EMBOSS an application suite for Bioinformatics v Shahid

EMBOSS – an application suite for Bioinformatics v Shahid Manzoor v Adnan Niazi SLU Global Bioinformatics Centre

E – European M – Molecular B – Biology O – Open S – Software S - Suite SLU Global Bioinformatics Centre

All Information Ø EMBOSS info at http: //emboss. sourceforge. net/. Ø w. EMBOSS info at http: //wemboss. sourceforge. net/. Ø E-mail martin. norling@slu. se to get a username and password for w. EMBOSS at http: //ebiokit. hgen. slu. se/. SLU Global Bioinformatics Centre

What is EMBOSS Ø Open Source molecular biology analysis package. Ø Handles a variety of common file formats. Ø Provides libraries for easy development Ø Software, licensed under GPL and LGPL Ø Developed by Martin Sarachu and Marc Colet Ø Available at http: //emboss. sourceforge. net SLU Global Bioinformatics Centre

Features of EMBOSS Ø A comprehensive set of sequence analysis programs. Ø All sequence and many alignment and structural formats are Handled. Ø It runs on practically every UNIX you can think of (and likely some that you can't), plus Windows and OS X. Ø Each application has the same style of interface so master one and you've mastered them all. SLU Global Bioinformatics Centre

Uses for EMBOSS Ø Sequence alignment. Ø Protein motif identification (including domain analysis) Ø Nucleotide sequence pattern analysis (for example to identify Cp. G islands or repeats). Ø Presentation tools for publications. SLU Global Bioinformatics Centre

Programs in EMBOSS Ø Ø Many small and large programs in package (>140). All programs share a common look and feel. Ø Easy to run from command line. Ø Retrieval of sequence data from the web. SLU Global Bioinformatics Centre

The one Argument Ø help the –help argument displays a short help for any EMBOSS program. SLU Global Bioinformatics Centre

The One Command Ø wossname searches the other programs short description for keywords. SLU Global Bioinformatics Centre
Large collection of gene and protein analysis tools Translation Protein domain searching Sequence retrieval Alignments Primer design Restriction Mapping SLU Global Bioinformatics Centre
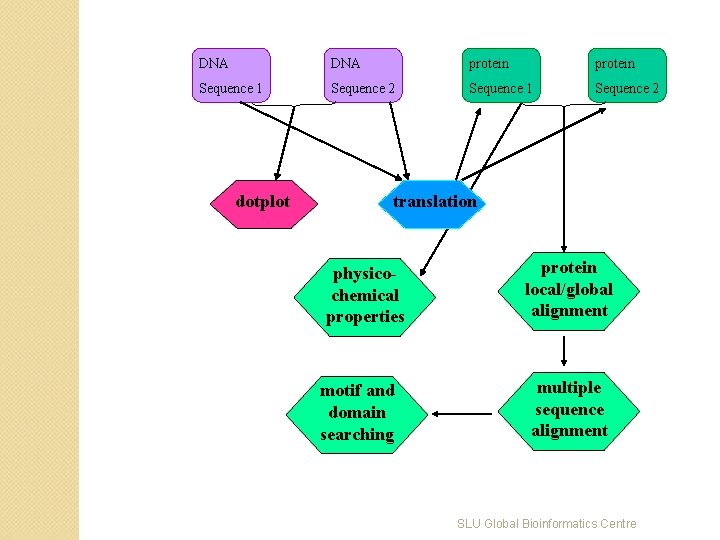
DNA protein Sequence 1 Sequence 2 dotplot translation physicochemical properties motif and domain searching protein local/global alignment multiple sequence alignment SLU Global Bioinformatics Centre

Dotplots >SEQ 1. fasta AGTGGTCGTGAAG AGAATGCTCCTCC TTTGGAATCTTAA >SEQ 2. fasta AGTGCTCCTCCCT TAGAATCTTAG For an exact match: Unix% dottup SEQ 1. fasta SEQ 2. fasta –window 10 & For a similarity match: Unix% dotmatcher SEQ 1. fasta SEQ 2. fasta –window 10 – threshold 17 & SLU Global Bioinformatics Centre

