Bioinformatics GBIO 0002 1 Biological Sequences Analysis GBIO

Bioinformatics GBIO 0002 1 Biological Sequences Analysis GBIO 00002 -1 Presented by Kirill Bessonov Oct 27, 2015 __________________________________________________________ Kirill Bessonov slide 1

Bioinformatics GBIO 0002 1 Biological Sequences Talk Structure 1. 2. 3. 4. Introduction Global Alignment: Needleman Wunsch Local Alignment: Smith Waterman Practical – Retrieval of sequences using R – Alignment of sequences using R – Finding ORF with R – Comparing two species based on genes __________________________________________________________ Kirill Bessonov slide 2

Bioinformatics GBIO 0002 1 Biological Sequences Gene structure and expression • Gene regions – 5’ untranslated region (5’UTR) • Directly upstream of start codon (AUG) • Regulates translation – 3’ untranslated region (3’UTR) • Right after the stop codon • Influences translation efficiency [protein] – Open Reading Frame (ORF) • Protein coding region (with introns / exons) m. RNA __________________________________________________________ Kirill Bessonov slide 3
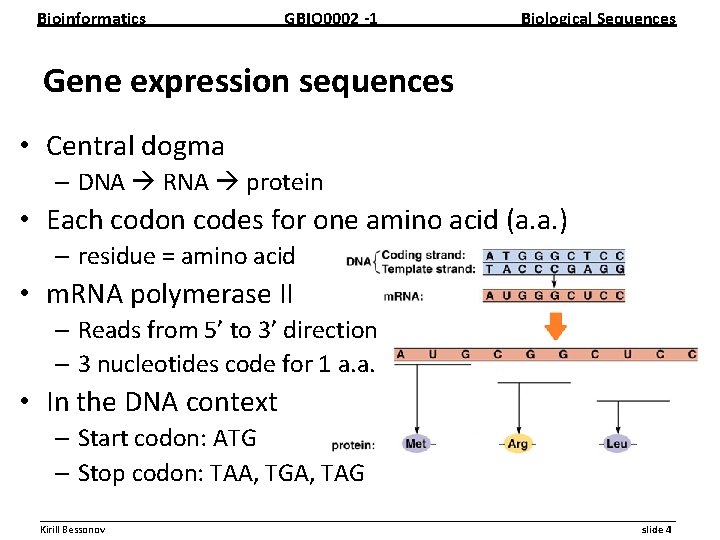
Bioinformatics GBIO 0002 1 Biological Sequences Gene expression sequences • Central dogma – DNA RNA protein • Each codon codes for one amino acid (a. a. ) – residue = amino acid • m. RNA polymerase II – Reads from 5’ to 3’ direction – 3 nucleotides code for 1 a. a. • In the DNA context – Start codon: ATG – Stop codon: TAA, TGA, TAG __________________________________________________________ Kirill Bessonov slide 4
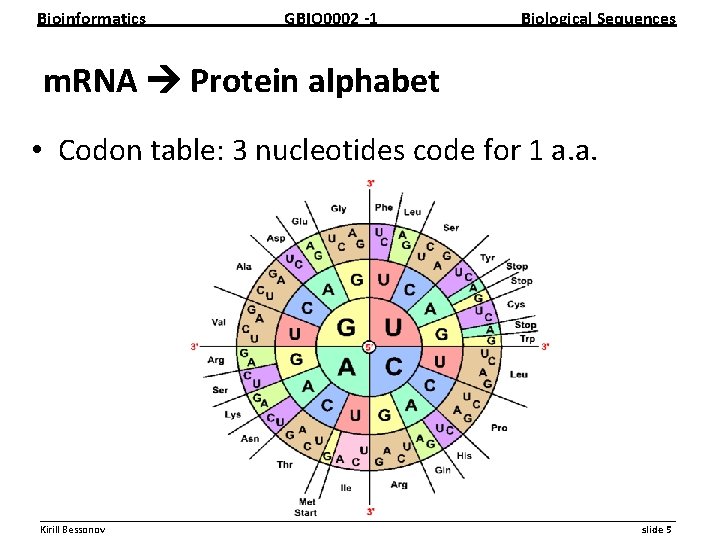
Bioinformatics GBIO 0002 1 Biological Sequences m. RNA Protein alphabet • Codon table: 3 nucleotides code for 1 a. a. __________________________________________________________ Kirill Bessonov slide 5

Bioinformatics GBIO 0002 1 Biological Sequences Biological sequence • Single continuous molecule – DNA ACGGCT – RNA ACGGCU – Protein TA or Thr. Ala __________________________________________________________ Kirill Bessonov slide 6

Bioinformatics GBIO 0002 1 Biological Sequences Biological problem • Given the DNA sequence AATCGGATGCGCGTAGGATCGGTAGGCTTT AAGATCATGCTATTTTCGAGATTCTAGCTA • Answer – Is it likely to be a gene? – What is its possible expression level? – What is the possible structure of the protein product? – Can we get the protein (i. e. express protein)? – Can we figure out the key residues of the protein? – Can we determine the organism from which this sequence came? __________________________________________________________ Kirill Bessonov slide 7
Bioinformatics GBIO 0002 1 Biological Sequences Biological alphabet – In the case of DNA • A, C, T, G – In the case of RNA • A, C, U, G – In the case of protein • 20 amino acids • Complete list is found here __________________________________________________________ Kirill Bessonov slide 8

Bioinformatics GBIO 0002 1 Biological Sequences Words • Short strings of letters from an alphabet • A word of length k is called a k word or k tuple • Examples: – 1 tuple: individual nucleotide – 2 tuple: dinucleotide – 3 tuple: codon __________________________________________________________ Kirill Bessonov slide 9

Bioinformatics GBIO 0002 1 Biological Sequences 2 words: dinucleotides • Composed of 2 nucleotides – Given DNA alphabet {A, T, C, G} • How many possible dinucleoties? • Total of 16: AA, AC, AG, AT … TG, TT • Cp. G islands are regions of DNA – Frequent repetition of Cp. G dinucleotides – Rich in ‘G’ and ‘C’ – Cp. G islands appear in some 70% of promoters of human genes __________________________________________________________ Kirill Bessonov slide 10

Bioinformatics GBIO 0002 1 Biological Sequences Cp. G islands • Cp. G sites could be methylated • Location and methylation state – Impact gene expression • If in promoter region and methylated – May inhibit expression – Present at the gene ‘start’ region • How many Cp. G sites in this DNA sequence? __________________________________________________________ Kirill Bessonov slide 11
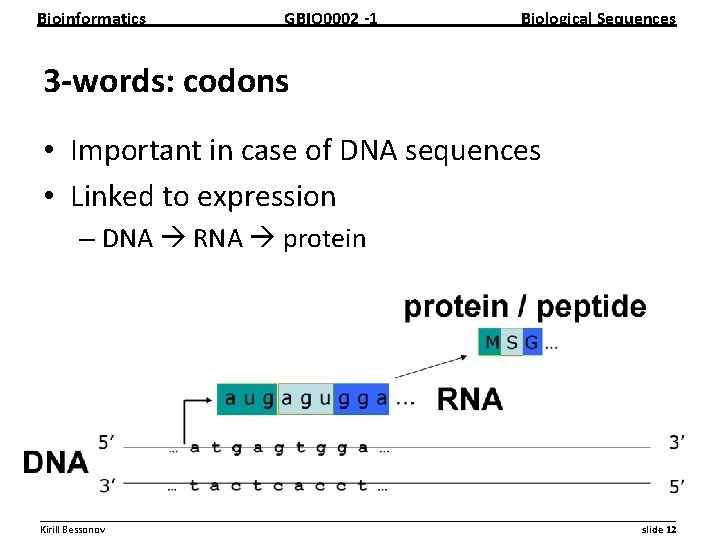
Bioinformatics GBIO 0002 1 Biological Sequences 3 words: codons • Important in case of DNA sequences • Linked to expression – DNA RNA protein __________________________________________________________ Kirill Bessonov slide 12

