Scope and Importance of Bioinformatics Do you have

Scope and Importance of Bioinformatics Do you have required skill? Type of bioinformaticians (Users, Analysis or Developer) Possible biological problems Protein/Peptide related bioinformatics Types of methods and use of Open Source How to start Linux, programming (PHP/perl) WAMP/LAMP (Linux/Windows, Apache, Mysql, PHP) Machine learning techniques (Weka) Beware with experimental biologist

Concept of Research in Bioinformatics • • • Bioinformatics vs Computational Biology Service vs Research Components vs System Biology Guidelines/Suggestions Role of technical expertise Our group (Service or Science? )

Bioinformatics Vs Computational Biology Examples • Ramachandran Plot • Proteins Structure Prediction (Ab-initio) • Dynamic Programming for Alignment Examples of Bioinformatics • BLAST search against sequence databases • Homology Modelling • Epitope Mapping

Service Vs Bioinformatics Research • BTIS Programme • NCBI and EBI Services • Web service / databases • New rules or models derived from known structures • New functional domain form experimental data • Discoveries based on data • • Service for better research Research for better service Range of service (Individual or Community) Collaboration with experimental scientist for validation

Suggestions Selection of Problem • • • Talk to biologist & understand their problem Raise question in your mind Read literature (self learning) for answer Raise more question and read again Work on important problem Select problem if large dataset is availble Develop method with high accuracy Validate using standard validation techniques Develop web server software for service Publish in appropriate journal (high IF)
Fingerprinting Technique Microarray Data Subcellular Localization Fingerprinting Technique Whole genome Sequencing Epitope based Vaccines Anotation of Proteomes Genome Annotation Drug Discover Drug targets Comparative Genomics RNA Interference Assembling of Genomes Development of Databses

Therapeutic Application of Proteins/Peptides Peptide based Drugs (Small Molecule, Peptide, Protein) • Anticancer peptides • Antibacterial peptides Peptide for delivering drug • Cell-penetrating peptides • Tumor-homing peptides Peptide as disease diagnostics • Disease specific Mimotopes Epitope/Peptide based Vaccines • T-cell epitopes • B-cell epitopes

