CS 5263 Bioinformatics Lecture 8 Multiple Sequence Alignment

CS 5263 Bioinformatics Lecture 8: Multiple Sequence Alignment

Roadmap • Homework? • Review of last lecture • Multiple sequence alignment
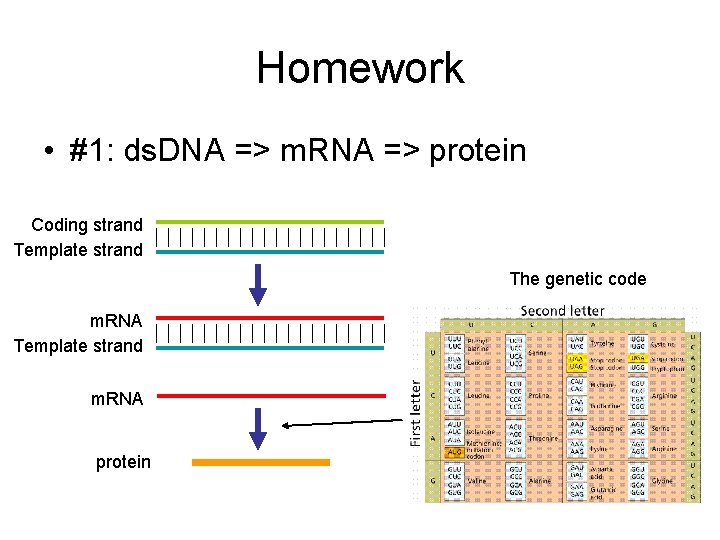
Homework • #1: ds. DNA => m. RNA => protein Coding strand Template strand The genetic code m. RNA Template strand m. RNA protein

Problem #2 • For two strings of lengths m and n, the number of alignment is equal to the number of paths from (0, 0) to (m, n) – How many ways we can get to (i, j) depend on how many ways we can get to its preceding neighbors
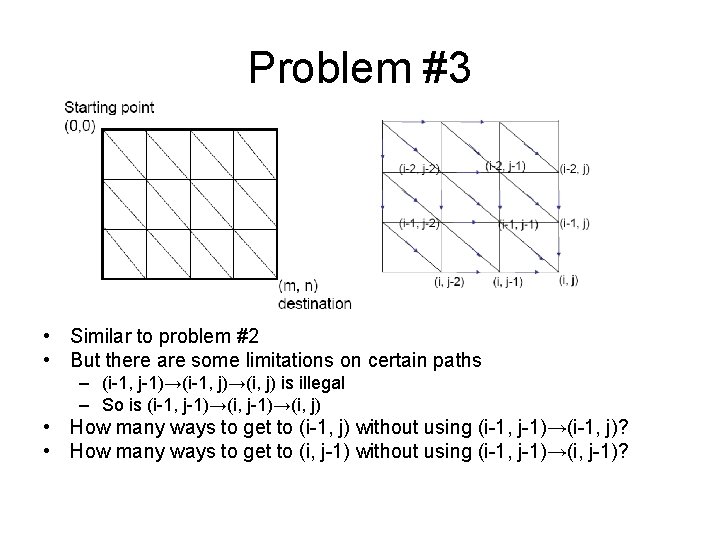
Problem #3 • Similar to problem #2 • But there are some limitations on certain paths – (i-1, j-1)→(i-1, j)→(i, j) is illegal – So is (i-1, j-1)→(i, j) • How many ways to get to (i-1, j) without using (i-1, j-1)→(i-1, j)? • How many ways to get to (i, j-1) without using (i-1, j-1)→(i, j-1)?

Problem #4 • Implementation is easy • Histogram: how you bin it may affect your results – bin for each discrete value you observed in your scores • Scores related to base frequency? • Scores differ between global and local alignments? • Score distribution?

