Introduction to Bioinformatics Ch Bi 406506 Ozlem Keskin

Introduction to Bioinformatics Ch. Bi 406/506 Ozlem Keskin For today’s lectures Many slides from gersteinlab. org/courses/452 And Bioinformatics and Functional Genomics by Jonathan Pevsner (ISBN 0 -471 -21004 -8).

EMails Ozlem Keskin okeskin@ku. edu. tr Engin Cukuroglu ecukuroglu@ku. edu. tr

Who is taking this course? • People with very diverse backgrounds in biology, chemical engineering (MS/BS) • People with diverse backgrounds in computer science -please visit Attila Hoca’s office! • Most people have a favorite gene, protein, or disease

What are the goals of the course? • To provide an introduction to bioinformatics with a focus on the National Center for Biotechnology Information (NCBI) and EBI • To focus on the analysis of DNA, RNA and proteins • To introduce you to the analysis of genomes • To combine theory and practice to help you solve research problems

Themes throughout the course Textbooks Web sites Literature references Gene/protein families Computer labs

Textbook The course textbook is J. Pevsner, Bioinformatics and Functional Genomics (Wiley, 2009). Several other bioinformatics texts are available: Baxevanis and Ouellette Mount Durbin et al. Lesk In our library you will find (e-book) Bioinformatics [electronic resource] : sequence and genome analysis / David W. Mount. Imprint. Cold Spring Harbor, N. Y. : Cold Spring Harbor Laboratory Press, c 2001. Bioinformatics : a practical guide to the analysis of genes and proteins / editedby Andreas D. Baxevanis, B. F. Francis Ouellette. Imprint. Hoboken, N. J. : John Wiley, 2005. (SOON)

Themes throughout the course: Literature references You are encouraged to read original source articles. Although articles are not required, they will enhance your understanding of the material. You can obtain articles through Pub. Med and Web of Science.

Web sites The course website is reached via: http: //pevsnerlab. kennedykrieger. org/bioinfo_course. htm (or Google “pevsnerlab” courses) This site contains the powerpoints for each lecture. The textbook website is: http: //www. bioinfbook. org This has 1000 URLs, organized by chapter This site also contains the same powerpoints. You will also find the lecture slides at F-folder.

Grading Midterm Final HWs Project 30% 35% 20% 15% (might change, the course will evolve)

Themes throughout the course: gene/protein families We will use beta globin and retinol-binding protein 4 (RBP 4) as model genes/proteins throughout the course. Globins including hemoglobin and myoglobin carry oxygen. RBP 4 is a member of the lipocalin family. It is a small, abundant carrier protein. We will study globins and lipocalins in a variety of contexts including • --sequence alignment • --gene expression • --protein structure • --phylogeny • --homologs in various species
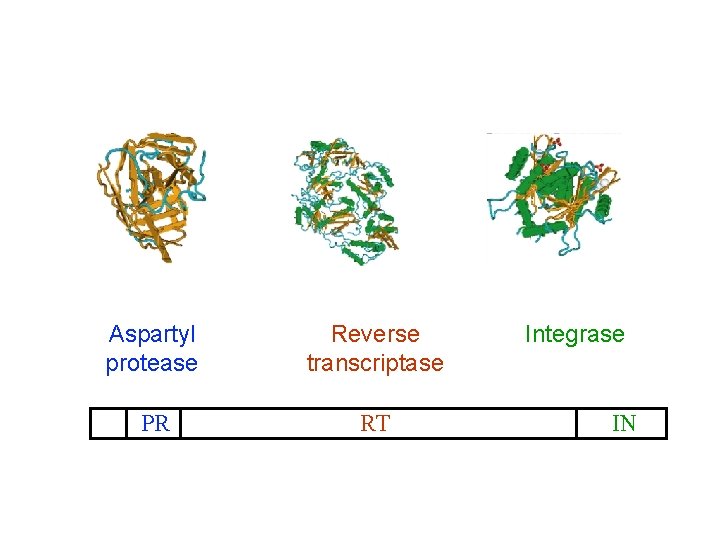
The HIV-1 pol gene encodes three proteins Aspartyl protease Reverse transcriptase PR RT Integrase IN

Outline for today (chapters 1 and 2) Definition of bioinformatics Overview of the NCBI website Accessing information about DNA and proteins --Definition of an accession number --Four ways to find information on proteins and DNA Access to biomedical literature

Bioinformatics Biological Data + Computer Calculations

What is bioinformatics? • Interface of biology and computers • Analysis of proteins, genes and genomes using computer algorithms and computer databases • Genomics is the analysis of genomes. The tools of bioinformatics are used to make sense of the billions of base pairs of DNA that are sequenced by genomics projects. Protein coordinates, DNA array data, annotated gene sequences Biological information is being generated now days in parallel. We can easily run 10, 000 simultaneous experiments on a single DNA microarray. To cope with this much data we really need computers. So Bioinformatics is that field that combines biology and computers.
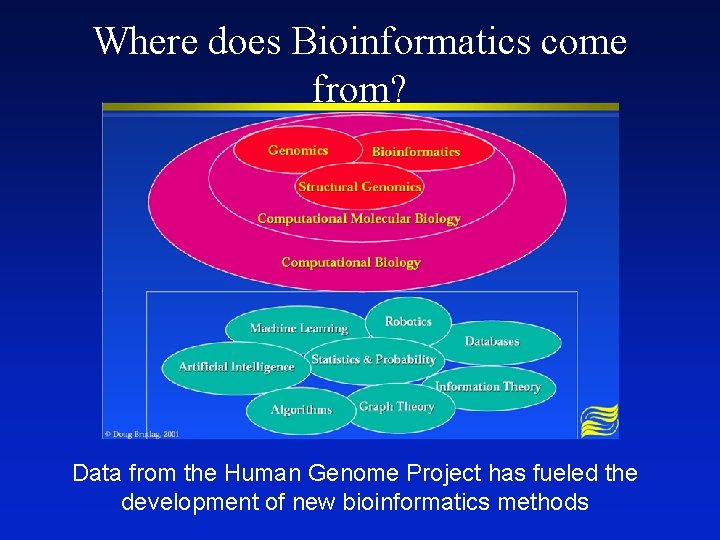
Where does Bioinformatics come from? Data from the Human Genome Project has fueled the development of new bioinformatics methods

HGP

What is Bioinformatics? • (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale.

