Introduction to Bioinformatics databases Nucleic Acid Databases Dinesh

Introduction to Bioinformatics databases: Nucleic Acid Databases Dinesh Gupta ICGEB

Biological databases: why? • Need for storing and communicating large datasets has grown • Make biological data available to scientists. • To make biological data available in computer-readable form.

Different classifications of databases • Type of data – nucleotide sequences – proteins sequence patterns or motifs – macromolecular 3 D structure – gene expression data – metabolic pathways

Different classifications of databases…. • Primary or derived databases – Primary databases: experimental results directly into database – Secondary databases: results of analysis of primary databases – Aggregate of many databases • Links to other data items • Combination of data • Consolidation of data

Different classifications of databases…. • Technical design – Flat-files – Relational database (SQL) – Exchange/publication technologies (FTP, HTML, CORBA, XML, . . . )

Different classifications of databases…. • Availability – Publicly available, no restrictions – Available, but with copyright – Accessible, but not downloadable – Academic, but not freely available – Proprietary, commercial; possibly free for academics

Where do I get DB of my interest ?
http: //www 3. oup. co. uk/nar/database/c/

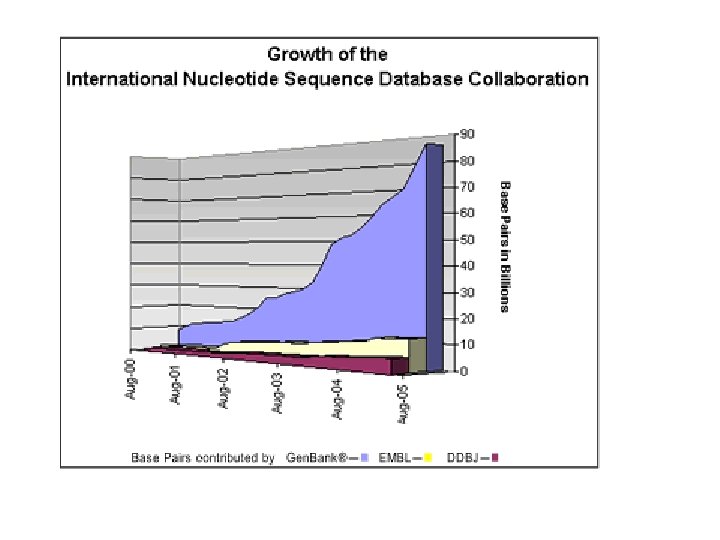
Nucleotide sequence databases • EMBL, Gen. Bank, and DDBJ are three primary nucleotide sequence databases • EMBL www. ebi. ac. uk/embl/ • Gen. Bank www. ncbi. nlm. nih. gov/Genbank/ • DDBJ www. ddbj. nig. ac. jp

Genbank • An annotated collection of all publicly available nucleotide and proteins • Set up in 1979 at the LANL (Los Alamos). • Maintained since 1992 NCBI (Bethesda). • http: //www. ncbi. nlm. nih. gov

EMBL Nucleotide Sequence Database • An annotated collection of all publicly available nucleotide and protein sequences • Created in 1980 at the European Molecular Biology Laboratory in Heidelberg. • Maintained since 1994 by EBI- Cambridge. • http: //www. ebi. ac. uk/embl. html
http: //www 3. ebi. ac. uk/Services/DBStats/

DDBJ–DNA Data Bank of Japan • An annotated collection of all publicly available nucleotide and protein sequences • Started, 1984 at the National Institute of Genetics (NIG) in Mishima. • Still maintained in this institute a team led by Takashi Gojobori. • http: //www. ddbj. nig. ac. jp

