CAP 5510 Bioinformatics Database Searches for Biological Sequences

CAP 5510 – Bioinformatics Database Searches for Biological Sequences Tamer Kahveci CISE Department University of Florida 1

Goals • Understand how major heuristic methods for sequence comparison work – FASTA – BLAST • Understand how search results are evaluated 2
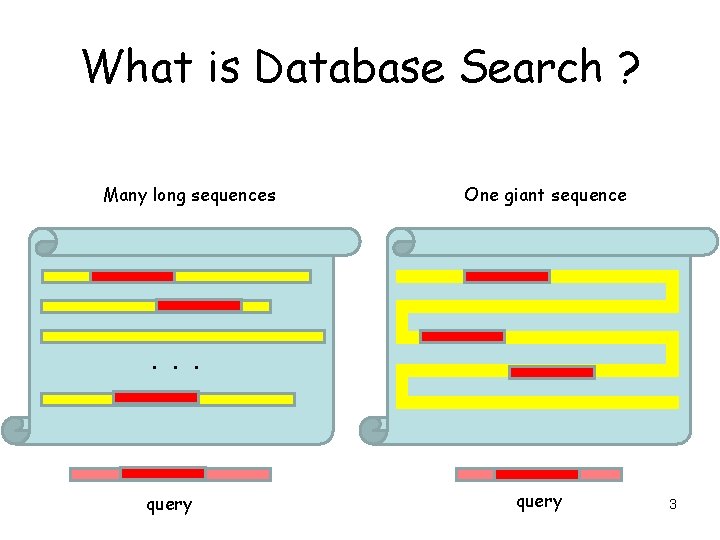
What is Database Search ? Many long sequences One giant sequence . . . query 3

What is Database Search ? Two giant sequences 4

What is Database Search ? • Find a particular (usually) short sequence in a database of sequences (or one huge sequence). • Problem is identical to local sequence alignment, but on a much larger scale. • We must also have some idea of the significance of a database hit. – Databases always return some kind of hit, how much attention should be paid to the result? • A similar problem is the global alignment of two large sequences • General idea: good alignments contain high scoring regions. 5

Database Search Issues • How can we search massive space quickly? • How can we evaluate the significance of the result? 6

Database Search Methods • Hash table based methods – FASTA family • FASTP, FASTA, TFASTA, FASTAX, FASTAY – BLAST family • BLASTP, BLASTN, TBLAST, BLASTX, BLAT, BLASTZ, Mega. BLAST, Psi. BLAST, Phi. BLAST – Others • FLASH, Pattern. Hunter, SSAHA, SENSEI, WABA, GLASS • Suffix tree based methods – Mummer, AVID, Reputer, MGA, QUASAR 7
Hash Table 8
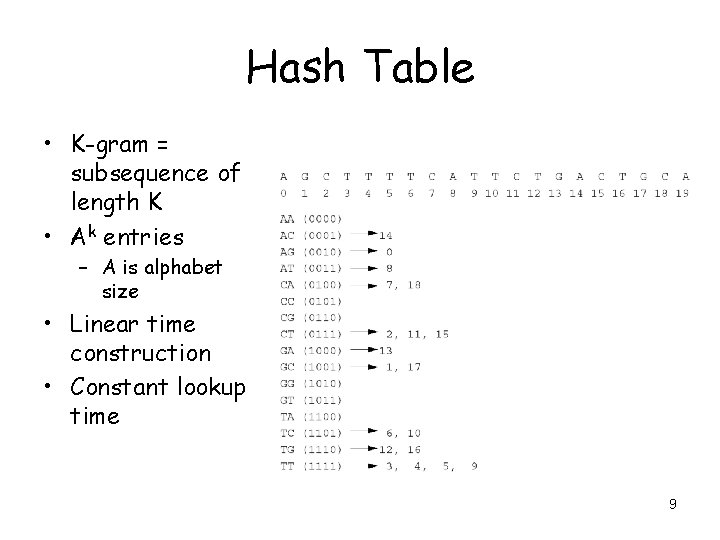
Hash Table • K-gram = subsequence of length K • Ak entries – A is alphabet size • Linear time construction • Constant lookup time 9