• Interface of biology and computers Analysis of proteins, genes and genomes using computer algorithms and computer databases Genomics is the analysis of genomes. The tools of bioinformatics are used to make sense of the billions of base pairs of DNA that are sequenced by genomics projects. •
![Top ten challenges for bioinformatics [1] Precise models of where and when transcription will Top ten challenges for bioinformatics [1] Precise models of where and when transcription will](http://slidetodoc.com/presentation_image_h/1e0498e9e5167c694810cacb0984c848/image-20.jpg)
Top ten challenges for bioinformatics [1] Precise models of where and when transcription will occur in a genome (initiation and termination) [2] Precise, predictive models of alternative RNA splicing [3] Precise models of signal transduction pathways; ability to predict cellular responses to external stimuli [4] Determining protein: DNA, protein: RNA, protein: protein recognition codes [5] Accurate ab initio protein structure prediction
![Top ten challenges for bioinformatics [6] Rational design of small molecule inhibitors of proteins Top ten challenges for bioinformatics [6] Rational design of small molecule inhibitors of proteins](http://slidetodoc.com/presentation_image_h/1e0498e9e5167c694810cacb0984c848/image-21.jpg)
Top ten challenges for bioinformatics [6] Rational design of small molecule inhibitors of proteins [7] Mechanistic understanding of protein evolution [8] Mechanistic understanding of speciation [9] Development of effective gene ontologies: systematic ways to describe gene and protein function [10] Education: development of bioinformatics curricula Source: Ewan Birney, Chris Burge, Jim Fickett
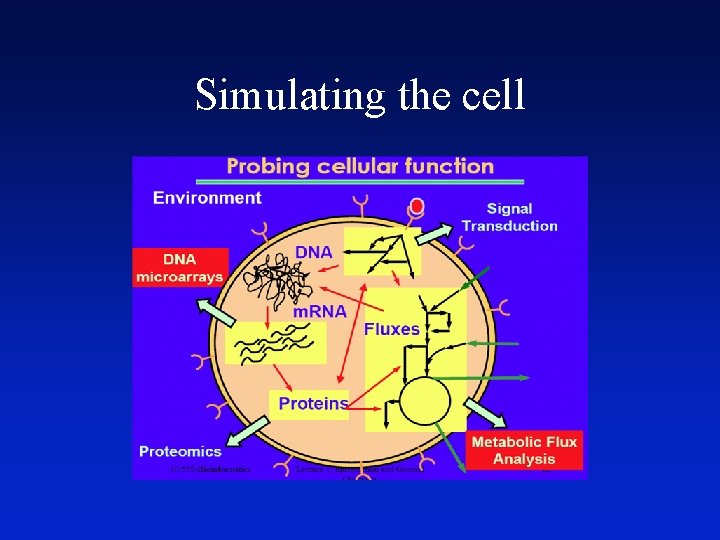
Simulating the cell

On bioinformatics “Science is about building causal relations between natural phenomena (for instance, between a mutation in a gene and a disease). The development of instruments to increase our capacity to observe natural phenomena has, therefore, played a crucial role in the development of science - the microscope being the paradigmatic example in biology. With the human genome, the natural world takes an unprecedented turn: it is better described as a sequence of symbols. Besides high-throughput machines such as sequencers and DNA chip readers, the computer and the associated software becomes the instrument to observe it, and the discipline of bioinformatics flourishes.

On bioinformatics However, as the separation between us (the observers) and the phenomena observed increases (from organism to cell to genome, for instance), instruments may capture phenomena only indirectly, through the footprints they leave. Instruments therefore need to be calibrated: the distance between the reality and the observation (through the instrument) needs to be accounted for. This issue of Genome Biology is about calibrating instruments to observe gene sequences; more specifically, computer programs to identify human genes in the sequence of the human genome. ” Martin Reese and Roderic Guigó, Genome Biology 2006 7(Suppl I): S 1, introducing EGASP, the Encyclopedia of DNA Elements (ENCODE) Genome Annotation Assessment Project
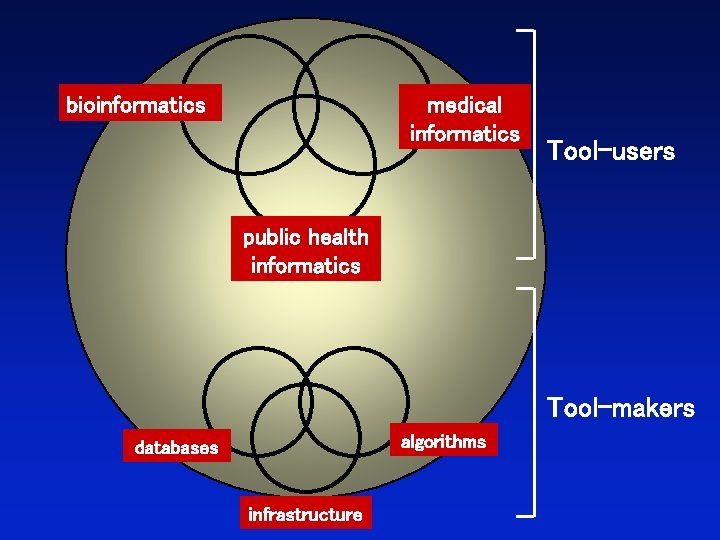
bioinformatics medical informatics Tool-users public health informatics Tool-makers algorithms databases infrastructure