Other NCBI nucleic acids DBs • EST database: A collection of expressed sequence tags, or short, single-pass sequence reads from m. RNA (c. DNA). • GSS database: A database of genome survey sequences, or short, single-pass genomic sequences. • Homolo. Gene: A gene homology tool that compares nucleotide sequences between pairs of organisms in order to identify putative orthologs. • HTG database: A collection of high-throughput genome sequences from large-scale genome sequencing centers, including unfinished and finished sequences. • SNPs database: A central repository for both single-base nucleotide substitutions and short deletion and insertion polymorphisms. • Ref. Seq: A database of non-redundant reference sequences standards, including genomic DNA contigs, m. RNAs, and proteins for known genes. Multiple collaborations, both within NCBI and with external groups, supports data-gathering efforts. • STS database: A database of sequence tagged sites, or short sequences that are operationally unique in the genome. • Uni. STS: A unified, non-redundant view of sequence tagged sites (STSs). • Uni. Gene: A collection of ESTs and full-length m. RNA sequences organized into clusters, each representing a unique known or putative human gene annotated with mapping and expression information and cross-references to other sources.

Sequence submission • Data mainly direct submissions from the authors. • Submissions through the Internet: – Web forms. – Email. • Sequences shared/exchanged between the 3 centers on a daily basis: – The sequence content of the banks is identical.

Derived databases • CUTG Codon usage tabulated from Gen. Bank http: //www. kazusa. or. jp/codon/ • Genetic Codes Deviations from the standard genetic code in various organisms and organelles http: //www. ncbi. nlm. nih. gov/Taxonomy/Utils/wprintgc. cgi? mode=c • TIGR Gene Indices Organism-specific databases of EST and gene sequences http: //www. tigr. org/tdb/tgi. shtml • Uni. Gene Unified clusters of ESTs and full-length m. RNA sequences http: //www. ncbi. nlm. nih. gov/Uni. Gene/ • ASAP Alternative spliced isoforms http: //www. bioinformatics. ucla. edu/ASAP • Intronerator Introns and alternative splicing in C. elegans and C. briggsae http: //www. cse. ucsc. edu/~kent/intronerator/

Nucleic acid structure databases • NDB Nucleic acid-containing structures http: //ndbserver. rutgers. edu/ • NTDB Thermodynamic data for nucleic acids http: //ntdb. chem. cuhk. edu. hk/ • RNABase RNA-containing structures from PDB and NDB http: //www. rnabase. org/ • SCOR Structural classification of RNA: RNA motifs by structure, function and tertiary interactions • http: //scor. lbl. gov/

Database searching tips • Look for links to Help or Examples • Try Boolean searches • Be careful with UK/US spelling differences – leukaemia vs leukemia – haemoglobin vs hemoglobin – colour vs color

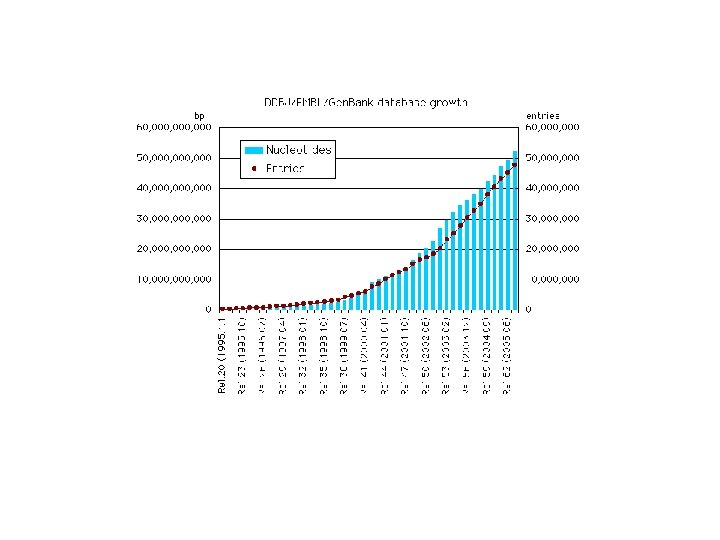
Exercises • Study the statistics of the three primary nucleic acid databases: Are they matching ? • Look for a gene of your interest in the three primary nucleic acid databases: compare the information given in each one of them. • Read NAR DB paper and NAR DB index site: search for different nucleic acid databases based on different search terms. • Self study: – http: //www 3. oup. co. uk/nar/database/c/ – Download NAR database paper (NARDB 2004) from: ftp: //cbag. sc. mahidol. ac. th/pub/Course_Materials/dinesh
- Slides: 37