Bioinformatics databases sequence retrieval Content of lecture I

Bioinformatics databases & sequence retrieval Content of lecture I. Introduction II. Bioinformatics data & databases III. Sequence Retrieval with MRS Celia van Gelder CMBI UMC Radboud September 2013

I. Bioinformatics questions Lookup • Is the gene known for my protein (or vice versa)? • What sequence patterns are present in my protein? • To what class or family does my protein belong? Compare • Are there sequences in the database which resemble the protein I cloned? • How can I optimally align the members of this protein family? Predict • Can I predict the active site residues of this enzyme? • Can I predict a (better) drug for this target? • How can I predict the genes located on this genome? ©CMBI 2009

Sequence similarity Image, you sequenced this human protein. MVVSGAPPAL GGGCLGTFTS LLLLASTAIL NAARIPVPPA CGKPQQLNRV VGGEDSTDSE WPWIVSIQKN GTHHCAGSLL TSRWVITAAH CFKDNLNKPY LFSVLLGAWQ LGNPGSRSQK VGVAWVEPHP VYSWKEGACA DIALVRLERS IQFSERVLPI CLPDASIHLP PNTHCWISGW GSIQDGVPLP HPQTLQKLKV PIIDSEVCSH LYWRGAGQGP ITEDMLCAGY LEGERDACLG DSGGPLMCQV DGAWLLAGII SWGEGCAERN RPGVYISLSA HRSWVEKIVQ GVQLRGRAQG You know it is a serine protease. Which residues belong to the active site? Is its sequence similar to the mouse serine protease? ©CMBI 2009
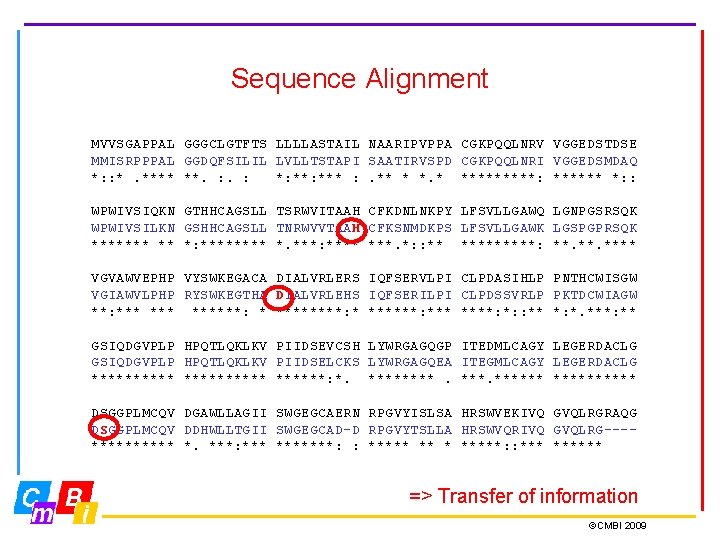