Three perspectives on bioinformatics The cell The organism The tree of life Page 4
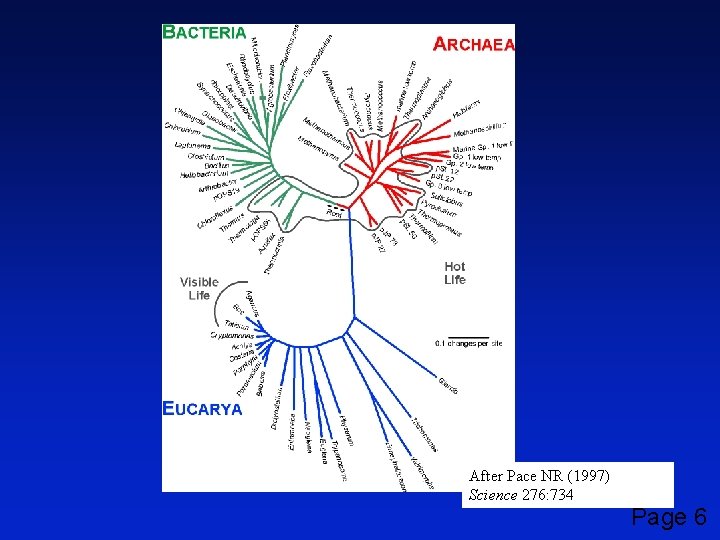
After Pace NR (1997) Science 276: 734 Page 6
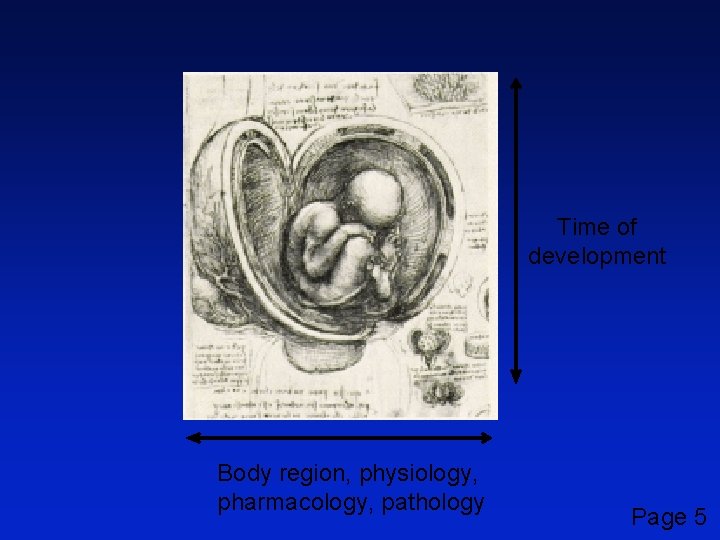
Time of development Body region, physiology, pharmacology, pathology Page 5
DNA RNA protein phenotype Page 5
DNA RNA protein phenotype
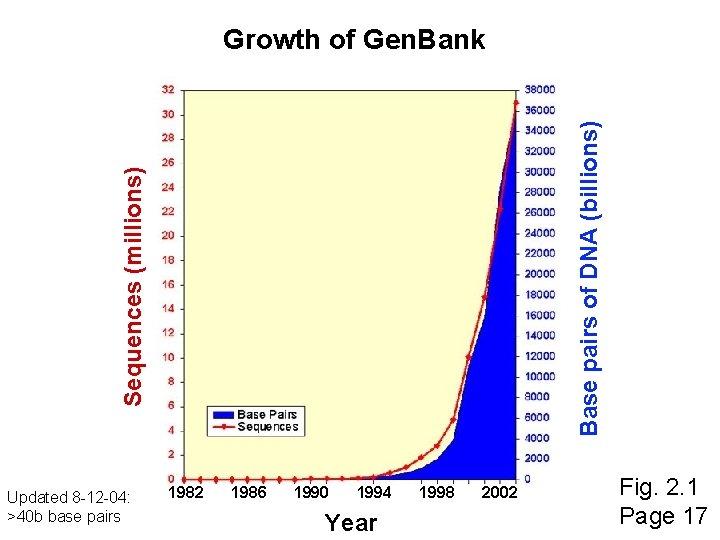
Sequences (millions) Base pairs of DNA (billions) Growth of Gen. Bank Updated 8 -12 -04: >40 b base pairs 1982 1986 1990 1994 Year 1998 2002 Fig. 2. 1 Page 17
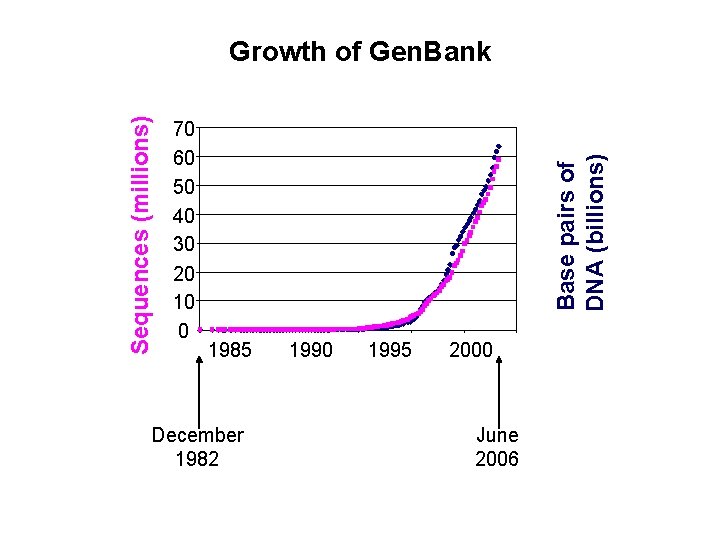
70 60 50 40 30 20 10 0 Base pairs of DNA (billions) Sequences (millions) Growth of Gen. Bank 1985 December 1982 1990 1995 2000 June 2006
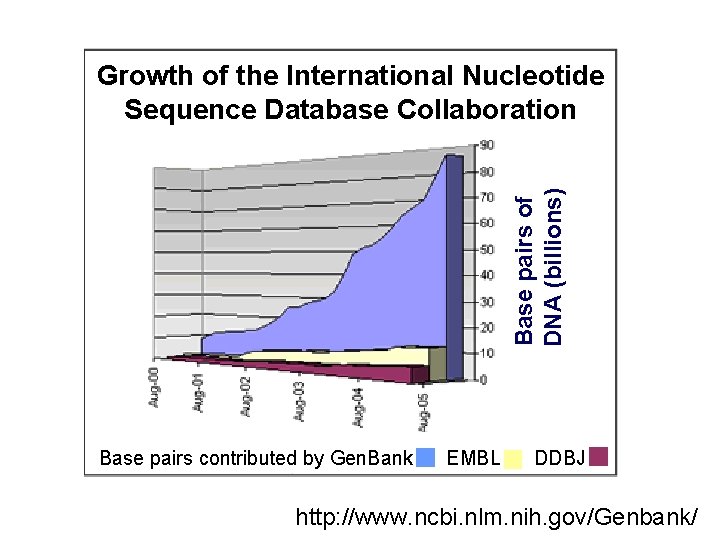
Base pairs of DNA (billions) Growth of the International Nucleotide Sequence Database Collaboration Base pairs contributed by Gen. Bank EMBL DDBJ http: //www. ncbi. nlm. nih. gov/Genbank/
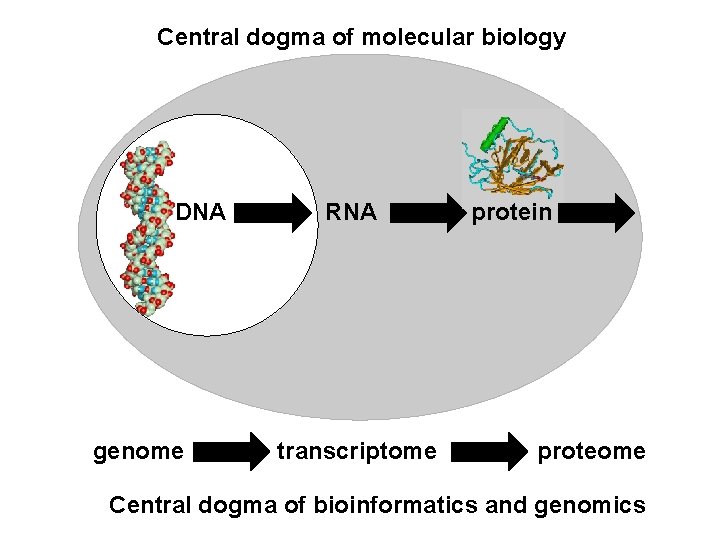
Central dogma of molecular biology DNA genome RNA transcriptome protein proteome Central dogma of bioinformatics and genomics

What is the Information? Molecular Biology as an Information Science • Central Dogma of Molecular Biology DNA -> RNA -> Protein -> Phenotype -> DNA • Molecules – Sequence, Structure, Function • Processes – Mechanism, Specificity, Regulation • Genetic material • Information transfer (m. RNA) • Protein synthesis (t. RNA/m. RNA) • Some catalytic activity • Central Paradigm for Bioinformatics Genomic Sequence Information -> m. RNA (level) -> Protein Sequence -> Protein Structure -> Protein Function -> Phenotype • Large Amounts of Information – Standardized – Statistical • Most cellular functions are performed or facilitated by proteins. • Primary biocatalyst • Cofactor transport/storage • Mechanical motion/support • Immune protection • Control of growth/differentiation (idea from D Brutlag, Stanford, graphics from S Strobel)

• • • Proteins fold into 3 D structures with specific functions which are reflected in a pheonotype. These functions are selected in a Darwinian sense by the environment of the phenotype. Which drives the evolution of the DNA sequence. • Many Bioinformatics techniques address this flow of molecular biology information inside the organism hoping to understand the organization and control of genes even predicting protein structure from sequence. • There is a second flow of information that bioinformatics seeks to address is the large amount of data generated by new high through methods. Bioinformatics owes its lively hood to the availability of large data sets that are too complex to allow manual analysis.
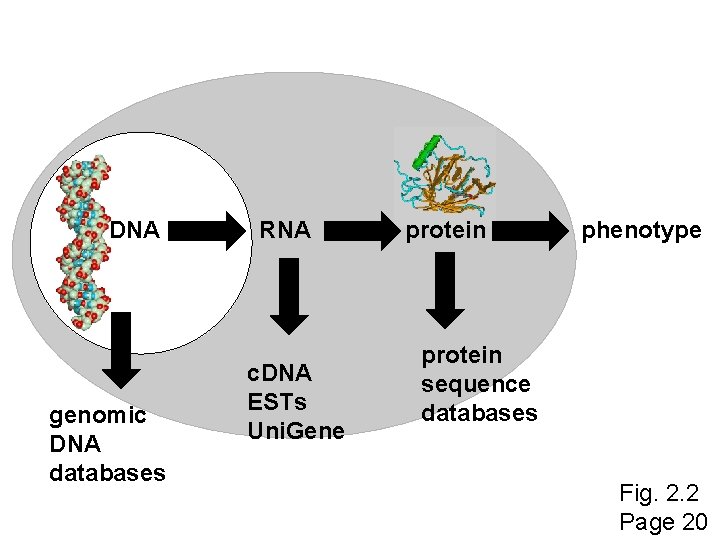
DNA genomic DNA databases RNA c. DNA ESTs Uni. Gene protein phenotype protein sequence databases Fig. 2. 2 Page 20
There are three major public DNA databases EMBL Gen. Bank DDBJ The underlying raw DNA sequences are identical Page 16
There are three major public DNA databases EMBL Housed at EBI European Bioinformatics Institute Gen. Bank DDBJ Housed at NCBI National Center for Biotechnology Information Housed in Japan Page 16