BLAST Main idea: Construct a dictionary of all the words in the query Alignment initiated between words of alignment score T …… query …… Alignment: Ungapped extensions until score below statistical threshold Output: All local alignments with score > statistical threshold scan DB query
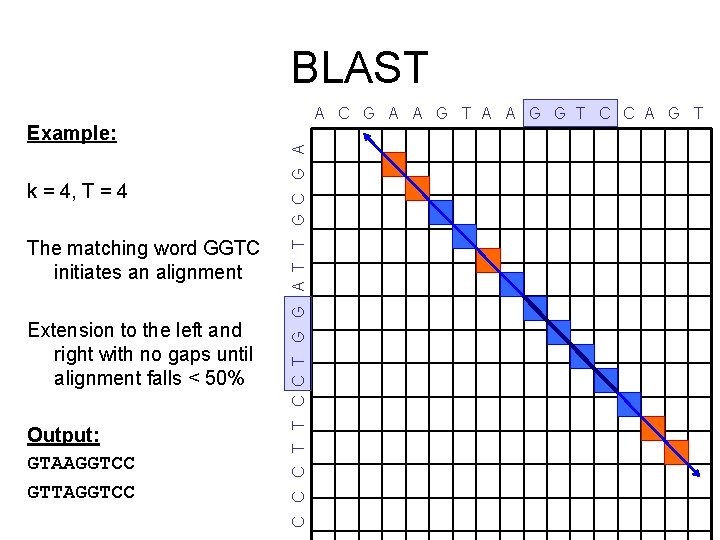
BLAST k = 4, T = 4 The matching word GGTC initiates an alignment Extension to the left and right with no gaps until alignment falls < 50% Output: GTAAGGTCC GTTAGGTCC C T T C C T G G A T T G C G A Example: A C G A A G T A A G G T C C A G T

Gapped BLAST • • Pairs of words can initiate alignment Extensions with gaps in a band around anchor Output: GTAAGGTCCAGT GTTAGGTC-AGT C T G A T C C T G G A T T G C G A Added features: A C G A A G T A A G G T C C A G T

• Advantages – Fast!!!! – A few minute to search a database of 1011 bases • Disadvantages – Sensitivity may be low – Often misses weak homologies

New improvement • Make it even faster – But even less sensitive – Mainly for aligning very similar sequences or really long sequences • E. g. whole genome vs whole genome • Make it more sensitive – PSI-BLAST: iteratively add more homologous sequences – Pattern. Hunter: discontinuous seeds

Things we’ve covered so far • Global alignment – Needleman-Wunsch and variants – Improvement on space and time • Local Alignment – Smith-Waterman • Heuristic algorithms – BLAST families • Statistics for sequence alignment – Extreme value distribution

Commonality • They all deal with aligning two sequences – Pair-wise sequence alignment

Today • Aligning multiple sequences all together – Multiple sequence alignment


Motivation • A faint similarity between two sequences becomes very significant if present in many • Protein domains • Motifs responsible for gene regulation

Definition • Given N sequences x 1, x 2, …, x. N: – Insert gaps (-) in each sequence xi, such that • All sequences have the same length L • Score of the global map is maximum • Pairwise alignment: a hypothesis on the evolutionary relationship between the letters of two sequences • Same for a multiple alignment!
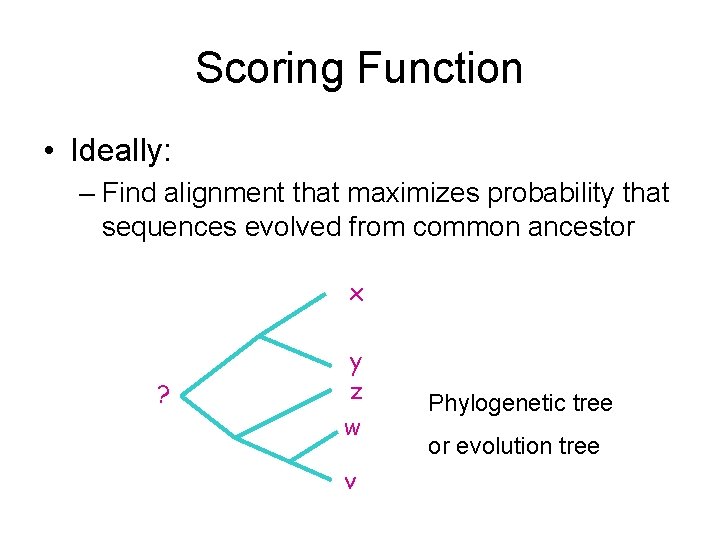
Scoring Function • Ideally: – Find alignment that maximizes probability that sequences evolved from common ancestor x ? y z w v Phylogenetic tree or evolution tree

