Introduction to Bioinformatics Sequence formats and databases in

Introduction to Bioinformatics, Sequence formats and databases in bioinformatics • Definitions/Basics • Sequence formats • Databases in Biology Dinesh Gupta Translational Bioinformatics ICGEB dinesh@icgeb. res. in http: //bioinfo. icgeb. res. in Image: http: //updates. engineeringwatch. in/cdac-to-host-symposium/

What is Bioinformatics? • Bioinformatics is the use of computers to solve biological and biomedical problems. • Bioinformatics is the application of information technology to mine, visualize, analyze, integrate, and manage biological and genetic information, which can then be applied in, among other things, accelerating drug discovery and development. • Biological Data management and analysis. • NIH definitions • Bioinformatics (http: //www. bisti. nih. gov/docs/compubiodef. pdf) Research, development, or application of computational tools and approaches for expanding the use of biological, medical, behavioral or health data, including those to acquire, store, organize, archive, analyze, or visualize such data. • Computational Biology: The development and application of dataanalytical and theoretical methods, mathematical modeling and computational simulation techniques to the study of biological, behavioral, and social systems.

Uses of Bioinformatics in modern biological research • DNA sequence analysis – Genome sequencing • • Sequence assembly Sequence/gene annotations Genefinding/Sequence translation tools Sequence Similarity searching (eg. BLAST, Clustal. W) • Comparison between genomes • Evolution of sequences (Phylogenetic analysis) • Gene expression

Use of Bioinformatics (. . contd. ) • Protein analysis – Structure • X-ray crystallography • Homology based models • Drug designing – Sequence • • • Sequence similarity Protein family assignments Conserved motifs Proteomics data analysis Protein Evolution

Uses of Bioinformatics (. . contd. ) • Other uses: – Vaccine development (eg. Reverse vaccinology) – Dairy technology – Forensics – Crop improvement – Designing enzymes for detergents – Genetic counseling
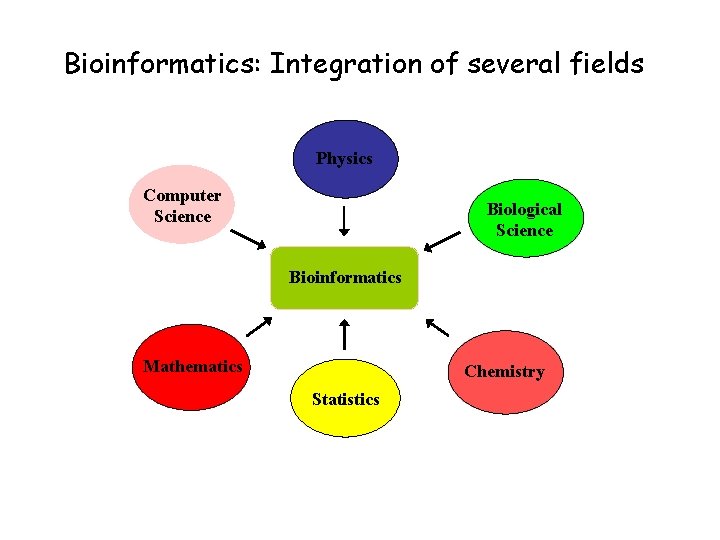
Bioinformatics: Integration of several fields Physics Computer Science Biological Science Bioinformatics Mathematics Chemistry Statistics