>100, 000 species are represented in Gen. Bank all species 128, 941 viruses 6, 137 bacteria 31, 262 archaea 2, 100 eukaryota 87, 147 Table 2 -1 Page 17

The most sequenced organisms in Gen. Bank Homo sapiens (6. 9 million entries) Mus musculus (5. 0 million) Zea mays (896, 000) Rattus norvegicus (819, 000) Gallus gallus (567, 000) Arabidopsis thaliana (519, 000) Danio rerio (492, 000) Drosophila melanogaster (350, 000) Oryza sativa (221, 000)

National Center for Biotechnology Information (NCBI) www. ncbi. nlm. nih. gov
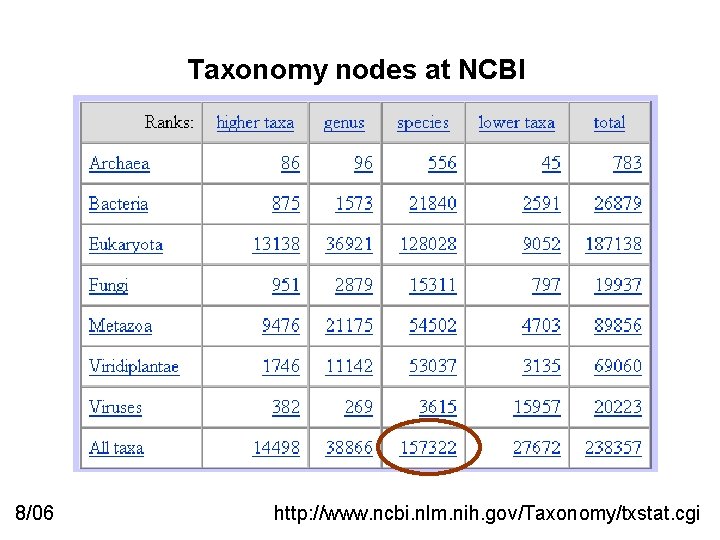
Taxonomy nodes at NCBI 8/06 http: //www. ncbi. nlm. nih. gov/Taxonomy/txstat. cgi

The most sequenced organisms in Gen. Bank Homo sapiens Mus musculus Rattus norvegicus Danio rerio Zea mays Oryza sativa Drosophila melanogaster Gallus gallus Arabidopsis thaliana Updated 8 -12 -04 Gen. Bank release 142. 0 10. 7 billion bases 6. 5 b 5. 6 b 1. 7 b 1. 4 b 0. 8 b 0. 7 b 0. 5 b Table 2 -2 Page 18

The most sequenced organisms in Gen. Bank Homo sapiens Mus musculus Rattus norvegicus Danio rerio Bos taurus Zea mays Oryza sativa (japonica) Xenopus tropicalis Canis familiaris Drosophila melanogaster Updated 8 -29 -05 Gen. Bank release 149. 0 11. 2 billion bases 7. 5 b 5. 7 b 2. 1 b 1. 9 b 1. 4 b 1. 2 b 0. 9 b 0. 8 b 0. 7 b Table 2 -2 Page 18

The most sequenced organisms in Gen. Bank Homo sapiens Mus musculus Rattus norvegicus Bos taurus Danio rerio Zea mays Oryza sativa (japonica) Strongylocentrotus purpurata Sus scrofa Xenopus tropicalis Updated 7 -19 -06 Gen. Bank release 154. 0 12. 3 billion bases 8. 0 b 5. 7 b 3. 5 b 2. 5 b 1. 8 b 1. 5 b 1. 2 b 1. 0 b Table 2 -2 Page 18

Molecular Biology Information DNA • Raw DNA Sequence – Coding or Not? – Parse into genes? – 4 bases: AGCT – ~1 K in a gene, ~2 M in genome – ~3 Gb Human atggcaattaaaattggtatcaatggttttggtcgtatcggccgtattccgtgca gcacaacaccgtgatgacattgaagttgtaggtattaacgacttaatcgacgttgaatac atggcttatatgttgaaatatgattcaactcacggtcgtttcgacggcactgttgaagtg aaagatggtaacttagtggttaatggtaaaactatccgtgtaactgcagaacgtgatcca gcaaacttaaactggggtgcaatcggtgttgatatcgctgttgaagcgactggtttattc ttaactgatgaaactgctcgtaaacatatcactgcaggcgcaaaaaaagttgtattaact ggcccatctaaagatgcaacccctatgttcgtggtgtaaacttcaacgcatacgca ggtcaagatatcgtttctaacgcatcttgtacaacaaactgtttagctcctttagcacgt gttgttcatgaaactttcggtatcaaagatggtttaatgaccactgttcacgcaacgact gcaactcaaaaaactgtggatggtccatcagctaaagactggcgcggcggccgcggtgca tcacaaaacatcattccatcttcaacaggtgcagcgaaagcagtaggtaaagtattacct gcattaaacggtaaattaactggtatggctttccgtgttccaacgccaaacgtatctgtt gttgatttaacagttaatcttgaaaaaccagcttcttatgatgcaatcaaacaagcaatc aaagatgcagcggaaggtaaaacgttcaatggcgaattaaaaggcgtattaggttacact gaagatgctgttgtttctactgacttcaacggttgtgctttaacttctgtatttgatgca gacgctggtatcgcattaactgattctttcgttaaattggtatc. . . caaaaatagggttaatatgaatctcgatctccattttgttcatcgtattcaa caacaagccaaaactcgtacaaatatgaccgcacttcgctataaagaacacggcttgtgg cgagatatctcttggaaaaactttcaagagcaactcaactttctcgagcattgctt gctcacaatattgacgtacaagataaaatcgccatttttgcccataatatggaacgttgg gttgttcatgaaactttcggtatcaaagatggtttaatgaccactgttcacgcaacgact acaatcgttgacattgcgaccttacaaattcgagcaatcacagtgcctatttacgcaacc aatacagcccagcagaatttatcctaaatcacgccgatgtaaaaattctcttcgtc ggcgatcaagagcaatacgatcaaacattggaaattgctcatcattgtccaaaattacaa aaaattgtagcaatgaaatccaccattcaattacaacaagatcctcttgcacttgg