Dotplots … Identity Matrix A A T G C 5 -4 -4 -4 T -4 5 -4 -4 G – 4 -4 5 -4 C -4 -4 -4 5 Window Size is number of bases in a sliding window that is moved along each sequence and compared to generate a single data point on the plot. Window size must be an odd number. Mismatch Limit determines how similar the two sequences in a window must be to "match". For example, if window size is 9 and mismatch limit is 2, then up to 2 mismatches in a 9 base window will still be classified as a match. SLU Global Bioinformatics Centre
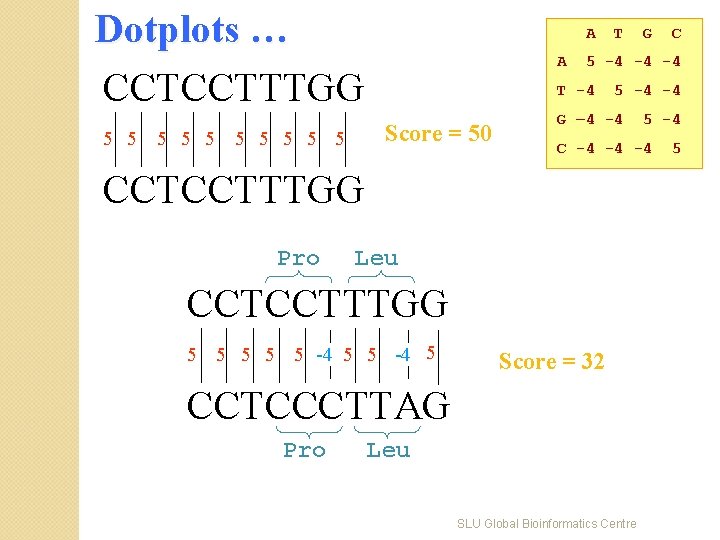
Dotplots … CCTCCTTTGG 5 5 5 A A G – 4 -4 CCTCCTTTGG Score = 32 CCTCCCTTAG Pro 5 -4 C -4 -4 -4 Leu 5 5 5 -4 5 C 5 -4 -4 CCTCCTTTGG Pro G 5 -4 -4 -4 T -4 Score = 50 5 5 5 T Leu SLU Global Bioinformatics Centre 5
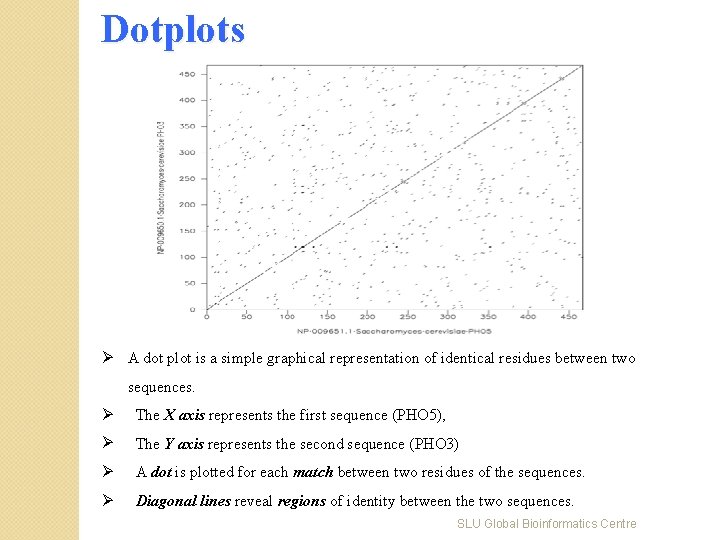