Recent events making bioinformatics more important • • Exponential expansion of biological information Expansion of multiple types of information Cheaper high throughput technologies Improvement in computation power (cost/capabilities) Lack of standards/quality Need for micro and macro analysis Need for better algorithms
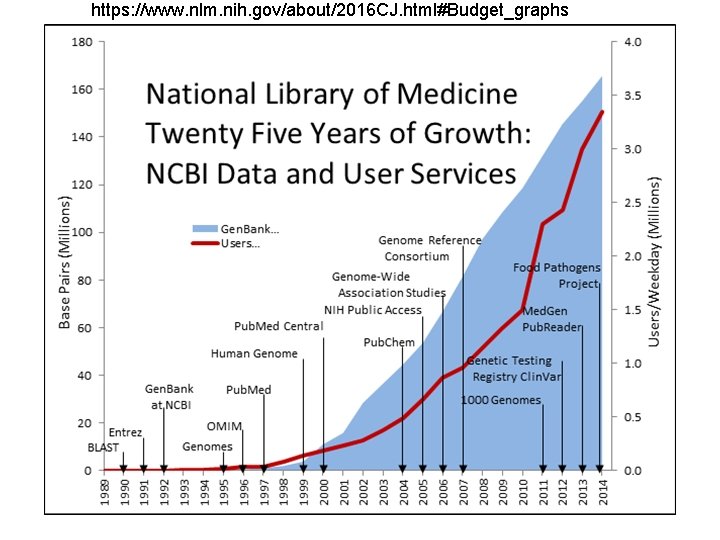
https: //www. nlm. nih. gov/about/2016 CJ. html#Budget_graphs
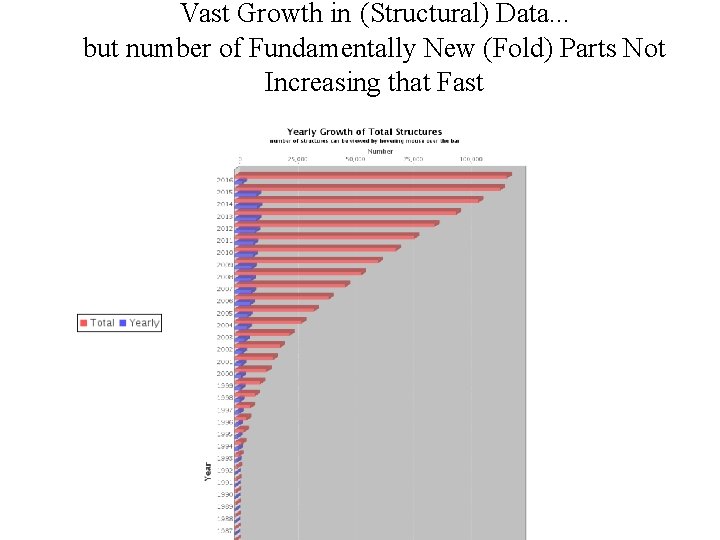
Vast Growth in (Structural) Data. . . but number of Fundamentally New (Fold) Parts Not Increasing that Fast

Bioinformatics Analysis? It is like any other lab analysis! • You need to know your data/input sources • You need to understand your methods and their assumptions • You need a plan to get from point A to point B • You need to understand your equipment • You need to be critical and understand potential sources of error • You need to interpret your results • Your results need to be reproducible • Your results should be testable

References, but not limited to: • • • http: //www. ncbi. nlm. nih. gov/About/primer/bioinformatics. html http: //icgeb. res. in/whotdr http: //en. wikipedia. org/wiki/Bioinformatics • http: //www. icgeb. res. in/cbioclass 2017/ • Baxevanis & Ouellette 2001. Bioinformatics: A Practical Guide to the Analysis of Genes and Proteins 2 nd Edition. John Wiley Publishing. • Gibas & Jambeck 2001. Developing Bioinformatics Computer Skills. O’Reilly. • Bioinformatics: Genome Sequence Analysis Mount 2001 • Bioinformatics For Dummies – Claverie & Notredame 2003 • Introduction to Bioinformatics – Lesk 2002

Sequence formats: Basics • Why different formats? – Type of information – Software requirements – Database requirements

Main file formats used in Bioinformatics • ASN. 1 • EMBL, Swiss Prot • FASTA • GCG • Gen. Bank/Gen. Pept • PHYLIP • PIR

ASN 1: Abstract Syntax Notation 1 used by NCBI Seq-entry : : = set { class phy-set , descr { pub { article { title { name "Cross-species infection of blood parasites between resident and migratory songbirds in Africa" } , authors { names std { { name { last "Waldenstroem" , first "Jonas" , initials "J. " } } , { name { last "Bensch" , first "Staffan" , initials "S. " } } , { name { last "Kiboi" , first "Sam" , initials "S. " } } , { name { last "Hasselquist" , first "Dennis" , initials "D. " } } , { name {

