Canadian Bioinformatics Workshops www bioinformatics ca Module Title

Canadian Bioinformatics Workshops www. bioinformatics. ca
Module #: Title of Module 2

Module 1 Introduction to Metagenomics Robert Beiko Analysis of Metagenomic Data June 24 -26, 2015 Rob Beiko rbeiko@dal. ca @rob_beiko

en. wikipedia. org Module 1 bioinformatics. ca
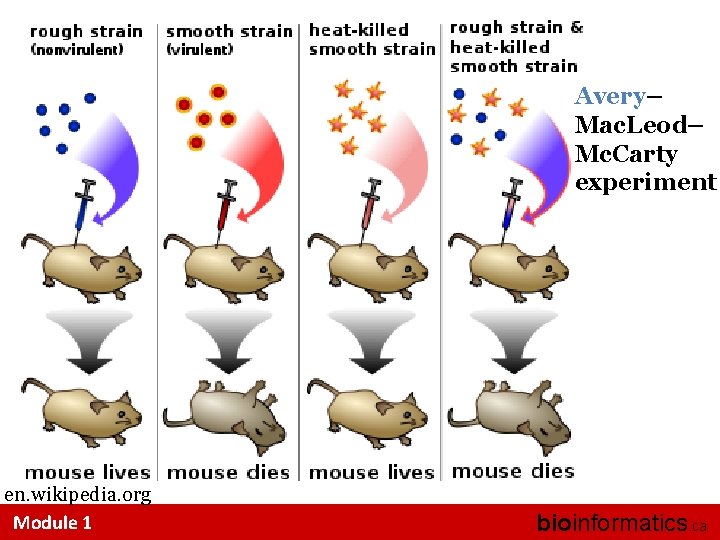
Avery– Mac. Leod– Mc. Carty experiment en. wikipedia. org Module 1 bioinformatics. ca

Course overview • Module 1: Introduction – definitions, approaches, considerations • Module 2: Marker genes – measuring community diversity • Module 3: Metagenome taxonomy – classifying and binning sequence reads • Module 4: Metagenome function – databases and pathways • Module 5: Metatranscriptomics: data, taxonomy, function • Module 6: Biomarker discovery Module 1 bioinformatics. ca

General Learning Objectives At the end of this workshop, you will be able to: • Define the objectives of different types of metagenomic projects • Process raw data files using appropriate quality control • Run standard pipelines for marker-gene, metagenome and metatranscriptome datasets • Analyze results using statistical and network approaches • Recognize the technical limitations of metagenomic studies Module 1 bioinformatics. ca

Learning objectives of Module 1 You will be able to: • Apply key terms in metagenomics, for example microbial communities, OTUs, metadata • Define the objectives of a metagenomic experiment, with appropriate choice of technology • Interpret the contents of sequence files • Acquire data from online resources and reference databases Module 1 bioinformatics. ca

Defining Metagenomics • Microbiome: Attributed to Joshua Lederberg by Hooper and Gordon (2001): “the collective genome of our indigenous microbes (microflora), the idea being that a comprehensive genetic view of Homo sapiens as a life-form should include the genes in our microbiome” • Is also used to mean microbiota, the set of microorganisms found in a particular setting • Metagenome: Handelsman et al. (1998) “…advances in molecular biology and eukaryotic genomics, which have laid the groundwork for cloning and functional analysis of the collective genomes of soil microflora, which we term the metagenome of the soil. ” • Does not encompass marker-gene surveys (e. g. , 16 S) This report says it does. Module 1 bioinformatics. ca

The big picture Explore the relationship between microbes and their habitat To accomplish this, we use a series of experimental and computational techniques to make inferences about the community: - Marker genes - Metagenomes - Metatranscriptomes - Metaproteomes - Metametabolomes - “Culturomes” Module 1 bioinformatics. ca

Why metagenomics? • The “great plate count anomaly”: <1% of organisms across many habitats are culturable (reviewed in Amann et al. , 1995: PMID 7535888) – CONTROVERSIAL; probably not true for habitats such as human body sites • In any event, it would be nearly impossible to culture ALL constituents of a given microbiome sample (apart from trivially simple ones) • Metagenomics offers an effective (if imperfect) way to profile the structure and function of microbial communities Module 1 bioinformatics. ca
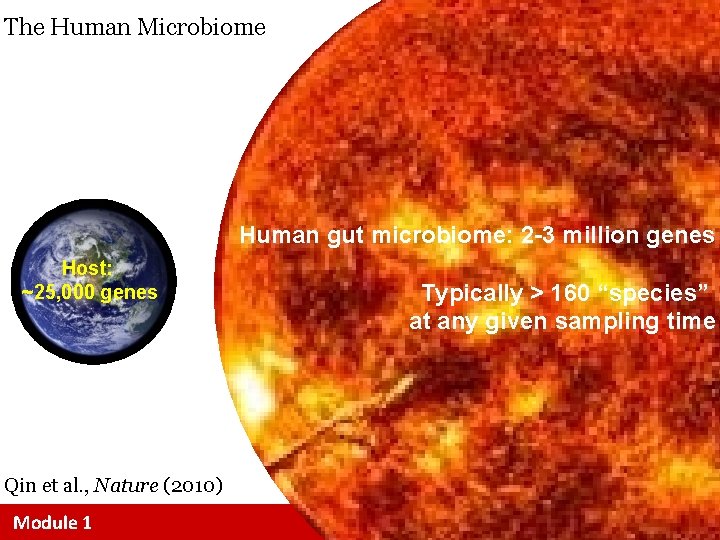
The Human Microbiome Human gut microbiome: 2 -3 million genes Host: ~25, 000 genes Typically > 160 “species” at any given sampling time Qin et al. , Nature (2010) Module 1 bioinformatics. ca

