CS 5263 Bioinformatics Lecture 5 Local Sequence Alignment

CS 5263 Bioinformatics Lecture 5: Local Sequence Alignment Algorithms

Poll • Who have learned and still remember Finite State Machine/Automata, regular grammar, and context-free grammar?

Roadmap • Review of last lecture • Local Sequence Alignment • Statistics of sequence alignment – Substitution matrix – Significance of alignment
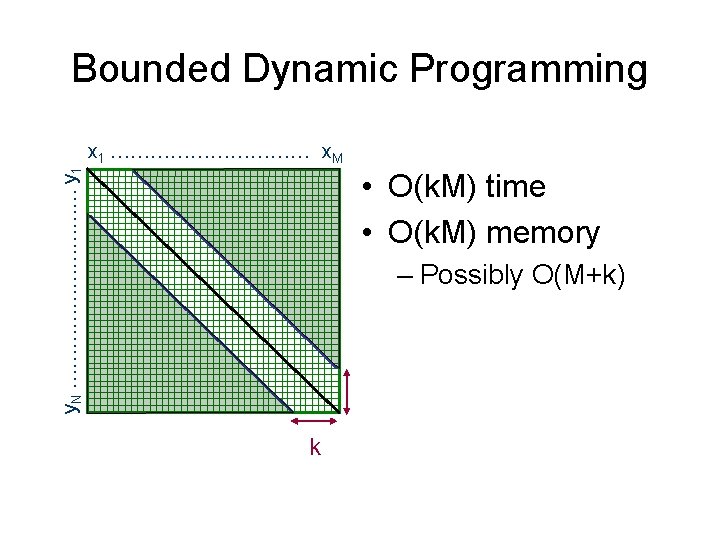
Bounded Dynamic Programming y. N …………… y 1 x 1 …………… x. M • O(k. M) time • O(k. M) memory – Possibly O(M+k) k
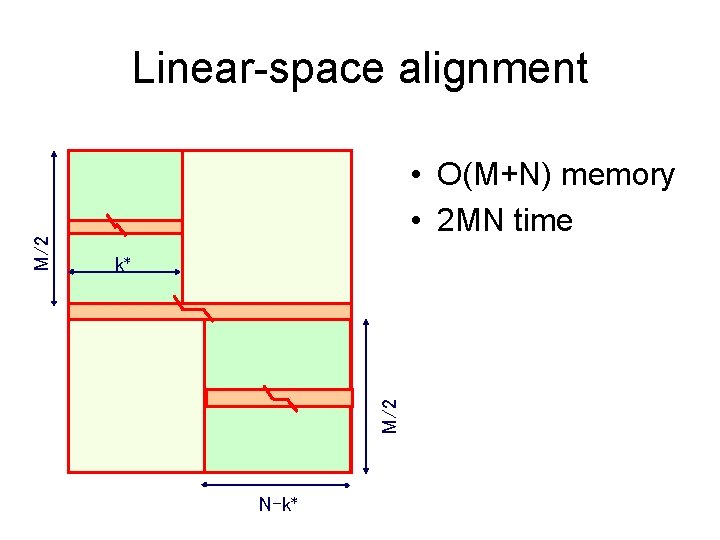
• O(M+N) memory • 2 MN time k* M/2 Linear-space alignment N-k*
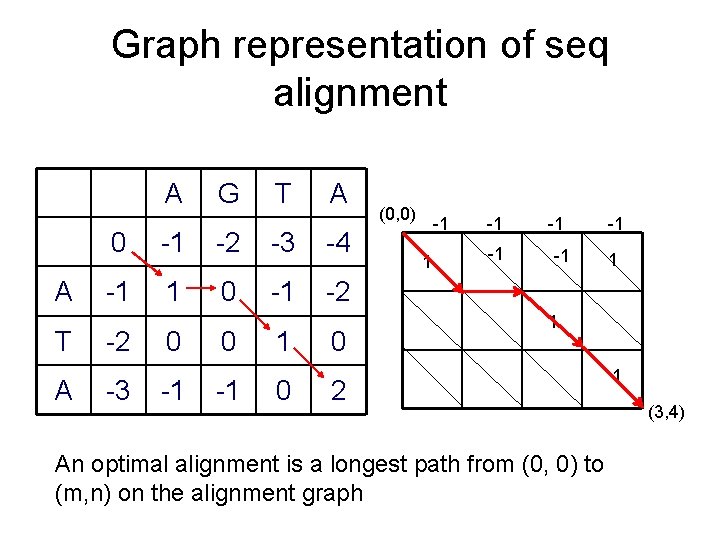
Graph representation of seq alignment A A T A G T A 0 -1 -2 -3 -4 -1 1 0 -1 -2 -2 -3 0 -1 1 0 0 (0, 0) -1 1 -1 -1 -1 1 1 2 An optimal alignment is a longest path from (0, 0) to (m, n) on the alignment graph 1 (3, 4)

Question • If I change the scoring scheme, will it change the alignment? – Match = 1, mismatch = gap = -2 || v – Match = 2, mismatch = gap = -1? • Answer: Yes

Proof • Let F 1 be the score of an optimal alignment under the scoring scheme – Match = m > 0 – Mismatch = s < 0 – Gap = d < 0 • Let a 1, b 1, c 1 be the number of matches, mismatches, and gaps in the alignment • F 1 = a 1 m + b 1 s + c 1 d

Proof (cont’) • Let F 2 be the score of a sub-optimal alignment under the same scoring scheme • Let a 2, b 2, c 2 be the number of matches, mismatches, and gaps in the alignment • F 2 = a 2 m + b 2 s + c 2 d • Let F 1 = F 2 + k, where k > 0

Proof (cont’) • Now we change the scoring scheme, so that – Match = m + 1 – Mismatch = s + 1 – Gap = d + 1

Proof (cont’) • The new scores for the two alignments become: F 1’= a 1 * (m+1) + b 1 * (s + 1) + c 1 * (d + 1) = a 1 m + b 1 s + c 1 d + (a 1+b 1+c 1) = F 1 + (a 1+b 1+c 1) length of alignment 1 = F 1 + L 1 F 2’ = a 2 * (m+1) + b 2 * (s + 1) + c 2 * (d + 1) = F 2 + (a 2+b 2+c 2) = F 2 + L 2 length of alignment 2

Proof (cont’) • F 1’ – F 2’ = F 1 – F 2 + (a 1+b 1+c 1) – (a 2+b 2+c 2) = k + L 1 – L 2 Length of alignment 1 Length of alignment 2 In order for F 1’ < F 2’, we need to have: k + L 1 – L 2 < 0, i. e. L 2 – L 1 > k