EMBL/Swiss Prot • The first line of each sequence entry is the ID definition line which contains entry name, dataclass, molecule, division and sequence length. • XX line contains no data, just a separator • The AC line lists the accession number. • DE line gives description about the sequence • FT precise annotation for the sequence • Sequence information SQ in the first two spaces. • The sequence information begins on the fifth line of the sequence entry. • The last line of each sequence entry in the file is a terminator line which has the two characters // in the first two spaces. ID XX AC XX DE DE DE RX RX XX FT FT FT SQ // AA 03518 standard; DNA; FUN; 237 BP. XX AC U 03518; Aspergillus awamori internal transcribed spacer 1 (ITS 1) and 18 S r. RNA and 5. 8 S r. RNA genes, partial sequence. MEDLINE; 94303342. PUBMED; 8030378. r. RNA <1. . 20 /product="18 S ribosomal RNA" misc_RNA 21. . 205 /standard_name="Internal transcribed spacer 1 (ITS 1)" r. RNA 206. . >237 /product="5. 8 S ribosomal RNA" Sequence 237 BP; 41 A; 77 C; 67 G; 52 T; 0 other; aacctgcgga aggatcatta ccgagtgcgg gtcctttggg cccaacctcc catccgtgtc 60 tattgtaccc tgttgcttcg gcgggcccgc cgcttgtcgg ccgccggggg ggcgcctctg 120 ccccccgggc ccgtgcccgc cggagacccc aacacgaaca ctgtctgaaa gcgtgcagtc 180 tgagttgatt gaatgcaatc agttaaaact ttcaacaatg gatctcttgg ttccggc 237

FASTA • A sequence in Fasta format begins with a single-line description, • followed by lines of sequence data. • The description line is distinguished from the sequence data by a greaterthan (">") symbol in the first column. • It is recommended that all lines of text be shorter than 80 characters in length. >U 03518 Aspergillus awamori internal transcribed spacer 1 (ITS 1) AACCTGCGGAAGGATCATTACCGAGTGCGGGTCCTTTGGGCCCAACCTCCCATCCGTGTCTATTGTACCC TGTTGCTTCGGCGGGCCCGCCGCTTGTCGGCCGCCGGGGGGGCGCCTCTGCCCCCCGGGCCCGTGCCCGC CGGAGACCCCAACACGAACACTGTCTGAAAGCGTGCAGTCTGAGTTGAATGCAATCAGTTAAAACT TTCAACAATGGATCTCTTGGTTCCGGC

GCG • Exactly one sequence • Begins with annotation lines • Start of the sequence is marked by a line ending with ". . “ • This line also contains the sequence identifier, the sequence length and a checksum ID XX AC XX DE DE XX AA 03518 standard; DNA; FUN; 237 BP. U 03518; Aspergillus awamori internal transcribed spacer 1 (ITS 1) and 18 S r. RNA and 5. 8 S r. RNA genes, partial sequence. SQ Sequence 237 BP; 41 A; 77 C; 67 G; 52 T; 0 other; AA 03518 Length: 237 Check: 4514 . . 1 61 121 181 aacctgcgga tattgtaccc ccccccgggc tgagttgatt aggatcatta tgttgcttcg ccgtgcccgc gaatgcaatc ccgagtgcgggcccgc cggagacccc agttaaaact gtcctttggg cgcttgtcgg aacacgaaca ttcaacaatg cccaacctcc ccgccggggg ctgtctgaaa gatctcttgg catccgtgtc ggcgcctctg gcgtgcagtc ttccggc

Gen. Bank/Gen. Pept The nucleotide (Gen. Bank) and protein (Gen Pept) database entries are available from Entrez in this format • Can contain several sequences • One sequence starts with: “LOCUS” • The sequence starts with: "ORIGIN“ • The sequence ends with: "//“ LOCUS AAU 03518 237 bp DNA PLN 04 -FEB-1995 DEFINITION Aspergillus awamori internal transcribed spacer 18 S r. RNA and 5. 8 S r. RNA genes, partial sequence. ACCESSION U 03518 BASE COUNT 41 a 77 c 67 g 52 t ORIGIN 1 aacctgcgga aggatcatta ccgagtgcgg gtcctttggg cccaacctcc 61 tattgtaccc tgttgcttcg gcgggcccgc cgcttgtcgg ccgccggggg 121 ccccccgggc ccgtgcccgc cggagacccc aacacgaaca ctgtctgaaa 181 tgagttgatt gaatgcaatc agttaaaact ttcaacaatg gatctcttgg // 1 (ITS 1) and catccgtgtc ggcgcctctg gcgtgcagtc ttccggc