Sequence Alignment MVVSGAPPAL GGGCLGTFTS LLLLASTAIL NAARIPVPPA CGKPQQLNRV VGGEDSTDSE MMISRPPPAL GGDQFSILIL LVLLTSTAPI SAATIRVSPD CGKPQQLNRI VGGEDSMDAQ *: : *. **** **. : *: *** : . ** * *****: ****** *: : WPWIVSIQKN GTHHCAGSLL TSRWVITAAH CFKDNLNKPY LFSVLLGAWQ LGNPGSRSQK WPWIVSILKN GSHHCAGSLL TNRWVVTAAH CFKSNMDKPS LFSVLLGAWK LGSPGPRSQK ******* ** *: **** *. ***: **** ***. *: : ** *****: **. **** VGVAWVEPHP VYSWKEGACA DIALVRLERS IQFSERVLPI CLPDASIHLP PNTHCWISGW VGIAWVLPHP RYSWKEGTHA DIALVRLEHS IQFSERILPI CLPDSSVRLP PKTDCWIAGW **: *** ******: ****: *: : ** *: *. ***: ** GSIQDGVPLP HPQTLQKLKV PIIDSEVCSH LYWRGAGQGP ITEDMLCAGY LEGERDACLG GSIQDGVPLP HPQTLQKLKV PIIDSELCKS LYWRGAGQEA ITEGMLCAGY LEGERDACLG **********: *. ****** DSGGPLMCQV DGAWLLAGII SWGEGCAERN RPGVYISLSA HRSWVEKIVQ GVQLRGRAQG DSGGPLMCQV DDHWLLTGII SWGEGCAD-D RPGVYTSLLA HRSWVQRIVQ GVQLRG---***** *. ***: *******: : ***** *****: : ****** => Transfer of information ©CMBI 2009
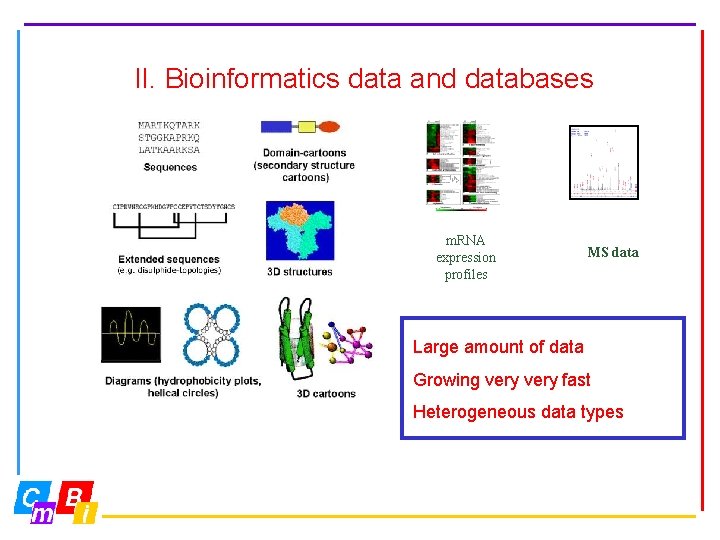
II. Bioinformatics data and databases m. RNA expression profiles MS data Large amount of data Growing very fast Heterogeneous data types
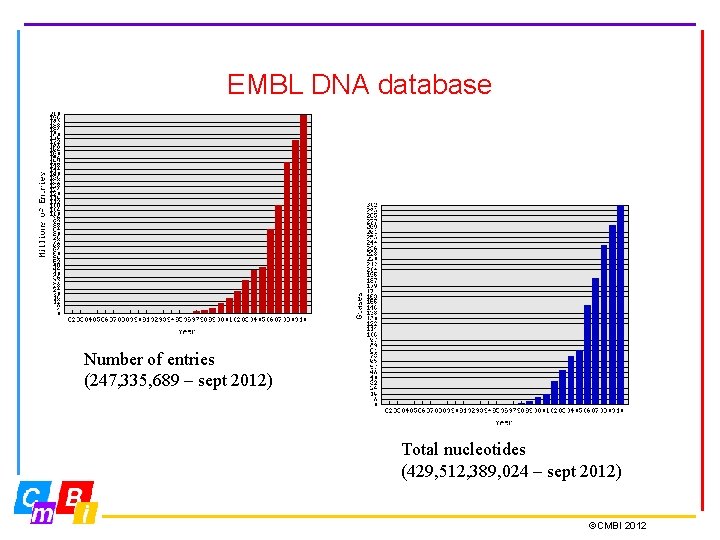
EMBL DNA database Number of entries (247, 335, 689 – sept 2012) Total nucleotides (429, 512, 389, 024 – sept 2012) ©CMBI 2012
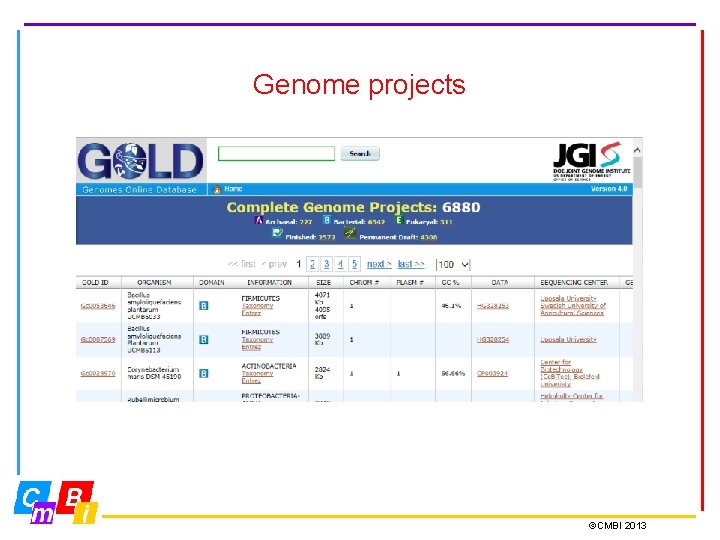
Genome projects ©CMBI 2013

