are highly connected (red ones) Bottlenecks Nodes on many shortest paths between other nodes (blue ones) Graph theory Degree centrality Betweenness centrality Junker et al. BMC Bioinformatics 2006 Module 5 bioinformatics. ca
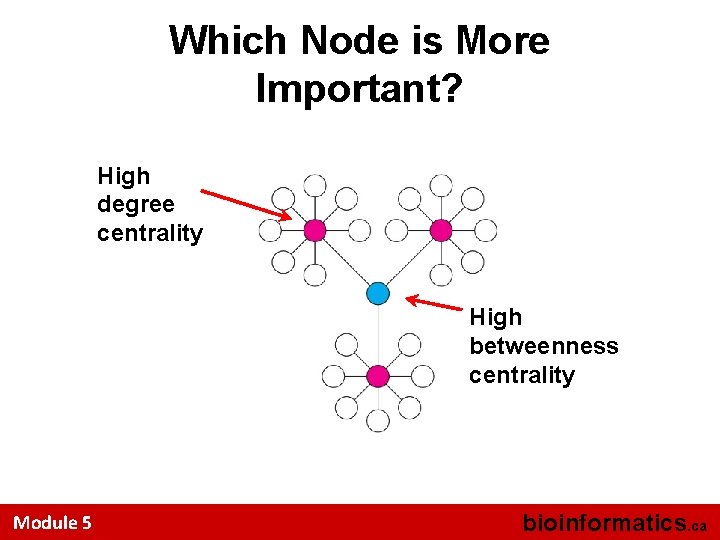
Which Node is More Important? High degree centrality High betweenness centrality Module 5 bioinformatics. ca
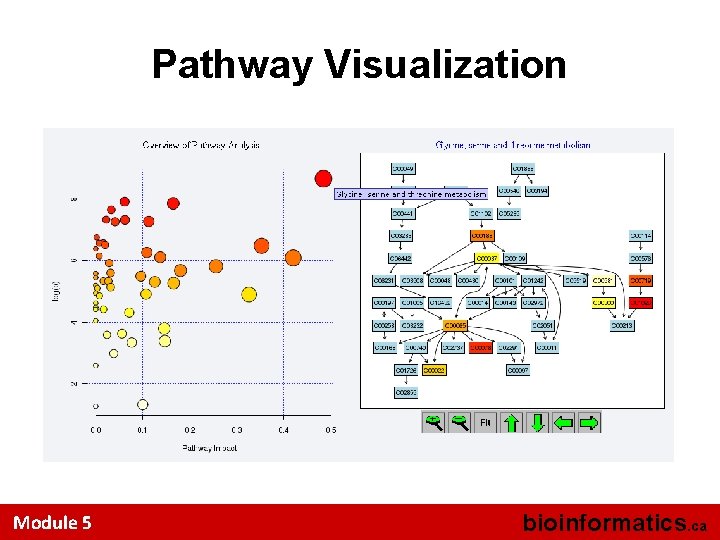
Pathway Visualization Module 5 bioinformatics. ca

Pathway Visualization (cont. ) Module 5 bioinformatics. ca

Pathway Impact • Incorporates parameters such as the log fold-change of the DE metabolites, the statistical significance of the set of pathway genes and the topology of the signaling pathway • Combines the pathway topology with the over-representation evidence Module 5 bioinformatics. ca
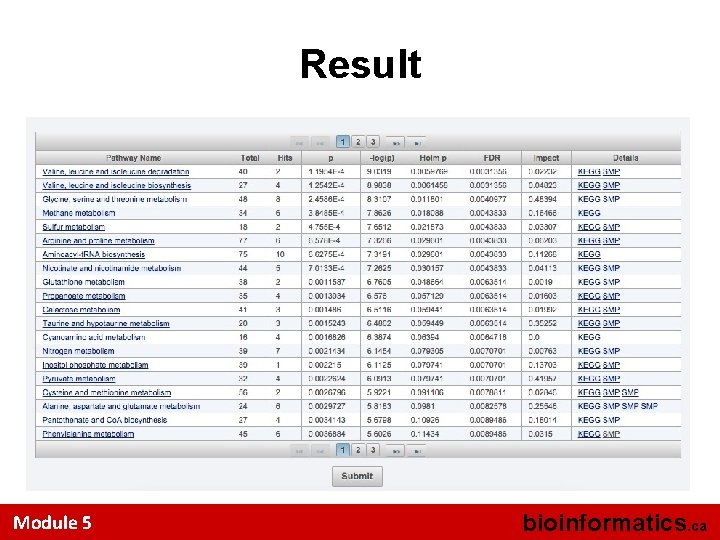
Result Module 5 bioinformatics. ca