Phylip format 2 2000 G 019 uabh ATACATCATA ACACTACTTC CTACCCATAA GCTCCTTTTA ACTTGTTAAA G 028 uaah CATAAGCTCC TTTTAACTTG TTAAAGTCTT GCTTGAATTA AAGACTTGTT GTCTTG AATTAAAGAC TTGTTTAAAC ACAAAAATTT AGAGTTTTAC TAAACACAAA ATTTAGACTT TTACTCAACA AAAGTGATTGAT TCAACAAAAG TGATTGAT TGATGG TTTACAGTAG TGATTG ATGGTTTACA GTAGGACTTC ATTCTAGTCA TTATAGCTGC • The first line of the input file contains the number of sequences and their length (all should have the same length) separated by blanks. • The next line contains a sequence name, next lines are the sequence itself in blocks of 10 characters. Then follow rest of sequences.

Other formats MEGA • • • • • #mega Title: infile. fasta #G 019 uabh ATACATCATAACACTACTTCCTACCCATAAGCTCCTTTTAACTTGTTAAAGTCTTG AATTAAAGACTTGTTTAAACACAAAAATTTAGAGTTTTACTCAACAAAAGTGATTGATTGATGGTTTACAGTAGGACTTCATTCTAGTCATTATAGCTGCTGGC AGTATAACTGGCCAGCCTTTAATACATTGCTGCTTAGAGTCAAAGCATGTACTTAGAGTT GGTATGATTTATCTTTTTGGTCTTCTATAGCCTCCTTCCCCATCAGTCTTAATC AGTCTTGTTACGTTATGACTAATCTTTGGGGATTGTGCAGAATGTTATTTTAGATAAGCA AAACGAGCAAAATGGGGAGTTACTTATATTTCTTTAAAGC #G 028 uaah CATAAGCTCCTTTTAACTTGTTAAAGTCTTGAATTAAAGACTTGTTTAAACACAAA ATTTAGACTTTTACTCAACAAAAGTGATTGATTGATTGATGGTTTACA GTAGGACTTCATTCTAGTCATTATAGCTGCTGGCAGTATAACTGGCCAGCCTTTAATACA TTGCTGCTTAGAGTCAAAGCATGTACTTAGAGTTGGTATGATTTATCTTTTTGGTCTTCT ATAGCCTCCTTCCCCATCAGTCT

FASTQ @SEQ_ID GATTTGGGGTTCAAAGCAGTATCGATCAAATAGTAAATCCATTTGTTCAACTCACA T + !''*((((***+))%%%++)(%%%%). 1***-+*''))**55 CCF>>>>>>CCCCCCC 65