Proof (cont’) • This means, if under the original scoring scheme, F 1 is greater than F 2 by k, but the length of alignment 2 is at least (k+1) greater than that of alignment 1, F 2’ will be greater than F 1’ under the new scoring scheme. • We only need to show one example that it is possible to find such two alignments
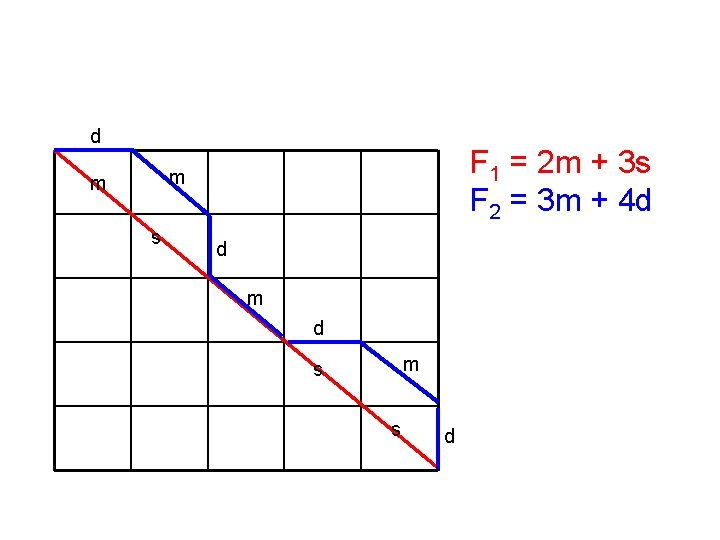
d F 1 = 2 m + 3 s F 2 = 3 m + 4 d m m s d m s s d
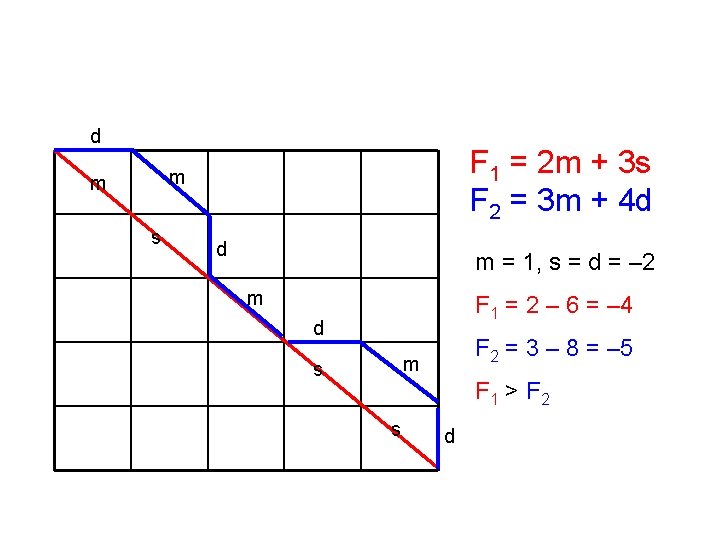
d F 1 = 2 m + 3 s F 2 = 3 m + 4 d m m s d m = 1, s = d = – 2 m F 1 = 2 – 6 = – 4 d F 2 = 3 – 8 = – 5 m s F 1 > F 2 s d
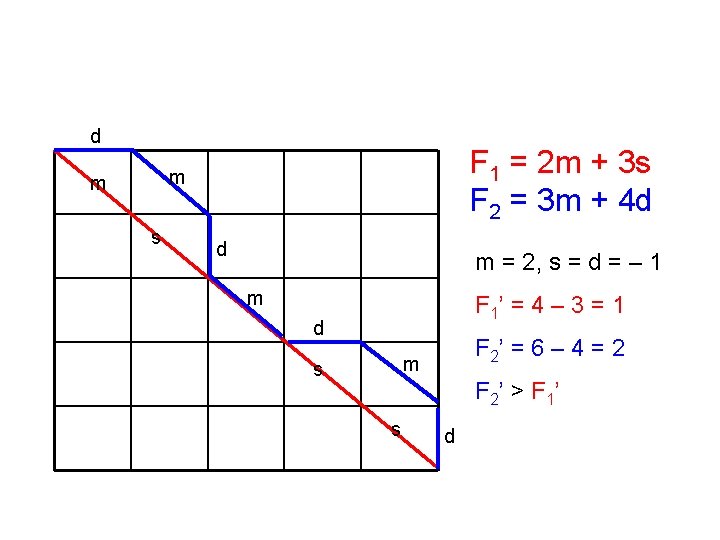
d F 1 = 2 m + 3 s F 2 = 3 m + 4 d m m s d m = 2, s = d = – 1 m F 1’ = 4 – 3 = 1 d F 2’ = 6 – 4 = 2 m s F 2’ > F 1’ s d
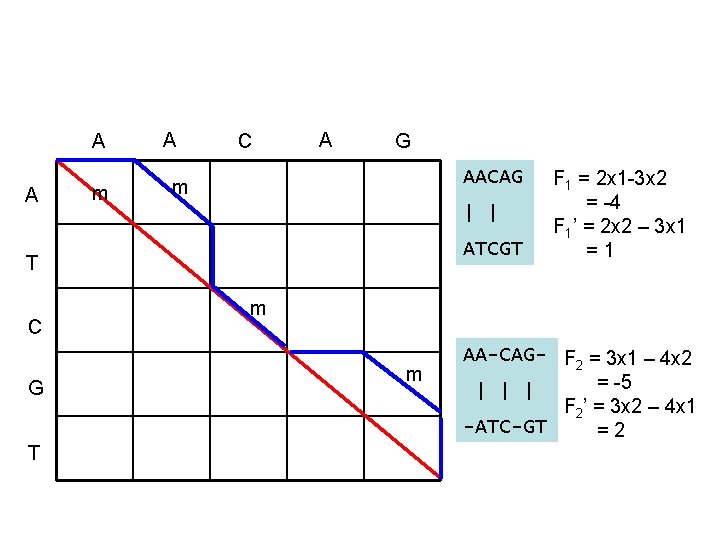
A A m A C A G AACAG m | | ATCGT T C G T F 1 = 2 x 1 -3 x 2 = -4 F 1’ = 2 x 2 – 3 x 1 =1 m m AA-CAG- F 2 = 3 x 1 – 4 x 2 = -5 | | | F 2’ = 3 x 2 – 4 x 1 -ATC-GT =2

• On the other hand, if we had doubled our scores, such that m’ = 2 m, s’ = 2 s d’ = 2 d • F 1’ = 2 F 1 • F 2’ = 2 F 2 • Our alignment won’t be changed

Today • How to model gaps more accurately? • Local sequence alignment • Statistics of alignment