Dotplots Ø A dot plot is a simple graphical representation of identical residues between two sequences. Ø The X axis represents the first sequence (PHO 5), Ø The Y axis represents the second sequence (PHO 3) Ø A dot is plotted for each match between two residues of the sequences. Ø Diagonal lines reveal regions of identity between the two sequences. SLU Global Bioinformatics Centre
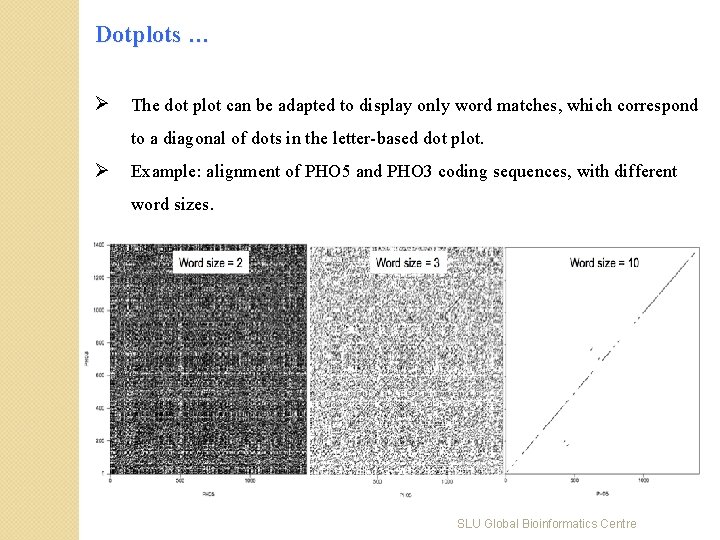
Dotplots … Ø The dot plot can be adapted to display only word matches, which correspond to a diagonal of dots in the letter-based dot plot. Ø Example: alignment of PHO 5 and PHO 3 coding sequences, with different word sizes. SLU Global Bioinformatics Centre
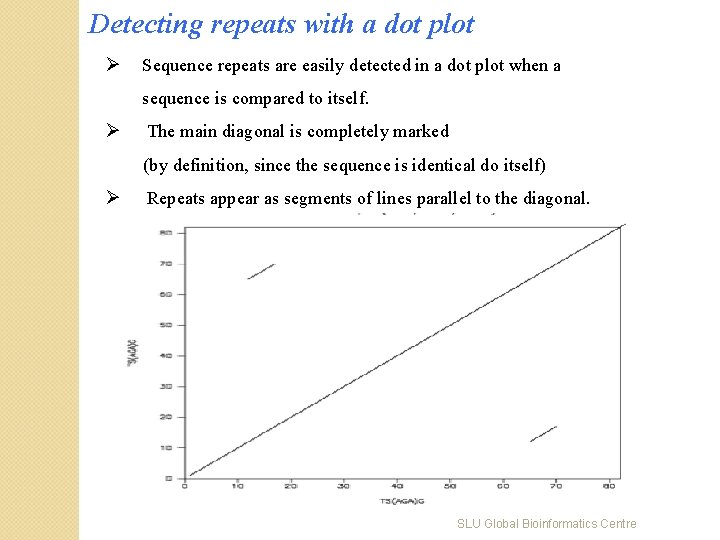
Detecting repeats with a dot plot Ø Sequence repeats are easily detected in a dot plot when a sequence is compared to itself. Ø The main diagonal is completely marked (by definition, since the sequence is identical do itself) Ø Repeats appear as segments of lines parallel to the diagonal. SLU Global Bioinformatics Centre