Select a Module (Biomarker Analysis) Module 5 bioinformatics. ca

Biomarker Analysis • Purpose is to find biomarkers using ROC (receiver operator characteristic) curves with high sensitivity and specificity • Maximize AUC under ROC curve while minimizing the number of metabolites used in the biomarker panel • 3 different modules (univariate – single marker at a time, multivariate – many combinations of biomarkers, manual – user choice) Module 5 bioinformatics. ca
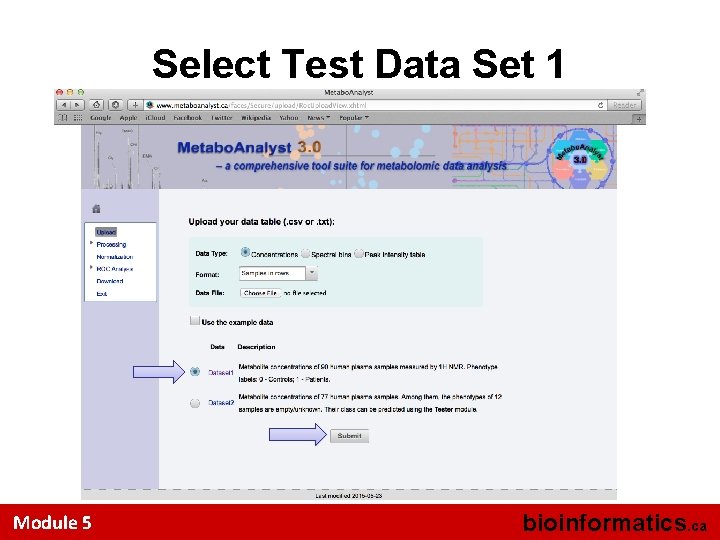
Select Test Data Set 1 Module 5 bioinformatics. ca

Data Set Selected • 90 patients (expectant mothers) at 3 months pregnancy • Serum samples • 45 patients went on to develop preeclampsia at 6 -7 months • 45 patients had normal pregancies • Trying to find biomarkers for predicting early pre-eclampsia Module 5 bioinformatics. ca
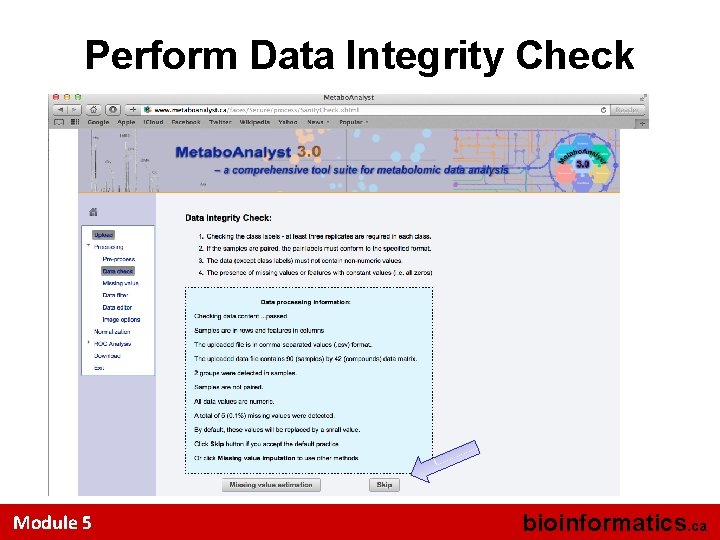
Perform Data Integrity Check Module 5 bioinformatics. ca

