Clustering The WWW of clustering in Bioinformatics or

Clustering The WWW of clustering in Bioinformatics. or, How homo-hokjens thinks ©CMBI 2006

Disclaimer I know nothing about the actual clustering techniques; for that you must ask Lutgarde, or Ron, or any of their colleagues. I will talk today about fields, mainly bioinformatics, where clustering is being used. And we mainly use clustering to classify things. ©CMBI 2006

My daily clustering ©CMBI 2006

Why clustering sequences? Remember bioinformatics 1? The main reason for aligning sequences is that allows us to transfer information. If there are many sequences available, clustering can help us figure out from which of those sequences we are going to transfer that information. ©CMBI 2006
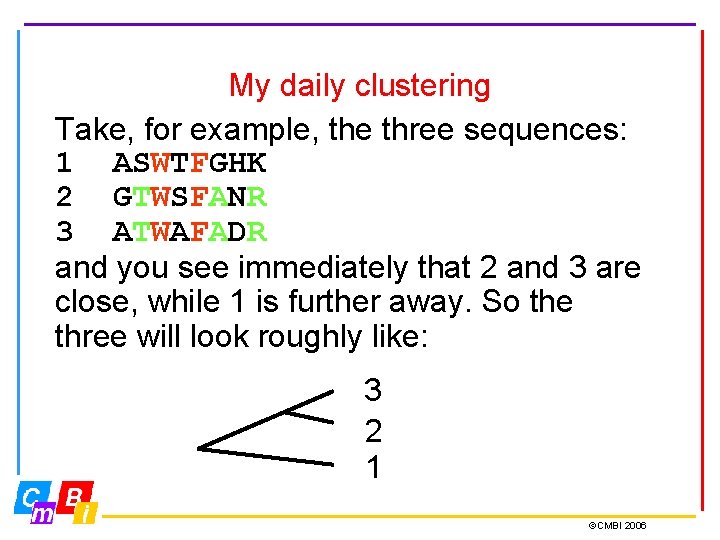
My daily clustering Take, for example, the three sequences: 1 ASWTFGHK 2 GTWSFANR 3 ATWAFADR and you see immediately that 2 and 3 are close, while 1 is further away. So the three will look roughly like: 3 2 1 ©CMBI 2006
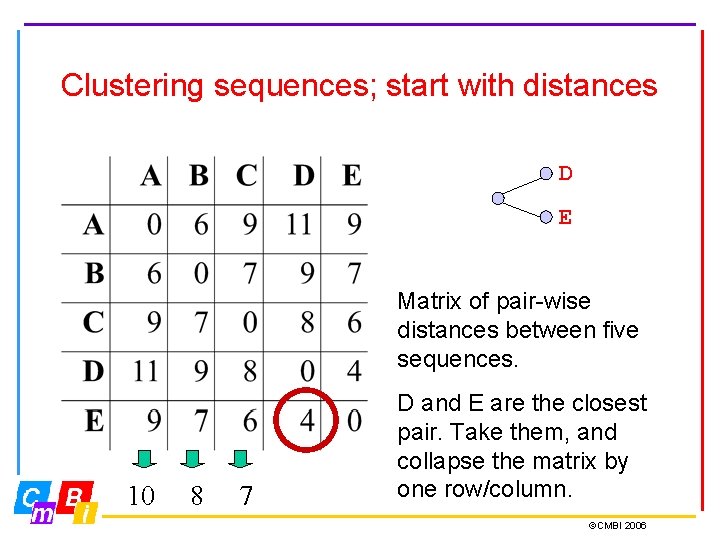
Clustering sequences; start with distances D E Matrix of pair-wise distances between five sequences. 10 8 7 D and E are the closest pair. Take them, and collapse the matrix by one row/column. ©CMBI 2006
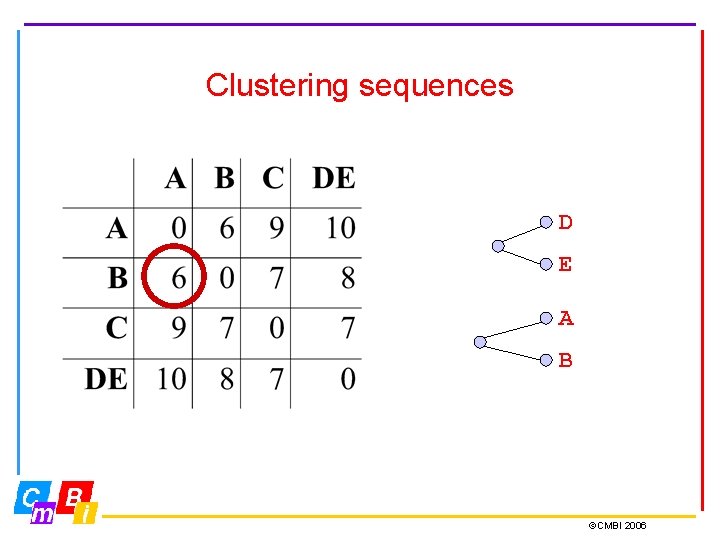
Clustering sequences D E A B ©CMBI 2006
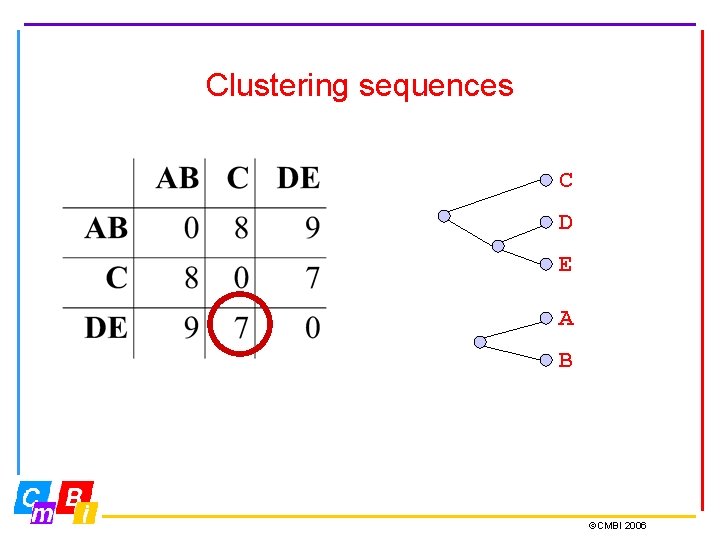
Clustering sequences C D E A B ©CMBI 2006
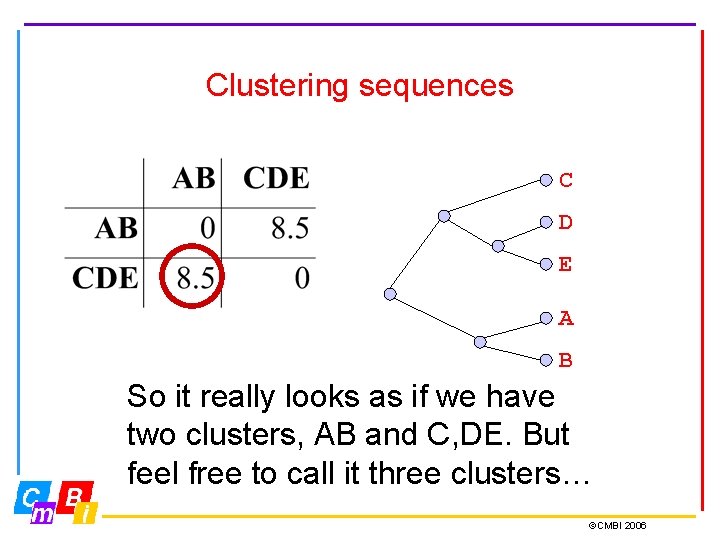
Clustering sequences C D E A B So it really looks as if we have two clusters, AB and C, DE. But feel free to call it three clusters… ©CMBI 2006