What’s a better alignment? GACGCCGAACG ||||| GACGC---ACG GACGCCGAACG |||| | | || GACG-C-A-CG Score = 8 x m – 3 x d However, gaps usually occur in bunches. - During evolution, chunks of DNA may be lost entirely - Aligning genomic sequence vs. c. DNA (reverse complimentary to m. RNA)
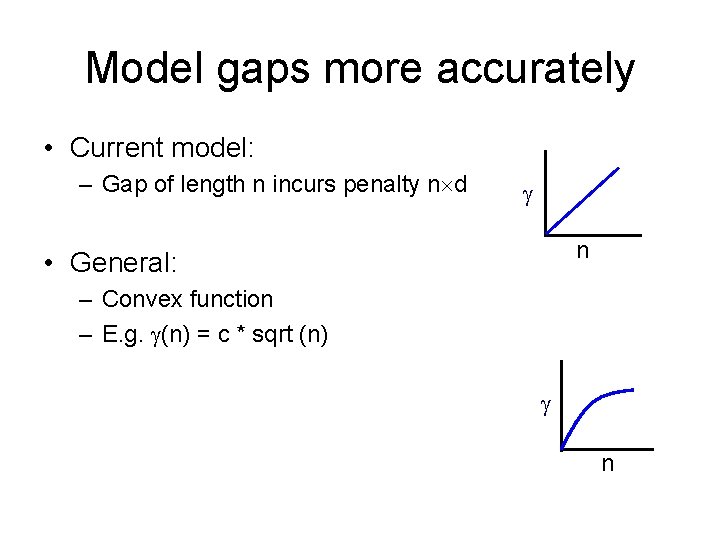
Model gaps more accurately • Current model: – Gap of length n incurs penalty n d n • General: – Convex function – E. g. (n) = c * sqrt (n) n
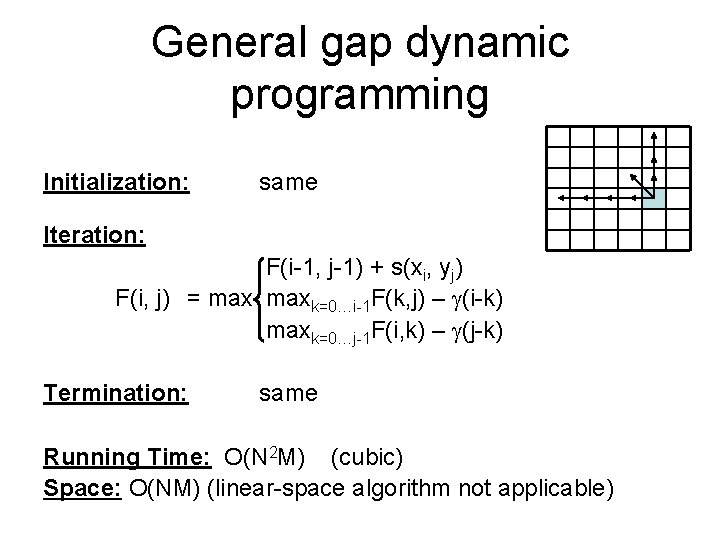
General gap dynamic programming Initialization: same Iteration: F(i-1, j-1) + s(xi, yj) F(i, j) = maxk=0…i-1 F(k, j) – (i-k) maxk=0…j-1 F(i, k) – (j-k) Termination: same Running Time: O(N 2 M) (cubic) Space: O(NM) (linear-space algorithm not applicable)
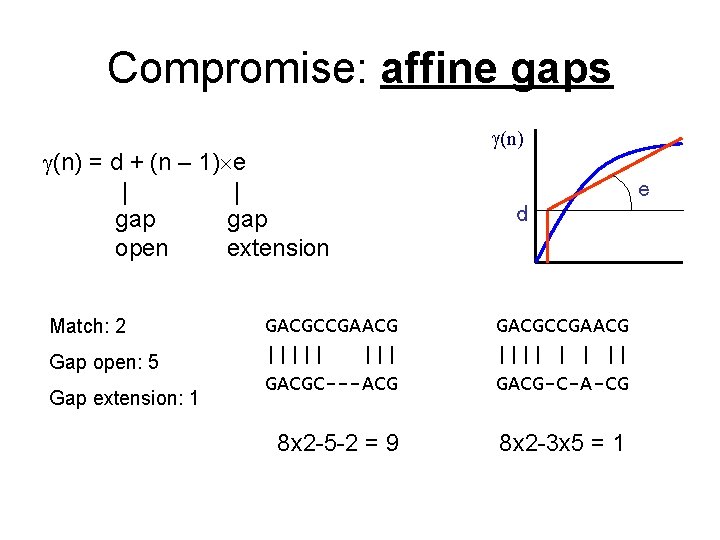
Compromise: affine gaps (n) = d + (n – 1) e | | gap open extension Match: 2 Gap open: 5 Gap extension: 1 (n) e d GACGCCGAACG ||||| GACGC---ACG GACGCCGAACG |||| | | || GACG-C-A-CG 8 x 2 -5 -2 = 9 8 x 2 -3 x 5 = 1

Additional states • The amount of state needed increases – In scoring a single entry in our matrix, we need remember an extra piece of information • Are we continuing a gap in x? (if no, start is more expensive) • Are we continuing a gap in y? (if no, start is more expensive) • Are we continuing from a match between xi and yj?
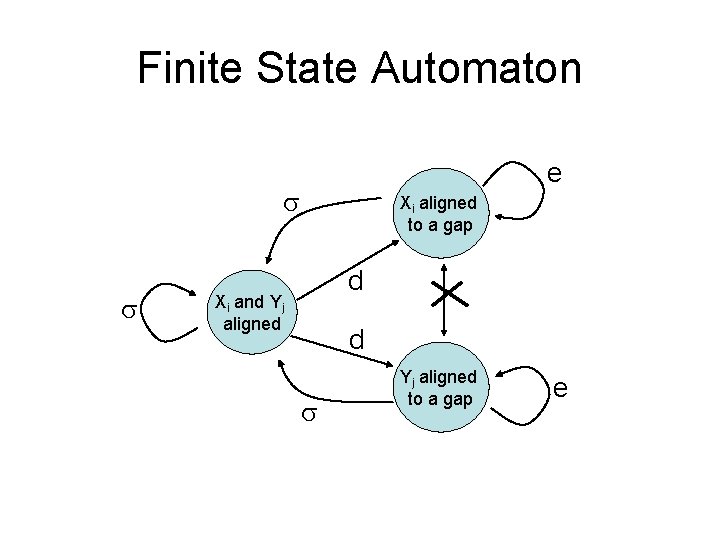
Finite State Automaton e Xi aligned to a gap d Xi and Yj aligned d Yj aligned to a gap e