Perform Log Normalization Module 5 bioinformatics. ca
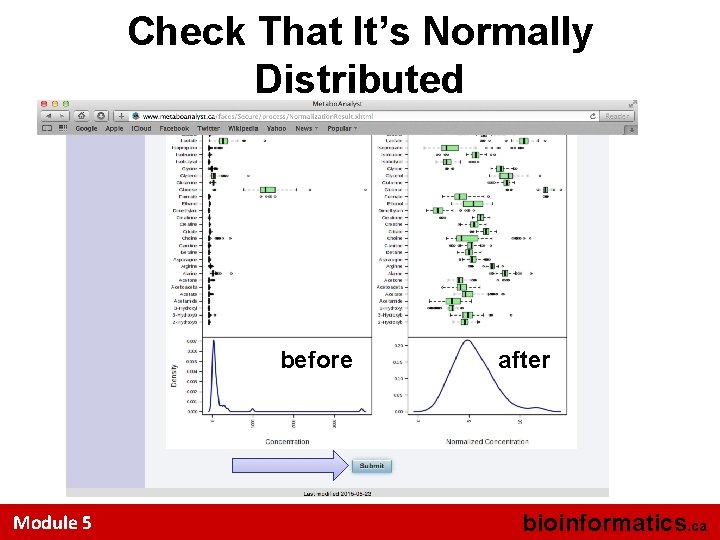
Check That It’s Normally Distributed before Module 5 after bioinformatics. ca

Select Multivariate Option Module 5 bioinformatics. ca

View ROC Curve Module 5 bioinformatics. ca

Choose a Model (95% conf. ) Select model Module 5 bioinformatics. ca

95% Confidence Interval Module 5 bioinformatics. ca

Select Sig. Features Tab Module 5 bioinformatics. ca
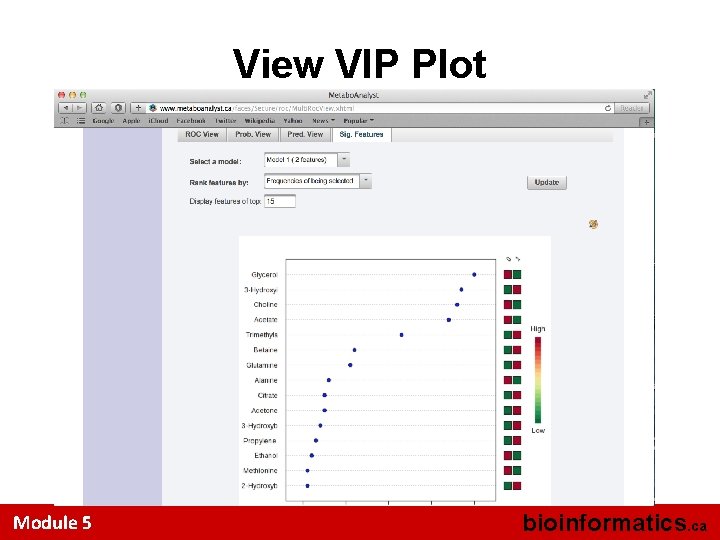
View VIP Plot Module 5 bioinformatics. ca

Select a Module (Power Analysis) Module 5 bioinformatics. ca

Statistical Power • Statistical power is the ability of a test to detect an effect, if the effect actually exists – A power of 0. 8 in a clinical trial means that the study has a 80% chance of ending up with a statistically significant treatment effect if there really was an important difference between treatments. • To answer research questions: – How powerful is my study? – How many samples do I need to have for what I want to get from the study? Module 5 bioinformatics. ca

Statistical Power (cont. ) • The statistical power of a test depends: 1. Sample size, 2. Significance criterion (alpha) 3. Effect size Increase power • Effect size • Sample size Decrease Power • Significance criterion Module 5 bioinformatics. ca