Bioinformatics GBIO 0002 1 Biological Sequences Patterns • Recognizing motifs, sites, signals, domains – functionally important regions – a conserved motif consensus sequence – Often words (in bold) are used interchangeably • Gene starts with an “ATG” codon – Identify # of potential gene start sites • 4 sites AATCGGATGCGCGTAGGATCGGTAGGCTTTAAGATCATGCTATTTTCGAGATT CGATTCTAGGTTTAGCTTAGTGCCAGAAATCGGATGCGCGTAGGATCGG TAGGGTAGGCTTTAAGATCATGCTATTTTCGAGATTCTAGCTAGGTTTTTAGT GCCAGAAATCGTTAGTGCCAGAAATCGATT __________________________________________________________ Kirill Bessonov slide 13

Bioinformatics GBIO 0002 1 Biological Sequences Bases distribution • The distribution of bases within a DNA – is not ordinarily uniform • i. e. not P(A) =P(C) = P(G) = P(T) = 0. 25 • There may be an excess of G over C on the leading strands – This can be described by the “GC skew”, __________________________________________________________ Kirill Bessonov slide 14
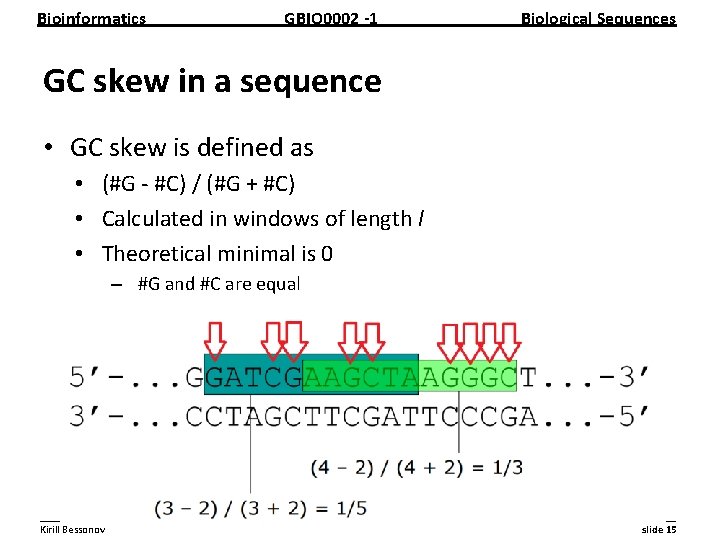
Bioinformatics GBIO 0002 1 Biological Sequences GC skew in a sequence • GC skew is defined as • (#G #C) / (#G + #C) • Calculated in windows of length l • Theoretical minimal is 0 – #G and #C are equal __________________________________________________________ Kirill Bessonov slide 15
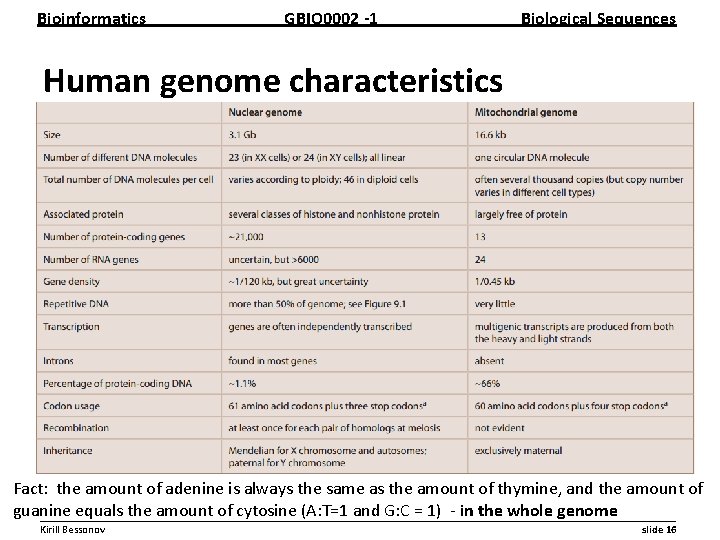
Bioinformatics GBIO 0002 1 Biological Sequences Human genome characteristics Fact: the amount of adenine is always the same as the amount of thymine, and the amount of guanine equals the amount of cytosine (A: T=1 and G: C = 1) in the whole genome __________________________________________________________ Kirill Bessonov slide 16

Bioinformatics GBIO 0002 1 Biological Sequences Alignments __________________________________________________________ Kirill Bessonov slide 17
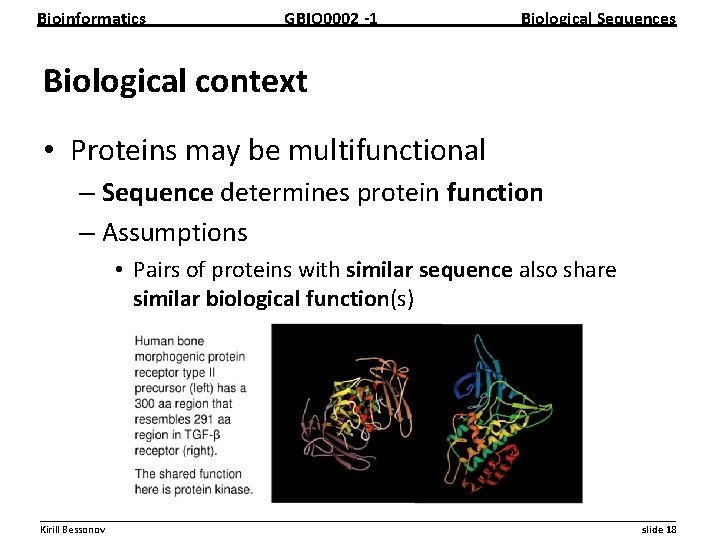
Bioinformatics GBIO 0002 1 Biological Sequences Biological context • Proteins may be multifunctional – Sequence determines protein function – Assumptions • Pairs of proteins with similar sequence also share similar biological function(s) __________________________________________________________ Kirill Bessonov slide 18
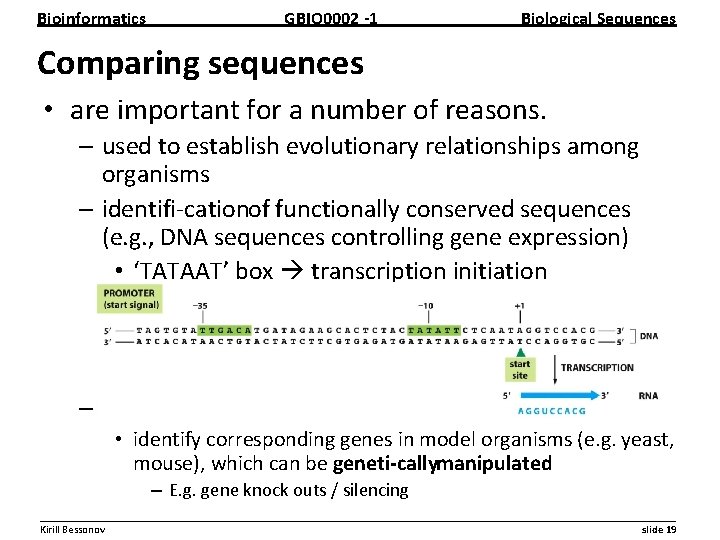
Bioinformatics GBIO 0002 1 Biological Sequences Comparing sequences • are important for a number of reasons. – used to establish evolutionary relationships among organisms – identifi cation of functionally conserved sequences (e. g. , DNA sequences controlling gene expression) • ‘TATAAT’ box transcription initiation – develop models for human diseases • identify corresponding genes in model organisms (e. g. yeast, mouse), which can be geneti cally manipulated – E. g. gene knock outs / silencing __________________________________________________________ Kirill Bessonov slide 19