Scoring Function (cont’d) • Unfortunately: too many parameters • Compromises: – Ignore phylogenetic tree • Compute from pair-wise scores – Based on sum of all pair-wise scores – Based on scores with a consensus sequence

First assumption • Columns are independent – Similar in pair-wise alignment • Therefore, the score of an alignment is the sum of all columns • Need to decide how to score a single column

Scoring Function: Sum Of Pairs Definition: Induced pairwise alignment A pairwise alignment induced by the multiple alignment Example: Induces: x: AC-GCGG-C y: AC-GC-GAG z: GCCGC-GAG x: ACGCGG-C; y: ACGC-GAC; x: AC-GCGG-C; z: GCCGC-GAG; y: AC-GCGAG z: GCCGCGAG

Sum Of Pairs (cont’d) • The sum-of-pairs score of an alignment is the sum of the scores of all induced pairwise alignments S(m) = k<l s(mk, ml): score of induced alignment (k, l)
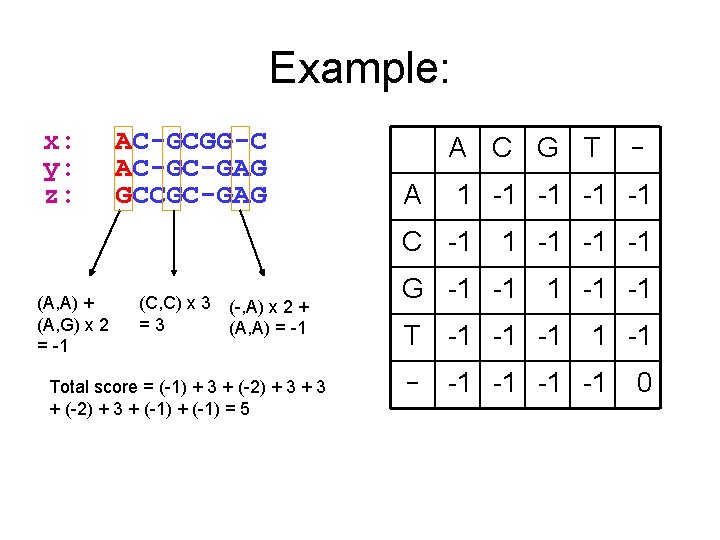
Example: x: y: z: AC-GCGG-C AC-GC-GAG GCCGC-GAG A C G T A 1 -1 -1 C -1 (A, A) + (A, G) x 2 = -1 (C, C) x 3 =3 (-, A) x 2 + (A, A) = -1 Total score = (-1) + 3 + (-2) + 3 + (-1) = 5 - 1 -1 -1 -1 G -1 -1 -1 T -1 -1 -1 -1 0

Sum Of Pairs (cont’d) • Drawback: no evolutionary characterization – Every sequence derived from all others • Heuristic way to incorporate evolution tree – Weighted Sum of Pairs: S(m) = k<l wkl s(mk, ml) wkl: weight decreasing with distance Human Mouse Duck Chicken

Consensus score -AGGCTATCACCTGACCTCCAGGCCGA--TGCCC--TAG-CTATCAC--GACCGC--GGTCGATTTGCCCGAC CAG-CTATCAC--GACCGC----TCGATTTGCTCGAC Consensus sequence: CAG-CTATCAC--GACCGC--GGTCGATTTGCCCGAC • Find optimal consensus string m* to maximize S(m) = i s(m*, mi) s(mk, ml): score of pairwise alignment (k, l)