Read. Seq Don Gilbert software@bio. indiana. edu, May 2001 Indiana University, Bloomington, Indiana WWW http: //www. ebi. ac. uk/Tools/sfc/emboss_seqret/ http: //www-bimas. cit. nih. gov/molbio/readseq/
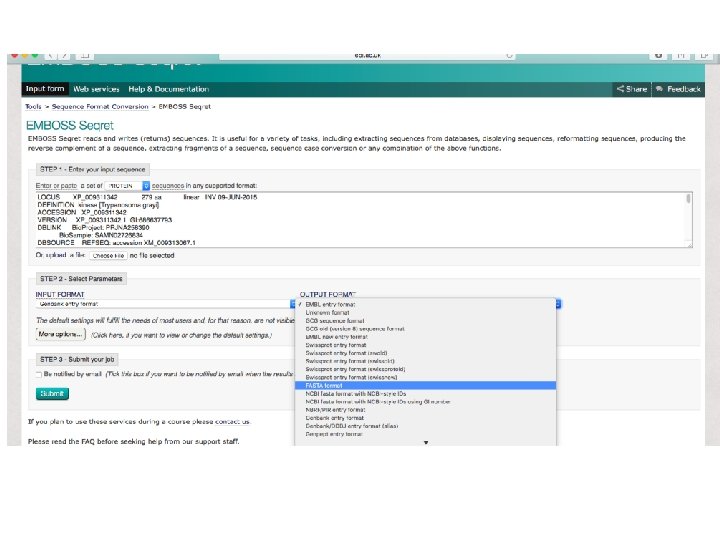
http: //www. ebi. ac. uk/Tools/sfc/emboss_seq ret/
![LOCUS XP_009311342 279 aa linear INV 09 -JUN-2015 DEFINITION kinase [Trypanosoma grayi]. ACCESSION XP_009311342 LOCUS XP_009311342 279 aa linear INV 09 -JUN-2015 DEFINITION kinase [Trypanosoma grayi]. ACCESSION XP_009311342](http://slidetodoc.com/presentation_image/311a4b4303d8053df770d5730abfda3d/image-24.jpg)
LOCUS XP_009311342 279 aa linear INV 09 -JUN-2015 DEFINITION kinase [Trypanosoma grayi]. ACCESSION XP_009311342 VERSION XP_009311342. 1 GI: 686637793 DBLINK Bio. Project: PRJNA 258390 Bio. Sample: SAMN 02726834 DBSOURCE REFSEQ: accession XM_009313067. 1 KEYWORDS Ref. Seq. SOURCE Trypanosoma grayi ORGANISM Trypanosoma grayi Eukaryota; Euglenozoa; Kinetoplastida; Trypanosomatidae; Trypanosoma. REFERENCE 1 (residues 1 to 279) AUTHORS Kelly, S. , Ivens, A. C. , Manna, P. T. , Gibson, W. and Field, M. C. TITLE A draft genome for the African crocodilian trypanosome Trypanosoma grayi JOURNAL Unpublished REFERENCE 2 (residues 1 to 279) CONSRTM NCBI Genome Project TITLE Direct Submission JOURNAL Submitted (09 -JUN-2015) National Center for Biotechnology Information, NIH, Bethesda, MD 20894, USA REFERENCE 3 (residues 1 to 279) AUTHORS Kelly, S. , Ivens, A. C. , Manna, P. T. , Gibson, W. and Field, M. C. TITLE Direct Submission JOURNAL Submitted (09 -JUN-2014) University of Edinburgh, Kings Buildings, Mayfield Rd, Edinburgh EH 9 3 JT, United Kingdom REFERENCE 4 (residues 1 to 279) AUTHORS Kelly, S. , Ivens, A. C. , Manna, P. T. , Gibson, W. and Field, M. C. TITLE Direct Submission JOURNAL Submitted (12 -MAY-2014) University of Edinburgh, Kings Buildings, Mayfield Rd, Edinburgh EH 9 3 JT, United Kingdom COMMENT PROVISIONAL REFSEQ: This record has not yet been subject to final NCBI review. The reference sequence is identical to KEG 10437. Method: conceptual translation. FEATURES Location/Qualifiers source 1. . 279 /organism="Trypanosoma grayi" /strain="ANR 4"
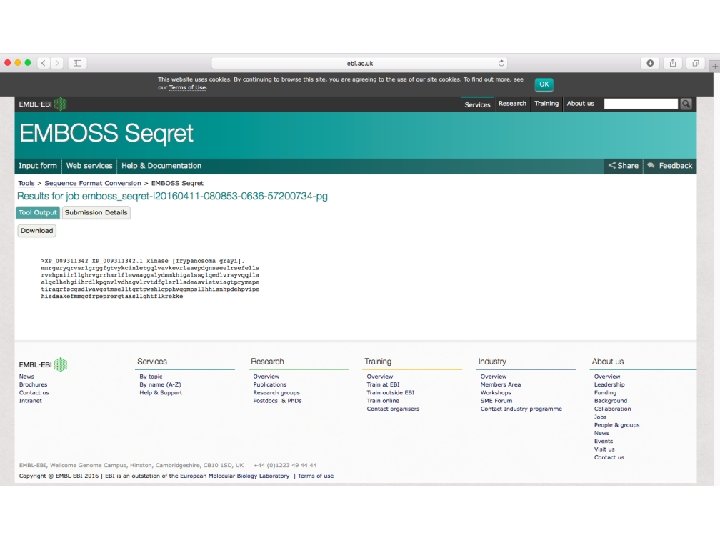