The Approach • How do we get these values? – Effect size can be estimated from a pilot data; – Significance criteria • Single metabolite - p value cutoff (i. e. 0. 05, 0. 01) • Metabolomics data – FDR (i. e. 0. 1) – Sample size is our interest – Power value is our interest • You need to upload a pilot data, and set the criteria, Metabo. Analyst will compute a power vs. sample size curve by computing power values for a range of sample sizes [3, 1000] Module 5 bioinformatics. ca
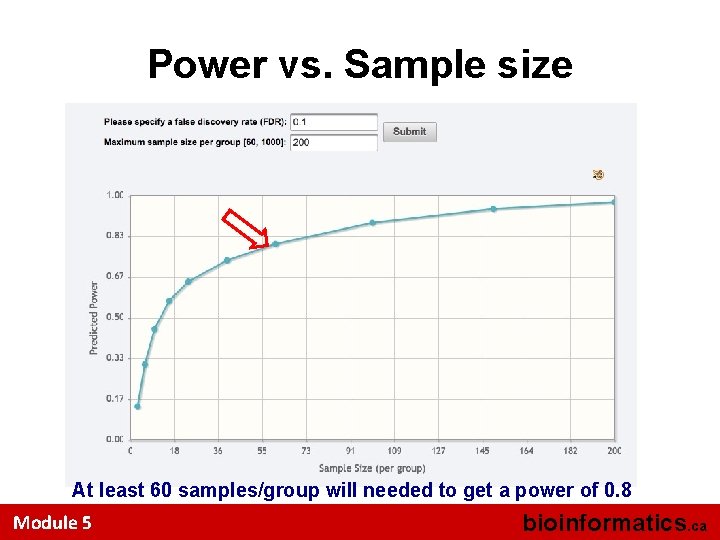
Power vs. Sample size At least 60 samples/group will needed to get a power of 0. 8 Module 5 bioinformatics. ca

Not Everything Was Covered • • • Clustering (K-means, SOM) Classification (SVM, random. Forests) Time-series data analysis Two factor data analysis Integrative pathway analysis (gene and metabolite) Module 5 bioinformatics. ca
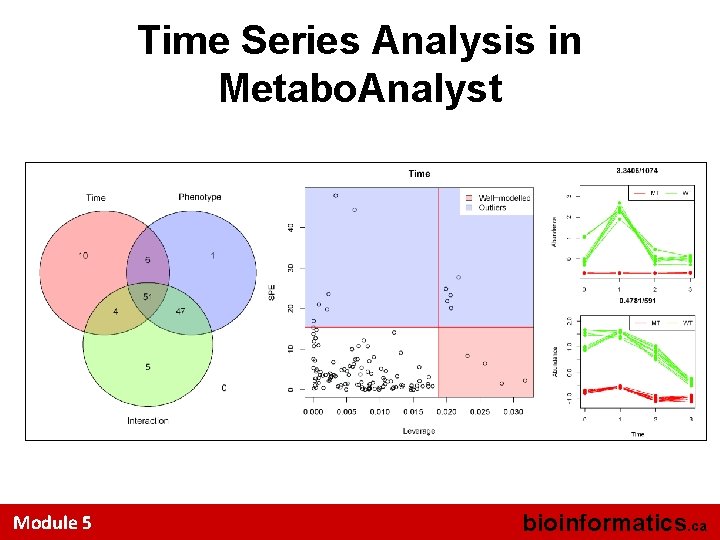
Time Series Analysis in Metabo. Analyst Module 5 bioinformatics. ca
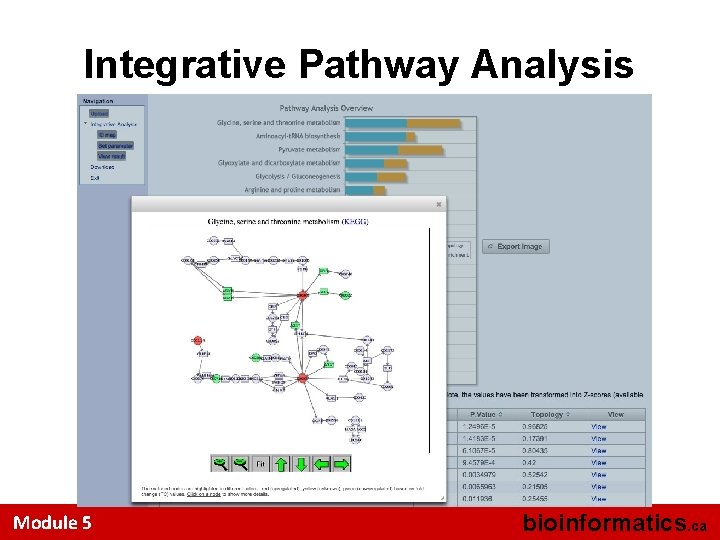
Integrative Pathway Analysis Module 5 bioinformatics. ca
- Slides: 124