Biological databases (1) Primary databases contain biomolecular sequences or structures (experimental data!) and associated annotation information Sequences Nucleic acid sequences Protein sequences EMBL, Genbank, DDBJ Swiss. Prot, tr. EMBL, Uni. Prot Structures Protein Structures PDB Structures of small compounds CSD Genomes Ensembl UCSC ©CMBI 2010

Biological databases (2) Secondary databases Contain data derived from primary database(s) Patterns, motifs, domains PROSITE, PFAM, PRINTS, INTERPRO, . . . Disease mutations OMIM / MIM SNPs db. SNP Pathways KEGG ©CMBI 2009

Databases Data must be in a certain format for software to recognize Every database can have its own format but some data elements are essential for every database: 1. Unique identifier, or accession code 2. Name of depositor 3. Literature references 4. Deposition date 5. The real data ©CMBI 2009

Quality of Data Swiss. Prot • Data is only entered by annotation experts EMBL, PDB • “Everybody” can submit data • No human intervention when submitted; some automatic checks ©CMBI 2009

Swiss. Prot database • Database of protein sequences • 537. 505 sequence entries (sept 2012) • Swissprot is curated annotated manually!! • Obligatory deposit of in Swiss. Prot before publication • Swiss. Prot is part of Uni. Prot • The other main part of Uni. Prot is Trembl is automatically annotated and is not reviewed. ©CMBI 2013

Important records in Swiss. Prot (1) ID AC DT DT DT HBA_HUMAN Reviewed; 142 AA. P 69905; P 01922; Q 3 MIF 5; Q 96 KF 1; Q 9 NYR 7; 21 -JUL-1986, integrated into Uni. Prot. KB/Swiss-Prot. 23 -JAN-2007, sequence version 2. 23 -SEP-2008, entry version 63. DE Rec. Name: Full=Hemoglobin subunit alpha; DE Alt. Name: Full=Hemoglobin alpha chain; DE Alt. Name: Full=Alpha-globin; ©CMBI 2009

Important records in Swiss. Prot (2) Cross references section: Hyperlinks to all entries in other databases which are relevant for the protein sequence HBA_HUMAN genes & m. RNA diseases protein domains structures ©CMBI 2009

Important records in Swiss. Prot (3) Features section: post-translational modifications, signal peptides, binding sites, enzyme active sites, domains, disulfide bridges, local secondary structure, sequence conflicts between references etc. ©CMBI 2011

And finally, the amino acid sequence! ©CMBI 2009

EMBL database Nucleotide database EMBL: 247 million sequence entries comprising 429 billion nucleotides (Sept 2012) EMBL records follows roughly same scheme as Swiss. Prot Obligatory deposit of sequence in EMBL before publication Most EMBL sequences never seen by a human ©CMBI 2013

Protein Data Bank (PDB) Databank for 3 -dimensional structures of biomolecules (by X-ray & NMR): • • Protein DNA RNA Ligands Obligatory deposit of coordinates in the PDB before publication ~ 84000 entries (Sep 2012) ( ~6000 “unique” structures) PDB file is a keyword-organised flat-file (80 column) 1) human readable 2) every line starts with a keyword (3 -6 letters) 3) platform independent ©CMBI 2011

PDB important records (1) PDB nomenclature Filename= accession number= PDB Code Filename is 4 positions (often 1 digit & 3 letters, e. g. 1 CRN) HEADER describes molecule & gives deposition date HEADER PLANT SEED PROTEIN 30 -APR-81 1 CRN CMPND name of molecule COMPND CRAMBIN SOURCE organism SOURCE ABYSSINIAN CABBAGE (CRAMBE ABYSSINICA) SEED ©CMBI 2009

PDB important records (2) SEQRES Sequence of protein; be aware: Not always all 3 d-coordinates are present for all the amino acids in SEQRES!! SEQRES 1 46 THR CYS PRO SER ILE VAL ALA ARG SER ASN PHE 1 CRN 51 SEQRES 2 46 ASN VAL CYS ARG LEU PRO GLY THR PRO GLU ALA ILE CYS 1 CRN 52 SEQRES 3 46 ALA THR TYR THR GLY CYS ILE ILE PRO GLY ALA THR 1 CRN 53 SEQRES 4 46 CYS PRO GLY ASP TYR ALA ASN 1 CRN 54 SSBOND disulfide bridges SSBOND 1 CYS 3 CYS 40 SSBOND 2 CYS 4 CYS 32 ©CMBI 2009