Bioinformatics GBIO 0002 1 Biological Sequences Deletions/insertions/substitution of nucleotides • Mutations are of 3 main types – Deletions – Insertions – Substitution – Cause a shift in the m. RNA reading frame __________________________________________________________ Kirill Bessonov slide 20
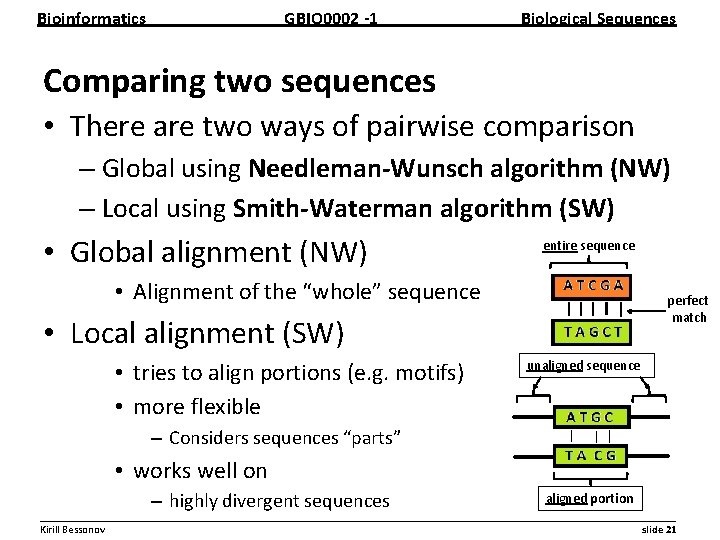
Bioinformatics GBIO 0002 1 Biological Sequences Comparing two sequences • There are two ways of pairwise comparison – Global using Needleman Wunsch algorithm (NW) – Local using Smith Waterman algorithm (SW) • Global alignment (NW) entire sequence • Alignment of the “whole” sequence perfect match • Local alignment (SW) • tries to align portions (e. g. motifs) • more flexible unaligned sequence – Considers sequences “parts” • works well on – highly divergent sequences aligned portion __________________________________________________________ Kirill Bessonov slide 21

Bioinformatics GBIO 0002 1 Biological Sequences Global alignment __________________________________________________________ Kirill Bessonov slide 22

Bioinformatics GBIO 0002 1 Biological Sequences Global alignment (NW) • Sequences are aligned end to end along their entire length • Many possible alignments are produced – The alignment with the highest score is chosen • Naïve algorithm is very inefficient (Oexp) – To align sequence of length 15, need to consider • (insertion, deletion, gap)15 = 315 = 1, 4*107 – Impractical for sequences of length >20 nt • Used to analyze homology/similarity of – genes and proteins – between species __________________________________________________________ Kirill Bessonov slide 23

Bioinformatics GBIO 0002 1 Biological Sequences Methodology of global alignment (1 of 4) • s 1: . . AATA. . s 2: . . AACA. . s 1: . . AAT-A. . s 2: . . AACA. . s 1: . . AATA. . s 2: . . AATA. . __________________________________________________________ Kirill Bessonov slide 24

Bioinformatics GBIO 0002 1 Biological Sequences Methodology of global alignment (2 of 4) • The matrix should have extra column and row – M+1 columns , where M is the length sequence M – N+1 rows, where N is the length of sequence N 1. Initialize the matrix – introduce gap penalty at every initial position along rows and columns – Scores at each cell are cumulative W H A T 2 0 2 2 2 4 2 6 2 8 W 2 H 4 2 Y 6 __________________________________________________________ Kirill Bessonov slide 25
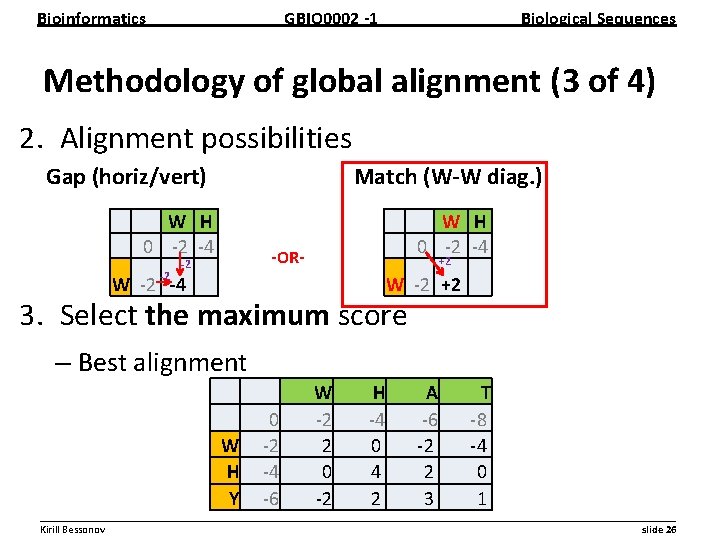
Bioinformatics GBIO 0002 1 Biological Sequences Methodology of global alignment (3 of 4) 2. Alignment possibilities Gap (horiz/vert) Match (W W diag. ) W H 0 2 4 2 W H 0 2 4 OR 2 +2 W 2 4 W 2 +2 3. Select the maximum score – Best alignment 0 W 2 H 4 Y 6 W 2 2 0 2 H 4 0 4 2 A 6 2 2 3 T 8 4 0 1 __________________________________________________________ Kirill Bessonov slide 26
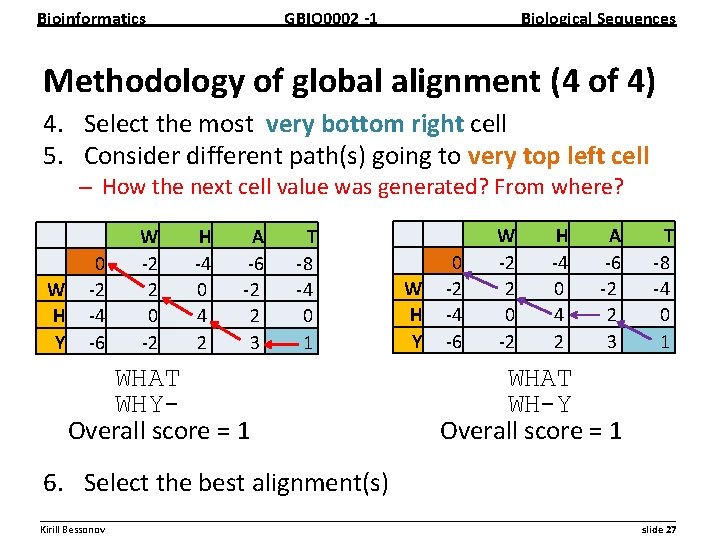
Bioinformatics GBIO 0002 1 Biological Sequences Methodology of global alignment (4 of 4) 4. Select the most very bottom right cell 5. Consider different path(s) going to very top left cell – How the next cell value was generated? From where? 0 W 2 H 4 Y 6 W 2 2 0 2 H 4 0 4 2 A 6 2 2 3 T 8 4 0 1 WHAT WHY Overall score = 1 0 W 2 H 4 Y 6 W 2 2 0 2 H 4 0 4 2 A 6 2 2 3 T 8 4 0 1 WHAT WH-Y Overall score = 1 6. Select the best alignment(s) __________________________________________________________ Kirill Bessonov slide 27
Bioinformatics GBIO 0002 1 Biological Sequences Local alignment __________________________________________________________ Kirill Bessonov slide 28

