An Introduction to Bioinformatics Algorithms www bioalgorithms info

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Molecular Biology Primer Angela Brooks, Raymond Brown, Calvin Chen, Mike Daly, Hoa Dinh, Erinn Hama, Robert Hinman, Julio Ng, Michael Sneddon, Hoa Troung, Jerry Wang, Che Fung Yung

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Outline: • • What Is Life Made of? What Is Genetic Material? What Do Genes Do? What Molecules Code for Genes? What Is the Structure of DNA? What Carries Information from DNA to Proteins How Are Proteins Made?

An Introduction to Bioinformatics Algorithms Outline Cont. • How Can We Analyze DNA • • Copying DNA Cutting and Pasting DNA Sequencing Probing DNA www. bioalgorithms. info
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Section 1: What is life made of?

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Cells • Fundamental working units of every living system. • Every organism is composed of one of two radically different types of cells: • prokaryotic cells. • eukaryotic cells. • Prokaryotes and eukaryotes are descended from the same primitive cell. • All extant prokaryotic and eukaryotic cells are the result of a total of 3. 5 billion years of evolution.
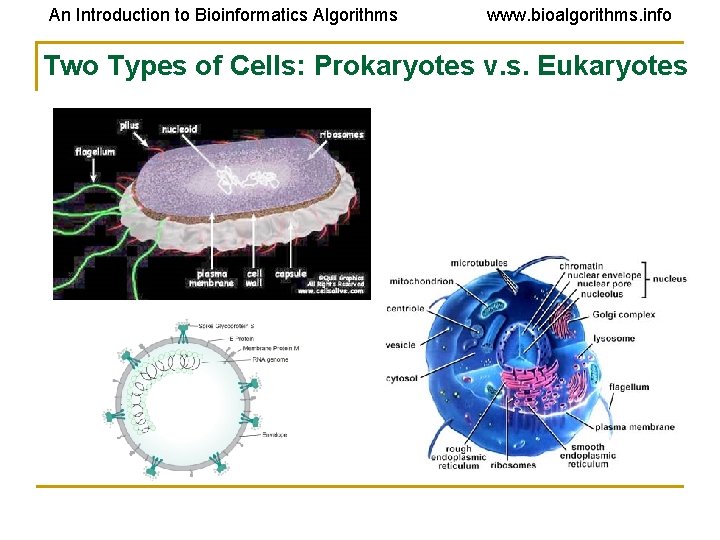
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Two Types of Cells: Prokaryotes v. s. Eukaryotes
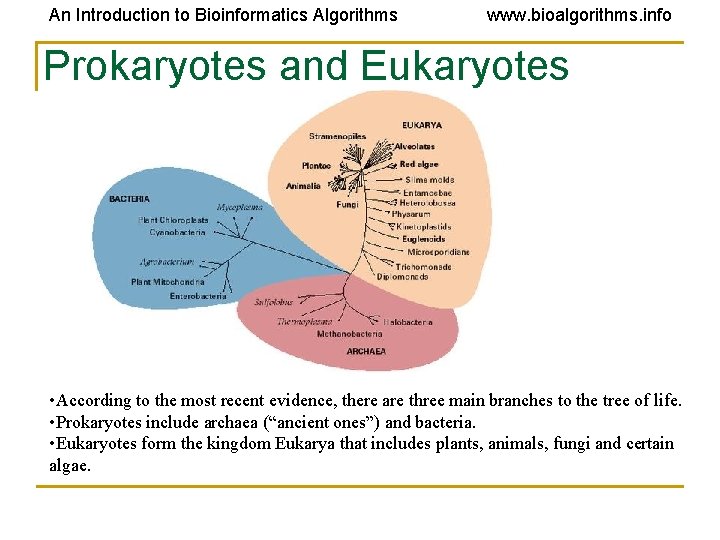
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Prokaryotes and Eukaryotes • According to the most recent evidence, there are three main branches to the tree of life. • Prokaryotes include archaea (“ancient ones”) and bacteria. • Eukaryotes form the kingdom Eukarya that includes plants, animals, fungi and certain algae.

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Prokaryotes and Eukaryotes, continued Prokaryotes Eukaryotes Single cell Single or multi cell No nucleus No organelles One piece of circular DNA Chromosomes No m. RNA post Exon/Intron splicing transcriptional modification

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Some Terminology • Genome: An organism’s genetic material • a bacteria contains about 600, 000 DNA base pairs • human and mouse genomes have some 3 billion • consists of one of more chromosomes • Gene: A discrete unit of hereditary information located on the chromosomes and consisting of DNA bases (or nucleotides). It is a basic physical and functional unit of heredity, and encodes instructions on how to make proteins. • Genotype: The genetic makeup of an organism • Phenotype: The physically expressed traits of an organism • Nucleic acid: Biological molecules (RNA and DNA) that allow organisms to reproduce