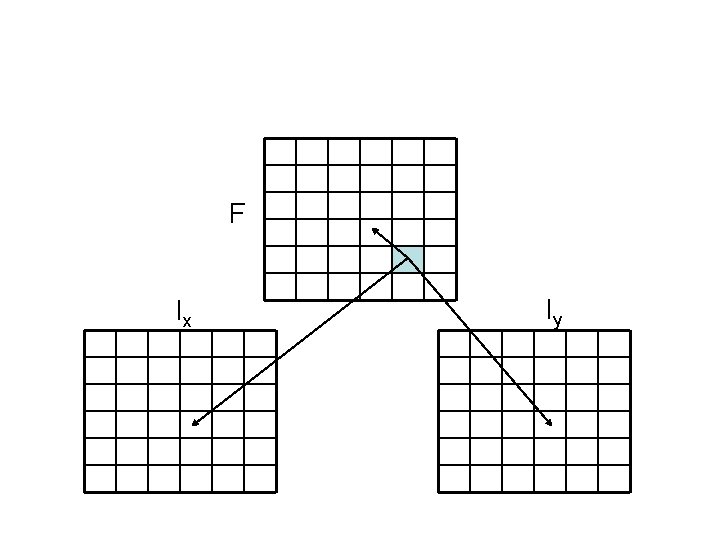
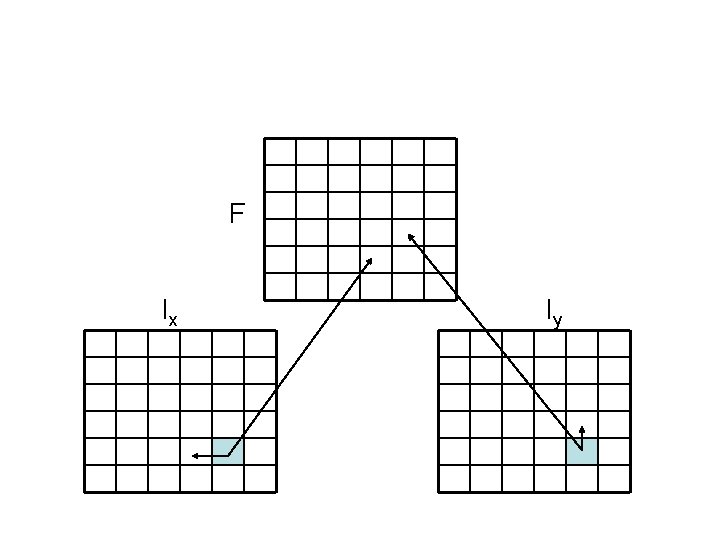
Dynamic programming • We encode this information in three different matrices • For each element (i, j) we use three variables – F(i, j): best alignment of x 1. . xi & y 1. . yj if xi aligns to yj – Ix(i, j): best alignment of x 1. . xi & y 1. . yj if yj aligns to gap – Iy(i, j): best alignment of x 1. . xi & y 1. . yj if xi aligns to gap
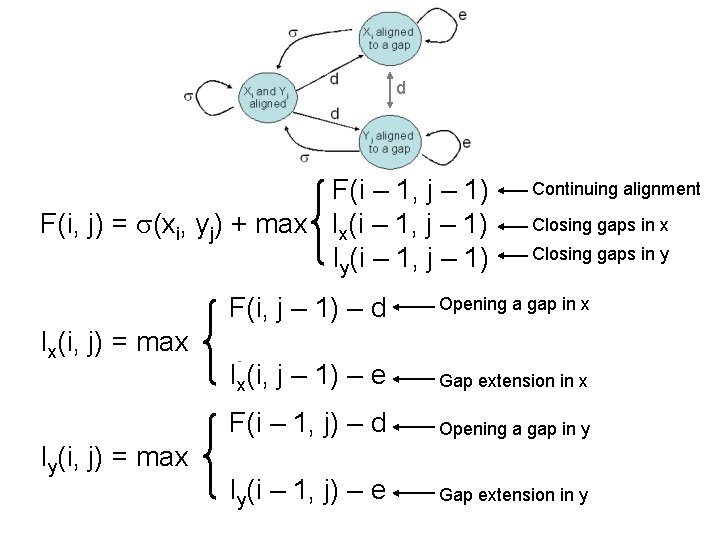
d F(i – 1, j – 1) F(i, j) = (xi, yj) + max Ix(i – 1, j – 1) Iy(i – 1, j – 1) Ix(i, j) = max Iy(i, j) = max F(i, j – 1) – d Iy(i, j – 1) – d Ix(i, j – 1) – e F(i – 1, j) – d Ix(i – 1, j) – d Iy(i – 1, j) – e Continuing alignment Closing gaps in x Closing gaps in y Opening a gap in x Gap extension in x Opening a gap in y Gap extension in y
F Ix Iy
F Ix Iy
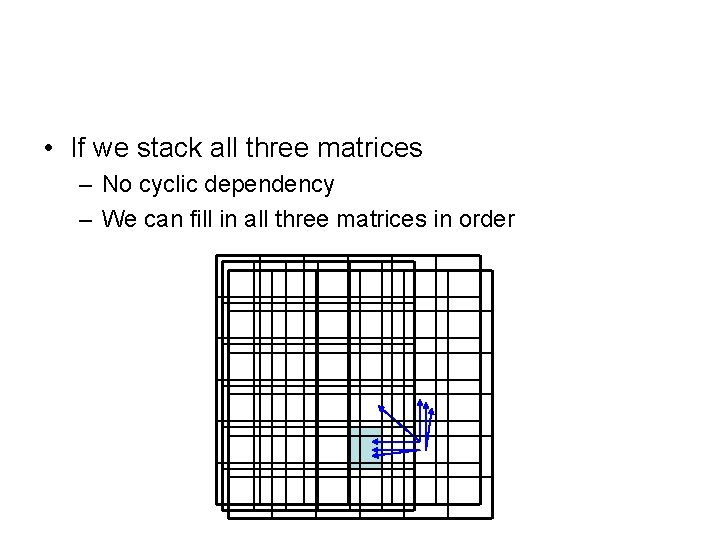
• If we stack all three matrices – No cyclic dependency – We can fill in all three matrices in order

Algorithm • for i = 1: m – for j = 1: n • Fill in F(i, j), Ix(i, j), Iy(i, j) – end • F(M, N) = max (F(M, N), Ix(M, N), Iy(M, N)) • Time: O(MN) • Space: O(MN) or O(N) when combine with the linear-space algorithm
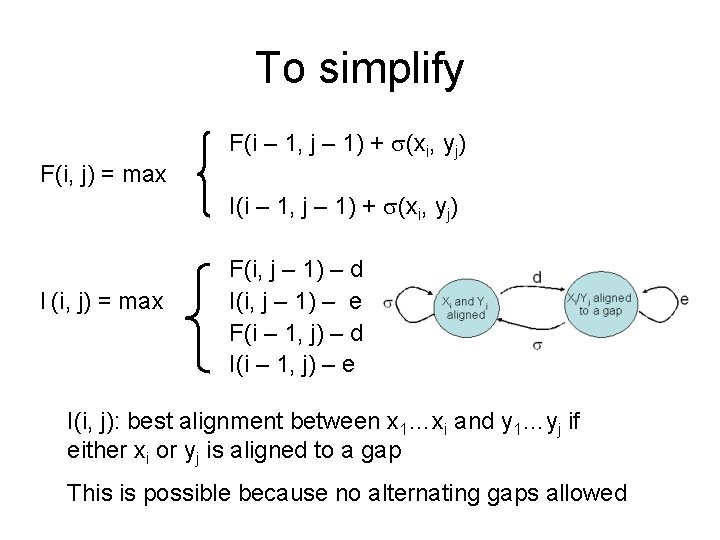
To simplify F(i – 1, j – 1) + (xi, yj) F(i, j) = max I(i – 1, j – 1) + (xi, yj) I (i, j) = max F(i, j – 1) – d I(i, j – 1) – e F(i – 1, j) – d I(i – 1, j) – e I(i, j): best alignment between x 1…xi and y 1…yj if either xi or yj is aligned to a gap This is possible because no alternating gaps allowed

To summarize • Global alignment – Basic algorithm: Needleman-Wunsch – Variants: • Overlapping detection • Longest common subsequences • Achieved by varying initial conditions or scoring – Bounded DP (pruning search space) – Linear space (divide-and-conquer) – Affine gap penalty

Local alignment

The local alignment problem Given two strings X = x 1……x. M, Y = y 1……y. N Find substrings x’, y’ whose similarity (optimal global alignment value) is maximum e. g. X = abcxdex Y = xxxcde x y X’ = cxde Y’ = c-de