FASTP Lipman & Pearson, 1985 10

FASTP • Three phase algorithm 1. Find short good matches using kgrams 1. K = 1 or 2 2. Find start and end positions for good matches 3. Use DP to align good matches 11

FASTP: Phase 1 (1) position 1 2 3 4 5 6 7 8 9 10 11 protein 1 n c s p t a. . . protein 2. . . a c s p r k position in offset amino acid protein A protein B pos A - pos. B --------------------------a 6 6 0 c 2 7 -5 k 11 n 1 p 4 9 -5 r 10 s 3 8 -5 t 5 --------------------------Note the common offset for the 3 amino acids c, s and p A possible alignment can be quickly found : protein 1 n c s p t a | | | protein 2 a c s p r k 12
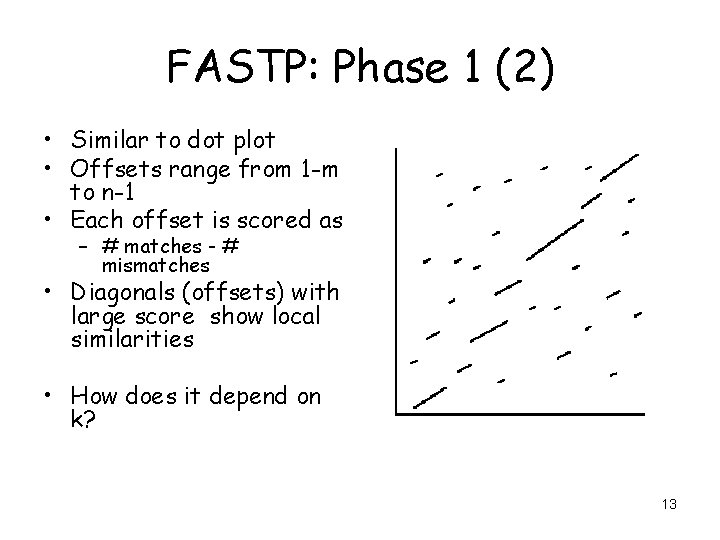
FASTP: Phase 1 (2) • Similar to dot plot • Offsets range from 1 -m to n-1 • Each offset is scored as – # matches - # mismatches • Diagonals (offsets) with large score show local similarities • How does it depend on k? 13
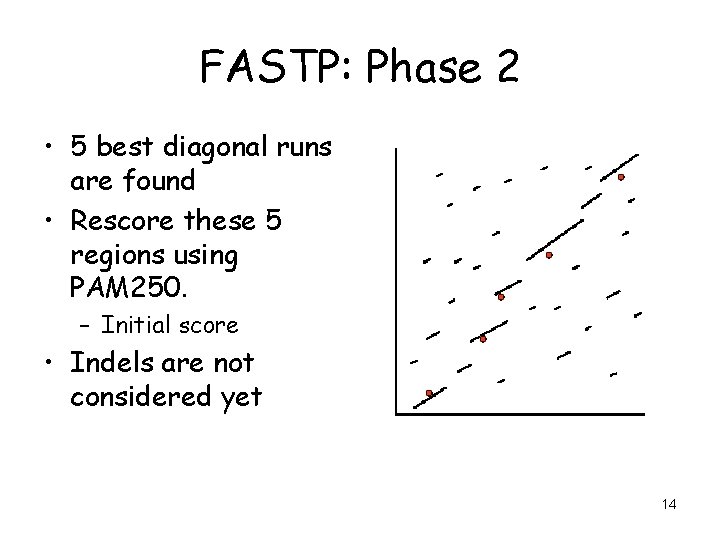
FASTP: Phase 2 • 5 best diagonal runs are found • Rescore these 5 regions using PAM 250. – Initial score • Indels are not considered yet 14