A Brief History of Metagenomics and Things Like It Module 1 bioinformatics. ca
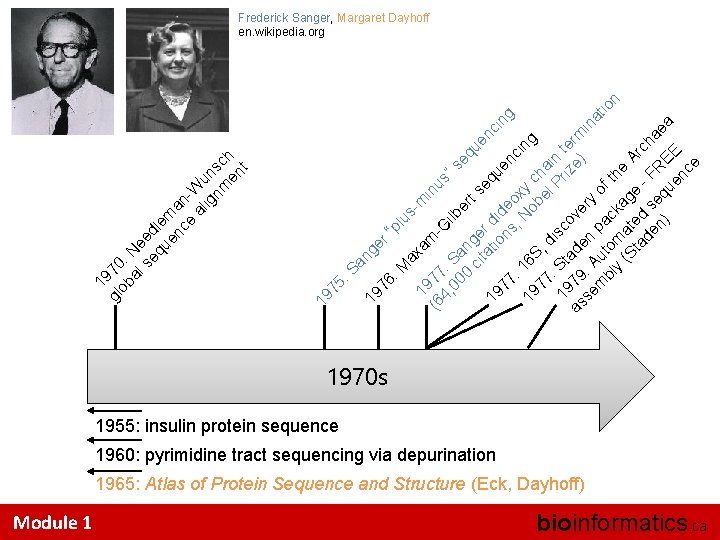
Module 1 er Sa ng M “p lu sax m 19 am in (6 77 us G 4, : S ”s ilb 00 a eq e 0 ng r t ue ci e se t r a nc q 19 tio did u in en 77 ns eo g ci : 1 ; N xy 19 6 S ob ch ng 77 , el ai : S dis Pr n t 19 t co iz er a as 79 de ve e) m in se : A n ry at m ut pa of io bl om ck th n y (S at ag e A ta ed e rc de s h F n) eq RE aea ue E nc e 76 : 19 75 : 19 1 gl 970 ob : al Ne se ed qu le en ma ce nal Wu ig n nm sc en h t Frederick Sanger, Margaret Dayhoff en. wikipedia. org 1970 s 1955: insulin protein sequence 1960: pyrimidine tract sequencing via depurination 1965: Atlas of Protein Sequence and Structure (Eck, Dayhoff) bioinformatics. ca

Staden (1979) “The continuing rapid fall in the cost of computer components is making it possible for most DNA sequencing laboratories to have their own small computer. The fact that DNA sequencing is now a fast procedure, and the availability of computers gives the possibility of more efficient overall strategies for sequence determination. ” Module 1 bioinformatics. ca

T 4 genome map: Wood and Revel, 1976 Phi. X 174 phage genome: Sanger et al. , 1977 Module 1 bioinformatics. ca
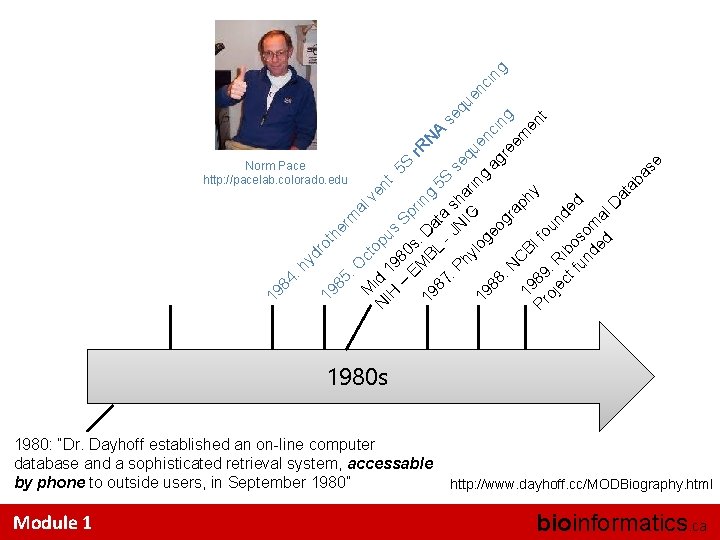
he 85 rm : O al ct M ve N id 1 opu nt IH 9 s 5 S S – 80 EM s: pri r. R 19 N BL Da ng A 5 87 t a S se : P JN sh se qu hy IG ari qu en ng lo en 19 ci g eo ag ci 88 ng gr re ng : N ap em 19 CB h en y If Pr 89 t o oj : R u n ec i t f bo ded un so de m d al D at ab as e 19 19 84 : hy dr ot Norm Pace http: //pacelab. colorado. edu 1980 s 1980: “Dr. Dayhoff established an on-line computer database and a sophisticated retrieval system, accessable by phone to outside users, in September 1980” Module 1 http: //www. dayhoff. cc/MODBiography. html bioinformatics. ca