Bioinformatics GBIO 0002 1 Biological Sequences Local alignment (SW) • Sequences are aligned to find regions where the best alignment occurs (i. e. highest score) • Assumes a local context (aligning parts of seq. ) • Ideal for finding short motifs, DNA binding sites – helix loop helix (b. HLH) motif – TATAAT box (a famous promoter region) – DNA binding site • Works well on highly divergent sequences __________________________________________________________ Kirill Bessonov slide 29

Bioinformatics GBIO 0002 1 Biological Sequences Methodology of local alignment (1 of 4) • __________________________________________________________ Kirill Bessonov slide 30
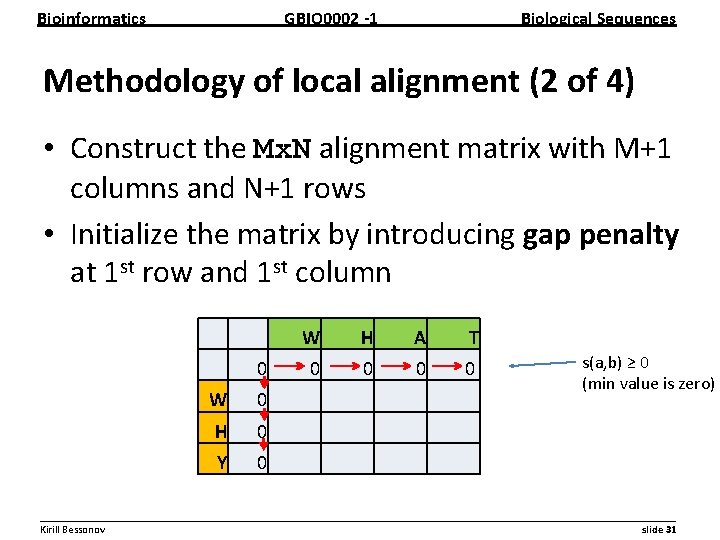
Bioinformatics GBIO 0002 1 Biological Sequences Methodology of local alignment (2 of 4) • Construct the Mx. N alignment matrix with M+1 columns and N+1 rows • Initialize the matrix by introducing gap penalty at 1 st row and 1 st column W H A T 0 0 0 W 0 H 0 Y 0 s(a, b) ≥ 0 (min value is zero) __________________________________________________________ Kirill Bessonov slide 31
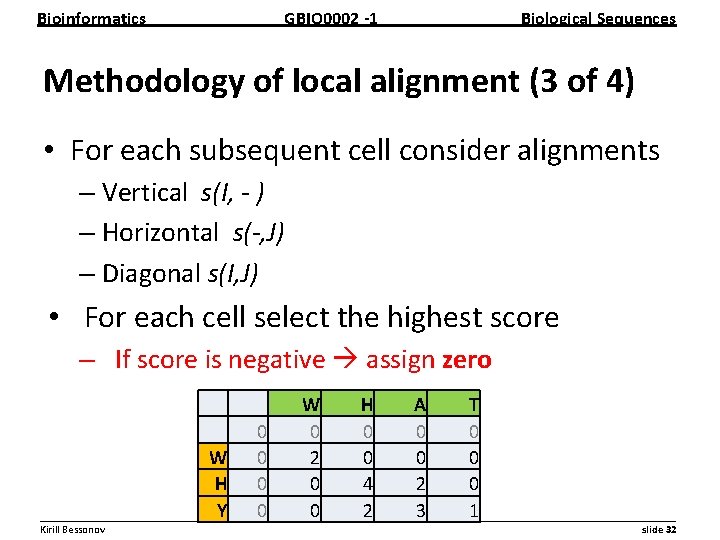
Bioinformatics GBIO 0002 1 Biological Sequences Methodology of local alignment (3 of 4) • For each subsequent cell consider alignments – Vertical s(I, - ) – Horizontal s(-, J) – Diagonal s(I, J) • For each cell select the highest score – If score is negative assign zero W H A T 0 0 0 W 0 2 0 0 0 H 0 0 4 2 0 Y 0 0 2 3 1 __________________________________________________________ Kirill Bessonov slide 32
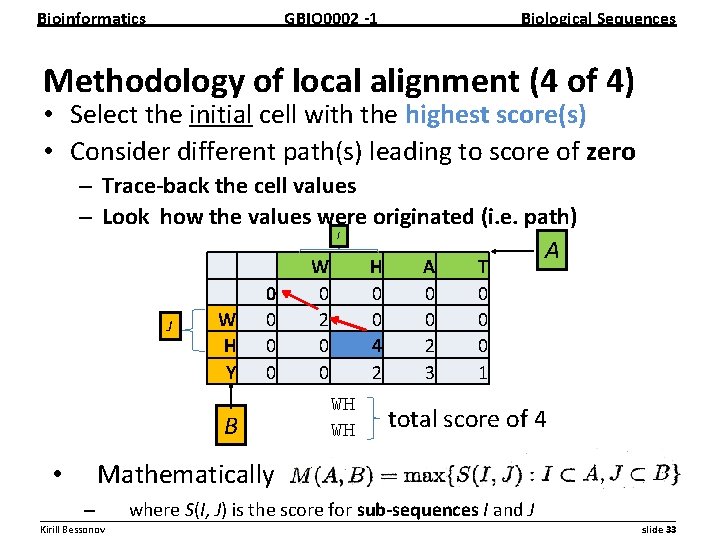
Bioinformatics GBIO 0002 1 Biological Sequences Methodology of local alignment (4 of 4) • Select the initial cell with the highest score(s) • Consider different path(s) leading to score of zero – Trace back the cell values – Look how the values were originated (i. e. path) I J W H Y 0 0 B W 0 2 0 0 H 0 0 4 2 WH WH A 0 0 2 3 T 0 0 0 1 A total score of 4 Mathematically • – where S(I, J) is the score for sub sequences I and J __________________________________________________________ Kirill Bessonov slide 33

Bioinformatics GBIO 0002 1 Biological Sequences Local alignment illustration (1 of 2) • __________________________________________________________ Kirill Bessonov slide 34