So, nice tree, but what did we actually do? 1)We determined a distance measure 2)We measured all pair-wise distances 3)We used an algorithm to visualize 4)We decided on the number of clusters And that, ladies and gentleman, is called clustering… ©CMBI 2006
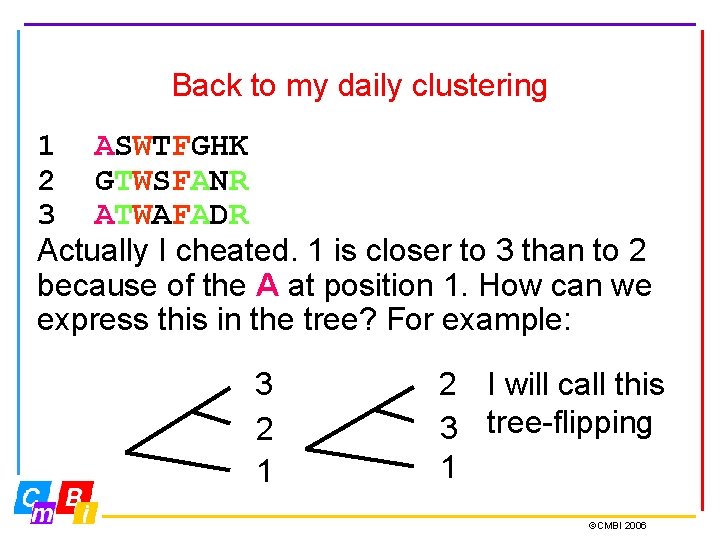
Back to my daily clustering 1 ASWTFGHK 2 GTWSFANR 3 ATWAFADR Actually I cheated. 1 is closer to 3 than to 2 because of the A at position 1. How can we express this in the tree? For example: 3 2 1 2 I will call this 3 tree-flipping 1 ©CMBI 2006