FASTP: Phase 3 • Sort the aligned regions in descending score • Optimize these alignments using Needleman-Wunsch • Report the results 15

FASTP - Discussion • Results are not optimal. Why ? • How does performance compare to Smith. Waterman? • What is the impact of k? • How does this idea work for DNAs ? – K = 4 or 6 for DNA 16

FASTA – Improvement Over FASTP Pearson 1995 17

FASTA (1) • Phase 2: Choose 10 best diagonal runs instead of 5 18
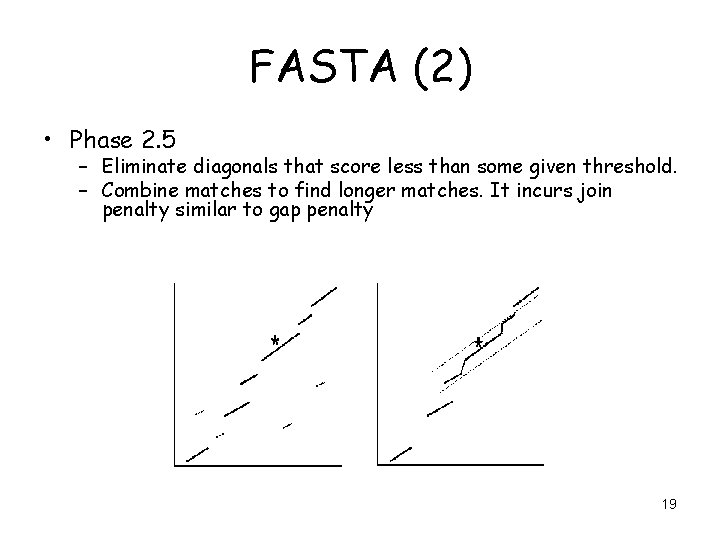
FASTA (2) • Phase 2. 5 – Eliminate diagonals that score less than some given threshold. – Combine matches to find longer matches. It incurs join penalty similar to gap penalty 19

BLAST Altschul, Gish, Miller, Myers, Lipman, 1990 20

BLAST (or BLASTP) • BLAST – Basic Local Alignment Search Tool • An approximation of Smith-Waterman • Designed for database searches – Short query sequence against long database sequence or a database of many sequences • Sacrifices search sensitivity for speed 21

BLAST Algorithm (1) • Eliminate low complexity regions from the query sequence. – Replace them with X (protein) or N (DNA) • Hash table on query sequence. – K = 3 for proteins MCGPFILGTYC CGP MCG 22
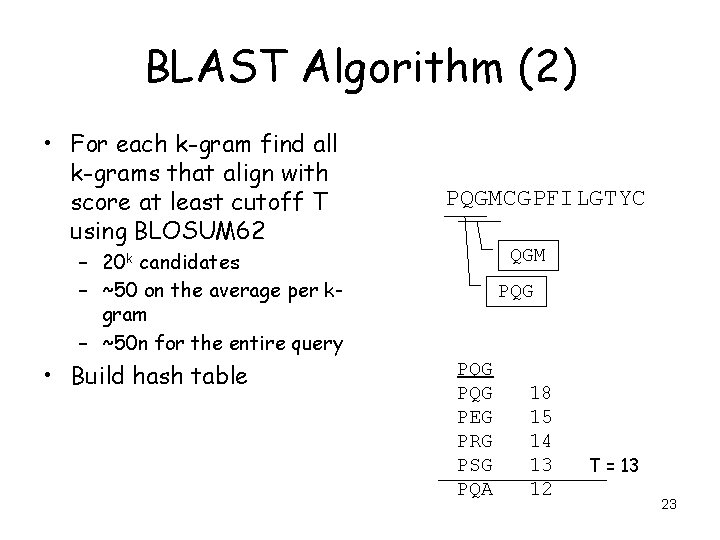
BLAST Algorithm (2) • For each k-gram find all k-grams that align with score at least cutoff T using BLOSUM 62 PQGMCGPFILGTYC QGM – 20 k candidates – ~50 on the average per kgram – ~50 n for the entire query • Build hash table PQG PQG PEG PRG PSG PQA 18 15 14 13 12 T = 13 23