Multiple Sequence Alignments Algorithms

Multidimensional Dynamic Programming (MDP) Generalization of Needleman-Wunsh: • Find the longest path in a high-dimensional cube – As opposed to a two-dimensional grid • Uses a N-dimensional matrix – As apposed to a two-dimensional array • Entry F(i 1, …, ik) represents score of optimal alignment for s 1[1. . i 1], … sk[1. . ik] F(i 1, i 2, …, i. N) = max(all neighbors of a cell) (F(nbr)+S(current))

Multidimensional Dynamic Programming (MDP) • Example: in 3 D (three sequences): (i-1, j-1, k-1) • 23 – 1 = 7 neighbors/cell F(i-1, j-1, k-1) + S(xi, xj, xk), F(i-1, j-1, k ) + S(xi, xj, -), F(i-1, j , k-1) + S(xi, -, xk), F(i, j, k) = max F(i , j-1, k-1) + S(-, xj, xk), F(i-1, j , k ) + S(xi, -, -), F(i , j-1, k ) + S(-, xj, -), F(i , j , k-1) + S(-, -, xk) (i-1, j-1, k) (i, j-1, k-1) (i, j-1, k) (i-1, j, k-1) (i-1, j, k) (i, j, k-1) (i, j, k)

Multidimensional Dynamic Programming (MDP) Running Time: 1. Size of matrix: LN; Where L = length of each sequence N = number of sequences 2. Neighbors/cell: 2 N – 1 Therefore…………… O(2 N LN)

Faster MDP • Carrillo & Lipman, 1988 – Branch and bound – Other heuristics • Practical for about 6 sequences of length about 200 -300.

Progressive Alignment • • Multiple Alignment is NP-hard Most used heuristic: Progressive Alignment Algorithm: 1. 2. 3. 4. Align two of the sequences xi, xj Fix that alignment Align a third sequence xk to the alignment xi, xj Repeat until all sequences are aligned Running Time: O(NL 2) Each alignment takes O(L 2) Repeat N times

Progressive Alignment x y z w • When evolutionary tree is known: – Align closest first, in the order of the tree Example: Order of alignments: 1. (x, y) 2. (z, w) 3. (xy, zw)

Progressive Alignment: CLUSTALW: most popular multiple protein alignment Algorithm: 1. Find all dij: alignment dist (xi, xj) • High alignment score => short distance 2. Construct a tree (Neighbor-joining hierarchical clustering. Will discuss in future) 3. Align nodes in order of decreasing similarity + a large number of heuristics

CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD
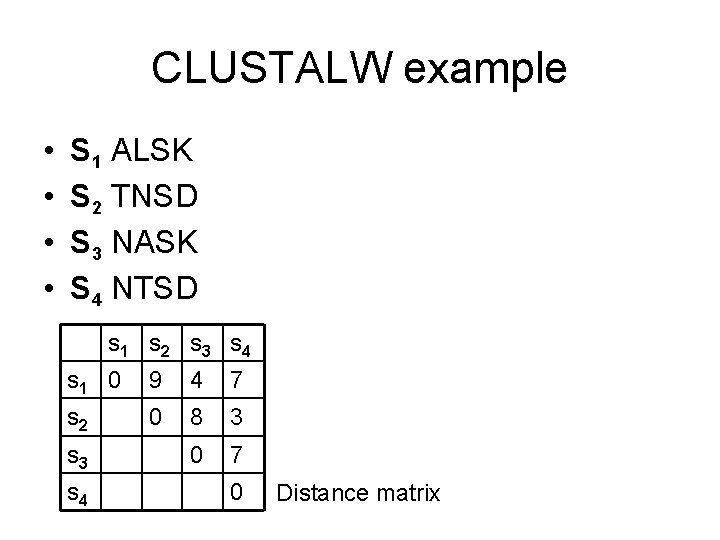
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 Distance matrix
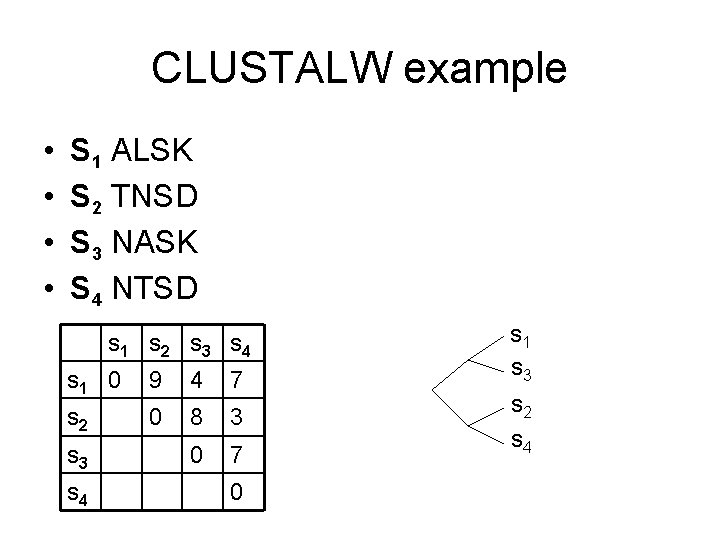
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 s 1 s 3 s 2 s 4
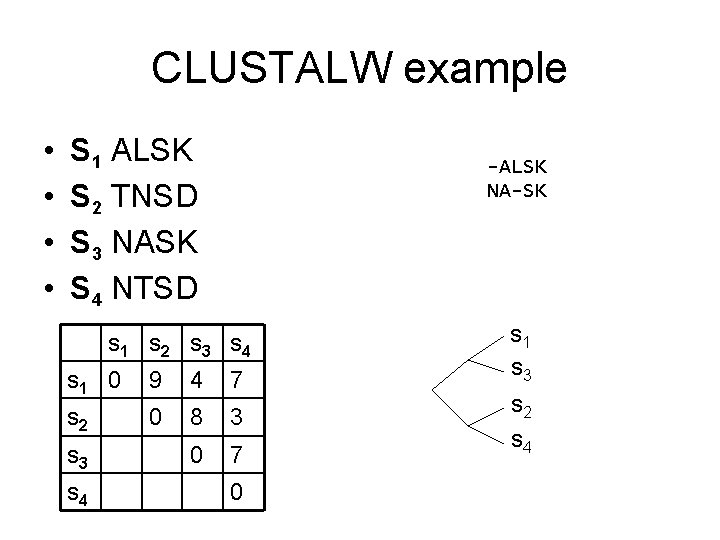
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD -ALSK NA-SK s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 s 1 s 3 s 2 s 4
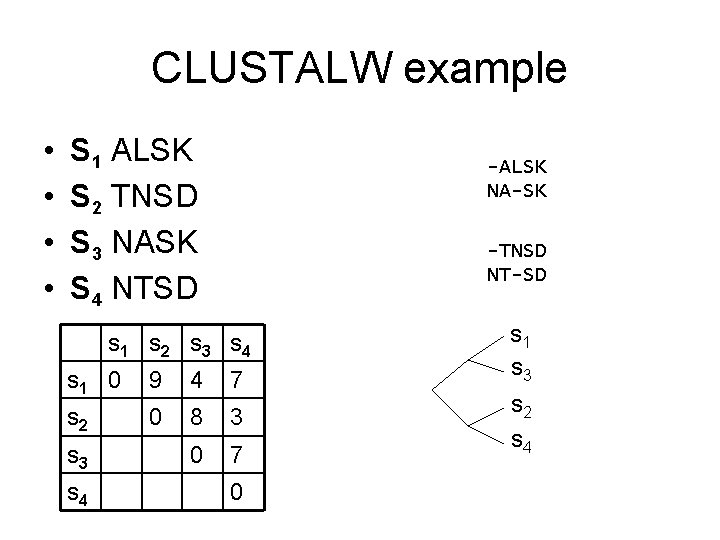
CLUSTALW example • • S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD -ALSK NA-SK -TNSD NT-SD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 s 1 s 3 s 2 s 4
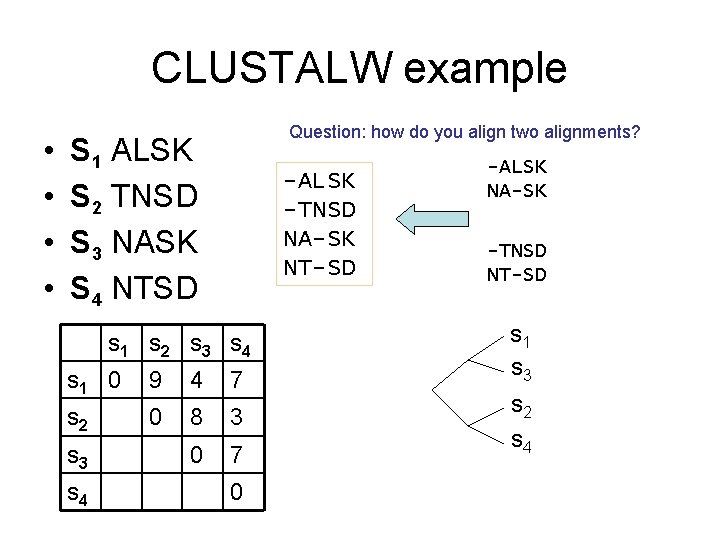
CLUSTALW example • • Question: how do you align two alignments? S 1 ALSK S 2 TNSD S 3 NASK S 4 NTSD -ALSK -TNSD NA-SK NT-SD s 1 s 2 s 3 s 4 s 1 0 9 4 7 s 2 0 8 3 0 7 s 3 s 4 0 -ALSK NA-SK -TNSD NT-SD s 1 s 3 s 2 s 4