Molecular Biology Information: Protein Sequence • 20 letter alphabet – ACDEFGHIKLMNPQRSTVWY but not BJOUXZ • Strings of ~300 aa in an average protein (in bacteria), ~200 aa in a domain • >1 M known protein sequences (uniprot) d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ LNCIVAVSQNMGIGKNGDLPWPPLRNEFRYFQRMTTTSSVEGKQ-NLVIMGKKTWFSI LNSIVAVCQNMGIGKDGNLPWPPLRNEYKYFQRMTSTSHVEGKQ-NAVIMGKKTWFSI ISLIAALAVDRVIGMENAMPWN-LPADLAWFKRNTL----NKPVIMGRHTWESI TAFLWAQDRDGLIGKDGHLPWH-LPDDLHYFRAQTV----GKIMVVGRRTYESF d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ LNCIVAVSQNMGIGKNGDLPWPPLRNEFRYFQRMTTTSSVEGKQ-NLVIMGKKTWFSI LNSIVAVCQNMGIGKDGNLPWPPLRNEYKYFQRMTSTSHVEGKQ-NAVIMGKKTWFSI ISLIAALAVDRVIGMENAMPW-NLPADLAWFKRNTLD----KPVIMGRHTWESI TAFLWAQDRNGLIGKDGHLPW-HLPDDLHYFRAQTVG----KIMVVGRRTYESF d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ VPEKNRPLKGRINLVLSRELKEPPQGAHFLSRSLDDALKLTEQPELANKVDMVWIVGGSSVYKEAMNHP VPEKNRPLKDRINIVLSRELKEAPKGAHYLSKSLDDALALLDSPELKSKVDMVWIVGGTAVYKAAMEKP ---G-RPLPGRKNIILS-SQPGTDDRV-TWVKSVDEAIAACGDVP------EIMVIGGGRVYEQFLPKA ---PKRPLPERTNVVLTHQEDYQAQGA-VVVHDVAAVFAYAKQHLDQ----ELVIAGGAQIFTAFKDDV d 1 dhfa_ d 8 dfr__ d 4 dfra_ d 3 dfr__ -PEKNRPLKGRINLVLSRELKEPPQGAHFLSRSLDDALKLTEQPELANKVDMVWIVGGSSVYKEAMNHP -PEKNRPLKDRINIVLSRELKEAPKGAHYLSKSLDDALALLDSPELKSKVDMVWIVGGTAVYKAAMEKP -G---RPLPGRKNIILSSSQPGTDDRV-TWVKSVDEAIAACGDVPE-----. IMVIGGGRVYEQFLPKA -P--KRPLPERTNVVLTHQEDYQAQGA-VVVHDVAAVFAYAKQHLD----QELVIAGGAQIFTAFKDDV
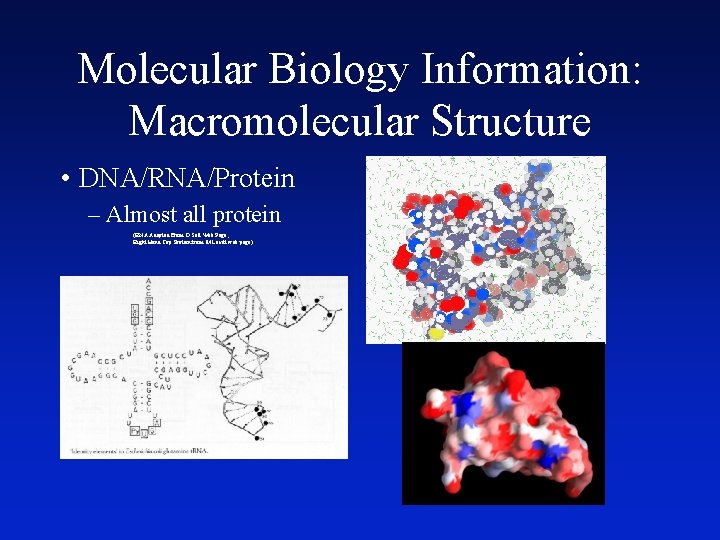
Molecular Biology Information: Macromolecular Structure • DNA/RNA/Protein – Almost all protein (RNA Adapted From D Soll Web Page, Right Hand Top Protein from M Levitt web page)

Molecular Biology Information: Whole Genomes • The Revolution Driving Everything Fleischmann, R. D. , Adams, M. D. , White, O. , Clayton, R. A. , Kirkness, E. F. , Kerlavage, A. R. , Bult, C. J. , Tomb, J. F. , Dougherty, B. A. , Merrick, J. M. , Mc. Kenney, K. , Sutton, G. , Fitzhugh, W. , Fields, C. , Gocayne, J. D. , Scott, J. , Shirley, R. , Liu, L. I. , Glodek, A. , Kelley, J. M. , Weidman, J. F. , Phillips, C. A. , Spriggs, T. , Hedblom, E. , Cotton, M. D. , Utterback, T. R. , Hanna, M. C. , Nguyen, D. T. , Saudek, D. M. , Brandon, R. C. , Fine, L. D. , Fritchman, J. L. , Fuhrmann, J. L. , Geoghagen, N. S. M. , Gnehm, C. L. , Mc. Donald, L. A. , Small, K. V. , Fraser, C. M. , Smith, H. O. & Venter, J. C. (1995). "Whole-genome random sequencing and assembly of Haemophilus influenzae rd. " Science 269: 496 -512. (Picture adapted from TIGR website, http: //www. tigr. org) • Integrative Data 1995, HI (bacteria): 1. 6 Mb & 1600 genes done 1997, yeast: 13 Mb & ~6000 genes for yeast 1998, worm: ~100 Mb with 19 K genes 1999: >30 completed genomes! 2003, human: 3 Gb & 100 K genes. . . Genome sequence now accumulate so quickly that, in less than a week, a single laboratory can produce more bits of data than Shakespeare managed in a lifetime, although the latter make better reading. -- G A Pekso, Nature 401: 115 -116 (1999)
![1995 Bacteria, 1. 6 Mb, ~1600 genes [Science 269: 496] 1997 Genomes highlight the 1995 Bacteria, 1. 6 Mb, ~1600 genes [Science 269: 496] 1997 Genomes highlight the](http://slidetodoc.com/presentation_image_h/1e0498e9e5167c694810cacb0984c848/image-52.jpg)
1995 Bacteria, 1. 6 Mb, ~1600 genes [Science 269: 496] 1997 Genomes highlight the Finiteness of the “Parts” in Biology Eukaryote, 13 Mb, ~6 K genes [Nature 387: 1] 1998 real thing, Apr ‘ 00 Animal, ~100 Mb, ~20 K genes [Science 282: 1945] 2000? Human, ~3 Gb, ~100 K genes [? ? ? ] ‘ 98 spoof Core

Other Types of Data • Gene Expression – Early experiments yeast – Now tiling array technology • 50 M data points to tile the human genome at ~50 bp res. – Can only sequence genome once but can do an infinite variety of array experiments • Phenotype Experiments • Protein Interactions – For yeast: 6000 x 6000 / 2 ~ 18 M possible interactions – maybe 30 K real

Weber Cartoon
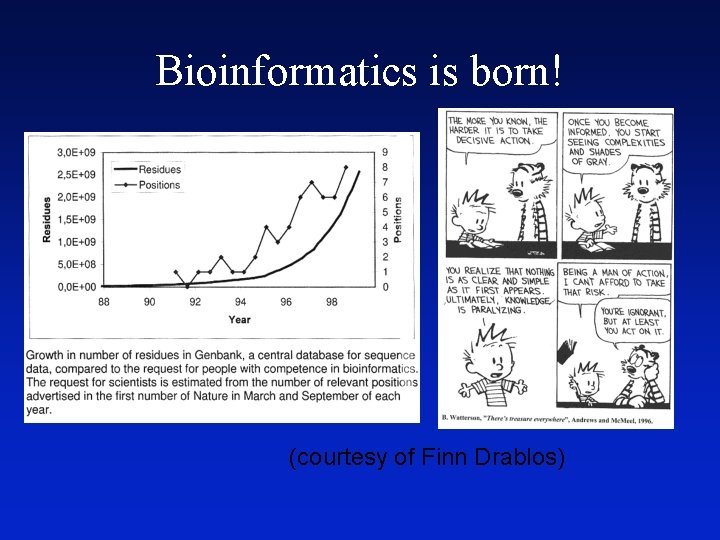
Bioinformatics is born! (courtesy of Finn Drablos)