Limitation of Peptides & Challenges for Bioinformaticians 1. Peptide Degradation (short half-life) • Large peptide degrading enzymes • Hepatic/renal Clearance (liver/kidenys) 2. Low Oral Bioavailability (injection) 3. High conformational flexibility • Environment dependent tertiary structure • Low selectivity 4. Immunogenicity and antigenicity 5. High cost of production & Storage • Modification/glycosylation of peptides
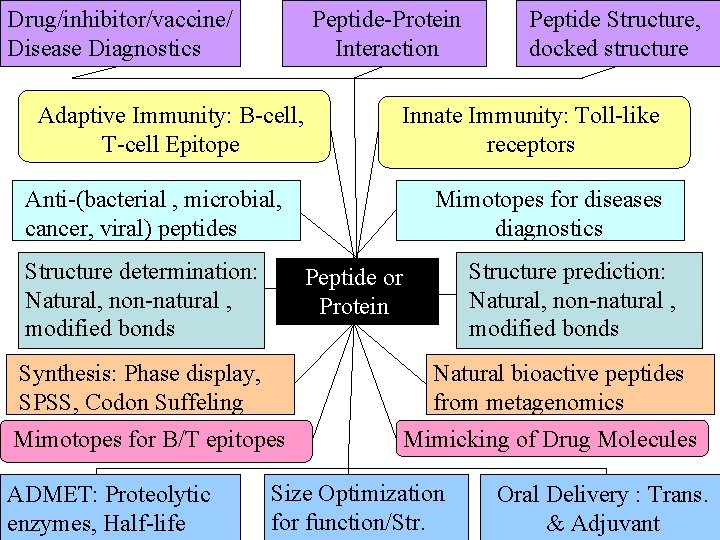
Drug/inhibitor/vaccine/ Disease Diagnostics Peptide-Protein Interaction Adaptive Immunity: B-cell, T-cell Epitope Innate Immunity: Toll-like receptors Anti-(bacterial , microbial, cancer, viral) peptides Structure determination: Natural, non-natural , modified bonds Structure prediction: Natural, non-natural , modified bonds Natural bioactive peptides from metagenomics Mimotopes for B/T epitopes ADMET: Proteolytic enzymes, Half-life Mimotopes for diseases diagnostics Peptide or Protein Synthesis: Phase display, SPSS, Codon Suffeling Peptide Structure, docked structure Mimicking of Drug Molecules Size Optimization for function/Str. Oral Delivery : Trans. & Adjuvant
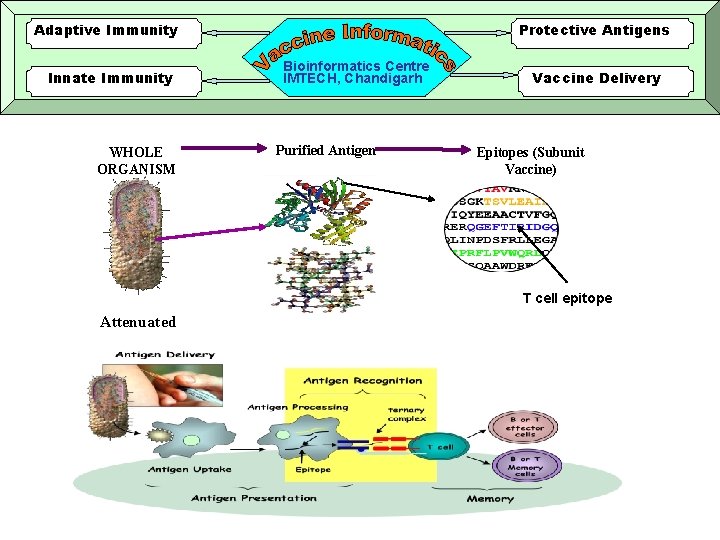
Adaptive Immunity Innate Immunity WHOLE ORGANISM Protective Antigens Bioinformatics Centre IMTECH, Chandigarh Purified Antigen Vaccine Delivery Epitopes (Subunit Vaccine) T cell epitope Attenuated
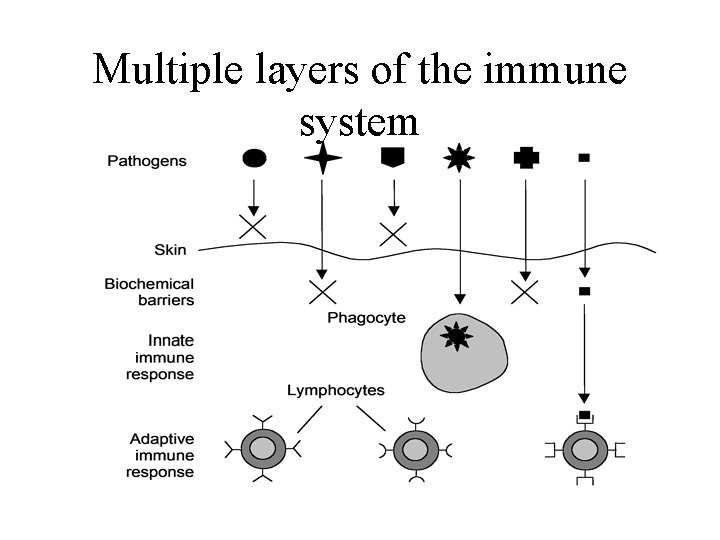
Multiple layers of the immune system
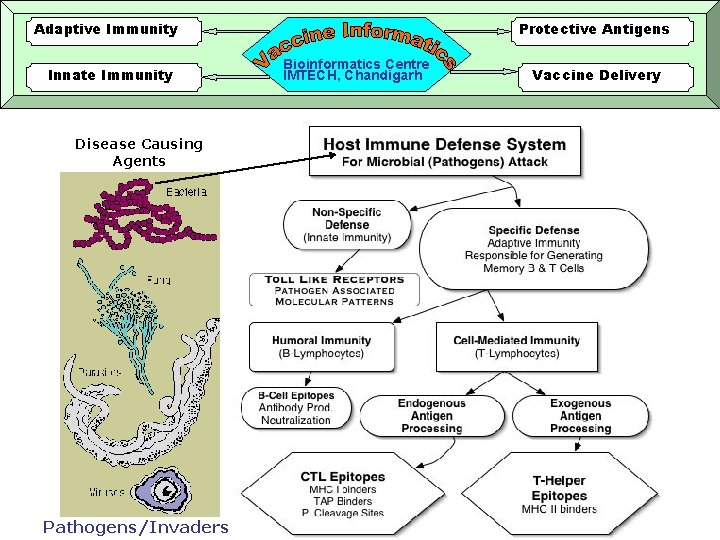
Adaptive Immunity Innate Immunity Disease Causing Agents Pathogens/Invaders Protective Antigens Bioinformatics Centre IMTECH, Chandigarh Vaccine Delivery
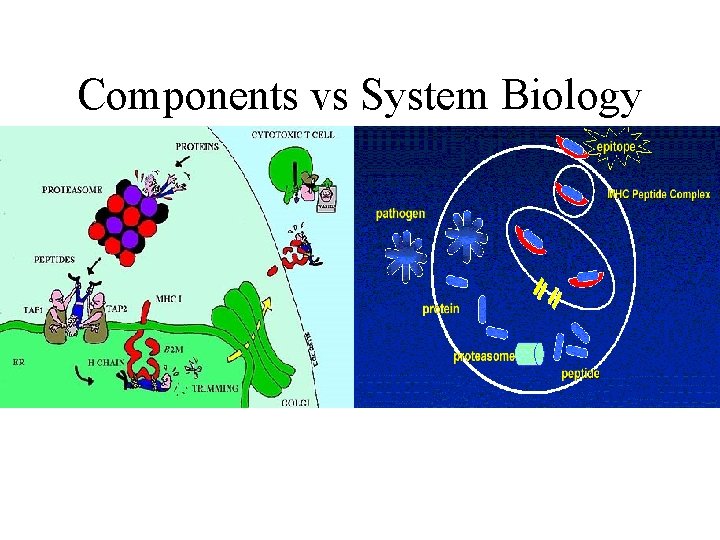
Components vs System Biology
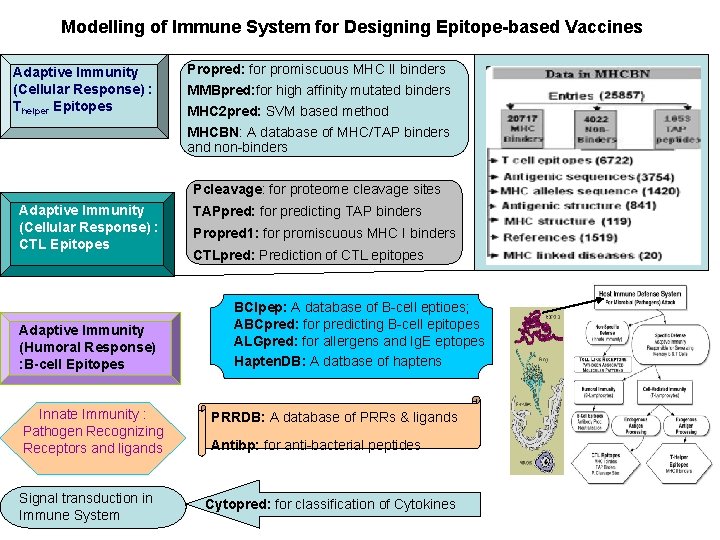
Modelling of Immune System for Designing Epitope-based Vaccines Adaptive Immunity (Cellular Response) : Thelper Epitopes Propred: for promiscuous MHC II binders MMBpred: for high affinity mutated binders MHC 2 pred: SVM based method MHCBN: A database of MHC/TAP binders and non-binders Pcleavage: for proteome cleavage sites Adaptive Immunity (Cellular Response) : CTL Epitopes Adaptive Immunity (Humoral Response) : B-cell Epitopes Innate Immunity : Pathogen Recognizing Receptors and ligands Signal transduction in Immune System TAPpred: for predicting TAP binders Propred 1: for promiscuous MHC I binders CTLpred: Prediction of CTL epitopes BCIpep: A database of B-cell eptioes; ABCpred: for predicting B-cell epitopes ALGpred: for allergens and Ig. E eptopes Hapten. DB: A datbase of haptens PRRDB: A database of PRRs & ligands Antibp: for anti-bacterial peptides Cytopred: for classification of Cytokines