Plotorf >SEQ 1. fasta ATGGGTCGTGAAG AGAATGCTCCTCC TTTGGAATCTTAA >SEQ 2. fasta ATGGCTCCTCCCT TAGAATCTTAG Unix% plotorf SEQ 1. fasta –stop TAA, TAG –out GA. plot & Unix% getorf SEQ 1. fasta –minsize 5 –table 0 –find 1 – out GA. getorf & SLU Global Bioinformatics Centre
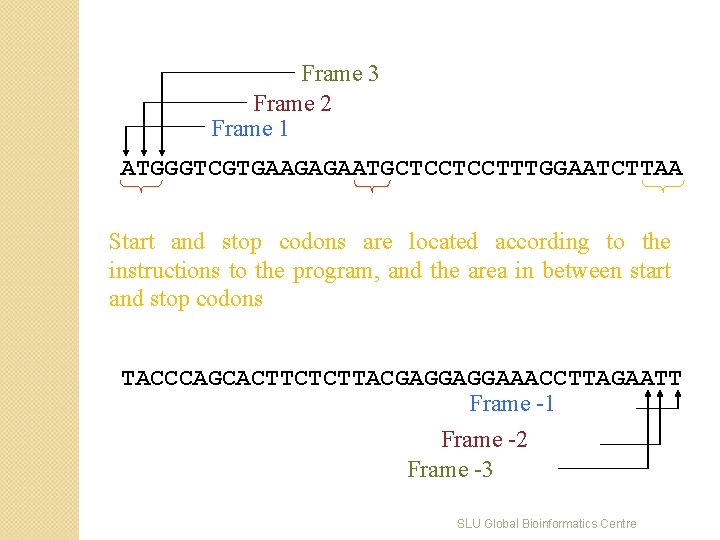
Frame 3 Frame 2 Frame 1 ATGGGTCGTGAAGAGAATGCTCCTCCTTTGGAATCTTAA Start and stop codons are located according to the instructions to the program, and the area in between start and stop codons TACCCAGCACTTCTCTTACGAGGAGGAAACCTTAGAATT Frame -1 Frame -2 Frame -3 SLU Global Bioinformatics Centre
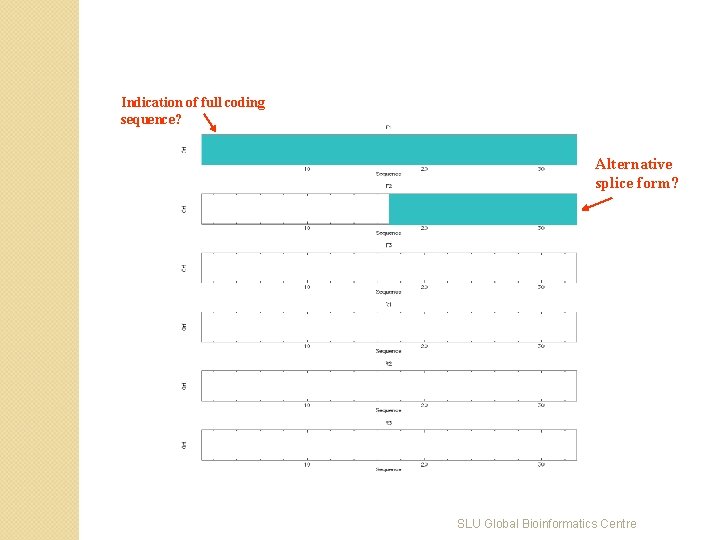
Indication of full coding sequence? Alternative splice form? SLU Global Bioinformatics Centre
![Using getorf: >_1 [17 - 37] MLLLWNL start methionine >_2 [1 - 36] stop Using getorf: >_1 [17 - 37] MLLLWNL start methionine >_2 [1 - 36] stop](http://slidetodoc.com/presentation_image_h/25a1855f7f06a970af53f9ea255a5f9c/image-21.jpg)
Using getorf: >_1 [17 - 37] MLLLWNL start methionine >_2 [1 - 36] stop codon MGREENAPPLES* SLU Global Bioinformatics Centre