Octopus Spring: Stahl et al. , 1985 Module 1 bioinformatics. ca
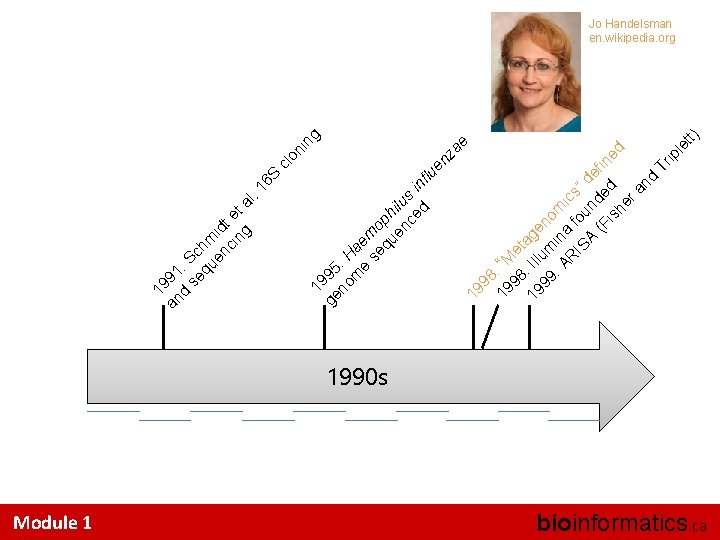
Module 1 6 S t) 98 19 : “M 98 e t 19 : Ill age 99 um no : A in m R a fo ics IS ” A und de fi (F is ed ned he ra nd Tr ip le t 19 ae nz in g cl on ge 95 no : H m ae e se mo qu ph en ilu ce s i d nfl ue 19 l. 1 an 91 d : S se c qu hm en id ci t e ng t a 19 Jo Handelsman en. wikipedia. org 1990 s bioinformatics. ca
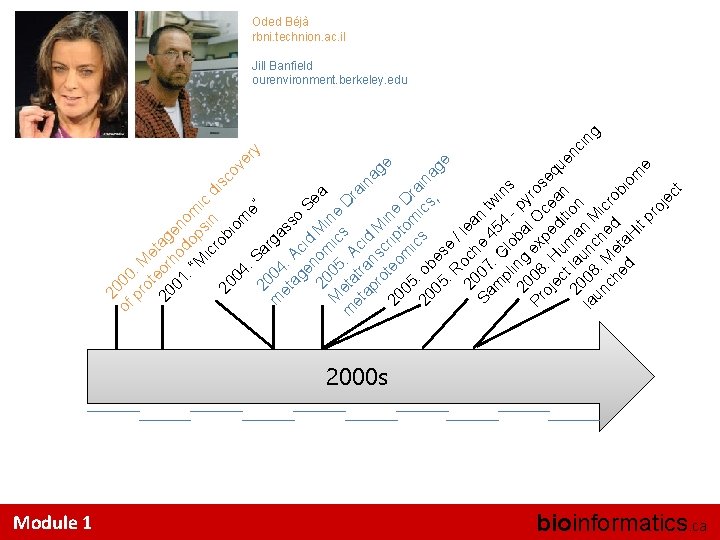
of 00 pr : M ot e 20 eor tag 01 ho en : “ do om M ps ic ic in d ro is 20 bi co om 04 ve : S e” ry 20 ar m 04 ga et : s ag Ac so Se en id M 2 om a M 005 ic ine D m eta : A s ra et tr c in ap an id ag M s ro c i e 20 teo rip ne 05 m tom Dr i 20 : ob cs ics aina , 05 es ge : R e / 20 och lea Sa 07 e n t m : G 454 win pl lo - s 20 ing ba py Pr 08 ex l O ros oj : H pe ce eq ec u d an u it en 20 t lau ma ion ci ng la 08 nc n M un : h i ch Me ed cro bi ed ta om H it e pr oj ec t 20 Oded Béjà rbni. technion. ac. il Jill Banfield ourenvironment. berkeley. edu 2000 s Module 1 bioinformatics. ca
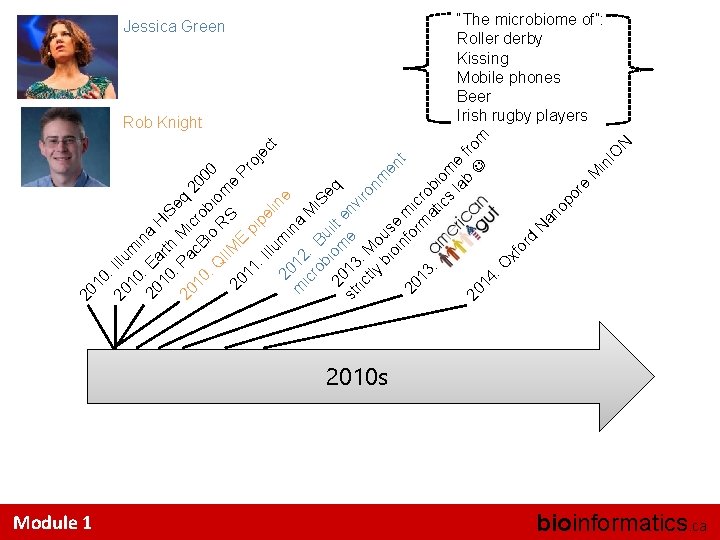
10 : 20 Illu 10 m in 20 : Ea a H 10 rth i. S : e 20 Pa Mic q 2 10 c. B ro 0 : Q io bio 00 IIM RS me 20 Pr 11 E p oj ip : I ec el llu t in 20 mi e m 12 na ic : ro B Mi. S b u e 20 iom ilt e q st 13 e nv ric : iro tly Mo nm bi us en oi e nf m t 20 or ic 13 m ro at bi : ic om s la e b fro 20 m 14 : O xf or d N an op or e M in IO N 20 Jessica Green Rob Knight Module 1 “The microbiome of”: Roller derby Kissing Mobile phones Beer Irish rugby players 2010 s bioinformatics. ca