BLAST Algorithm (3) • Sequentially scan the database and locate each k-gram in the hash table • Each match is a seed for an ungapped alignment. 24

BLAST Algorithm (4) • HSP (High Scoring Pair) = A match between a query word and the database • Find a “hit”: Two nonoverlapping HSP’s on a diagonal within distance A • Extend the hit until the score falls below a threshold value, X 25

BLAST Algorithm (5) • Keep only the extended matches that have a score at least S. • Determine the statistical significance of the result 26

What is Statistical Significance? • Two one-on-one games, two scores. 13 : 15 • Which result is more significant? • Expected: maybe a random result. • Unexpected: significant, may have significant meanings. 13 : 15 27

Statistical Significance • E-value: The expected number of matches with score at least S • • E = Kmne-lambda. S m, n : sequence lengths S : alignment score K, lambda: normalization parameters • P-value: The probability of having at least one match with score at least S • 1 – e-E • The smaller these values are, the more significant the result • http: //www. ncbi. nlm. nih. gov/Education/BLASTinfo/glossary 2. ht ml 28

BLAST - Analysis • K (k-gram) – Lower: more sensitive. Slower. • T (neighbor cutoff) – Lower: Find distant neighbors. Introduces noise • X (extension cutoff) – Higher: lower chances of getting into a local minima. Slower. 29

Sample Query • http: //www. ncbi. nlm. nih. gov/BLAST/ Dhal_ecoli IDRAMSAARGVFERGDWSLSSPAKRKAVLNKLADLMEAH AEELALLETLDTGKPIRHSLRDDIPGAARAIRWYAEAIDK VYGEVATTSSHELAMIVREPVGVIAAIVPWNFPLLLTCW KLGPALAAGNSVILKPSEKSPLSAIRLAGLAKEAGLPDGVL NVVTGFGHEAGQALSRHNDIDAIAFTGSTRTGKQLLKDA GDSNMKRVWLEAGGKSANIVFADCPDLQQAASATAAGI FYNQGQVCIAGTRLLLEESIADEFLALLKQQAQNWQPG HPLDPATTMGTLIDCAHADSVHSFIREGESKGQLLLDGR NAGLAAAIGPTIFVDVDPNASLSREEIFGPVLVVTRFTSE EQALQLANDSQYGLGAAVWTRDLSRAHRMSRRLKAGSV FVNNYNDGDMTVPFGGYKQSGNGRDKSLHALEKFTELKT IWI 30

BLASTN • BLAST for nucleic acids • K = 11 • Exact match instead of neighborhood search. 31
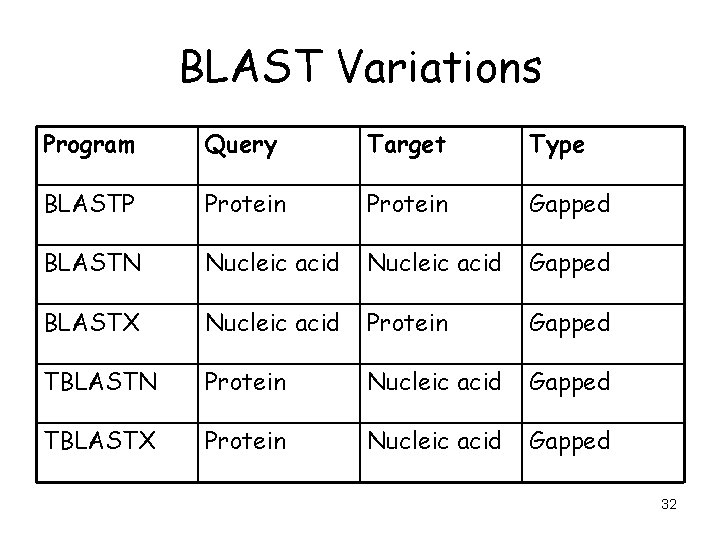
BLAST Variations Program Query Target Type BLASTP Protein Gapped BLASTN Nucleic acid Gapped BLASTX Nucleic acid Protein Gapped TBLASTN Protein Nucleic acid Gapped TBLASTX Protein Nucleic acid Gapped 32