Can we generalize tree-flipping? To generalize tree flipping, sequences must be placed ‘distance-correct’ in 1 dimension: And then connect them, So, now most info as we did before: sits in the horizontal dimension. Can we use the vertical dimension usefully? 2 3 1 ©CMBI 2006
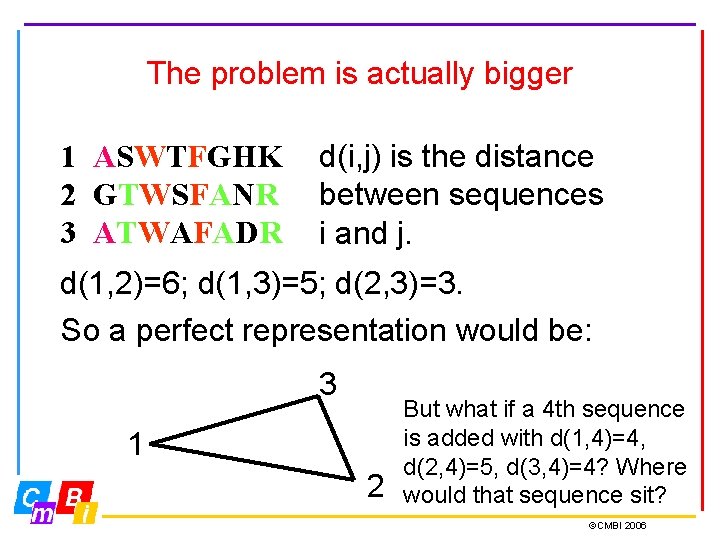
The problem is actually bigger 1 ASWTFGHK 2 GTWSFANR 3 ATWAFADR d(i, j) is the distance between sequences i and j. d(1, 2)=6; d(1, 3)=5; d(2, 3)=3. So a perfect representation would be: 3 1 2 But what if a 4 th sequence is added with d(1, 4)=4, d(2, 4)=5, d(3, 4)=4? Where would that sequence sit? ©CMBI 2006
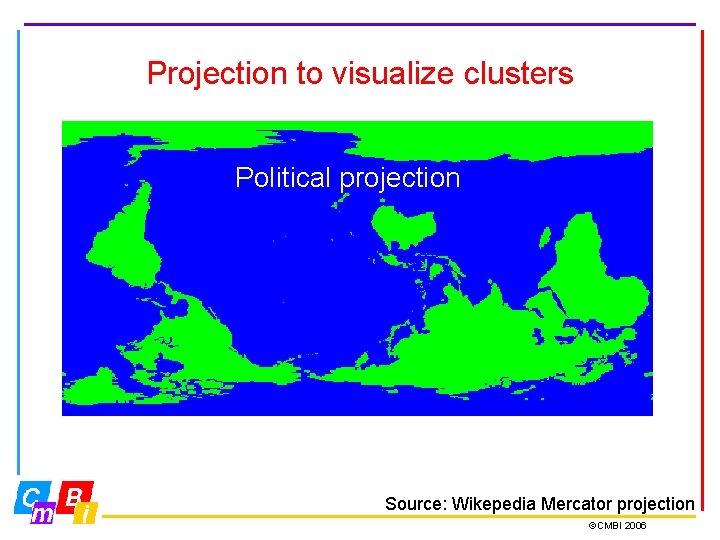
Projection to visualize clusters Gnomonic projection: Correct distances Fuller projection; Unfolded Dymaxion map Political projection Source: Wikepedia Mercator projection ©CMBI 2006
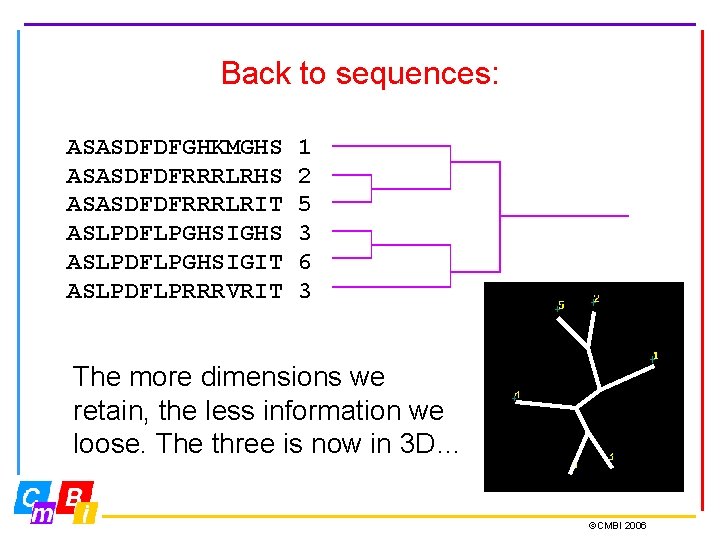
Back to sequences: ASASDFDFGHKMGHS ASASDFDFRRRLRIT ASLPDFLPGHSIGHS ASLPDFLPGHSIGIT ASLPDFLPRRRVRIT 1 2 5 3 6 3 The more dimensions we retain, the less information we loose. The three is now in 3 D… ©CMBI 2006