PDB important records (3) and at the end of the PDB file the “real” data: ATOM one line for each atom with its unique name and its x, y, z coordinates ATOM 1 N THR 1 17. 047 14. 099 3. 625 1. 00 13. 79 1 CRN 70 ATOM 2 CA THR 1 16. 967 12. 784 4. 338 1. 00 10. 80 1 CRN 71 ATOM 3 C THR 1 15. 685 12. 755 5. 133 1. 00 9. 19 1 CRN 72 ATOM 4 O THR 1 15. 268 13. 825 5. 594 1. 00 9. 85 1 CRN 73 ATOM 5 CB THR 1 18. 170 12. 703 5. 337 1. 00 13. 02 1 CRN 74 ATOM 6 OG 1 THR 1 19. 334 12. 829 4. 463 1. 00 15. 06 1 CRN 75 ATOM 7 CG 2 THR 1 18. 150 11. 546 6. 304 1. 00 14. 23 1 CRN 76 ATOM 8 N THR 2 15. 115 11. 555 5. 265 1. 00 7. 81 1 CRN 77 ATOM 9 CA THR 2 13. 856 11. 469 6. 066 1. 00 8. 31 1 CRN 78 ATOM 10 C THR 2 14. 164 10. 785 7. 379 1. 00 5. 80 1 CRN 79 ATOM 11 O THR 2 14. 993 9. 862 7. 443 1. 00 6. 94 1 CRN 80 ©CMBI 2009

Structure Visualization Structures from PDB can be visualized with: 1. Yasara (www. yasara. org) 2. Swiss. PDBViewer (http: //spdbv. vital-it. ch/) 3. Protein Explorer (http: //www. umass. edu/microbio/rasmol/) 4. Cn 3 D (http: //www. ncbi. nlm. nih. gov/Structure/CN 3 D/cn 3 d. shtml) ©CMBI 2009

Part III: Sequence Retrieval with MRS Google Thé best generic search and retrieval system Google searches everywhere for everything MRS Maarten’s Retrieval System (http: //mrs. cmbi. ru. nl ) MRS searches in selected data environments MRS is the Google of the biological database world Search engine (like Google) • Input/Query = word(s) • Output = entry/entries from database Other programs exist: Entrez, SRS, . . ©CMBI 2009

MRS Search Steps • Select database(s) of choice • Formulate your query • Hit “Search” • The result is a “query set” or “hitlist” • Analyze the results ©CMBI 2009