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info All life depends on 3 critical molecules • DNAs • Hold information on how cell works • Made of 4 types of nucleotides • RNAs • Act to transfer short pieces of information to different parts of cell • Provide templates to synthesize into proteins • Also made of 4 types of nucleotides • Proteins • Make up the cellular structure • large, complex molecules made up of 20 types of smaller subunits called amino acids • Form enzymes that send signals to other cells and regulate gene activity • Form body’s major components (e. g. , hair, skin, etc. )
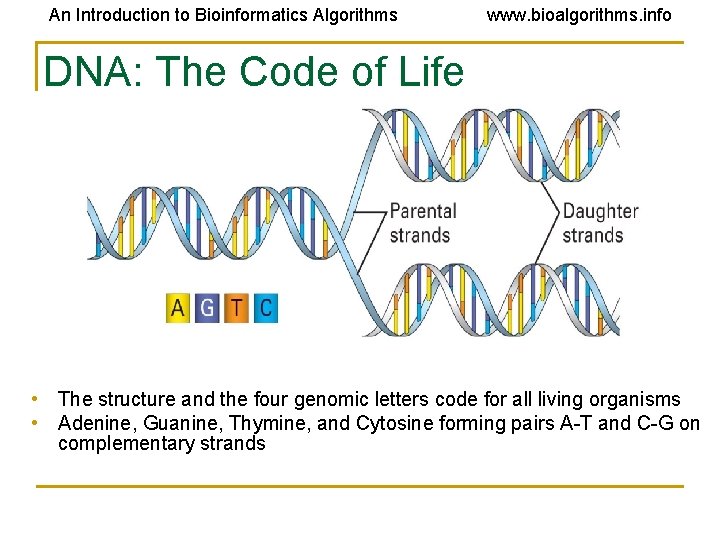
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info DNA: The Code of Life • The structure and the four genomic letters code for all living organisms • Adenine, Guanine, Thymine, and Cytosine forming pairs A-T and C-G on complementary strands
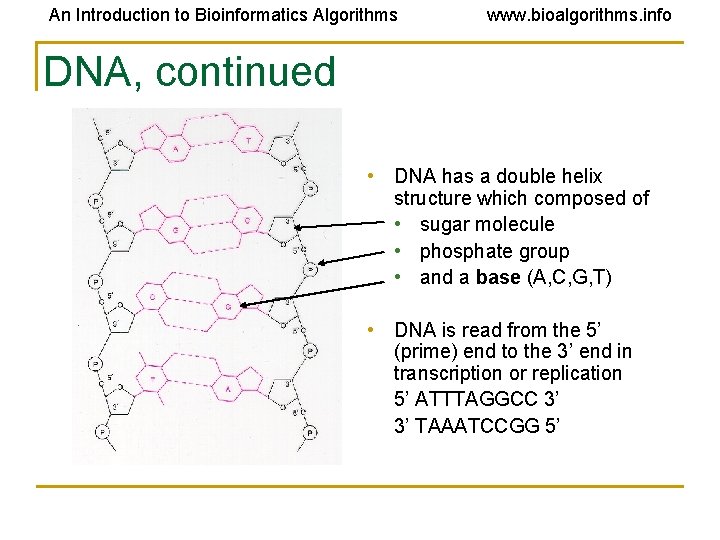
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info DNA, continued • DNA has a double helix structure which composed of • sugar molecule • phosphate group • and a base (A, C, G, T) • DNA is read from the 5’ (prime) end to the 3’ end in transcription or replication 5’ ATTTAGGCC 3’ 3’ TAAATCCGG 5’
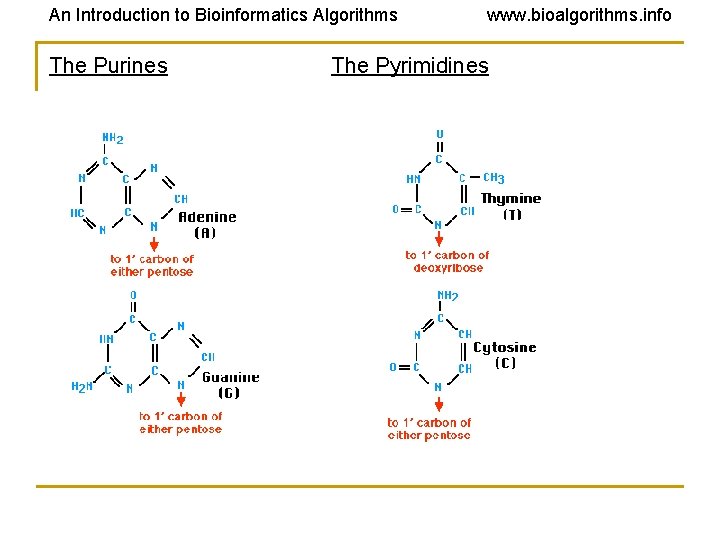
An Introduction to Bioinformatics Algorithms The Purines www. bioalgorithms. info The Pyrimidines
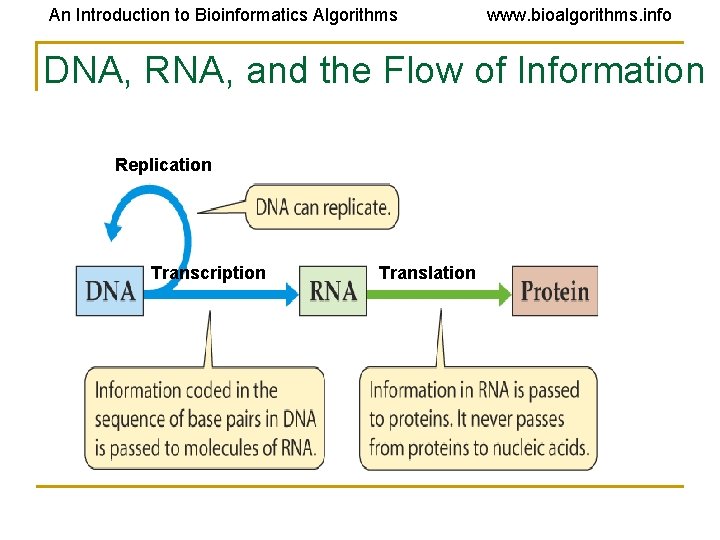
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info DNA, RNA, and the Flow of Information Replication Transcription Translation