Projection to visualize clusters We want to reduce the dimensionality with minimal distortion of the pair-wise distances. One way is Eigenvector determination, or PCA. ©CMBI 2006

PCA to the rescue Now we have made the data one-dimensional, while the second, vertical, dimension is noise. If we did this correctly, we kept as much data as possible. ©CMBI 2006
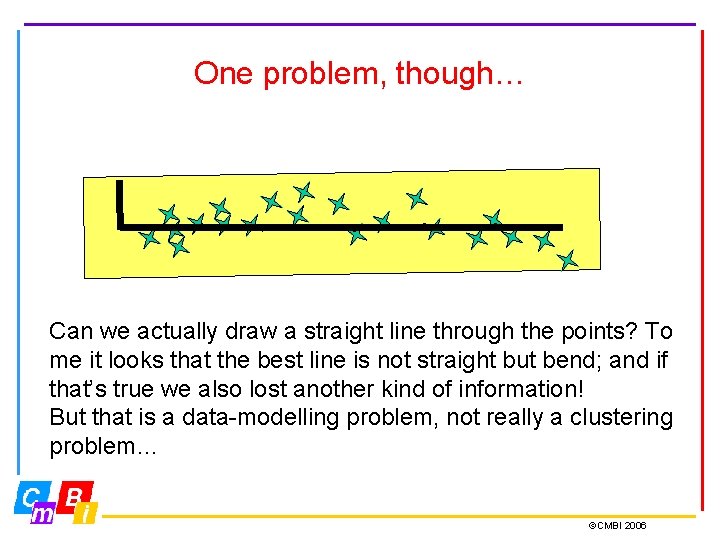
One problem, though… Can we actually draw a straight line through the points? To me it looks that the best line is not straight but bend; and if that’s true we also lost another kind of information! But that is a data-modelling problem, not really a clustering problem… ©CMBI 2006

Back to sequences: In we have N sequences, we can only draw their distance matrix in an N-1 dimensional space. By the time it is a tree, how many dimensions, and how much information have we lost? Perhaps we should cluster in a different way? ©CMBI 2006