(Very) high-level workflows Module 1 bioinformatics. ca
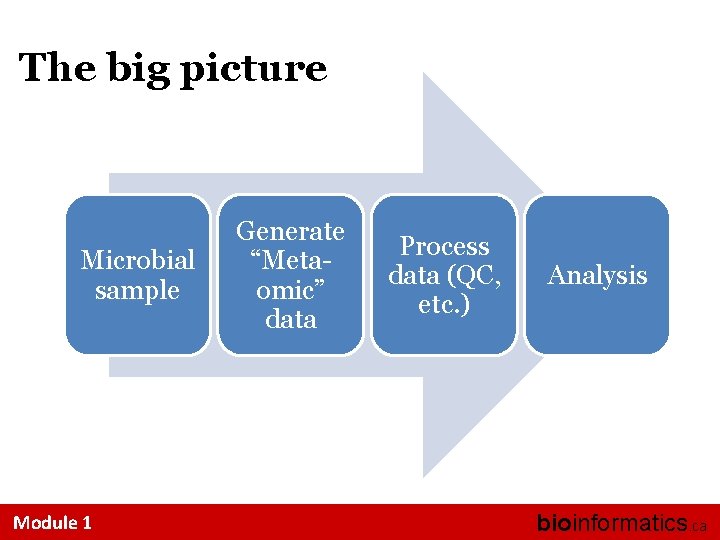
The big picture Microbial sample Module 1 Generate “Metaomic” data Process data (QC, etc. ) Analysis bioinformatics. ca
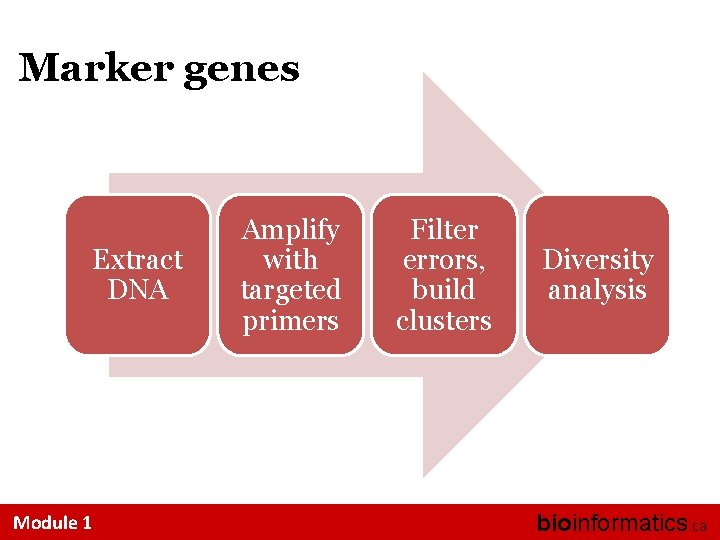
Marker genes Extract DNA Module 1 Amplify with targeted primers Filter errors, build clusters Diversity analysis bioinformatics. ca
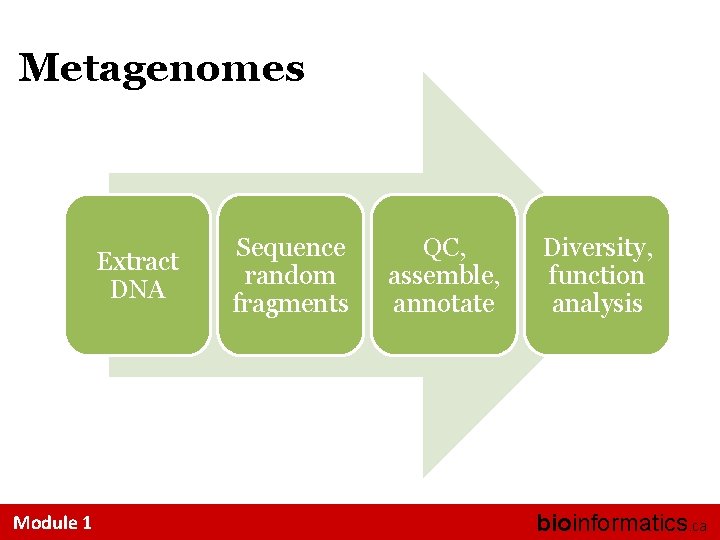
Metagenomes Extract DNA Module 1 Sequence random fragments QC, assemble, annotate Diversity, function analysis bioinformatics. ca
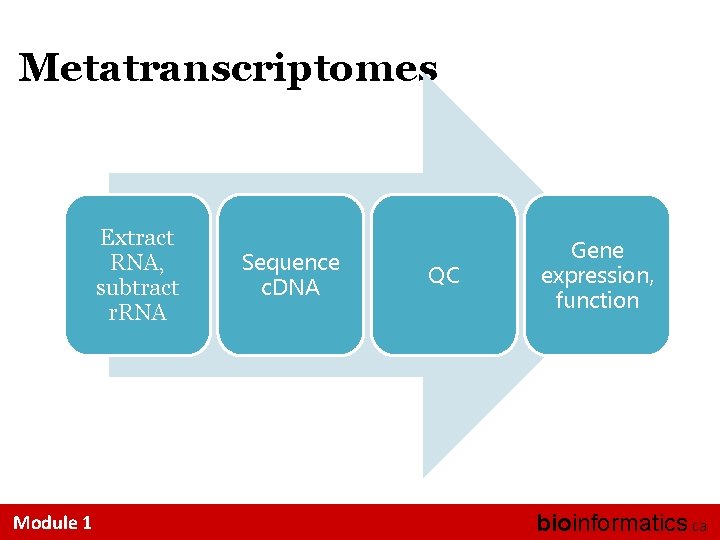
Metatranscriptomes Extract RNA, subtract r. RNA Module 1 Sequence c. DNA QC Gene expression, function bioinformatics. ca
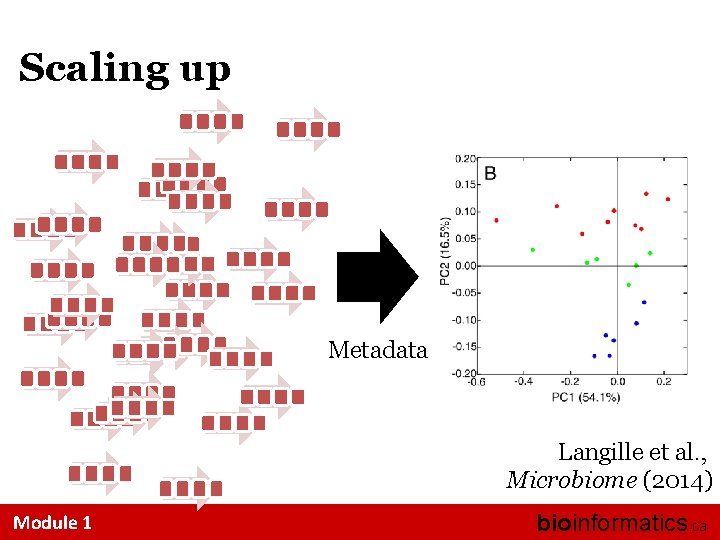
Scaling up Metadata Langille et al. , Microbiome (2014) Module 1 bioinformatics. ca