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Cell Information: Instruction book of life • DNA, RNA, and proteins are examples of strings written in either the four-letter nucleotide alphabet for DNAs and RNAs (A, C, G, T/U) • or the twenty-letter amino acid alphabet for proteins. Each amino acid is coded by 3 nucleotides called a codon

An Introduction to Bioinformatics Algorithms What is genetic material? • Mendel’s experiments • Pea plant experiments • Mutations in DNA • Good, bad, silent • Chromosomes • Linked genes www. bioalgorithms. info

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info The Pea Plant Experiments • Mendel discovered that genes were passed on to offspring by both parents in two forms: dominant and recessive. • The dominant form would decide the phenotypic characteristic of the offspring

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info DNA: The building blocks of genetic material • DNA was later discovered to be the molecule that makes up the inherited genetic material. • DNA provides a code, consisting of 4 letters, for all cellular functions.

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Mutation • The DNA can be thought of as a sequence of the nucleotides: C, A, G, or T. • What happens to genes when the DNA sequence is mutated?

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info The Good, the Bad, and the Silent • Mutations may serve an organism in three ways: • The Good : A mutation cause a trait that enhances the organism’s function: Mutation in the sickle cell gene provides resistance to malaria. • The Bad : A mutation cause a trait that is harmful, sometimes fatal to the organism: Huntington’s disease, a symptom of gene mutations, is a degenerative disease of the nervous system. • The Silent: A mutation may simply cause no difference in the functions of the organism. Campbell, Biology, 5 th edition, p. 255

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Genes (encoded in DNA) are organized into chromosomes • What are chromosomes? It is a threadlike structure found in the nucleus of the cell which is made from a long strand of DNA. Different organisms have different numbers of chromosomes in their cells. • Human genome has 24 distinct chromosomes.
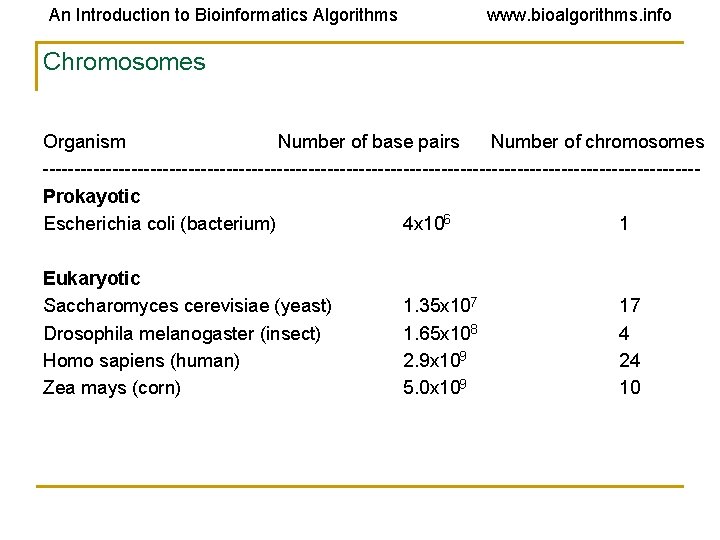
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Chromosomes Organism Number of base pairs Number of chromosomes ----------------------------------------------------Prokayotic Escherichia coli (bacterium) 4 x 106 1 Eukaryotic Saccharomyces cerevisiae (yeast) 1. 35 x 107 17 Drosophila melanogaster (insect) 1. 65 x 108 4 Homo sapiens (human) 2. 9 x 109 24 Zea mays (corn) 5. 0 x 109 10

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info What do genes do? • Design of life (genes -> proteins) • Protein synthesis -- central dogma of molecular biology Closer genes are more likely inherited together from the same parent!
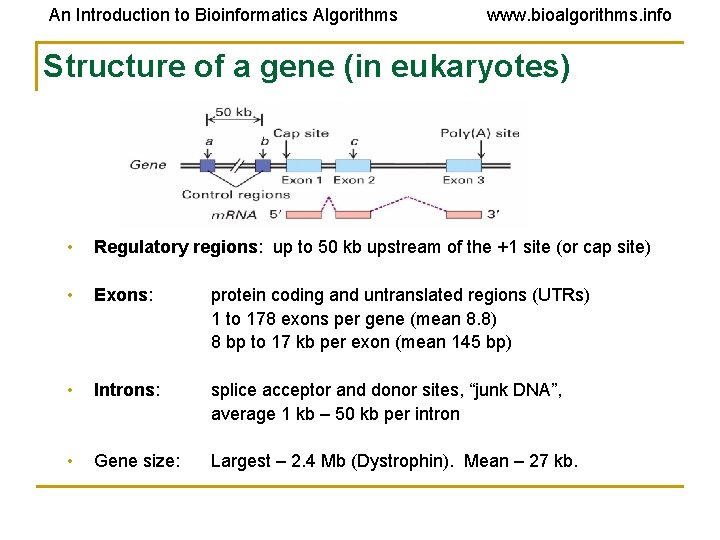
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Structure of a gene (in eukaryotes) • • Regulatory regions: up to 50 kb upstream of the +1 site (or cap site) Exons: protein coding and untranslated regions (UTRs) 1 to 178 exons per gene (mean 8. 8) 8 bp to 17 kb per exon (mean 145 bp) • Introns: splice acceptor and donor sites, “junk DNA”, average 1 kb – 50 kb per intron • Gene size: Largest – 2. 4 Mb (Dystrophin). Mean – 27 kb.