Bioinformatics GBIO 0002 1 Biological Sequences Local alignment illustration (2 of 2) G G C T C A A T C A 0 0 0 A 0 0 0 2 2 0 0 2 C 0 0 0 2 0 2 0 1 1 2 0 C 0 0 0 2 1 2 1 0 0 3 2 1 T 0 0 0 4 2 1 0 2 1 A 0 0 0 2 3 4 3 1 1 3 2 A 0 0 1 5 6 4 2 3 G 0 0 2 2 0 0 0 3 4 5 3 1 G 0 0 2 4 2 0 0 1 2 3 4 2 __________________________________________________________ Kirill Bessonov slide 35
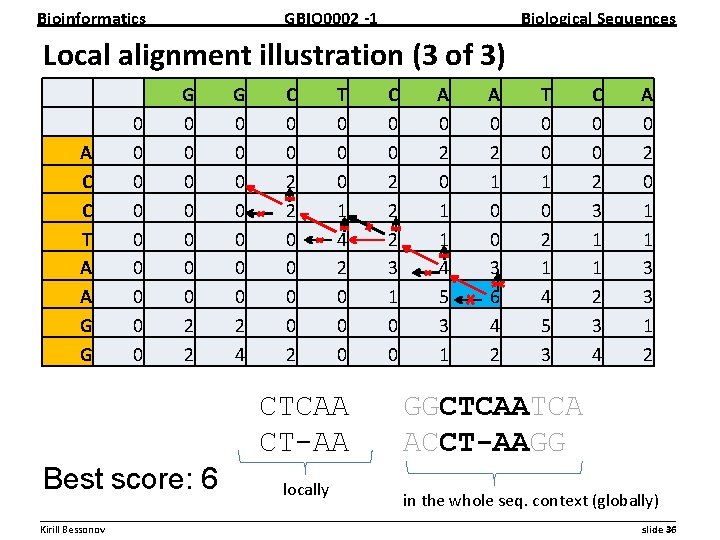
Bioinformatics GBIO 0002 1 Biological Sequences Local alignment illustration (3 of 3) A C C T A A G G 0 0 0 0 0 G 0 0 0 0 2 2 G 0 0 0 0 2 4 C 0 0 2 2 0 0 2 T 0 0 0 1 4 2 0 0 0 CTCAA CT-AA Best score: 6 locally C 0 0 2 2 2 3 1 0 0 A 0 2 0 1 1 4 5 3 1 A 0 2 1 0 0 3 6 4 2 T 0 0 1 0 2 1 4 5 3 C 0 0 2 3 1 1 2 3 4 A 0 2 0 1 1 3 3 1 2 GGCTCAATCA ACCT-AAGG in the whole seq. context (globally) __________________________________________________________ Kirill Bessonov slide 36

Bioinformatics GBIO 0002 1 Biological Sequences Aligning proteins Globally and Locally __________________________________________________________ Kirill Bessonov slide 37
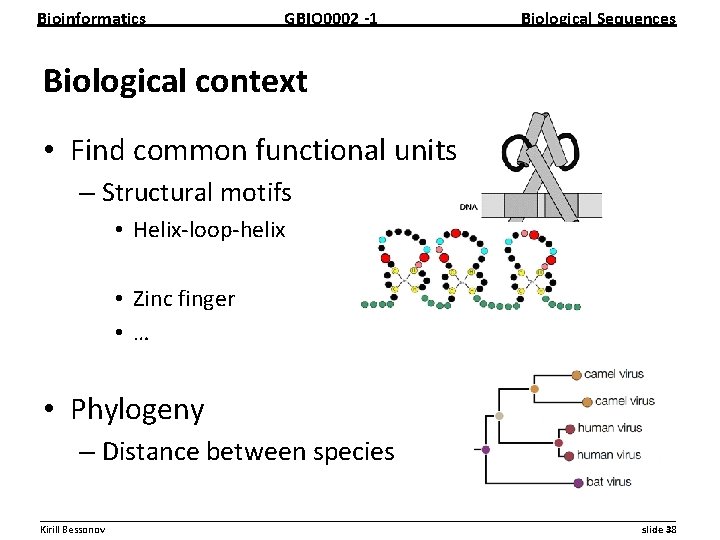
Bioinformatics GBIO 0002 1 Biological Sequences Biological context • Find common functional units – Structural motifs • Helix loop helix • Zinc finger • … • Phylogeny – Distance between species __________________________________________________________ Kirill Bessonov slide 38

Bioinformatics GBIO 0002 1 Biological Sequences Protein Alignment • Protein local and global alignment – follows the same rules as we saw with DNA/RNA • Differences (∆) – alphabet of proteins is 22 residues (aa) long – scoring/substitution matrices used (BLOSUM) • protein proprieties are taken into account – residues that are totally different due to charge such as polar Lysine and apolar Glycine are given a low score __________________________________________________________ Kirill Bessonov slide 39

Bioinformatics GBIO 0002 1 Biological Sequences Substitution matrices • Protein sequences are more complex – matrices = collection of scoring rules • Matrices over events such as – mismatch and perfect match • Need to define gap penalty separately • E. g. BLOcks SUbstitution Matrix (BLOSUM) __________________________________________________________ Kirill Bessonov slide 40

Bioinformatics GBIO 0002 1 Biological Sequences BLOSUM x matrices • Constructed from aligned sequences with specific x% similarity – matrix built using sequences with no more then 50% similarity is called BLOSUM 50 • For highly mutating / dissimilar sequences use – BLOSUM 45 and lower • For highly conserved / similar sequences use – BLOSUM 62 and higher __________________________________________________________ Kirill Bessonov slide 41
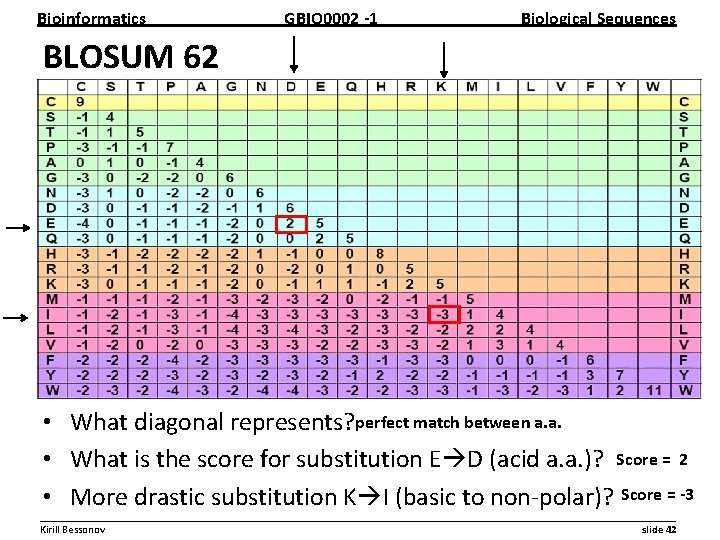
Bioinformatics GBIO 0002 1 Biological Sequences BLOSUM 62 • What diagonal represents? perfect match between a. a. • What is the score for substitution E D (acid a. a. )? Score = 2 • More drastic substitution K I (basic to non polar)? Score = 3 __________________________________________________________ Kirill Bessonov slide 42