Why local alignment • Conserved regions may be a small part of the whole – “Active site” of a protein – Scattered genes or exons among “junks” – Don’t have whole sequence • Global alignment might miss them if flanking “junk” outweighs similar regions
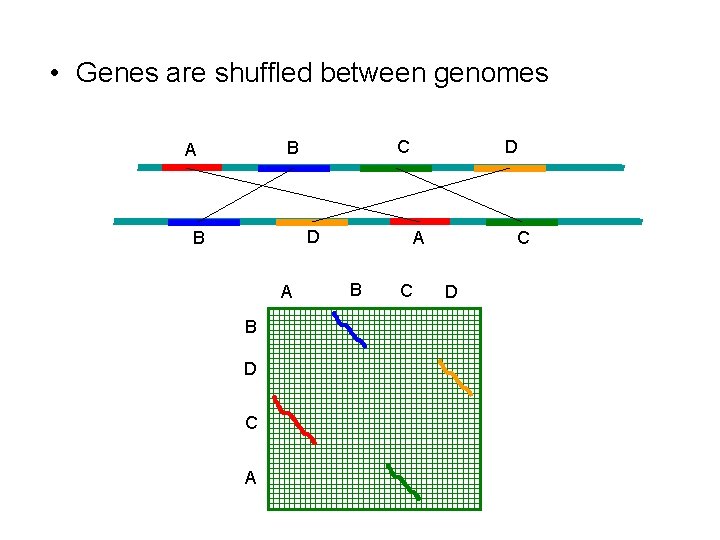
• Genes are shuffled between genomes C B A D B A B D C A D A B C C D

Naïve algorithm for all substrings X’ of X and Y’ of Y Align X’ & Y’ via dynamic programming Retain pair with max value end ; Output the retained pair • Time: O(n 2) choices for A, O(m 2) for B, O(nm) for DP, so O(n 3 m 3 ) total.
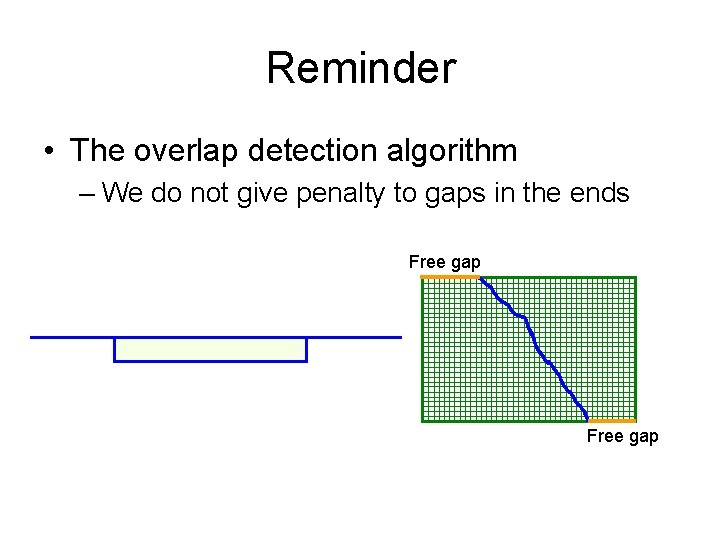
Reminder • The overlap detection algorithm – We do not give penalty to gaps in the ends Free gap
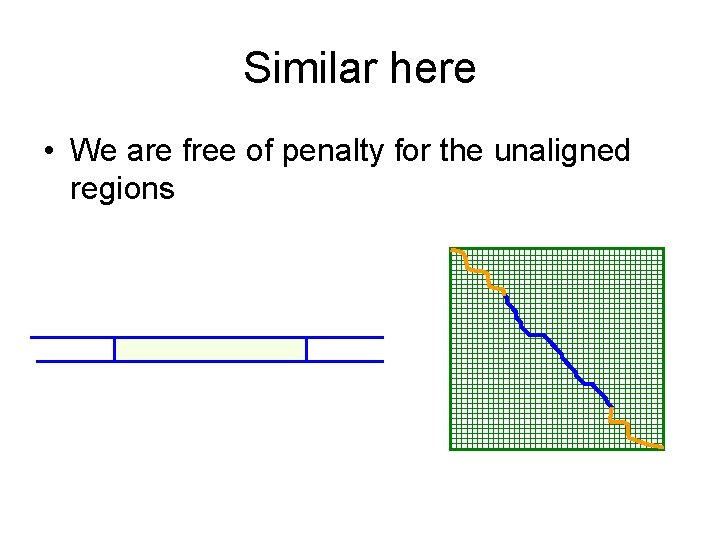
Similar here • We are free of penalty for the unaligned regions

The big idea • Whenever we get to some bad region (negative score), we ignore the previous alignment – Reset score to zero

The Smith-Waterman algorithm Initialization: F(0, j) = F(i, 0) = 0 0 F(i – 1, j) – d Iteration: F(i, j) = max F(i, j – 1) – d F(i – 1, j – 1) + (xi, yj)