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Proteins: Workhorses of the Cell • Made of 20 different amino acids • different chemical properties cause the protein chains to fold up into specific three-dimensional structures that define their particular functions in the cell. • Proteins do all essential work for the cell • • build cellular structures digest nutrients execute metabolic functions Mediate information flow within a cell and among cellular communities. • Proteins work together with other proteins or nucleic acids as "molecular machines" • structures that fit together and function in highly specific, lock-and -key ways. • examples include ribosomes, RNA polymerase, etc.
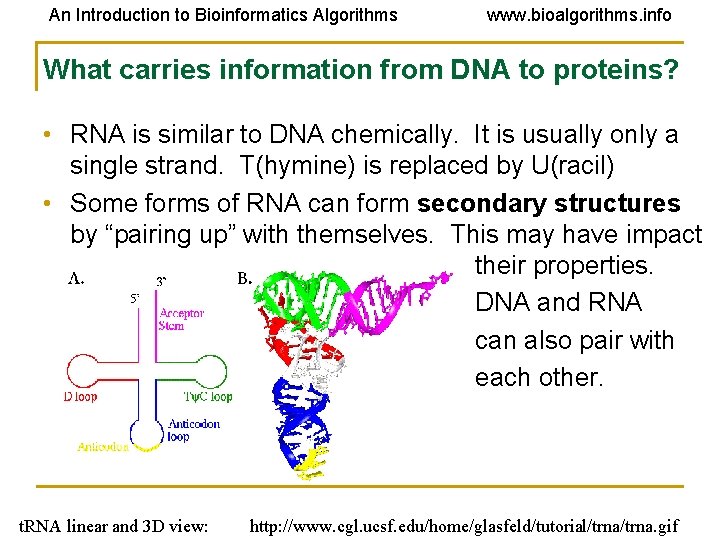
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info What carries information from DNA to proteins? • RNA is similar to DNA chemically. It is usually only a single strand. T(hymine) is replaced by U(racil) • Some forms of RNA can form secondary structures by “pairing up” with themselves. This may have impact on their properties. DNA and RNA can also pair with each other. t. RNA linear and 3 D view: http: //www. cgl. ucsf. edu/home/glasfeld/tutorial/trna. gif
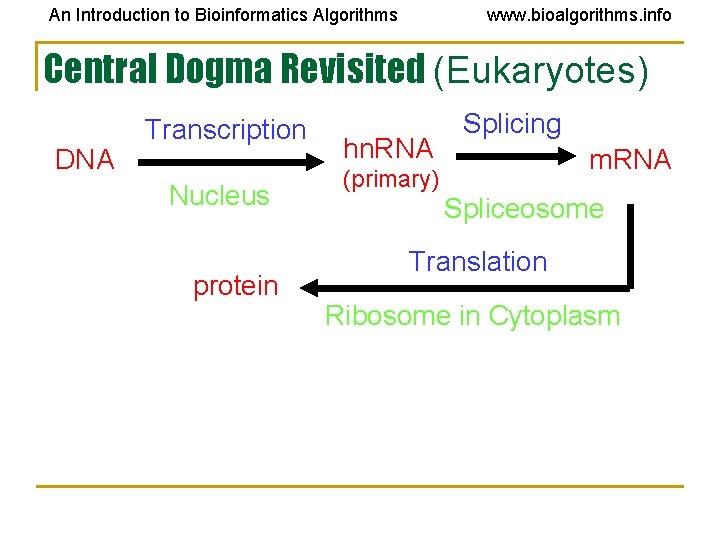
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Central Dogma Revisited (Eukaryotes) DNA Transcription Nucleus protein hn. RNA Splicing m. RNA (primary) Spliceosome Translation Ribosome in Cytoplasm