Cluster on critical residues? QWERTYAKDFGRGH AWTRTYAKDFGRPM SWTRTNMKDTHRKC QWGRTNMKDTHRVW Gray = conserved Red = variable Green = correlated ©CMBI 2006

Cluster based on correlated residues ©CMBI 2006
Conclusion about sequence clustering Important topics: 1. Distance measure (~ data selection) 2. Data-modelling 3. Visualization 4. Dimensionality reduction ©CMBI 2006

Other sciences that cluster: Determination of a structure from massive averaging of very low resolution data. ©CMBI 2006
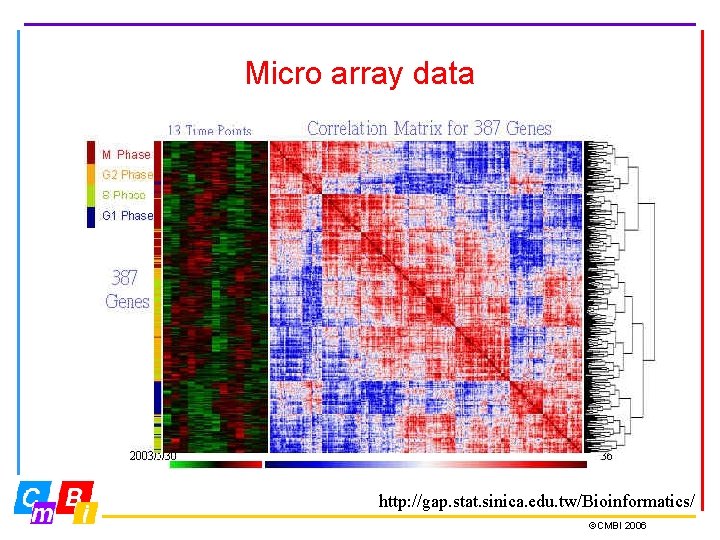
Micro array data http: //gap. stat. sinica. edu. tw/Bioinformatics/ ©CMBI 2006
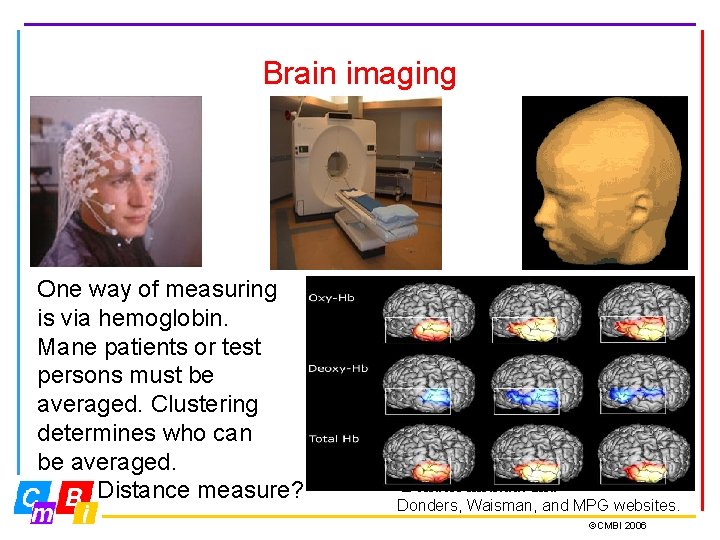
Brain imaging One way of measuring is via hemoglobin. Mane patients or test persons must be averaged. Clustering determines who can be averaged. Distance measure? Donders instituut and Donders, Waisman, and MPG websites. ©CMBI 2006

Molecular dynamics These actually are many superposed motions acting at the same time. Essential dynamics, eigenvector analysis in distance space, separates those motions. Bert de Groot ©CMBI 2006
Essential dynamics Motion along eigenvector 1 Daan van Aalten ©CMBI 2006

Domain knowledge is needed These examples make clear that domain knowledge is needed to cluster biological data. Everybody can cluster this But this one? ©CMBI 2006