Even More Variations – Psi. BLAST (iterative) – BLAT, BLASTZ, Mega. BLAST – FLASH, Pattern. Hunter, SSAHA, SENSEI, WABA, GLASS – Main differences are • Seed choice (k, gapped seeds) • Additional data structures 33

Suffix Trees 34

Suffix Tree • Tree structure that contains all suffixes of the input sequence • • • TGAGTGCGA TGCGA CGA GA A 35
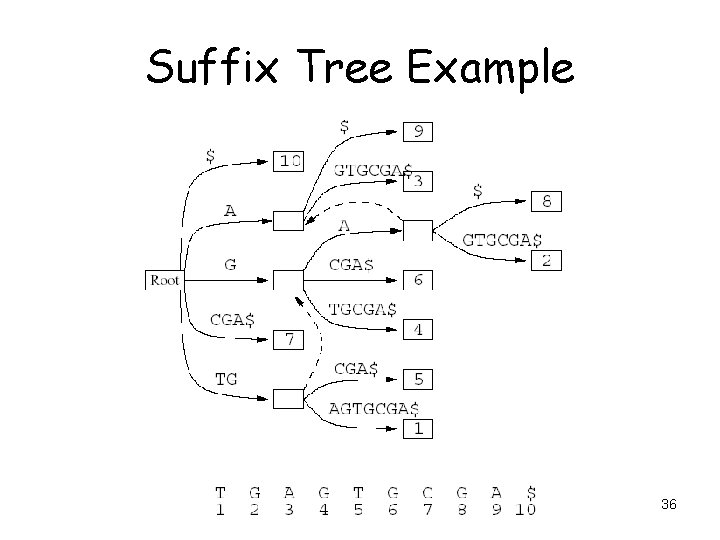
Suffix Tree Example 36

Suffix Tree Analysis • O(n) space and construction time – 10 n to 70 n space usage reported • O(m) search time for m-letter sequence • Good for – Small data – Exact matches 37
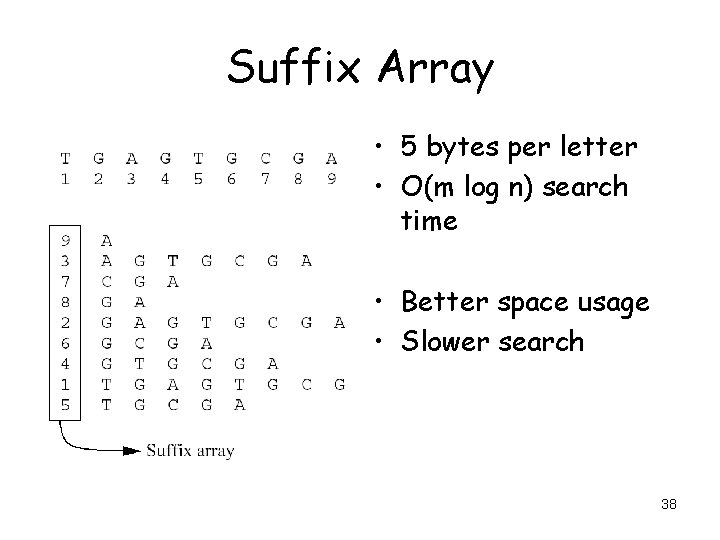
Suffix Array • 5 bytes per letter • O(m log n) search time • Better space usage • Slower search 38

Mummer 39

Other Sequence Comparison Tools • Reputer, MGA, AVID • QUASAR (suffix array) 40
- Slides: 40