Examples of “Metagenomics” Module 1 bioinformatics. ca
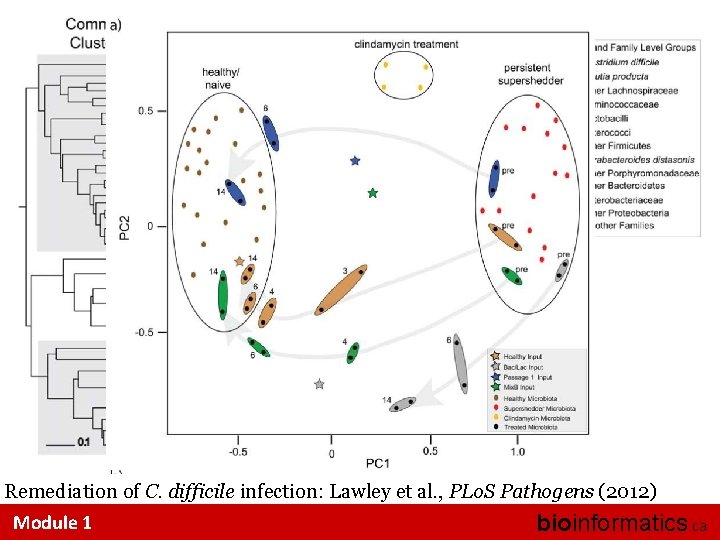
Remediation of C. difficile infection: Lawley et al. , PLo. S Pathogens (2012) Module 1 bioinformatics. ca
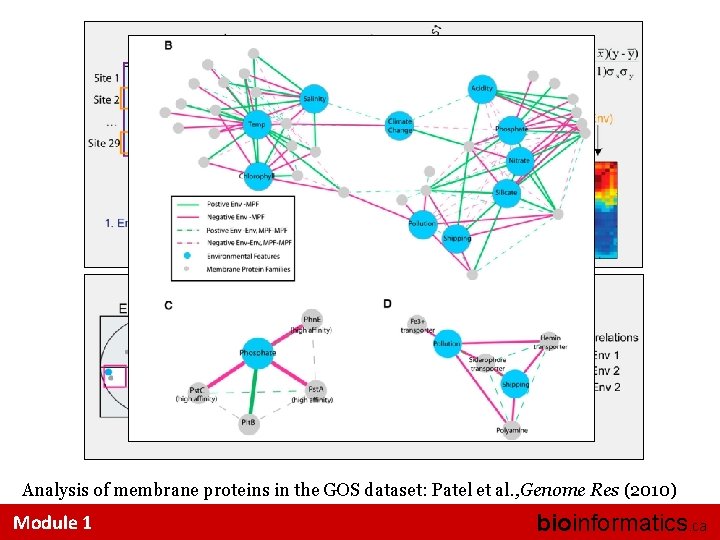
Analysis of membrane proteins in the GOS dataset: Patel et al. , Genome Res (2010) Module 1 bioinformatics. ca
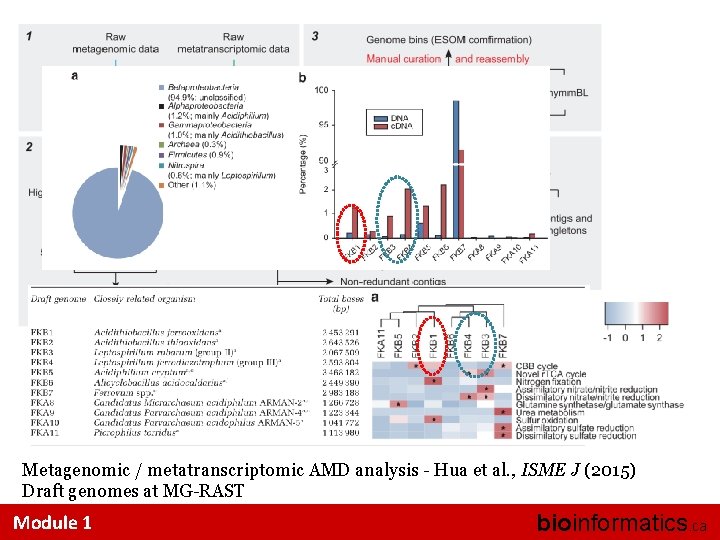