Bioinformatics GBIO 0002 1 Biological Sequences Practical problem: Align following sequences both globally and locally using BLOSUM 62 matrix with gap penalty of -8 Sequence A: AAEEKKLAAA Sequence B: AARRIA __________________________________________________________ Kirill Bessonov slide 43
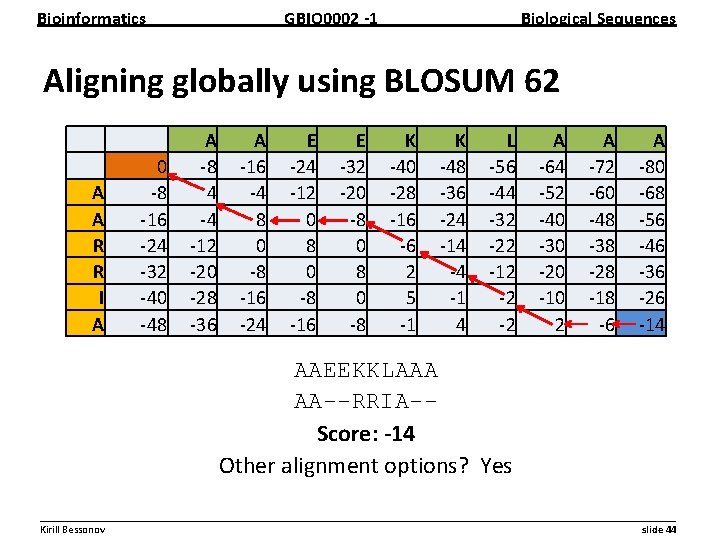
Bioinformatics GBIO 0002 1 Biological Sequences Aligning globally using BLOSUM 62 A A R R I A 0 8 16 24 32 40 48 A 8 4 4 12 20 28 36 A 16 4 8 0 8 16 24 E 24 12 0 8 16 E 32 20 8 0 8 K 40 28 16 6 2 5 1 K 48 36 24 14 4 1 4 L 56 44 32 22 12 2 2 A 64 52 40 30 20 10 2 A 72 60 48 38 28 18 6 A 80 68 56 46 36 26 14 AAEEKKLAAA AA--RRIA-Score: 14 Other alignment options? Yes __________________________________________________________ Kirill Bessonov slide 44
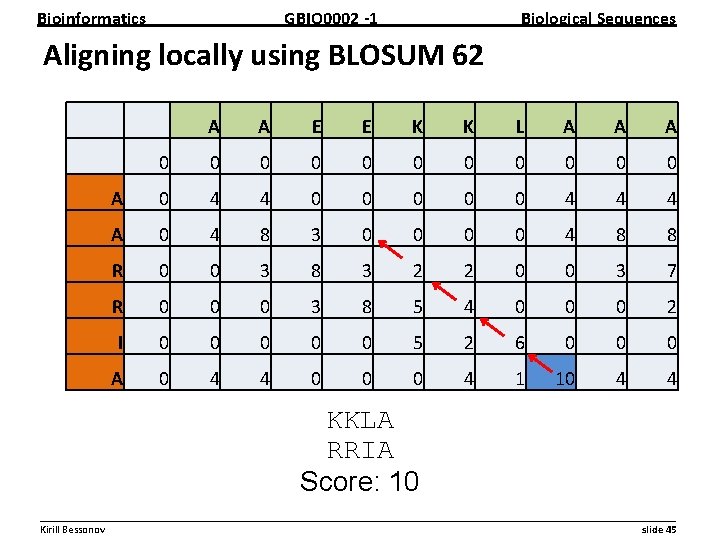
Bioinformatics GBIO 0002 1 Biological Sequences Aligning locally using BLOSUM 62 A A E E K K L A A A 0 0 0 A 0 4 4 0 0 0 4 4 4 A 0 4 8 3 0 0 4 8 8 R 0 0 3 8 3 2 2 0 0 3 7 R 0 0 0 3 8 5 4 0 0 0 2 I 0 0 0 5 2 6 0 0 0 A 0 4 4 0 0 0 4 1 10 4 4 KKLA RRIA Score: 10 __________________________________________________________ Kirill Bessonov slide 45

Bioinformatics GBIO 0002 1 Biological Sequences Practical 1 of 4 : Sequence Retrieval and Analysis via R __________________________________________________________ Kirill Bessonov slide 46
Bioinformatics GBIO 0002 1 Biological Sequences Protein database • Uni. Prot database (http: //www. uniprot. org/) has high quality protein data manually curated • It is manually curated • Each protein is assigned Uni. Prot ID __________________________________________________________ Kirill Bessonov slide 47

Bioinformatics GBIO 0002 1 Biological Sequences Retrieving data from • In search field one can enter either use Uni. Prot ID or common protein name – example: myelin basic protein Uniprot ID • We will use retrieve data for P 02686 __________________________________________________________ Kirill Bessonov slide 48

Bioinformatics GBIO 0002 1 Biological Sequences FASTA format • FASTA format is widely used and has the following parameters – Sequence name start with > sign – The fist line corresponds to protein name Actual protein sequence starts from 2 nd line __________________________________________________________ Kirill Bessonov slide 49

Bioinformatics GBIO 0002 1 Biological Sequences Retrieving protein data with R • Can “talk” programmatically to Uni. Prot database using R and seqin. R library – seqin. R library is suitable for • “Biological Sequences Retrieval and Analysis” • Detailed manual could be found here – Install this library in your R environment install. packages("seqinr") library("seqinr") – Choose database to retrieve data from choosebank("swissprot") – Download data object for target protein (P 02686) MBP_HUMAN = query("MBP_HUMAN", "AC=P 02686") – See sequence of the object MBP_HUMAN_seq = get. Sequence(MBP_HUMAN); MBP_HUMAN_seq __________________________________________________________ Kirill Bessonov slide 50

Bioinformatics GBIO 0002 1 Biological Sequences Dot Plot (comparison of 2 sequences) (1 of 2) • Each sequence plotted on vertical or horizontal dimension – If two a. a. from two sequences at given positions are identical the dot is plotted – matching sequence segments appear as diagonal lines (that could be parallel to the absolute diagonal line if insertion or gap is present) __________________________________________________________ Kirill Bessonov slide 51
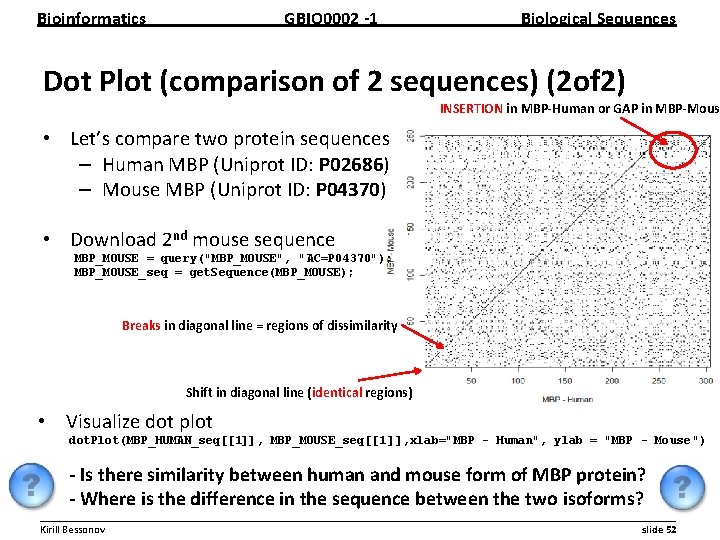
Bioinformatics GBIO 0002 1 Biological Sequences Dot Plot (comparison of 2 sequences) (2 of 2) INSERTION in MBP Human or GAP in MBP Mous • Let’s compare two protein sequences – Human MBP (Uniprot ID: P 02686) – Mouse MBP (Uniprot ID: P 04370) • Download 2 nd mouse sequence MBP_MOUSE = query("MBP_MOUSE", "AC=P 04370"); MBP_MOUSE_seq = get. Sequence(MBP_MOUSE); Breaks in diagonal line = regions of dissimilarity Shift in diagonal line (identical regions) • Visualize dot plot dot. Plot(MBP_HUMAN_seq[[1]], MBP_MOUSE_seq[[1 ]], xlab="MBP - Human", ylab = "MBP - Mouse ") Is there similarity between human and mouse form of MBP protein? Where is the difference in the sequence between the two isoforms? __________________________________________________________ Kirill Bessonov slide 52