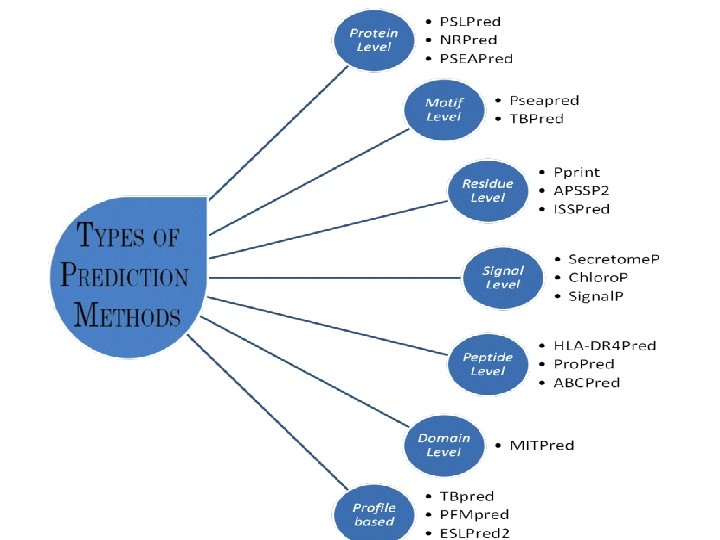
Important Information in Manual for Develpers
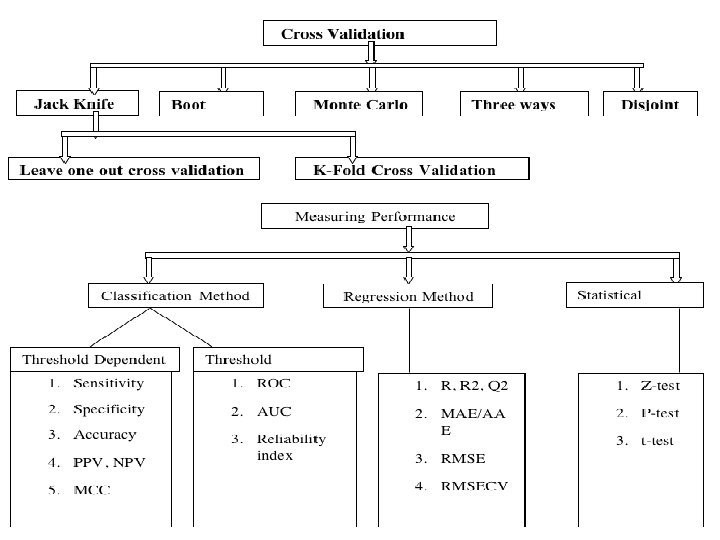
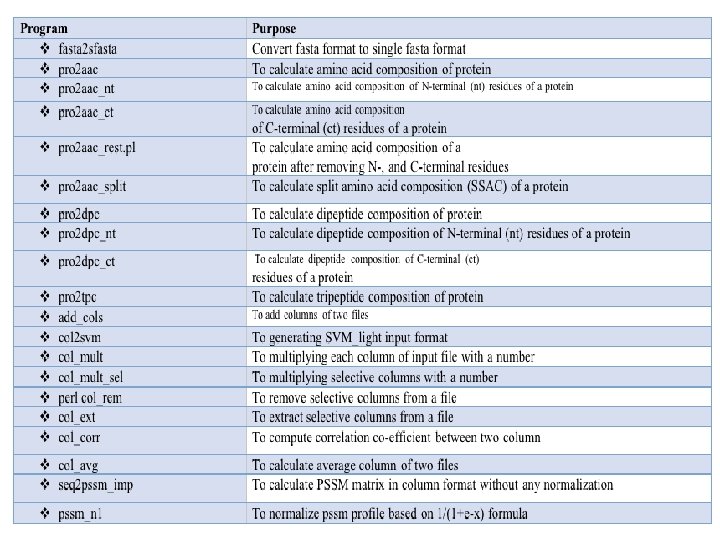