Metagenomic / metatranscriptomic AMD analysis - Hua et al. , ISME J (2015) Draft genomes at MG-RAST Module 1 bioinformatics. ca
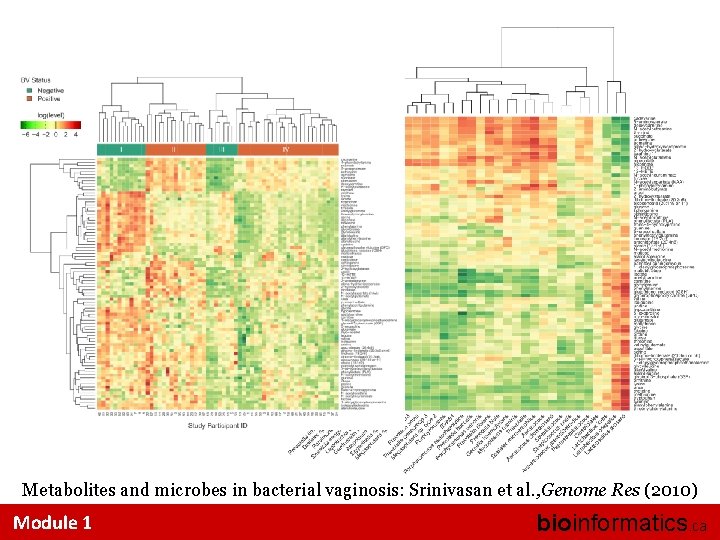
Metabolites and microbes in bacterial vaginosis: Srinivasan et al. , Genome Res (2010) Module 1 bioinformatics. ca
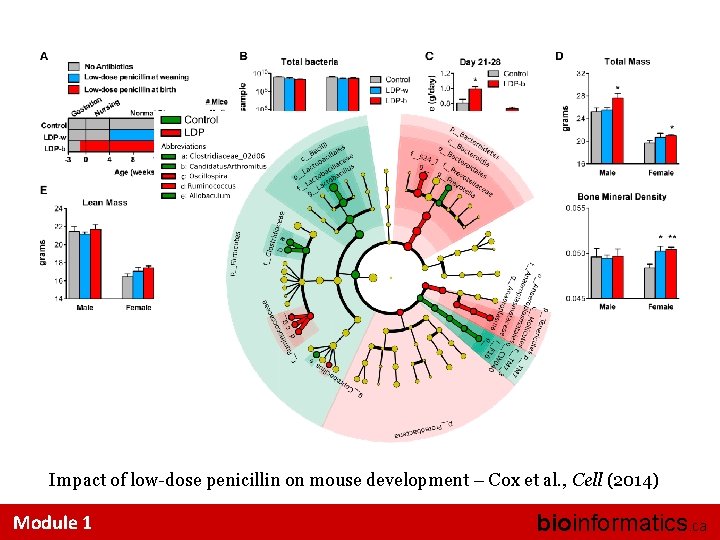
Impact of low-dose penicillin on mouse development – Cox et al. , Cell (2014) Module 1 bioinformatics. ca