Unix% transeq SEQ 1. fasta –frame 1 –table 0 –sbegin 4 – send 33 -out GA. fasta & >GA. fasta GREENAPPLES SLU Global Bioinformatics Centre
Alignments >GA. fasta GREENAPPLES >A. fasta APPLES For a global alignment: Unix% needle GA. fasta –gapopen 10 –gapextend 0. 5 –matrix EPAM 250 & For a local alignment: Unix% water GA. fasta –gapopen 10 –gapextend 0. 5 –matrix EPAM 250 & SLU Global Bioinformatics Centre

Alignments … To align two or more sequences in a biologically significant way. GREENAPPLES Gap penalty = 10; Extension penalty = 0. 5 Local (water) Global (needle) APPLES GREENAPPLES SLU Global Bioinformatics Centre
GREENAPPLES looks like the “apples” motif may be part of a larger domain APPLES physicochemical properties pattern searching SLU Global Bioinformatics Centre

Physico-chemical properties Isoelectric point Unix% iep GA. fasta –plot -step 0. 5 –out GA. IEP & General properties Unix% pepinfo GA. fasta –hwindow 8 –generalplot –hydropathyplot & SLU Global Bioinformatics Centre
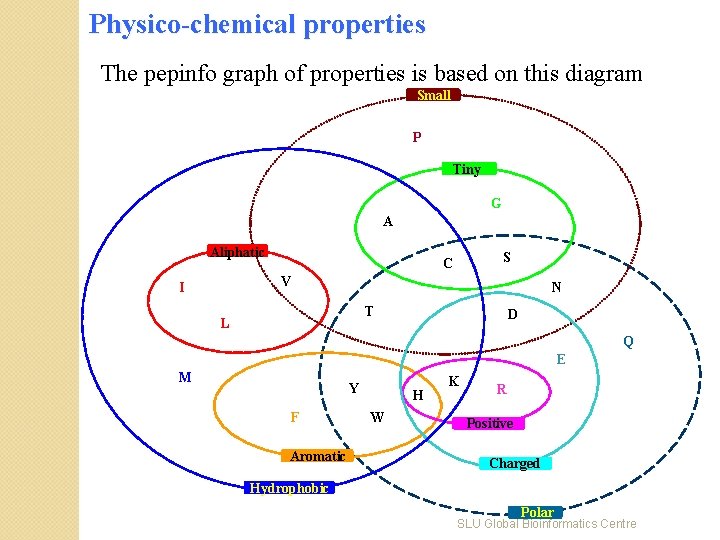
Physico-chemical properties The pepinfo graph of properties is based on this diagram Small P Tiny G A Aliphatic S C V I N T L D Q E M Y F Aromatic H W K R Positive Charged Hydrophobic Polar SLU Global Bioinformatics Centre
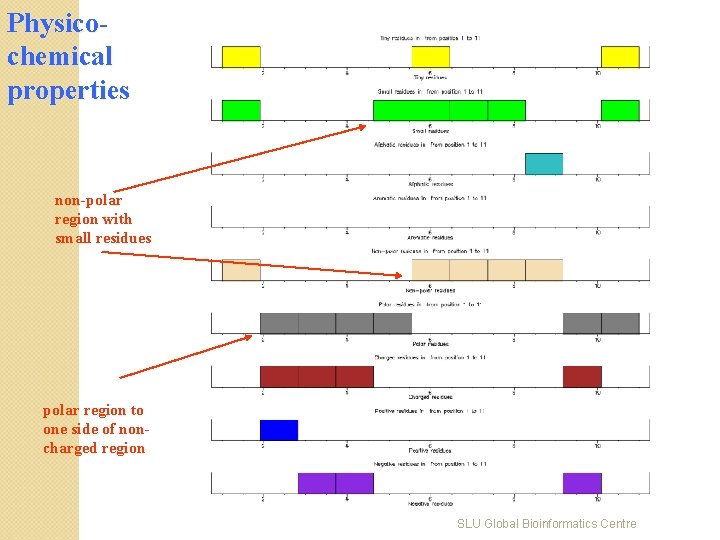
Physicochemical properties non-polar region with small residues polar region to one side of noncharged region SLU Global Bioinformatics Centre
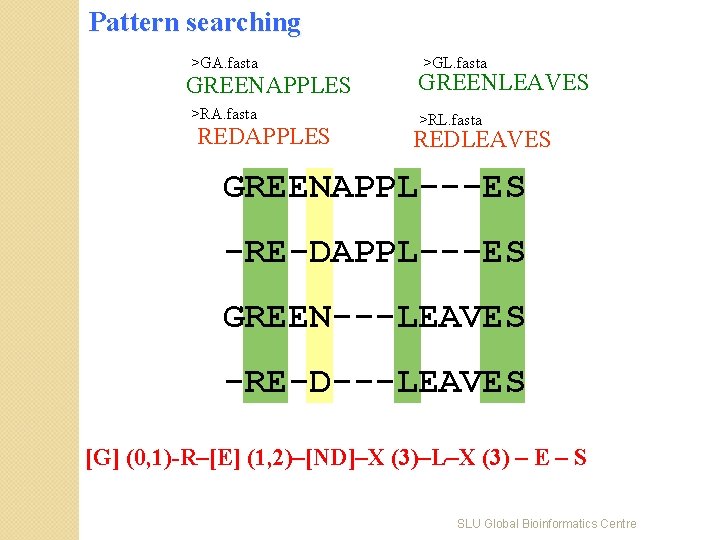
Pattern searching >GA. fasta GREENAPPLES >RA. fasta REDAPPLES >GL. fasta GREENLEAVES >RL. fasta REDLEAVES GREENAPPL---ES -RE-DAPPL---ES GREEN---LEAVES -RE-D---LEAVES [G] (0, 1)-R–[E] (1, 2)–[ND]–X (3)–L–X (3) – E – S SLU Global Bioinformatics Centre
![Pattern searching pattern. fruit [G] (0, 1) - [R] – [E] (1, 2) – Pattern searching pattern. fruit [G] (0, 1) - [R] – [E] (1, 2) –](http://slidetodoc.com/presentation_image_h/25a1855f7f06a970af53f9ea255a5f9c/image-30.jpg)
Pattern searching pattern. fruit [G] (0, 1) - [R] – [E] (1, 2) – [ND] –x (3) – [L] –x (3) – [E] – [S] Search a protein database: Unix% fuzzpro sptr: * pattern. fruit –mismatch 0 –out GA. fuzzpro & Nothing resembling this pattern is found in the database - But we could try scanning PRINTS (pscan) and PROSTIE (patmatmotifs) with one of our sequences. SLU Global Bioinformatics Centre

Some Programs SLU Global Bioinformatics Centre

Some Programs … SLU Global Bioinformatics Centre

More Information SLU Global Bioinformatics Centre
- Slides: 33