Aligning two alignments • You can treat each column in an alignment as a single letter – Remember in the case of gene finder, we aligned three nucleic acids at a time • How do we score it? – Naïve: compute Sum of Pair • Better: only compute the cross terms – We already have (K, K) and (D, D) – Need to add 2 x(K, D) -ALSK NA-SK -TNSD NT-SD
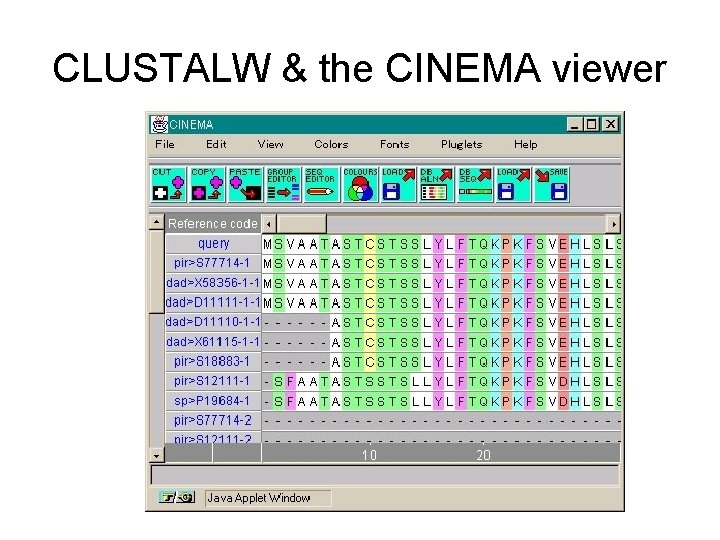
CLUSTALW & the CINEMA viewer

Iterative Refinement Problems with progressive alignment: • Depend on pair-wise alignments • If sequences are very distantly related, much higher likelihood of errors • Initial alignments are “frozen” even when new evidence comes Example: x: y: GAAGTT GAC-TT Frozen! z: w: GAACTG GTACTG Now clear: correct y should be GA-CTT