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Terminology for Splicing • Exon: A portion of the gene that appears in both the primary and the mature m. RNA transcripts. • Intron: A portion of the gene that is transcribed but excised prior to translation.
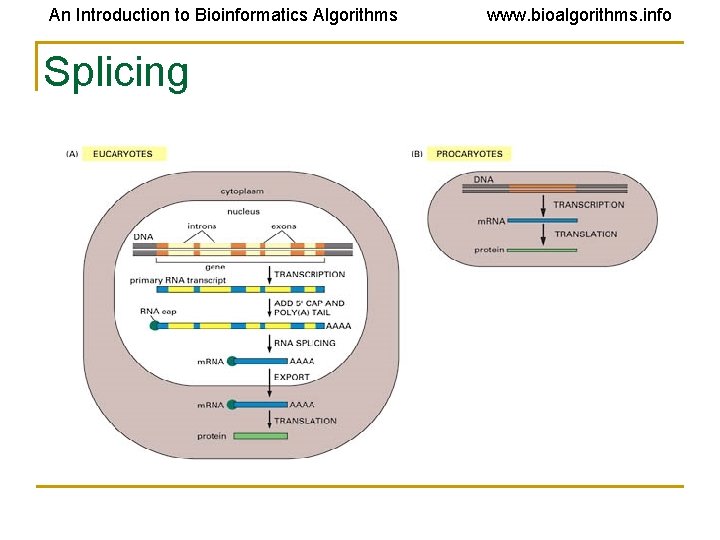
An Introduction to Bioinformatics Algorithms Splicing www. bioalgorithms. info

An Introduction to Bioinformatics Algorithms Splicing • Sometimes alternative splicing can create different valid proteins. • A typical eukaryotic gene has 4 -20 introns. Locating them by analytical means is not easy. www. bioalgorithms. info

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info RNA Protein: Translation • Ribosomes and transfer-RNAs (t. RNAs) run along the length of a newly synthesized m. RNA, decoding it one codon at a time to build a growing chain of amino acids (“polypeptide”) • The t. RNAs have anti-codons, which complementarily match the codons of m. RNA to know what amino acids get added next

An Introduction to Bioinformatics Algorithms Translation • The process of going from RNA to polypeptide (protein). • Three bases of RNA (called a codon) correspond to one amino acid based on a fixed table. • Always starts with Methionine (start codon) and ends with a stop codon. www. bioalgorithms. info
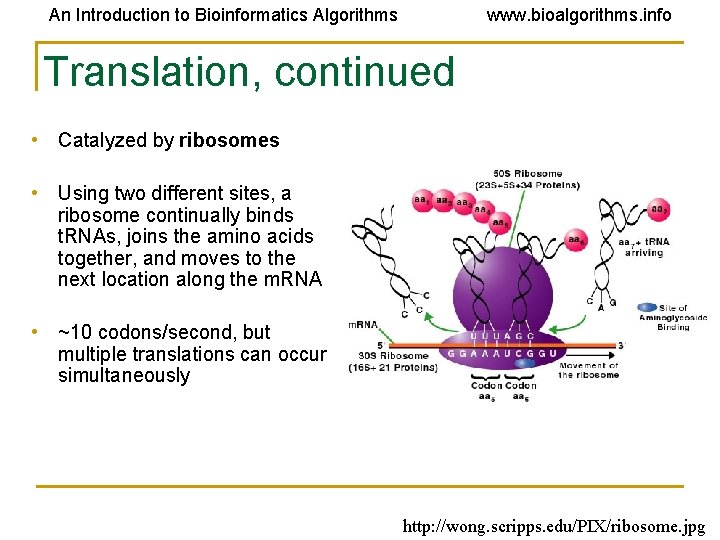
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Translation, continued • Catalyzed by ribosomes • Using two different sites, a ribosome continually binds t. RNAs, joins the amino acids together, and moves to the next location along the m. RNA • ~10 codons/second, but multiple translations can occur simultaneously http: //wong. scripps. edu/PIX/ribosome. jpg

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info What are proteins made of? • Complex organic molecules made up of amino acid subunits. • 20* different kinds of amino acids. Each has a 1 -letter and 3 -letter abbreviations. • A protein adopts a 3 D structure specific to its amino acid sequence and function. • Also called “polypeptides”. *Some other amino acids exist but not in humans.

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Protein Folding • • Proteins are not linear structures, though they are built that way. Proteins tend to fold into the lowest free energy conformation. Proteins begin to fold while the polypeptide is still being translated. The amino acids have very different chemical properties; they interact with each other after the protein has been formed. • This causes the protein to start folding and adopting its functional structure. • Proteins may fold in reaction to some ions, and several separate chains of peptides may join together through their hydrophobic and hydrophilic amino acids to form a polymer.
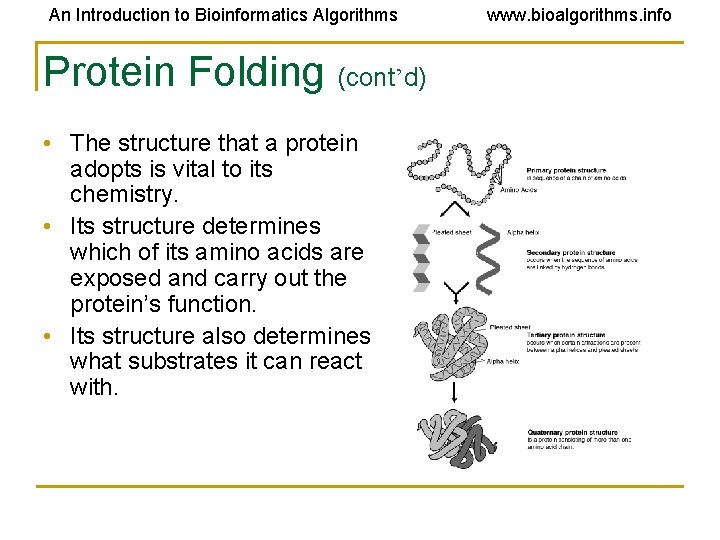
An Introduction to Bioinformatics Algorithms Protein Folding (cont’d) • The structure that a protein adopts is vital to its chemistry. • Its structure determines which of its amino acids are exposed and carry out the protein’s function. • Its structure also determines what substrates it can react with. www. bioalgorithms. info