The Readseq package can read most common formats: examples of all these formats are included in the readseq directory. The formats include: • • • • IG/Stanford, used by Intelligenetics and others Gen. Bank/GB, genbank flatfile format NBRF format (SAM modifications cause this to break when sequences do not have a terminating asterix) EMBL, EMBL flatfile format GCG, single sequence format of GCG software DNAStrider, for common Mac program Fitch format, limited use Pearson/Fasta, a common format used by Fasta programs and others Zuker format, limited use. Input only. Olsen, format printed by Olsen VMS sequence editor. Input only. Phylip 3. 2, sequential format for Phylip programs Plain/Raw, sequence data only (no name, document, numbering) MSF multi sequence format used by GCG software PAUP's multiple sequence (NEXUS) format PIR/CODATA format used by PIR

Databases in Biology
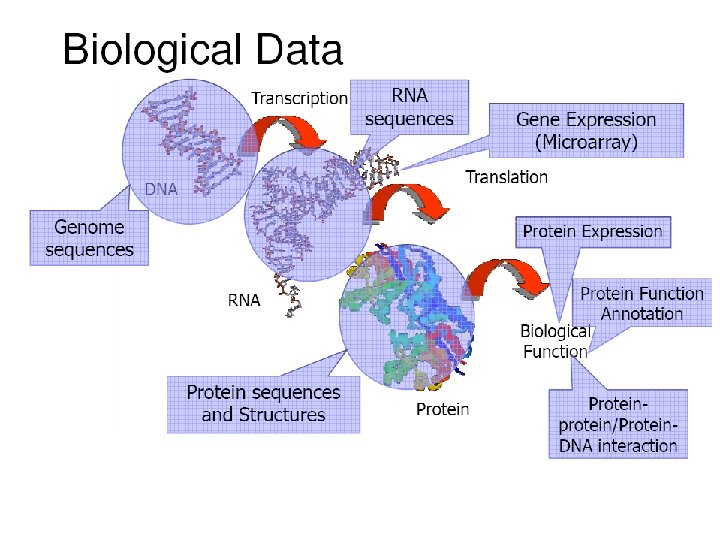

Need for databases in Biology? • Need for storing and communicating large datasets has grown. • Need to disseminate biological information. • Provide Organized data for analysis friendly retrieval. • Need to make biological data available in computerreadable form.

Different classifications of databases • Type of data – nucleotide sequences – proteins sequence patterns or motifs – macromolecular 3 D structure – gene expression data – metabolic pathways – proteomics data

Big Data • Big data is a broad term for data sets so large or complex that traditional data processing applications are inadequate. • Analysis challenges include capture, analysis, data curation, search, sharing, storage, transfer, visualization, and information privacy. • Help answer questions that were previously unanswered or considered impossible.
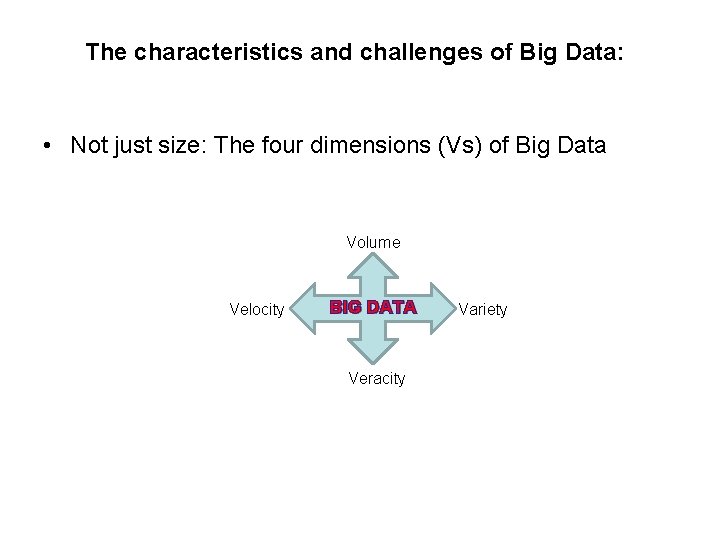
The characteristics and challenges of Big Data: • Not just size: The four dimensions (Vs) of Big Data Volume Velocity BIG DATA Veracity Variety