Iterative Refinement Algorithm (Barton-Stenberg): 1. 2. 3. 4. Align most similar xi, xj Align xk most similar to (xixj) Repeat 2 until (x 1…x. N) are aligned For j = 1 to N, Remove xj, and realign to x 1…xj-1 xj+1…x. N 5. Repeat 4 until convergence Note: Guaranteed to converge Running time: O(k. NL 2), k: number of iterations
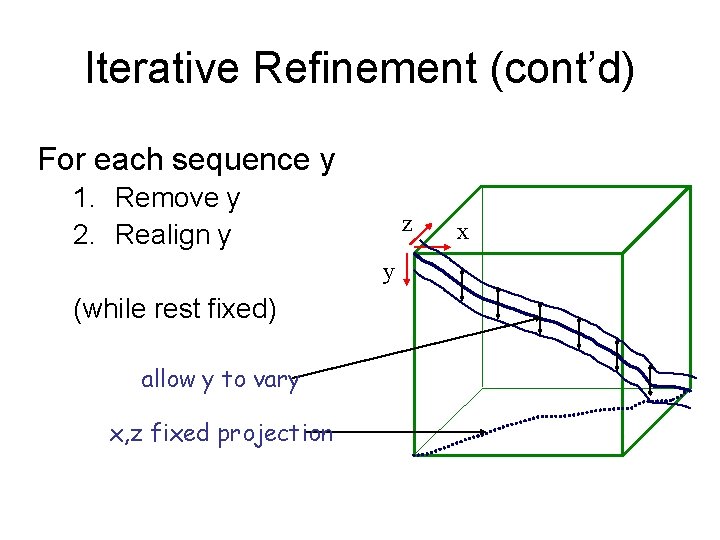
Iterative Refinement (cont’d) For each sequence y 1. Remove y 2. Realign y z y (while rest fixed) allow y to vary x, z fixed projection x

Iterative Refinement Example: align (x, y), (z, w), (xy, zw): x: y: z: w: GAAGTTA GAC-TTA GAACTGA GTACTGA After realigning y: x: y: z: w: GAAGTTA G-ACTTA GAACTGA GTACTGA + 3 matches

Iterative Refinement • Example not handled well: x: y 1: y 2: y 3: GAAGTTA GAC-TTA z: w: GAACTGA GTACTGA Realigning any single yi changes nothing
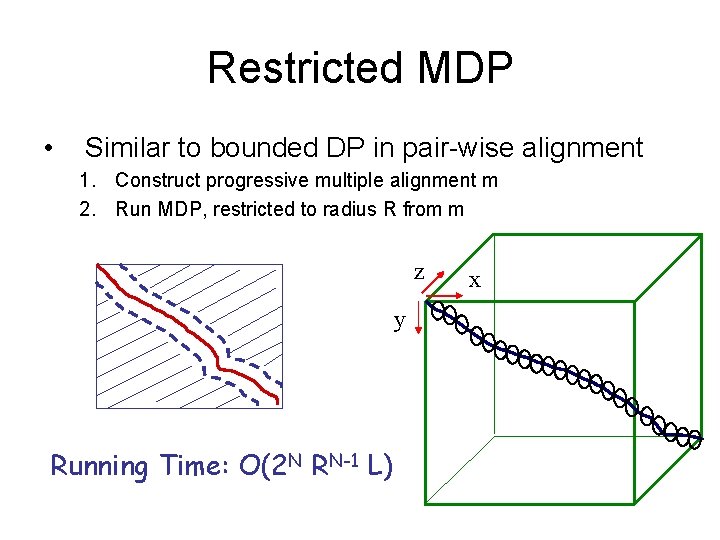
Restricted MDP • Similar to bounded DP in pair-wise alignment 1. Construct progressive multiple alignment m 2. Run MDP, restricted to radius R from m z y Running Time: O(2 N RN-1 L) x

Restricted MDP x: y 1: y 2: y 3: GAAGTTA GAC-TTA z: w: GAACTGA GTACTGA • Within radius 1 of the optimal Restricted MDP will fix it.

Other approaches • Profile Hidden Markov Models – Statistical learning methods – Will discuss in future

Multiple alignment tools • Clustal W (Thompson, 1994) – Most popular • • PRRP (Gotoh, 1993) HMMT (Eddy, 1995) DIALIGN (Morgenstern, 1998) T-Coffee (Notredame, 2000) MUSCLE (Edgar, 2004) Align-m (Walle, 2004) PROBCONS (Do, 2004)

In summary • Multiple alignment algorithms: – MDP (too slow) • B&B doesn’t solve the problem entirely – Progressive alignment: clustal. W – Iterative refinement – Restricted MDP
- Slides: 51