Sequencing technologies Sanger Illumina *Seq Module 1 Ion Torrent Pacific Biosciences Roche 454 Nanopore bioinformatics. ca

Resources Module 1 bioinformatics. ca

16 S rrn. DB: Stoddard et al. NAR (2014) RDP II: Cole et al. NAR (2013) SILVA: Quast et al. NAR (2013) Module 1 Green. Genes: Mac. Donald et al. ISME J (2012) bioinformatics. ca
Genomes Gen. Bank Genomes GOLD Module 1 PATRIC Ensembl Genomes bioinformatics. ca
“Metagenomes” EBI metagenomics MG-RAST HMP DACC Module 1 bioinformatics. ca
Function KEGG CARD Uni. Prot. KB Gene Ontology Module 1 bioinformatics. ca

Major concerns in metagenomic analysis Module 1 bioinformatics. ca

Data Quality • Sequencing errors – Introduced in workup – Error rates, error type (Pac. Bio: 10% random, Illumina – 0. 1% substitution) • Chimeras – Amplification artifacts, cloning of restriction fragments Module 1 bioinformatics. ca

Comparability / Reproducibility • 16 S: different V regions give different results • Different sequencing platforms / sampling conditions ALSO give different results – Eisen paper about different recoveries under different conditions • Workflow complexity / plethora of tools Module 1 bioinformatics. ca
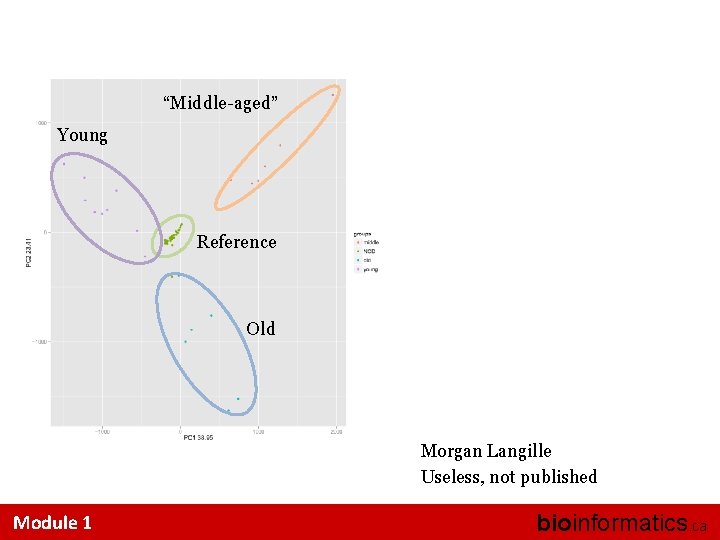
“Middle-aged” Young Reference Old Morgan Langille Useless, not published Module 1 bioinformatics. ca