Major Application I: Designing Drugs Core • Understanding How Structures Bind Other Molecules (Function) • Designing Inhibitors • Docking, Structure Modeling (From left to right, figures adapted from Olsen Group Docking Page at Scripps, Dyson NMR Group Web page at Scripps, and from Computational Chemistry Page at Cornell Theory Center).
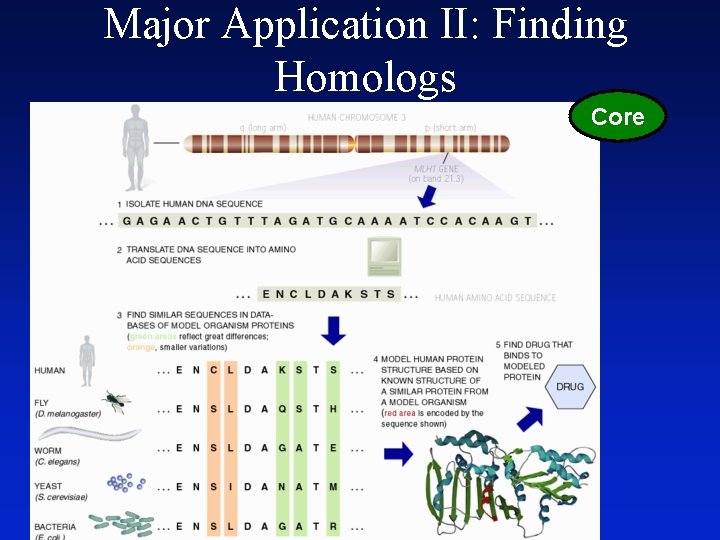
Major Application II: Finding Homologs Core

Major Application II: Finding Homologues • Find Similar Ones in Different Organisms • Human vs. Mouse vs. Yeast – Easier to do Expts. on latter! Best Sequence Similarity Matches to Date Reproduced Between Positionally Cloned (Section from NCBI Disease Genes Database Below. ) Human Genes and S. cerevisiae Proteins Human Disease MIM # Human Gene Gen. Bank BLASTX Acc# for P-value Human c. DNA Yeast Gene Gen. Bank Yeast Gene Acc# for Description Yeast c. DNA Hereditary Non-polyposis Colon Cancer Cystic Fibrosis Wilson Disease Glycerol Kinase Deficiency Bloom Syndrome Adrenoleukodystrophy, X-linked Ataxia Telangiectasia Amyotrophic Lateral Sclerosis Myotonic Dystrophy Lowe Syndrome Neurofibromatosis, Type 1 120436 219700 277900 307030 210900 300100 208900 105400 160900 309000 162200 MSH 2 MLH 1 CFTR WND GK BLM ALD ATM SOD 1 DM OCRL NF 1 U 03911 U 07418 M 28668 U 11700 L 13943 U 39817 Z 21876 U 26455 K 00065 L 19268 M 88162 M 89914 9. 2 e-261 6. 3 e-196 1. 3 e-167 5. 9 e-161 1. 8 e-129 2. 6 e-119 3. 4 e-107 2. 8 e-90 2. 0 e-58 5. 4 e-53 1. 2 e-47 2. 0 e-46 MSH 2 MLH 1 YCF 1 CCC 2 GUT 1 SGS 1 PXA 1 TEL 1 SOD 1 YPK 1 YIL 002 C IRA 2 M 84170 U 07187 L 35237 L 36317 X 69049 U 22341 U 17065 U 31331 J 03279 M 21307 Z 47047 M 33779 DNA repair protein Metal resistance protein Probable copper transporter Glycerol kinase Helicase Peroxisomal ABC transporter PI 3 kinase Superoxide dismutase Serine/threonine protein kinase Putative IPP-5 -phosphatase Inhibitory regulator protein Choroideremia Diastrophic Dysplasia Lissencephaly Thomsen Disease Wilms Tumor Achondroplasia Menkes Syndrome 303100 222600 247200 160800 194070 100800 309400 CHM DTD LIS 1 CLC 1 WT 1 FGFR 3 MNK X 78121 U 14528 L 13385 Z 25884 X 51630 M 58051 X 69208 2. 1 e-42 7. 2 e-38 1. 7 e-34 7. 9 e-31 1. 1 e-20 2. 0 e-18 2. 1 e-17 GDI 1 SUL 1 MET 30 GEF 1 FZF 1 IPL 1 CCC 2 S 69371 X 82013 L 26505 Z 23117 X 67787 U 07163 L 36317 GDP dissociation inhibitor Sulfate permease Methionine metabolism Voltage-gated chloride channel Sulphite resistance protein Serine/threoinine protein kinase Probable copper transporter

Core Major Application I|I: Overall Genome Characterization • Overall Occurrence of a Certain Feature in the Genome – e. g. how many kinases in Yeast • Compare Organisms and Tissues – Expression levels in Cancerous vs Normal Tissues • Databases, Statistics (Clock figures, yeast v. Synechocystis, adapted from Gene. Quiz Web Page, Sander Group, EBI)
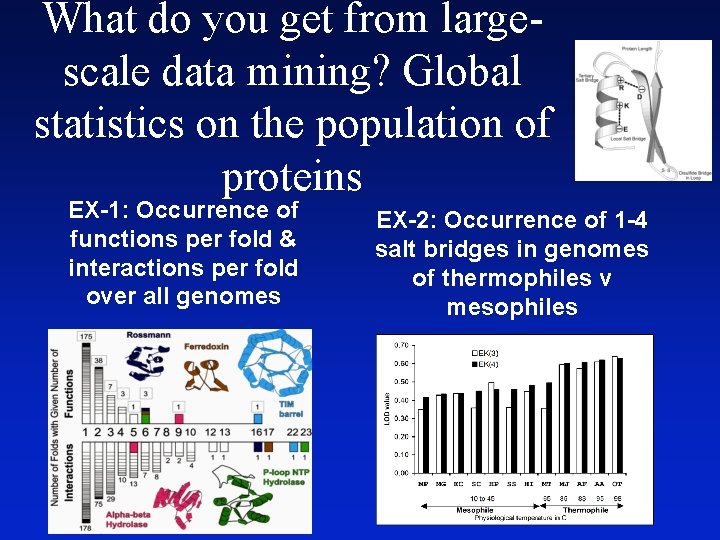
What do you get from largescale data mining? Global statistics on the population of proteins EX-1: Occurrence of functions per fold & interactions per fold over all genomes EX-2: Occurrence of 1 -4 salt bridges in genomes of thermophiles v mesophiles

http: //www. nature. com/nature/journal/vaop/ncurrent/pdf/

End of First Lecture 2009 Remaining Slides Lecture 2
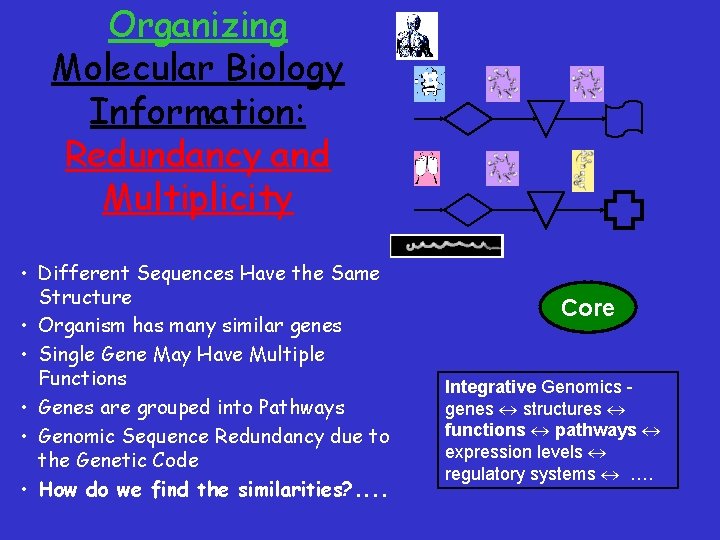
Organizing Molecular Biology Information: Redundancy and Multiplicity • Different Sequences Have the Same Structure • Organism has many similar genes • Single Gene May Have Multiple Functions • Genes are grouped into Pathways • Genomic Sequence Redundancy due to the Genetic Code • How do we find the similarities? . . Core Integrative Genomics genes structures functions pathways expression levels regulatory systems ….