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Copying DNA - Polymerase Chain Reaction (PCR) • PCR is used to massively replicate DNA sequences. • How it works: • • Separate the two strands with low heat. Add some bases, primer sequences (or oligos), and DNA Polymerase. • Creates double stranded DNA from a single strand. • Primer sequences create a seed from which double stranded DNA grows. Now you have two copies. Repeat. Amount of DNA grows exponentially. • 1→ 2→ 4→ 8→ 16→ 32→ 64→ 128→ 256…
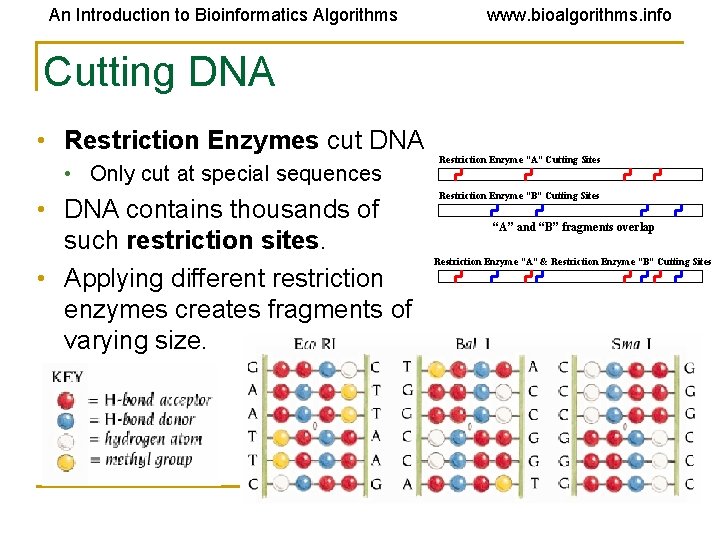
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Cutting DNA • Restriction Enzymes cut DNA • Only cut at special sequences • DNA contains thousands of such restriction sites. • Applying different restriction enzymes creates fragments of varying size. Restriction Enzyme “A” Cutting Sites Restriction Enzyme “B” Cutting Sites “A” and “B” fragments overlap Restriction Enzyme “A” & Restriction Enzyme “B” Cutting Sites
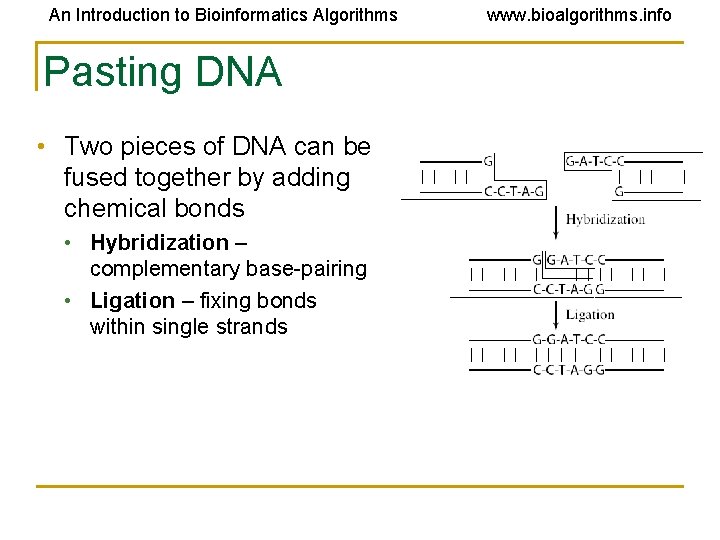
An Introduction to Bioinformatics Algorithms Pasting DNA • Two pieces of DNA can be fused together by adding chemical bonds • Hybridization – complementary base-pairing • Ligation – fixing bonds within single strands www. bioalgorithms. info
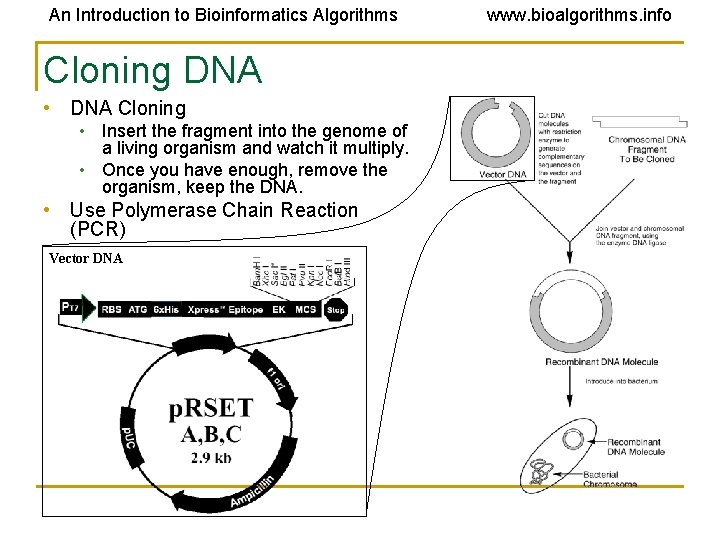
An Introduction to Bioinformatics Algorithms Cloning DNA • DNA Cloning • Insert the fragment into the genome of a living organism and watch it multiply. • Once you have enough, remove the organism, keep the DNA. • Use Polymerase Chain Reaction (PCR) Vector DNA www. bioalgorithms. info