http: //mrs. cmbi. ru. nl ©CMBI 2011
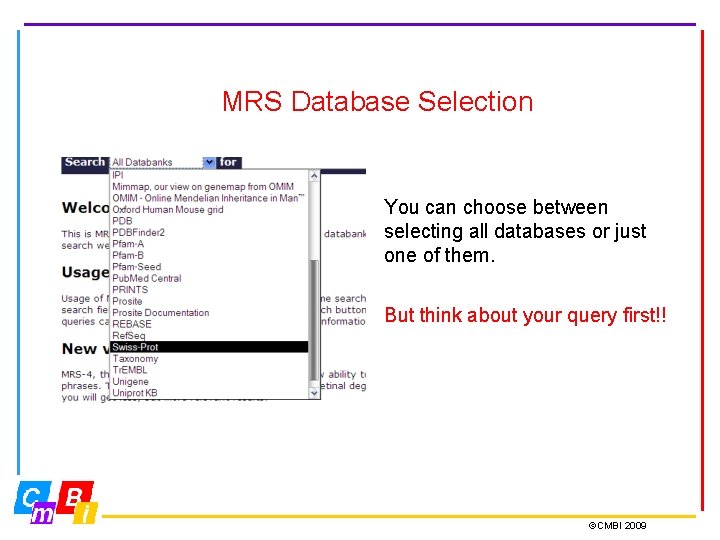
MRS Database Selection You can choose between selecting all databases or just one of them. But think about your query first!! ©CMBI 2009
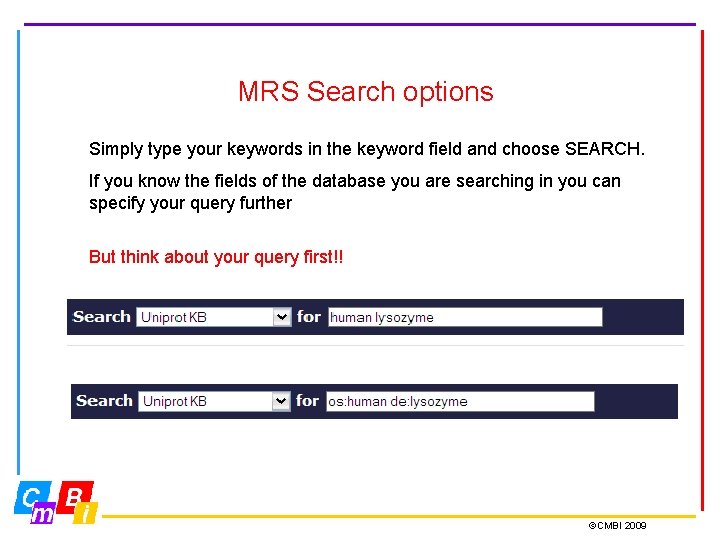
MRS Search options Simply type your keywords in the keyword field and choose SEARCH. If you know the fields of the database you are searching in you can specify your query further But think about your query first!! ©CMBI 2009
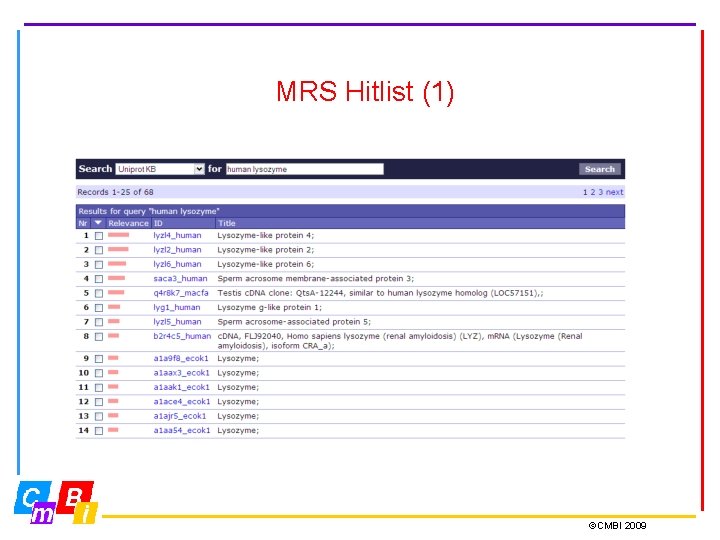
MRS Hitlist (1) ©CMBI 2009
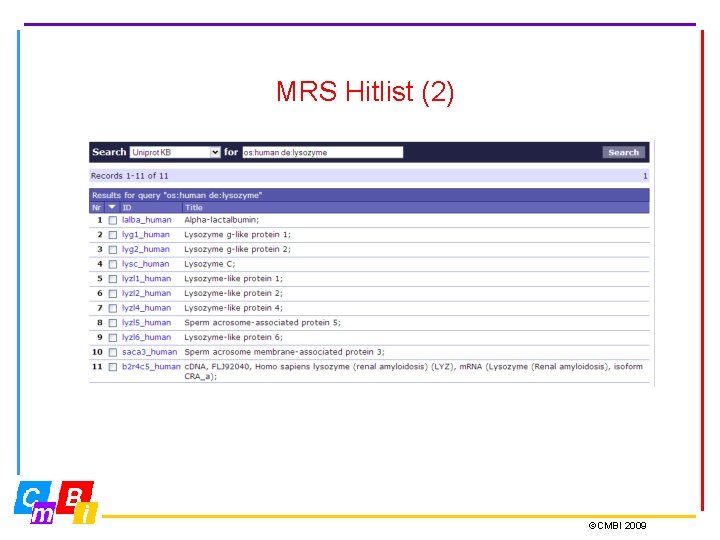
MRS Hitlist (2) ©CMBI 2009