Linkage and resolution • Strain-level diversity in metagenomes will often be missed by amplicon (esp. short-read) and shotgun approaches • This may be especially important between samples • Should you assemble metagenomic reads? What are the assumptions? Module 1 bioinformatics. ca
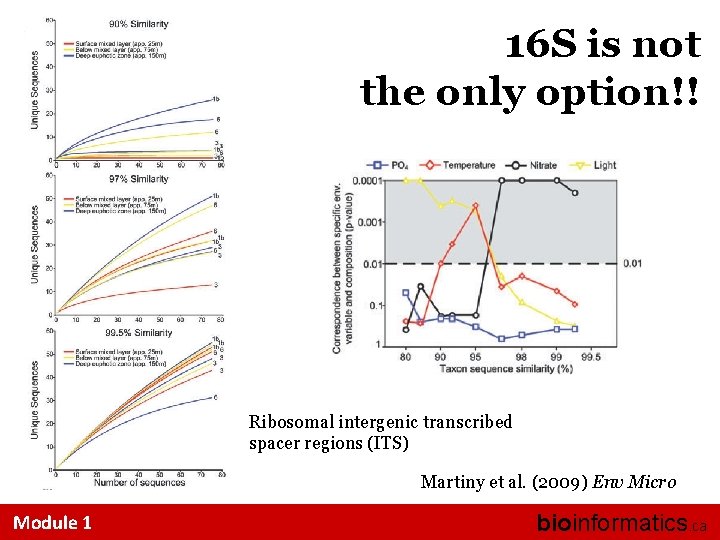
16 S is not the only option!! Ribosomal intergenic transcribed spacer regions (ITS) Martiny et al. (2009) Env Micro Module 1 bioinformatics. ca
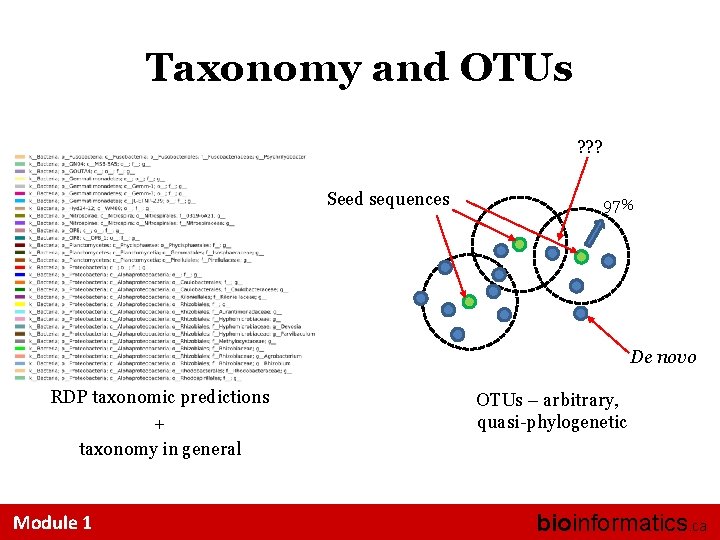
Taxonomy and OTUs ? ? ? Seed sequences 97% De novo RDP taxonomic predictions + taxonomy in general Module 1 OTUs – arbitrary, quasi-phylogenetic bioinformatics. ca
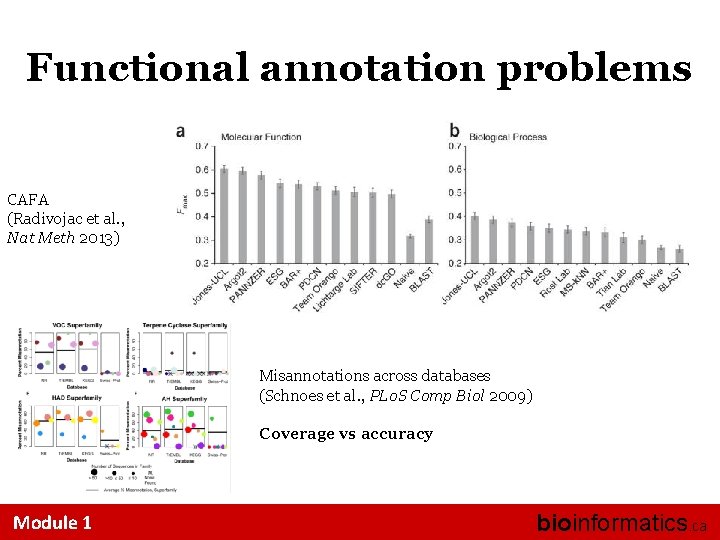
Functional annotation problems CAFA (Radivojac et al. , Nat Meth 2013) Misannotations across databases (Schnoes et al. , PLo. S Comp Biol 2009) Coverage vs accuracy Module 1 bioinformatics. ca

We are on a Coffee Break & Networking Session Module 1 bioinformatics. ca
- Slides: 48