Bioinformatics GBIO 0002 1 Biological Sequences Practical 2 of 4: Pairwise global and local alignments via R and Biostrings __________________________________________________________ Kirill Bessonov slide 53

Bioinformatics GBIO 0002 1 Biological Sequences Installing Biostrings library • DNA_subst_matrix __________________________________________________________ Kirill Bessonov slide 54

Bioinformatics GBIO 0002 1 Biological Sequences Global alignment using R and Biostrings • Create two sting vectors (i. e. sequences) seq. A = "GATTA" seq. B = "GTTA" • Use pairwise. Alignment() and the defined rules global. Align. AB = pairwise. Alignment(seq. A, seq. B, substitution. Matrix = DNA_subst_matrix, gap. Opening = -2, score. Only = FALSE, type="global") • Visualize best paths (i. e. alignments) global. Align. AB Global Pairwise. Aligned. Fixed. Subject (1 of 1) pattern: [1] GATTA subject: [1] G-TTA score: 2 __________________________________________________________ Kirill Bessonov slide 55

Bioinformatics GBIO 0002 1 Biological Sequences Local alignment using R and Biostrings • Input two sequences seq. A = "AGGATTTTAAAA" seq. B = "TTTT" • The scoring rules will be the same as we used for global alignment local. Align. AB = pairwise. Alignment(seq. A, seq. B, substitution. Matrix = DNA_subst_matrix, gap. Opening = -2, score. Only = FALSE, type="local") • Visualize alignment global. Align. AB Local Pairwise. Aligned. Fixed. Subject (1 of 1) pattern: [5] TTTT subject: [1] TTTT score: 8 __________________________________________________________ Kirill Bessonov slide 56

Bioinformatics GBIO 0002 1 Biological Sequences Aligning protein sequences • Protein sequences alignments are very similar except the substitution matrix is specified data(BLOSUM 62) BLOSUM 62 • Will align sequences seq. A = "PAWHEAE" seq. B = "HEAGAWGHEE" • Execute the global alignment global. Align. AB <- pairwise. Alignment(seq. A, seq. B, substitution. Matrix = "BLOSUM 62", gap. Opening = -2, gap. Extension = -8, score. Only = FALSE) __________________________________________________________ Kirill Bessonov slide 57

Bioinformatics GBIO 0002 1 Biological Sequences Practical 3 of 4: DNA sequence statistics and Seqinr __________________________________________________________ Kirill Bessonov slide 58

Bioinformatics GBIO 0002 1 Biological Sequences Retrieving genome sequence data Can retrieve sequence data from NCBI 1. Manually via web. GUI 2. Programmatically via R • DEN 1 Dengue virus genome sequence, which has NCBI accession NC_001477 • Gain in speed compared to manual retrieval • More complex queries __________________________________________________________ Kirill Bessonov slide 59
Bioinformatics GBIO 0002 1 Biological Sequences Manually • NCBI Sequence Database via its website www. ncbi. nlm. nih. gov • Dengue DEN 1 DNA sequence is a viral DNA sequence • NCBI accession is NC_001477 __________________________________________________________ Kirill Bessonov slide 60
Bioinformatics GBIO 0002 1 Biological Sequences NCBI database __________________________________________________________ Kirill Bessonov slide 61
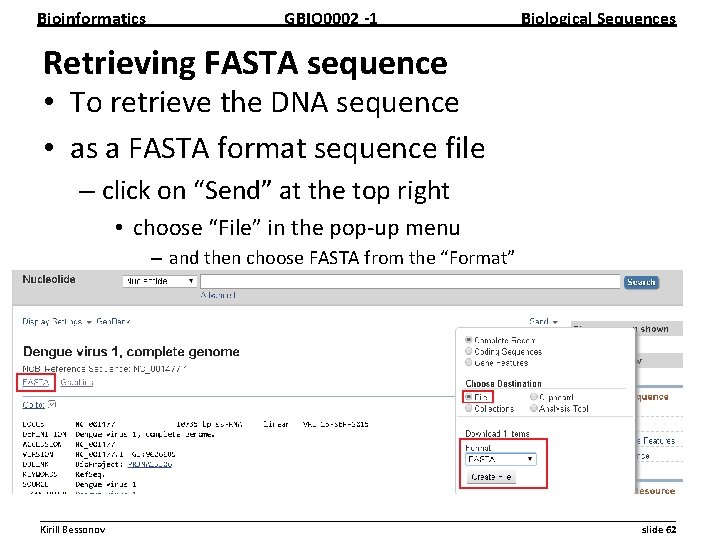
Bioinformatics GBIO 0002 1 Biological Sequences Retrieving FASTA sequence • To retrieve the DNA sequence • as a FASTA format sequence file – click on “Send” at the top right • choose “File” in the pop up menu – and then choose FASTA from the “Format” » click on “Create file”. __________________________________________________________ Kirill Bessonov slide 62

Bioinformatics GBIO 0002 1 Biological Sequences Retrieving genome sequence data using Seqin. R • Can retrieve sequences much faster programmatically getncbiseq <- function(accession) { require("seqinr"); # this function requires the Seqin. R R package # first find which ACNUC database the accession is stored in: dbs <- c("genbank", "refseq. Viruses", "bacterial"); numdbs <- length(dbs); for (i in 1: numdbs) { db <- dbs[i]; choosebank(db); # check if the sequence is in ACNUC database 'db': resquery <- try(query(". tmpquery", paste("AC=", accession)), silent = TRUE); if (!(inherits(resquery, "try-error"))) { queryname <- "query 2"; thequery <- paste("AC=", accession, sep=""); query 2 <- query(`queryname`, `thequery`); # see if a sequence was retrieved: seq <- get. Sequence(query 2$req[[1]]); closebank(); return(seq); } closebank(); } print(paste("ERROR: accession", accession, "was not found")); } dengueseq <- getncbiseq("NC_001477"); dengueseq[1: 50]; length(dengueseq); __________________________________________________________ Kirill Bessonov slide 63

Bioinformatics GBIO 0002 1 Biological Sequences Base composition of a DNA sequence • Count frequencies of the 4 nucleotides table(dengueseq); dengueseq; a c g t 3426 2240 2770 2299 • This means that the DEN 1 Dengue virus genome sequence has – 3426 As, 2240 Cs, 2770 Gs and 2299 Ts __________________________________________________________ Kirill Bessonov slide 64

Bioinformatics GBIO 0002 1 Biological Sequences GC Content of DNA • the fraction of the sequence that consists of Gs and Cs, ie. the %(G+C). – the percentage of the bases in the genome that are – Gs or Cs GC(dengueseq) [1] 0. 4666977 (2240+2770)*100/(3426+2240+2770+2299) [1] 46. 66977 __________________________________________________________ Kirill Bessonov slide 65