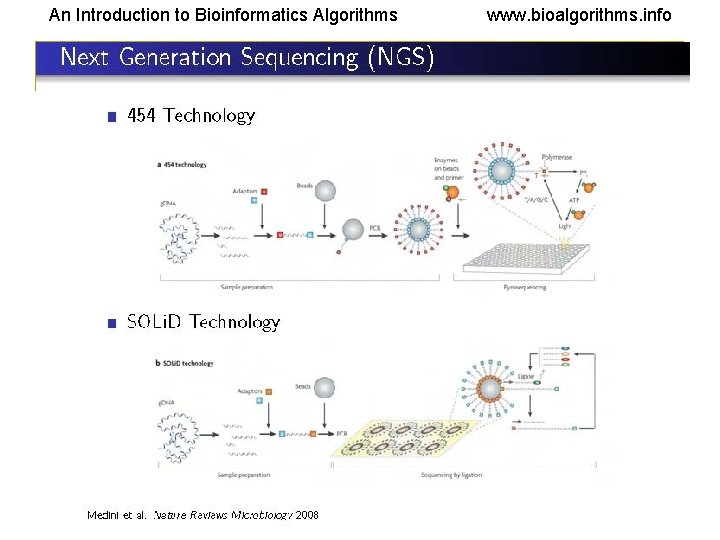
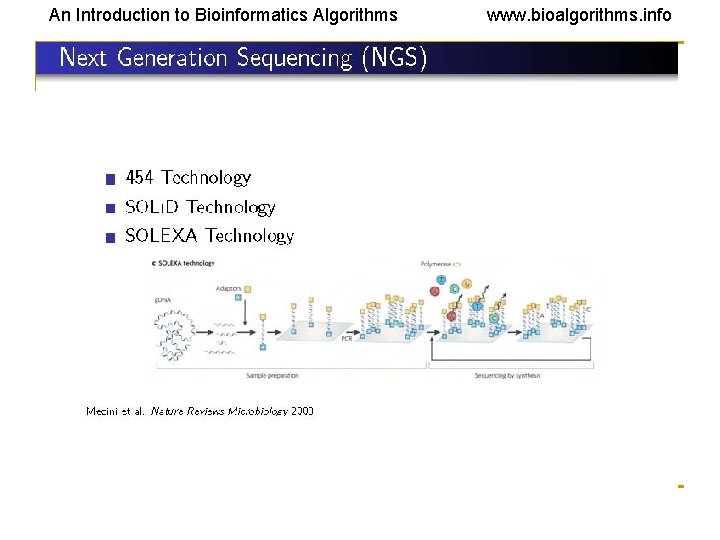
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Reading (Sequencing) DNA • Electrophoresis • • • Reading is done mostly by using this technique. This is based on separation of molecules by their sizes (and in 2 D gel by size and charge). DNA or RNA molecules are charged in aqueous solution and move to a definite direction by the action of an electric field. The DNA molecules are either labeled with radioisotopes or tagged with fluorescent dyes. In the latter, a laser beam can trace the dyes and send information to a computer. Given a DNA molecule, it is then possible to obtain all fragments from it that end in either A, or T, or G, or C and these can be sorted in a gel experiment. This (Sanger technique) usually produces reads of lengths between 500 bps and 1000 bps. • Another route to sequencing is direct sequencing using gene chips or NGS technologies, which have much higher throughputs but produce shorter reads (30 bps – 500 bps).
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info 10
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info 10

An Introduction to Bioinformatics Algorithms Assembling Genome • Cut it into many pieces • Sequence each random fragment and put them back together • Not as easy as it sounds • SCS problem (Shortest Common Superstring) • Some fragments overlap • Fit overlapping sequences together to get the shortest possible sequence (superstring) that contains all fragment sequences www. bioalgorithms. info

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Assembling Genome • DNA fragments contain sequencing errors. • Two complementary strands of DNA. • Need to take into account both orientations of DNA. • The human genome is a diploid. • Repeat problem • 50% of human DNA is just repeats. • If you have repeating DNA, how do you know where it goes? Hint: Repeats are usually different due to mutations. You could probably figure it out if you know the mutation rates between repeats and sequencing error rates.