MRS Options MRS creates a result, or a “query set”, or “hitlist”. With the result you can do different things in MRS: – View the hits – Blast single hit sequences – Clustal multiple hit sequences ©CMBI 2009
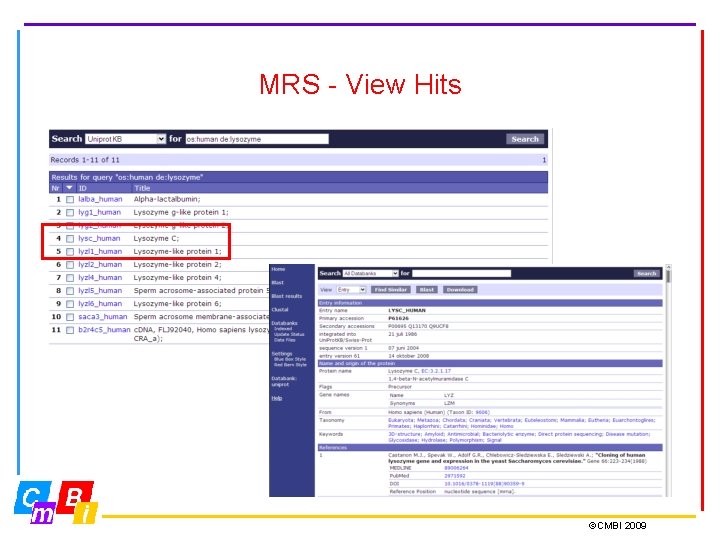
MRS - View Hits ©CMBI 2009
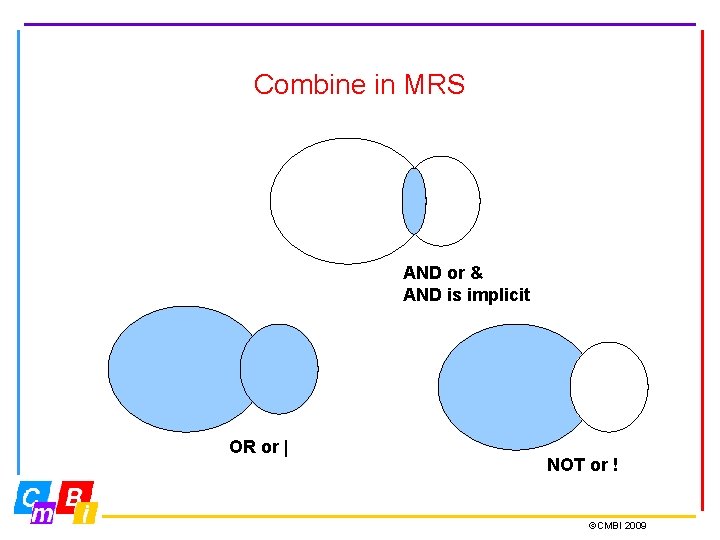
Combine in MRS AND or & AND is implicit OR or | NOT or ! ©CMBI 2009
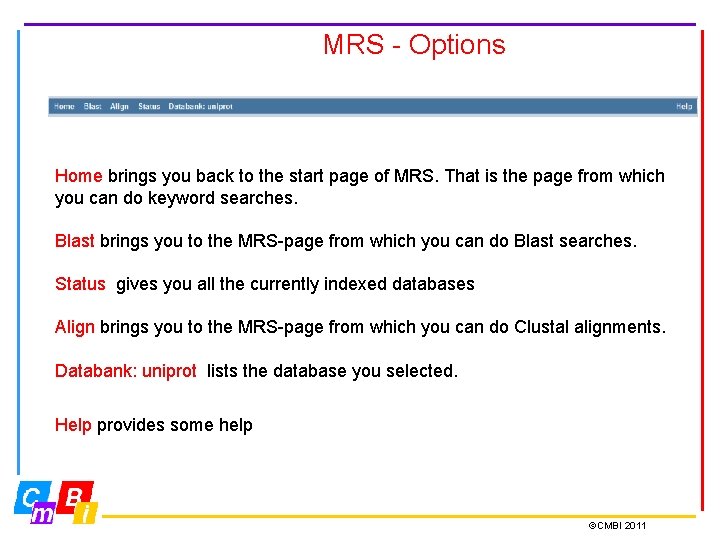
MRS - Options Home brings you back to the start page of MRS. That is the page from which you can do keyword searches. Blast brings you to the MRS-page from which you can do Blast searches. Status gives you all the currently indexed databases Align brings you to the MRS-page from which you can do Clustal alignments. Databank: uniprot lists the database you selected. Help provides some help ©CMBI 2011

Try it yourself with the exercises! Ground rules for bioinformatics Don't always believe what programs tell you - they're often misleading & sometimes wrong! Don't always believe what databases tell you - they're often misleading & sometimes wrong! Don't always believe what lecturers tell you - they're sometimes wrong! Don't be a naive user, computers don’t do biology & bioinformatics, you do! free after Terri Attwood ©CMBI 2009
- Slides: 34