The Smith-Waterman algorithm Termination: 1. If we want the best local alignment… FOPT = maxi, j F(i, j) 2. If we want all local alignments scoring > t For all i, j find F(i, j) > t, and trace back
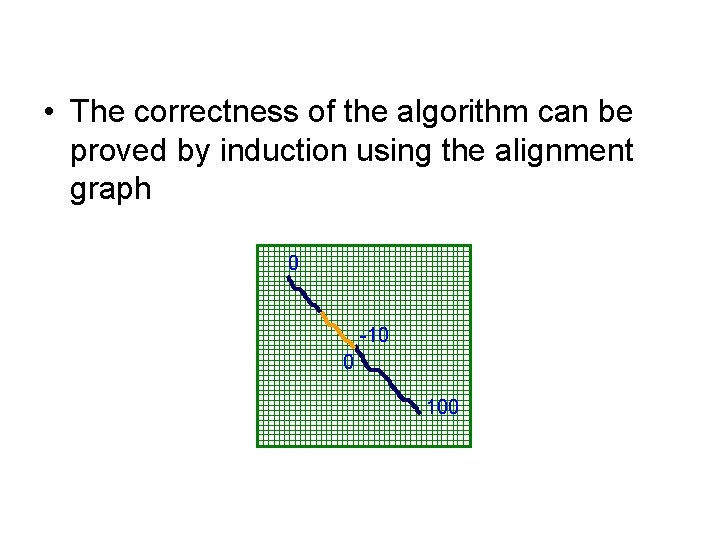
• The correctness of the algorithm can be proved by induction using the alignment graph 0 -10 0 100
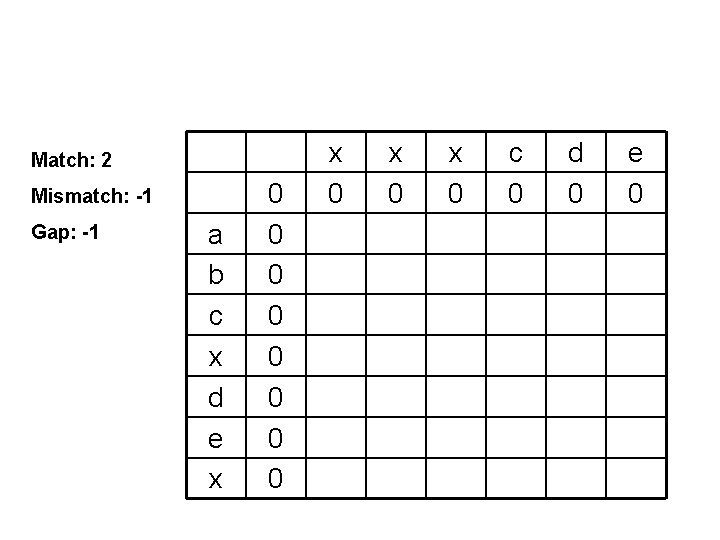
Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 x 0 x 0 c 0 d 0 e 0
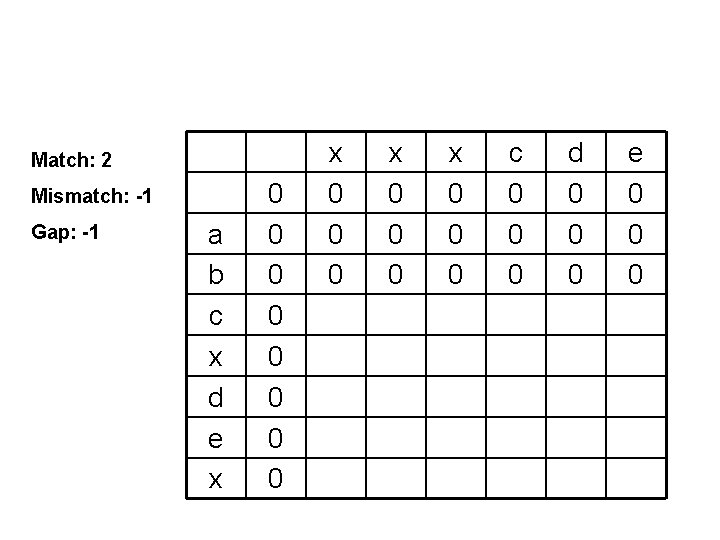
Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 x 0 0 0 c 0 0 0 d 0 0 0 e 0 0 0
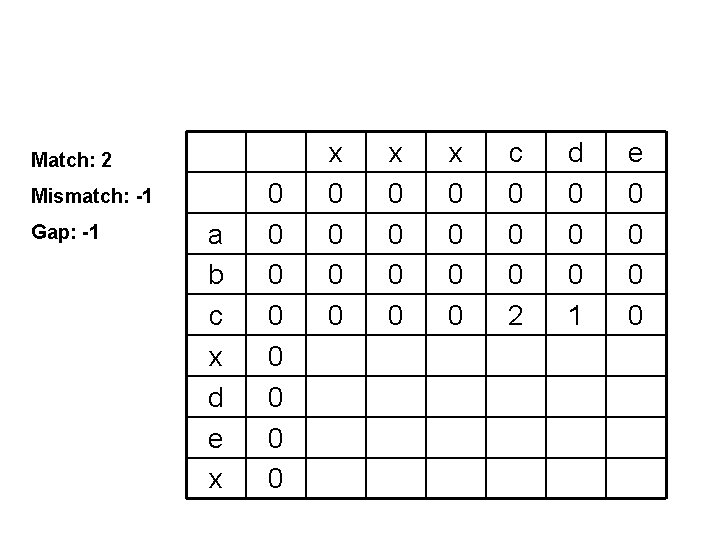
Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 x 0 0 c 0 0 0 2 d 0 0 0 1 e 0 0
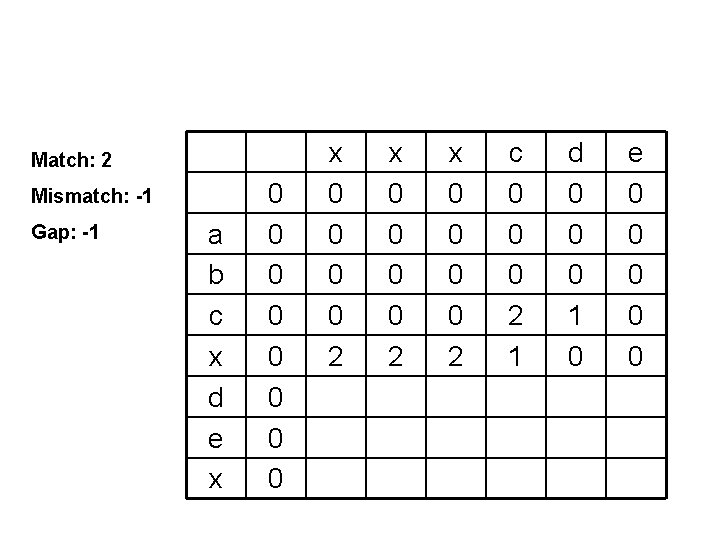
Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 2 x 0 0 2 c 0 0 0 2 1 d 0 0 0 1 0 e 0 0 0
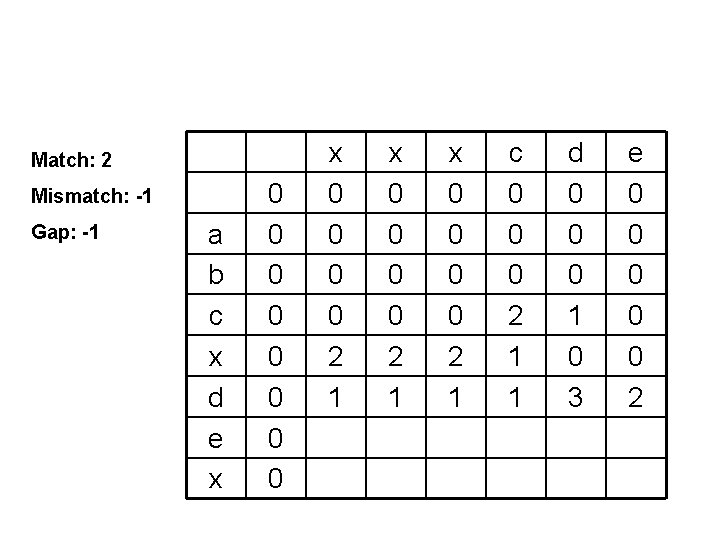
Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 2 1 x 0 0 2 1 c 0 0 0 2 1 1 d 0 0 0 1 0 3 e 0 0 0 2
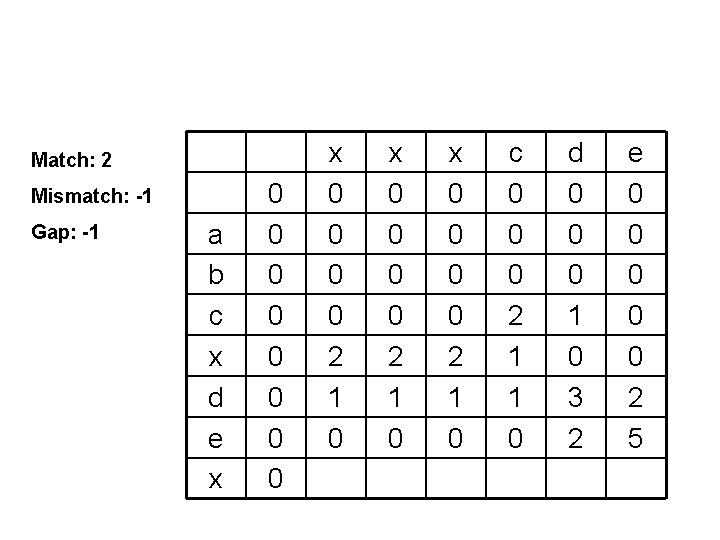
Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 2 1 0 x 0 0 2 1 0 c 0 0 0 2 1 1 0 d 0 0 0 1 0 3 2 e 0 0 0 2 5
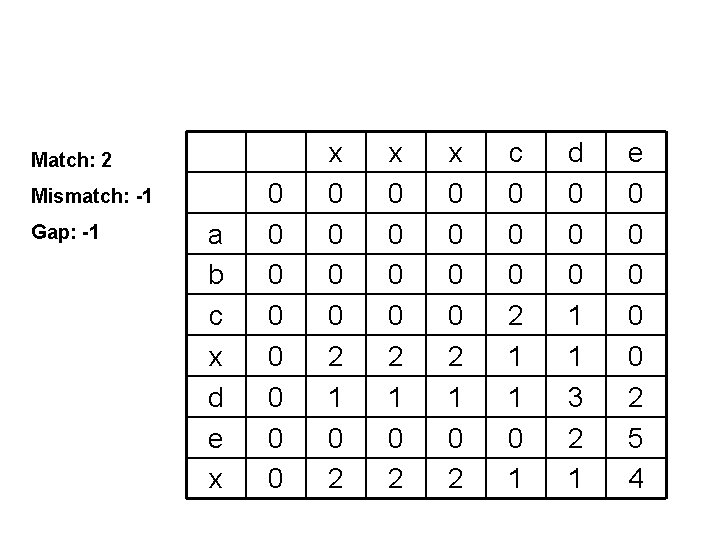
Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 2 1 0 2 x 0 0 2 1 0 2 c 0 0 0 2 1 1 0 1 d 0 0 0 1 1 3 2 1 e 0 0 0 2 5 4