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Probing DNA • DNA probes • • Oligonucleotide: single-stranded DNA of 20 -30 nucleotides long Oligonucleotides are used to find complementary DNA segments. Made by working backwards: AA sequence m. RNA c. DNA. Made with automated DNA synthesizers and tagged with a radioactive isotope. 60
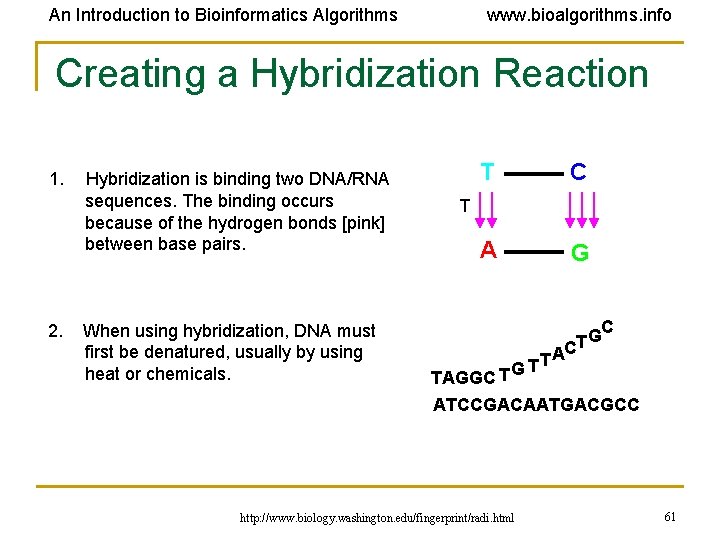
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Creating a Hybridization Reaction 1. Hybridization is binding two DNA/RNA sequences. The binding occurs because of the hydrogen bonds [pink] between base pairs. T C A G T 2. When using hybridization, DNA must GC T first be denatured, usually by using AC T T heat or chemicals. TAGGC T G ATCCGACAATGACGCC http: //www. biology. washington. edu/fingerprint/radi. html 61
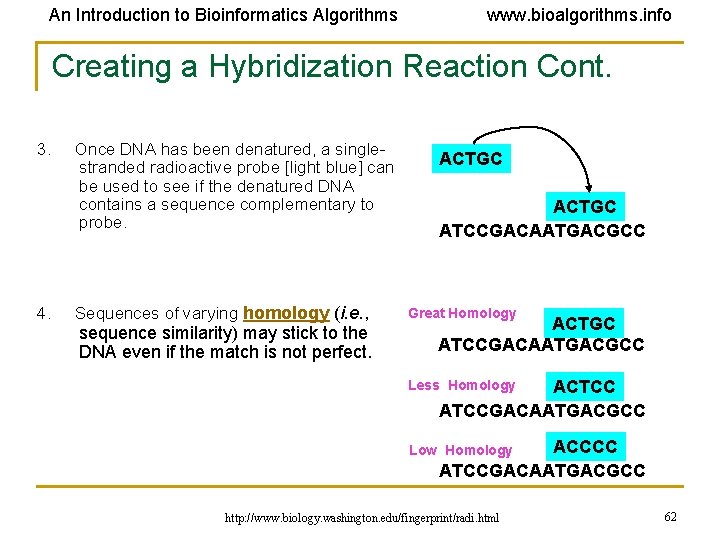
An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Creating a Hybridization Reaction Cont. 3. Once DNA has been denatured, a singlestranded radioactive probe [light blue] can be used to see if the denatured DNA contains a sequence complementary to probe. 4. Sequences of varying homology (i. e. , ACTGC ATCCGACAATGACGCC Great Homology ACTGC ATCCGACAATGACGCC sequence similarity) may stick to the DNA even if the match is not perfect. ACTCC ATCCGACAATGACGCC Less Homology ACCCC ATCCGACAATGACGCC Low Homology http: //www. biology. washington. edu/fingerprint/radi. html 62

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info DNA (Micro) Arrays --Technical Foundation • An array works by exploiting the ability of a given m. RNA molecule to hybridize to the DNA template. • Using an array containing many DNA probes (corresponding to different genes) in an experiment, the expression levels of hundreds or thousands genes within a cell is obtained by measuring the amount of m. RNA bound to each site on the array. • With the aid of a computer, the amount of m. RNA bound to the spots on the microarray is “precisely” measured, generating a profile of gene expression in the cell. • Microarrays suffer from high noise and are being quickly replaced by NGS methods (RNA-Seq). http: //www. ncbi. nih. gov/About/primer/microarrays. html 64

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info An Experiment on Microarray In this schematic: GREEN represents Control RNA RED represents Case RNA YELLOW represents a combination of Control and Case RNA BLACK represents areas where neither the Control nor Case RNA Each color in an array represents either healthy (control) or diseased (case) tissue. The location and intensity of a color tell us whether a gene, or its mutation, is expressed in the control and/or case RNA. http: //www. ncbi. nih. gov/About/primer/microarrays. html

An Introduction to Bioinformatics Algorithms www. bioalgorithms. info Sources Cited • • • Daniel Sam, “Greedy Algorithm” presentation. Glenn Tesler, “Genome Rearrangements in Mammalian Evolution: Lessons from Human and Mouse Genomes” presentation. Ernst Mayr, “What evolution is”. Neil C. Jones, Pavel A. Pevzner, “An Introduction to Bioinformatics Algorithms”. Alberts, Bruce, Alexander Johnson, Julian Lewis, Martin Raff, Keith Roberts, Peter Walter. Molecular Biology of the Cell. New York: Garland Science. 2002. Mount, Ellis, Barbara A. List. Milestones in Science & Technology. Phoenix: The Oryx Press. 1994. Voet, Donald, Judith Voet, Charlotte Pratt. Fundamentals of Biochemistry. New Jersey: John Wiley & Sons, Inc. 2002. Campbell, Neil. Biology, Third Edition. The Benjamin/Cummings Publishing Company, Inc. , 1993. Snustad, Peter and Simmons, Michael. Principles of Genetics. John Wiley & Sons, Inc, 2003.
- Slides: 51