Even informatics now knows: Michele Ceccarelli and Antonio Maratea (2006) Improving Fuzzy Clustering of Biological Data by International Journal on Approximate Reasoning : . Abstract Semi Supervised methods use a small amount of auxiliary information as a guide in the learning process in presence of unlabeled data. When using a clustering algorithm, the auxiliary information has the form of Side Information, that is a list of co-clustered points. Recent literature shows better performance of these methods with respect to totally unsupervised ones even with a small amount of Side Information. This fact suggests that the use of Semi Supervised methods may be useful especially in very difficult and noisy tasks where little a priori information is available, as is the case of data deriving from biological experiments. http: //www. scoda. unisannio. it/Papers/Publications/JAR_2006 ©CMBI 2006

Cluster techniques: K-means But how many clusters? Simply speaking k-means C clustering is an algorithm to D classify or to group your objects E based on attributes/features into A K number of group. K is positive B Elbow rule-of-thumb integer number. The grouping is done by minimizing the sum of squares of distances between data and the corresponding cluster centroid. Thus the purpose of K-mean clustering is to classify the data. We use TEAO…. ©CMBI 2006

My daily clustering and TEAO ©CMBI 2006

Entropy Sequence entropy Ei at position i is calculated from the frequency pi of the twenty amino acid types (p) at position i: 20 Ei = S i=1 pi ln(pi)

Variability Sequence variability Vi is the number of amino acid types observed at position i in more than 0. 5% of all sequences.
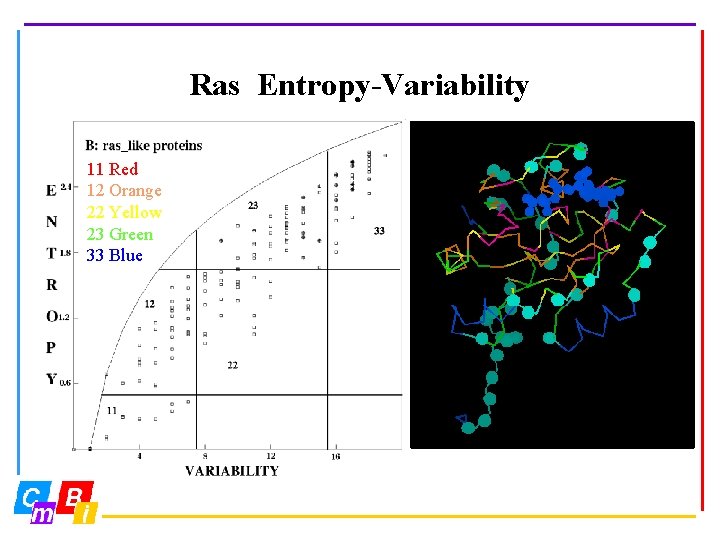
Ras Entropy-Variability 11 Red 12 Orange 22 Yellow 23 Green 33 Blue
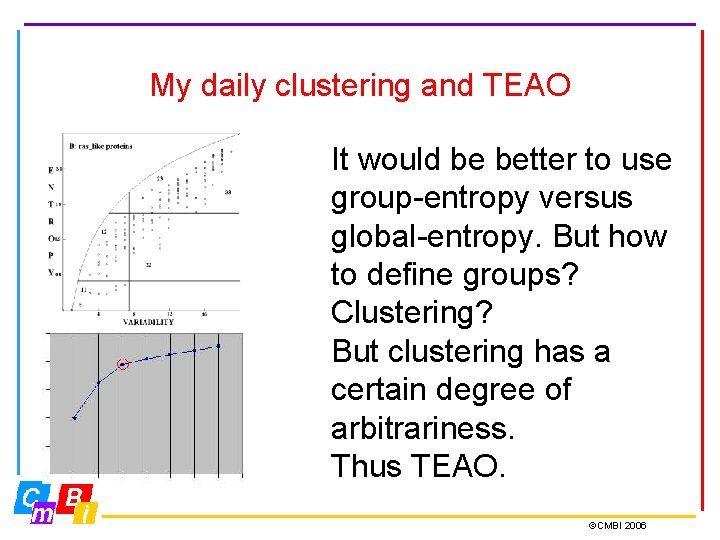
My daily clustering and TEAO It would be better to use group-entropy versus global-entropy. But how to define groups? Clustering? But clustering has a certain degree of arbitrariness. Thus TEAO. ©CMBI 2006

. ©CMBI 2006

. ©CMBI 2006

. ©CMBI 2006

. ©CMBI 2006

One problem, though… ©CMBI 2006
- Slides: 40