Trace back Match: 2 Mismatch: -1 Gap: -1 a b c x d e x 0 0 0 0 2 1 0 2 x 0 0 2 1 0 2 c 0 0 0 2 1 1 0 1 d 0 0 0 1 1 3 2 1 e 0 0 0 2 5 4
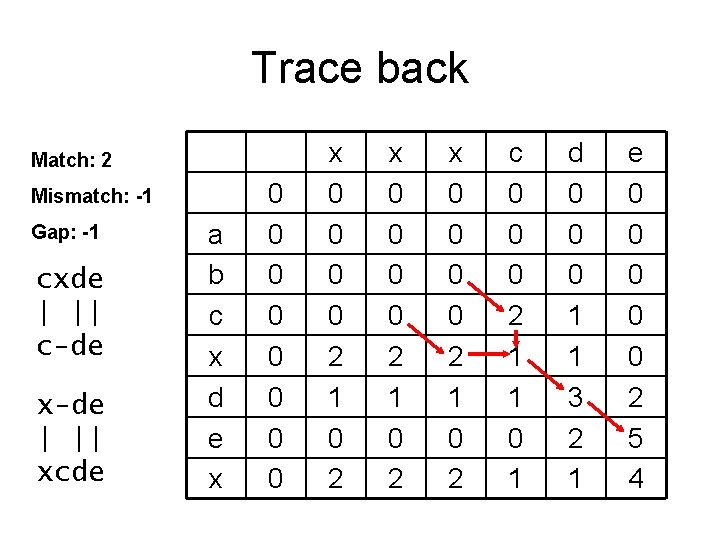
Trace back Match: 2 Mismatch: -1 Gap: -1 cxde | || c-de x-de | || xcde a b c x d e x 0 0 0 0 2 1 0 2 x 0 0 2 1 0 2 c 0 0 0 2 1 1 0 1 d 0 0 0 1 1 3 2 1 e 0 0 0 2 5 4

• No negative values in local alignment DP array • Optimal local alignment will never have a gap on either end • Local alignment: “Smith-Waterman” • Global alignment: “Needleman-Wunsch”

Analysis • Time: – O(MN) for finding the best alignment – Depending on the number of sub-opt alignments • Memory: – O(MN) – O(M+N) possible

The statistics of alignment Where does (xi, yj) come from? Are two aligned sequences actually related?

Probabilistic model of alignments • We’ll focus on protein alignments without gaps • Given an alignment, we can consider two possibilities – R: the sequences are related by evolution – U: the sequences are unrelated • How can we distinguish these possibilities? • How is this view related to amino-acid substitution matrix?

Model for unrelated sequences • Assume each position of the alignment is independently sampled from some distribution of amino acids • ps: probability of amino acid s in the sequences • Probability of seeing an amino acid s aligned to an amino acid t by chance is – Pr(s, t | U) = ps * pt • Probability of seeing an ungapped alignment between x = x 1…xn and y = y 1…yn randomly is

Model for related sequences • Assume each pair of aligned amino acids evolved from a common ancestor • Let qst be the probability that amino acid s in one sequence is related to t in another sequence • The probability of an alignment of x and y is give by

Probabilistic model of Alignments • How can we decide which possibility (U or R) is more likely? • One principled way is to consider the relative likelihood of the two possibilities (the odd ratios) – A higher ratio means that R is more likely than U

Log odds ratio • Taking the log, we get • Recall that the score of an alignment is given by

• Therefore, if we define This is indeed how biologists have defined the substitution matrices for proteins • We are actually defining the alignment score as the log odds ratio (log likelihood) between the two models R and U

• ps can be counted from the available protein sequences • But how do we get qst? (the probability that s and t have a common ancestor) • Counted from trusted alignments of related sequences

Protein Substitution Matrices • Two popular sets of matrices for protein sequences – PAM matrices [Dayhoff et al, 1978] • Better for aligning closely related sequences – BLOSUM matrices [Henikoff & Henikoff, 1992] • For both closely or remotely related sequences
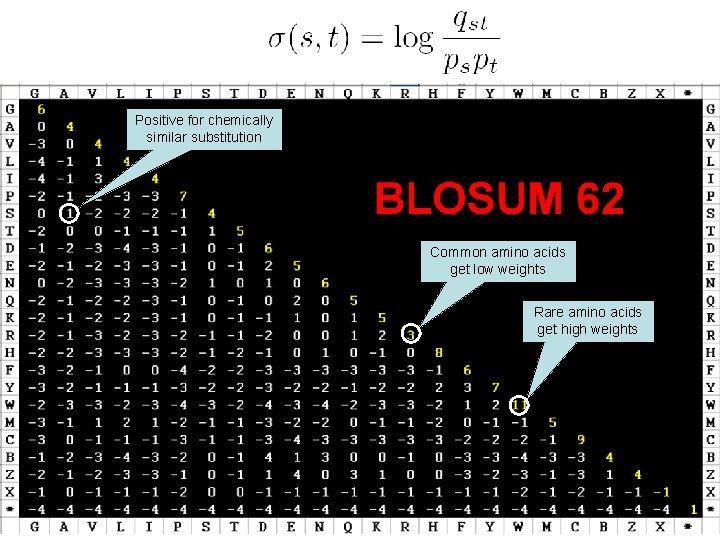
Positive for chemically similar substitution Common amino acids get low weights Rare amino acids get high weights

BLOSUM-N matrices • Constructed from a database called BLOCKS • Contain many closely related sequences – Conserved amino acids may be over-counted • N = 62: the probabilities qst were computed using trusted alignments with no more than 62% identity – identity: % of matched columns • Using this matrix, the Smith-Waterman algorithm is most effective in detecting real alignments with a similar identity level (i. e. ~62%)

• If you want to detect homologous genes with high identify, you may want a BLOSUM matrix with higher N. say BLOSUM 75 • On the other hand, if you want to detect remote homology, you may want to use lower N, say BLOSUM 50 • BLOSUM 62 is the standard