Ome molecular group Genome Proteome Transcriptome Phenome Interactome Metabolome Physiome Orfeome Secretome Morphome Glycome Regulome Functome Cellome Transportome Ribonome Operome 'Omics: studying populations of molecules in a database framework
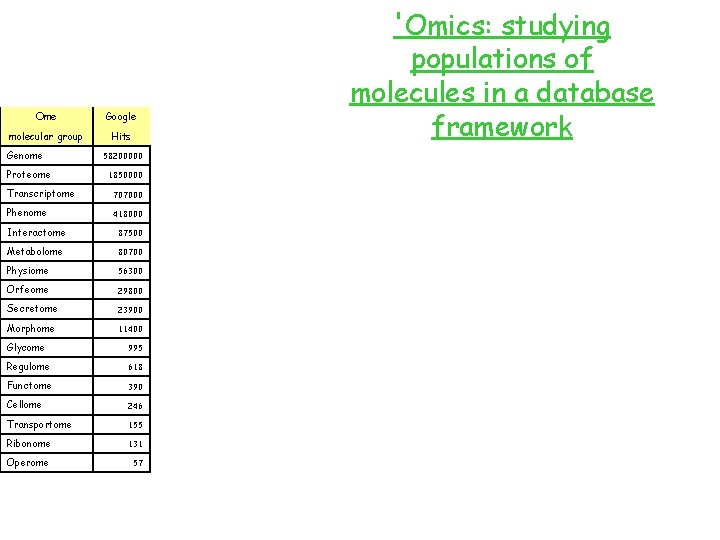
Ome Google molecular group Hits Genome Proteome 58200000 1850000 Transcriptome 707000 Phenome 418000 Interactome 87500 Metabolome 80700 Physiome 56300 Orfeome 29800 Secretome 23900 Morphome 11400 Glycome 995 Regulome 618 Functome 390 Cellome 246 Transportome 155 Ribonome 131 Operome 57 'Omics: studying populations of molecules in a database framework
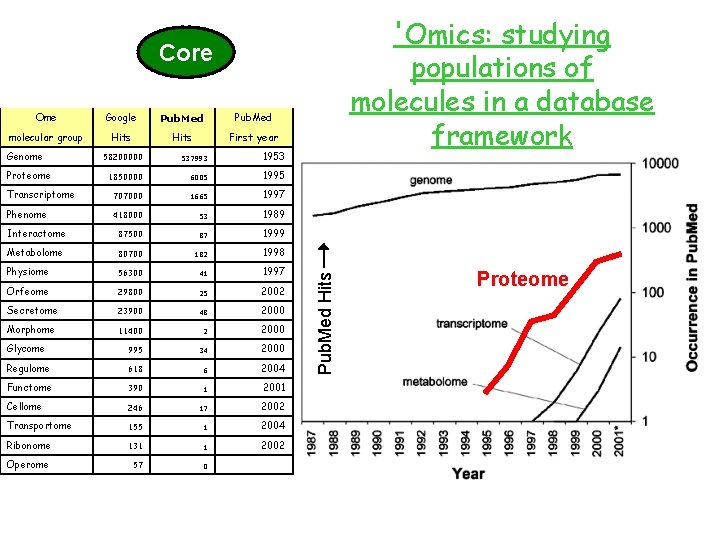
'Omics: studying populations of molecules in a database framework Ome Google Pub. Med molecular group Hits First year Genome 58200000 537993 1953 1850000 6005 1995 Transcriptome 707000 1665 1997 Phenome 418000 53 1989 Interactome 87500 87 1999 Metabolome 80700 182 1998 Physiome 56300 41 1997 Orfeome 29800 25 2002 Secretome 23900 48 2000 Morphome 11400 2 2000 Glycome 995 34 2000 Regulome 618 6 2004 Functome 390 1 2001 Cellome 246 17 2002 Transportome 155 1 2004 Ribonome 131 1 2002 Operome 57 0 Proteome Pub. Med Hits Core Proteome
A Parts List Approach to Bike Maintenance Extra
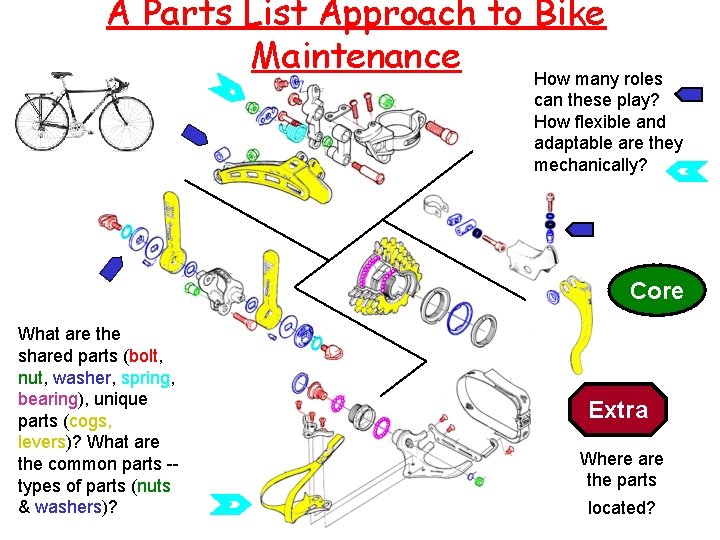
A Parts List Approach to Bike Maintenance How many roles can these play? How flexible and adaptable are they mechanically? Core What are the shared parts (bolt, nut, washer, spring, bearing), unique parts (cogs, levers)? What are the common parts -types of parts (nuts & washers)? Extra Where are the parts located?
Molecular Parts = Conserved Domains, Folds, &c
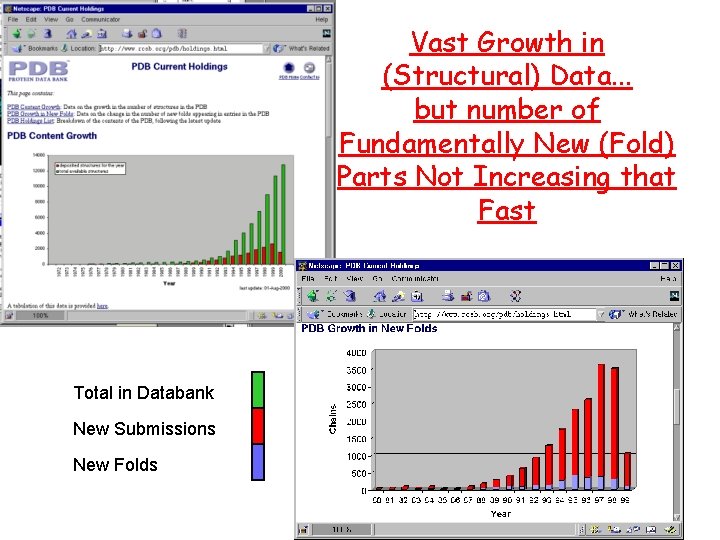
Vast Growth in (Structural) Data. . . but number of Fundamentally New (Fold) Parts Not Increasing that Fast Total in Databank New Submissions New Folds
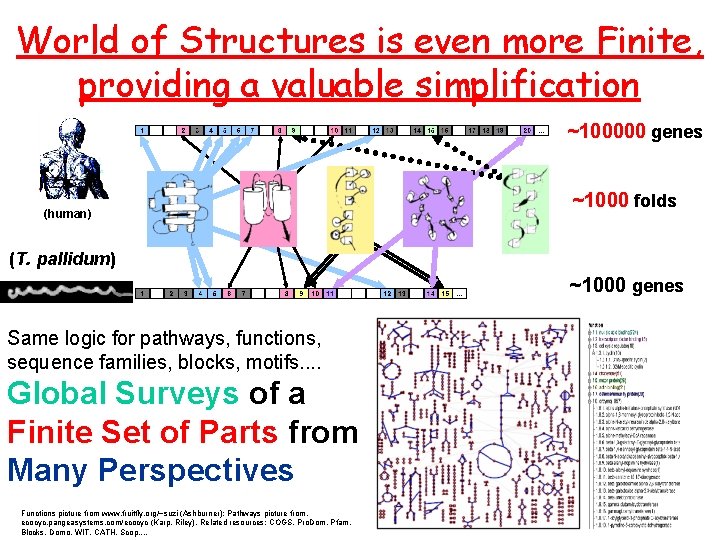
World of Structures is even more Finite, providing a valuable simplification ~100000 genes (human) ~1000 folds (T. pallidum) ~1000 genes Same logic for pathways, functions, sequence families, blocks, motifs. . Global Surveys of a Finite Set of Parts from Many Perspectives Functions picture from www. fruitfly. org/~suzi (Ashburner); Pathways picture from, ecocyc. pangeasystems. com/ecocyc (Karp, Riley). Related resources: COGS, Pro. Dom, Pfam, Blocks, Domo, WIT, CATH, Scop. .