Bioinformatics GBIO 0002 1 Biological Sequences Di nucleotides • it is also interesting to know – the frequency of longer DNA “words” – dinucleotides • ie. “AA”, “AG”, “AC”, “AT”, “CA”, “CG”, “CC”, “CT”, “GA”, “GG”, “GC”, “GT”, “TA”, “TG”, “TC”, and “TT” count(dengueseq, 2) aa ac ag at 1108 720 890 708 ca 901 cc 523 cg 261 ct 555 ga 976 gc 500 gg 787 gt 507 ta 440 tc 497 tg 832 tt 529 __________________________________________________________ Kirill Bessonov slide 66

Bioinformatics GBIO 0002 1 Biological Sequences Practical 4 of 4: Using BLAST for sequence identification __________________________________________________________ Kirill Bessonov slide 67

Bioinformatics GBIO 0002 1 Biological Sequences BLAST • Basic Local Alignment Search Tool • Many different types • http: //blast. ncbi. nlm. nih. gov/Blast. cgi __________________________________________________________ Kirill Bessonov slide 68

Bioinformatics GBIO 0002 1 Biological Sequences Types • blastn – nucleotide query vs nucleotide database • blastp – protein query vs protein DB • blastx – translated in 6 frames nucleotide query vs protein DB __________________________________________________________ Kirill Bessonov slide 69

Bioinformatics GBIO 0002 1 Biological Sequences Sequence identity • Want to know which genes are coded by the genomic sequence • >human_genomic_seq TGGACTCTGCTTCCCAGACAGTACCCCTGACAGAACTGCCACTCTCCCCACCTG ACCCTGTTAGGAAGGTACAACCTATGAAGAAAAAGCCAGAATACAGGGGACATGTGAGC C ACAGACAACACAAGTGTGCACAACACCTCTGAGCTTTTCTTGATTCAAGGGCTAG TGAGAACGCCCCGCCAGAGATTTACCTCTGGTCTTCTGAGGTTGAGGGCTCGTTCTCTCT TCCTGAATGTAAAGGTCAAGATGCTGGGCCTCAGTTTCCTCTTACATACTCACCAAAAGG CTCTCCTGATCAGAGAAGCAGGATGCTGCACTTGTCCTCCTGTCGATGCTCTTGGCTATG ACAAAATCTGAGCTTACCTTCTCTTGCCCACCTCTAAACCCCATAAGGGCTTCGTTCTGT GTCTCTTGAGAATGTCCCTATCTCCAACTCTGTCATACGGGGGAGAGCGAGTGGGAAGG A TCCAGGGCTCAGACCCCGGCGCATGGACCTAGTCGGGGGCGCTGGCTCAGCCCCGCGCGCCCCCGTCGCAGCCGACGCGCGCTCCCGGGAGGCGGCGGCAGAGGCAG CATCCACAGCATCAGCAGCCTCAGCTTCATCCCCGGGCGGTCTCCGGCGGGGAAGGCCG GTGGGACAAACGGACAGAAGGCAAAGTGCCCGCAATGGAGCATCCTTTGGCGCG GGCCGTGCGGGAGCTGCCTTTGATCCCGTGAGCTTTGCGCGGCGGCCCCAGACCCTGTT GCGGGTCGTGTCCTGG __________________________________________________________ Kirill Bessonov slide 70
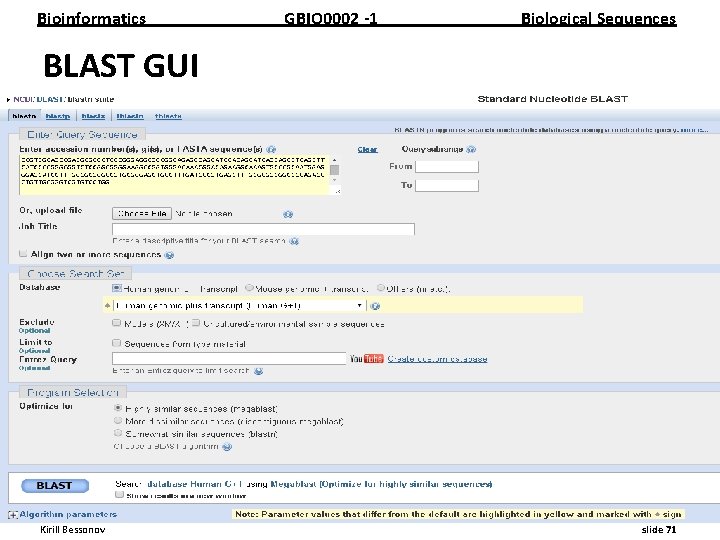
Bioinformatics GBIO 0002 1 Biological Sequences BLAST GUI __________________________________________________________ Kirill Bessonov slide 71
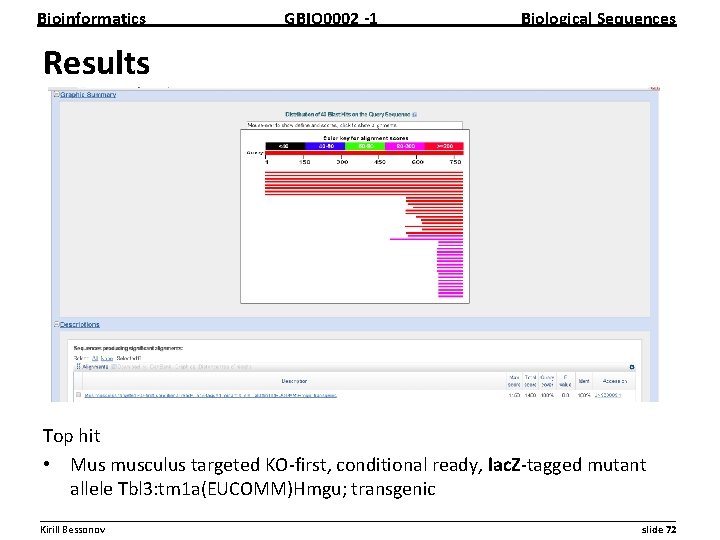
Bioinformatics GBIO 0002 1 Biological Sequences Results Top hit • Mus musculus targeted KO first, conditional ready, lac. Z tagged mutant allele Tbl 3: tm 1 a(EUCOMM)Hmgu; transgenic __________________________________________________________ Kirill Bessonov slide 72

Bioinformatics GBIO 0002 1 Biological Sequences __________________________________________________________ Kirill Bessonov slide 73

Bioinformatics GBIO 0002 1 Biological Sequences Resources • Online Tutorial on Sequence Alignment – http: //a little book of r for bioinformatics. readthedocs. org/en/latest/src/chapter 4. html • Graphical alignment of proteins – http: //www. itu. dk/~sestoft/bsa/graphalign. html • Pairwise alignment of DNA and proteins using your rules: – http: //www. bioinformatics. org/sms 2/pairwise_align_dna. html • Documentation on libraries – Biostings: http: //www. bioconductor. org/packages/2. 10/bioc/manuals/Biostrings/man/Biostrings. pdf – Seqin. R: http: //seqinr. r forge. r project. org/seqinr_2_0 7. pdf __________________________________________________________ Kirill Bessonov slide 74
- Slides: 74