For DNAs • No database of trusted alignments to start with • Specify the percentage identity you would like to detect • You can then get the substitution matrix by some calculation

For example • Suppose p. A = p. C = p. T = p. G = 0. 25 • We want 88% identity • q. AA = q. CC = q. TT = q. GG = 0. 22, the rest = 0. 12/12 = 0. 01 • (A, A) = (C, C) = (G, G) = (T, T) = log (0. 22 / (0. 25*0. 25)) = 1. 26 • (s, t) = log (0. 01 / (0. 25*0. 25)) = -1. 83 for s ≠ t.
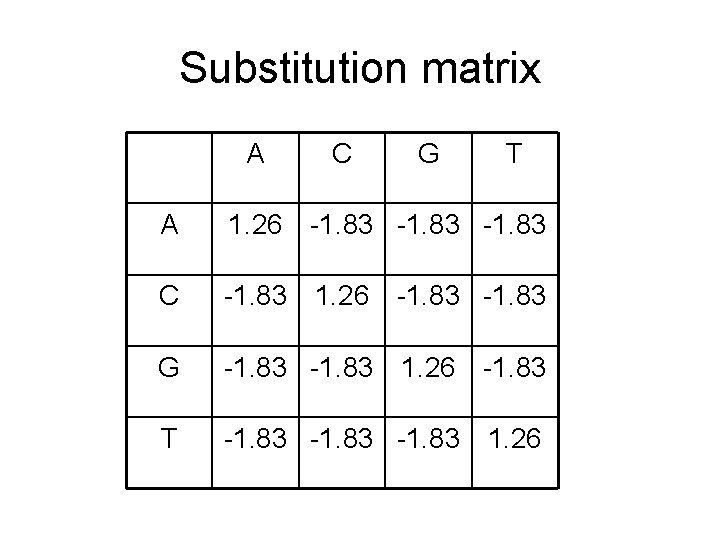
Substitution matrix A C G T A 1. 26 -1. 83 C -1. 83 1. 26 -1. 83 G -1. 83 1. 26 -1. 83 T -1. 83 1. 26
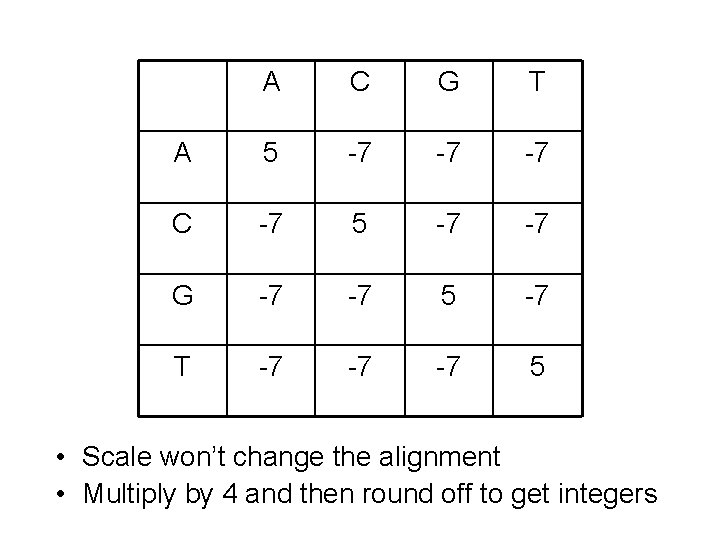
A C G T A 5 -7 -7 -7 C -7 5 -7 -7 G -7 -7 5 -7 T -7 -7 -7 5 • Scale won’t change the alignment • Multiply by 4 and then round off to get integers

Arbitrary substitution matrix • Say you have a substitution matrix provided by someone • It’s important to know what you are actually looking for when you use the matrix

A C G T A 1 -2 -2 -2 A C G T A 5 -4 -4 -4 C -2 1 -2 -2 C -4 5 -4 -4 G -2 -2 1 -2 G -4 -4 5 -4 T -2 -2 -2 1 T -4 -4 -4 5 • What’s the difference? • Which one should I use?
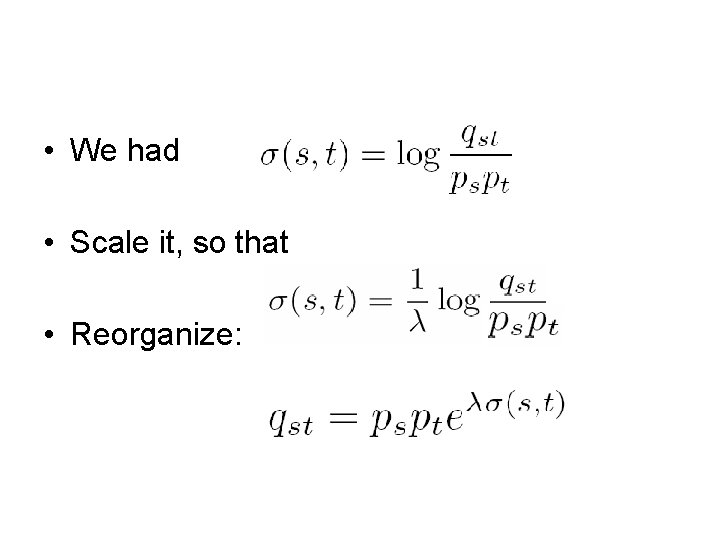
• We had • Scale it, so that • Reorganize:

• Since all probabilities must sum to 1, • We have • Suppose again ps = 0. 25 for any s • We know (s, t) from the substitution matrix • We can solve the equation for λ • Plug λ into to get qst
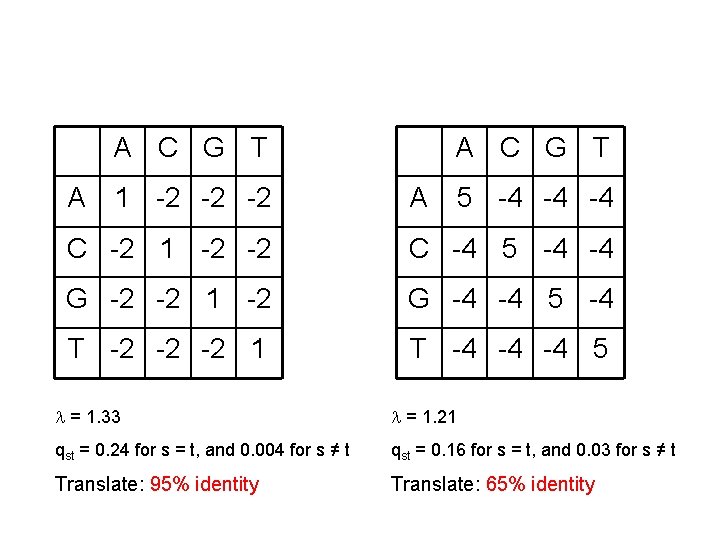
A C G T A 1 -2 -2 -2 A C G T A 5 -4 -4 -4 C -2 1 -2 -2 C -4 5 -4 -4 G -2 -2 1 -2 G -4 -4 5 -4 T -2 -2 -2 1 T -4 -4 -4 5 = 1. 33 = 1. 21 qst = 0. 24 for s = t, and 0. 004 for s ≠ t qst = 0. 16 for s = t, and 0. 03 for s ≠ t Translate: 95% identity Translate: 65% identity
- Slides: 76