Computer-Aided Drug Discovery Searching Drug Targets: Bioinformatics Comparative genomics Genome Annotation FTGpred: Prediction of Prokaryotic genes EGpred: Prediction of eukaryotic genes Gene. Bench: Benchmarking of gene finders SRF: Spectral Repeat finder Subcellular Localization Methods PSLpred: localization of prokaryotic proteins ESLpred: localization of Eukaryotic proteins HSLpred: localization of Human proteins MITpred: Prediction of Mitochndrial proteins TBpred: Localization of mycobacterial proteins GWFASTA: Genome-Wide FASTA Search GWBLAST: Genome wide BLAST search COPID: Composition based similarity search LGEpred: Gene from protein sequence Prediction of drugable proteins Nrpred: Classification of nuclear receptors GPCRpred: Prediction of G-protein-coupled receptors GPCRsclass: Amine type of GPCR VGIchan: Voltage gated ion channel Pprint: RNA interacting residues in proteins GSTpred: Glutathione S-transferases proteins Protein Structure Prediction APSSP 2: protein secondary structure prediction Betatpred: Consensus method for -turns prediction Bteval: Benchmarking of -turns prediction Beta. Turns: Prediction of -turn types in proteins Turn Predictions: Prediction of / / -turns in proteins Gamma. Pred: Prediction of-turns in proteins Bhair. Pred: Prediction of Beta Hairpins TBBpred: Prediction of trans membrane beta barrel proteins SARpred: Prediction of surface accessibility (real accessibility) Pep. Str: Prediction of tertiary structure of Bioactive peptides

Suggestions Technical Learning Linux/Unix system • • Programming language PERL/PHP/C • Example Scripts (http: //www. imtech. res. in/raghava/gpsr/ ) • HTML and CGI script (Java) • LAMP learning • SVM_Light for SVM, SNNNS for ANN • TREMBL for nearest neighbour • Weka for wide range • Rapidminer

Important URLs http: //www. imtech. res. in/raghava/gpsr/ http: //www. imtech. res. in/raghava/reprints/ http: //www. imtech. res. in/raghava/slides/ http: //crdd. osdd. net/raghava/ccpdb/
- Slides: 24