What is Bioinformatics? • (Molecular) Bio - informatics • One idea for a definition? Bioinformatics is conceptualizing biology in terms of molecules (in the sense of physical-chemistry) and then applying “informatics” techniques (derived from disciplines such as applied math, CS, and statistics) to understand organize the information associated with these molecules, on a large-scale. • Structural Bioinformatics is a practical discipline with many applications that deals with biological three dimensional structural data.

General Types of “Informatics” techniques in Bioinformatics • Databases à Building, Querying à Object DB • Text String Comparison à à Text Search 1 D Alignment Significance Statistics Google, grep • Finding Patterns à AI / Machine Learning à Clustering à Datamining • Geometry à Robotics à Graphics (Surfaces, Volumes) à Comparison and 3 D Matching (Vision, recognition) • Physical Simulation à à Newtonian Mechanics Electrostatics Numerical Algorithms Simulation

Bioinformatics Topics -Genome Sequence • Finding Genes in Genomic DNA à introns à exons à promotors • Characterizing Repeats in Genomic DNA à Statistics à Patterns • Duplications in the Genome à Large scale genomic alignment • Whole-Genome Comparisons • Finding Structural RNAs

• Sequence Alignment à non-exact string matching, gaps à How to align two strings optimally via Dynamic Programming à Local vs Global Alignment à Suboptimal Alignment à Hashing to increase speed (BLAST, FASTA) à Amino acid substitution scoring matrices • Multiple Alignment and Consensus Patterns à How to align more than one sequence and then fuse the result in a consensus representation à Transitive Comparisons à HMMs, Profiles à Motifs Bioinformatics Topics -Protein Sequence • Scoring schemes and Matching statistics à How to tell if a given alignment or match is statistically significant à A P-value (or an e-value)? à Score Distributions (extreme val. dist. ) à Low Complexity Sequences • Evolutionary Issues à Rates of mutation and change

Bioinformatics Topics -Sequence / Structure • Secondary Structure “Prediction” à via Propensities à Neural Networks, Genetic Alg. à Simple Statistics à TM-helix finding à Assessing Secondary Structure Prediction • Structure Prediction: Protein v RNA • Tertiary Structure Prediction à Fold Recognition à Threading à Ab initio • Function Prediction à Active site identification • Relation of Sequence Similarity to Structural Similarity

Problems in Protein Bioinformatics • Prediction of structure from sequence à Fold recognition à Fragment construction • Proteome annotation • Protein-protein docking
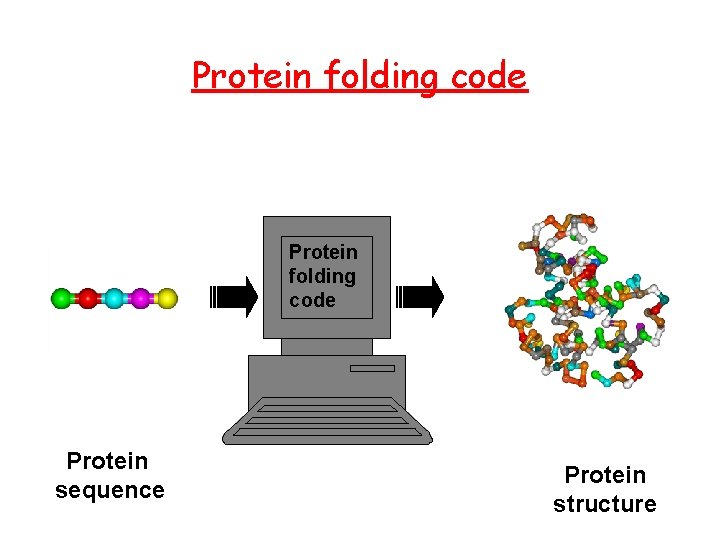
Protein folding code Protein sequence Protein structure
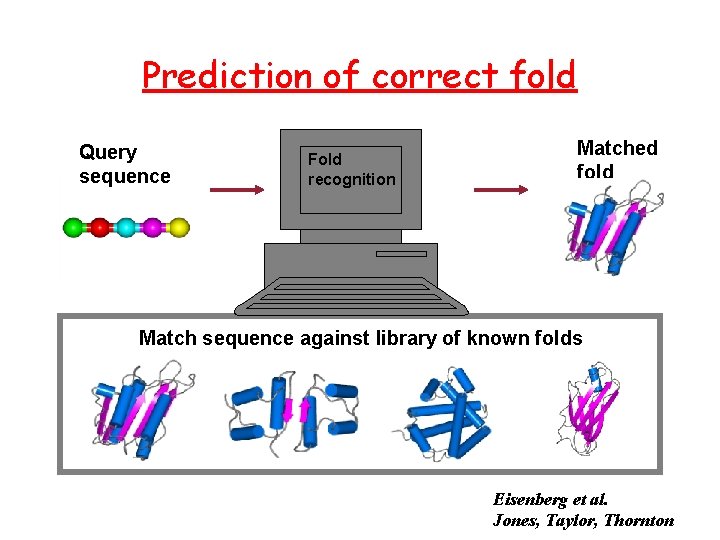
Prediction of correct fold Query sequence Fold recognition Matched fold Match sequence against library of known folds Eisenberg et al. Jones, Taylor, Thornton

Computational Requirements • 1 sequence search takes 12 mins (3 Ghz) • Benchmarking on 100 proteins with 100 runs for a simplex search of parameter space = 80 days • 30 approaches explored = 7 years (on 1 cpu)
- Slides: 80