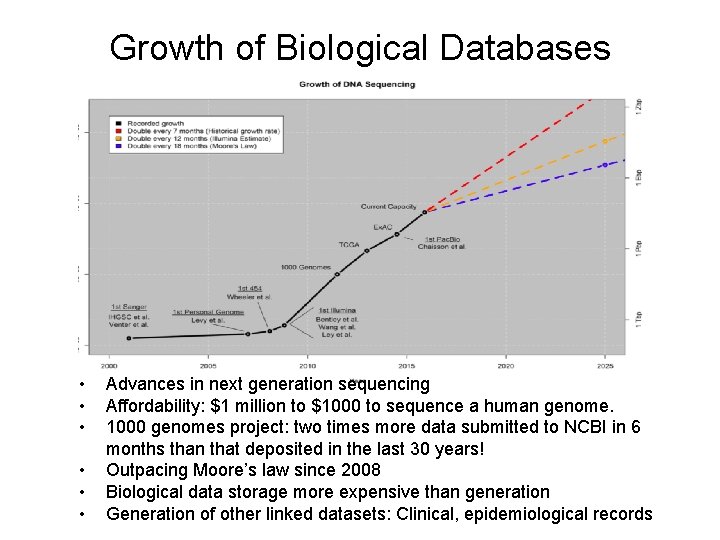
Growth of Biological Databases • • • Advances in next generation sequencing Affordability: $1 million to $1000 to sequence a human genome. 1000 genomes project: two times more data submitted to NCBI in 6 months than that deposited in the last 30 years! Outpacing Moore’s law since 2008 Biological data storage more expensive than generation Generation of other linked datasets: Clinical, epidemiological records

Different classifications of databases…. • Primary or derived databases – Primary databases: experimental results directly into database – Secondary databases: results of analysis of primary databases – Aggregate of many databases • Links to other data items • Combination of data • Consolidation of data

Different classifications of databases…. • Technical design – Flat-files – Relational database (SQL) – Exchange/publication technologies (HTML, CORBA, XML, . . . ) • Each one of the above are inter convertible

Different classifications of databases…. • Availability – Publicly available, no restrictions – Available, but with copyright – Accessible, but not downloadable – Academic, but not freely available – Proprietary, commercial; possibly free for academics

Different classifications of databases…. • Content – Protein/DNA/RNA/mi. RNA etc. – Family: kinases – Common physical properties: membrane bound, mitochondrial proteins – Common chemical properties: Proteases, reductases etc. – Sequences of a particular genome/species: e. g. Influenza sequences, plasmodium sequences etc. – Motifs/domains

Where to look for databases? • Search Engines • Journals related to Bioinformatics • Websites like: – Wikipedia https: //en. wikipedia. org/wiki/List_of_biological _databases – www. expasy. ch – Several others websites


NAR DB issue 2015 • • 54 (62 in prev. yr. ) new dbs since last year! Total 1685 databases (http: //www. oxfordjournals. org/nar/database/a/ Complete list – 15 categories and 41 sub-categories – Searchable – http: //www. oxfordjournals. org/nar/database/a/ (html format), also as downloadable word file)
http: //www 3. oup. co. uk/nar/database/c/

Database searching tips • • • Look for links to Help or Examples Always check updates Level of curation Try Boolean searches Be careful with UK/US spelling differences – leukaemia vs leukemia – haemoglobin vs hemoglobin – colour vs color

Exercises • Retrieve sequences from sequence databases • Convert sequence formats • Study different formats and flow of information • Study a database from NAR collection and write one page report about the database, with critical evaluation. Submit by 30 th January 2017.
- Slides: 45