Computational Molecular Biology Multiple Sequence Alignment Sequence Alignment

Computational Molecular Biology Multiple Sequence Alignment

Sequence Alignment Ø Problem Definition: q Given: 2 DNA or protein sequences q Find: Best match between them Ø What is an Alignment: q Given: 2 Strings S and S’ q Goal: The lengths of S and S’ are the same by inserting spaces (--; sometimes denote as ∆) into these strings A -- T C -- A -- C T C A A My T. Thai mythai@cise. ufl. edu 2

Matches, Mismatches and Indels Ø Match: two aligned, identical characters in an alignment Ø Mismatch: two aligned, unequal characters Ø Indel: A character aligned with a space A A C T -- C C T A A C T -- -- C T C C T A C C T -- -- T A C T T T 10 matches, 2 mismatches, 7 indels My T. Thai mythai@cise. ufl. edu 3

Basic Algorithmic Problem Ø Find the alignment of the two strings that: q max m where m = (# matches – mismatches – indels) q Or min m where m is the SP-score of an alignment Ø m defines the similarity of the two strings, also called Optimal Global Alignment Ø Biologically: a mismatch represents a mutation, whereas an indel represents a historical insertion or deletion of a single character My T. Thai mythai@cise. ufl. edu 4

Multiple Sequence Alignment Ø Problem Definition: q Similar to the sequence alignment problem but the input has more than 2 strings Ø Challenges: q NP-hard, MAX-SNP q Guarantee factor: 2 – 2/k where k is the number of the input sequences. q More work to reduce the time and space complexity My T. Thai mythai@cise. ufl. edu 5

Sum of Pairs Score (SP-Score) Ø Given a finite alphabet and where ∆ denotes a space Ø Consider k sequences over that we want to align. After an alignment, each sequence has length l Ø A score d is assigned to each pair of letters: My T. Thai mythai@cise. ufl. edu 6

SP-Score Ø The SP-Score of an alignment A is defined as: q Consider a matrix of l columns and k rows where the rows represents the sequences and columns represent the letters q SP-Score is the sum of the scores of all columns: § Score of each column is the sum of the scores of all distinct unordered pairs of letters in the column q Or we can view as sum of pairwise sequence alignment values. Ø Find an (optimal) alignment to minimize the SP -Score value My T. Thai mythai@cise. ufl. edu 7

Proving MSA with SP-Score that is a Metric is NP-hard My T. Thai mythai@cise. ufl. edu 8

Some Notations My T. Thai mythai@cise. ufl. edu 9

Some Basic Properties Ø Lemma 1: Let s 1, s 2 be two sequences over Σ such that l 1=|s 1|, l 2=|s 2|, l 2≥l 1 and there are m symbols of s 1 that are not in s 2. Then every alignment of the set {s 1, s 2} has at least m+l 2 -l 1 mismatches My T. Thai mythai@cise. ufl. edu 10

My T. Thai mythai@cise. ufl. edu 11

The construction Ø Reduce the vertex cover (or node cover) to MSA. Ø Vertex cover: q Instance: A graph G=(V, E) and an integer k≤|V| q Question: Is there a vertex cover V 1 of G of size k or less? Ø MSA: q Instance: A set S={s 1, …, sn} of finite sequences over a fixed alphabet Σ, an SP-score and an integer C q Question: Is there a multiple alignment of the sequences in S that is of value C or less? My T. Thai mythai@cise. ufl. edu 12

SP-Score (alphabet of size 6) My T. Thai mythai@cise. ufl. edu 13

The Reduction So, we have , T is a set of C 2 sequences t and X contains C 1 sequences x(k), where C 1 and C 2 will be determined later My T. Thai mythai@cise. ufl. edu 14

An Example My T. Thai mythai@cise. ufl. edu 15

Intuition Ø By the above construction, an optimal alignment A of S is obtained when A satisfies certain properties (called standard alignment) Ø The value of standard alignment is bounded by a given threshold C only where G has a vertex cover of size k Ø How to obtain: q Force d’s of the test sequences to be aligned with b’s of the edge sequences q Only one b of each edge sequence can be aligned to a d q The number of such alignment determines the value of the alignment My T. Thai mythai@cise. ufl. edu 16

Standard Alignemnt My T. Thai mythai@cise. ufl. edu 17

My T. Thai mythai@cise. ufl. edu 18

My T. Thai mythai@cise. ufl. edu 19

My T. Thai mythai@cise. ufl. edu 20

My T. Thai mythai@cise. ufl. edu 21

Ø Let US and US, X denote the upper bounds of D(AS) and D(AS, X) respectively Ø By Corollary 8 and Lemma 9, we have the standard alignment has value not greater than DSD + US, X q where DSD = D(AX) + D(AT) + D(AX, T) + D(AS, T) over a standard alignment A Ø Now, let C 1 > US and C 2 > US + US, X, we can prove that an optimal alignment must be a standard one My T. Thai mythai@cise. ufl. edu 22

My T. Thai mythai@cise. ufl. edu 23

My T. Thai mythai@cise. ufl. edu 24

Show the NP-hardness of any scoring matrix in a broad class M Show that there is a scoring matrix M 0 such that MSA for M 0 is MAX-SNP hard My T. Thai mythai@cise. ufl. edu 25

Interesting Observation Ø Via the brute force, optimal MSA contains very few gaps Ø Suggesting the study of gap limitations: q Have an upper bound of the number of gaps one can insert during the alignment Ø Special case: q Gap-0: No gap allows, but we can shift the strings for an alignment (insert gaps at the beginning or at the end of a string) q Gap-0 -1: a gap-0 alignment such that the gaps at the beginning or at the end of each string is exactly one space My T. Thai mythai@cise. ufl. edu 26

Problem Definition Ø Given a finite alphabet Ø Scoring matrix q For i, j > 0, si, j represents the penalty for aligning ai with aj q For i > 0, s 0, i and si, 0 are called indel penalites q Gap opening penalties (in addition to the indel penalties) for aligning ai with the first or last ∆ in the string of ∆’s My T. Thai mythai@cise. ufl. edu 27

Generic Scoring Matrix Where Σ={A, T}, x, y, x are fixed nonnegative numbers and u > max{0, v. A, v. T} holds • Let M 2 be the class of all scoring matrices that contain a generic submatrix M • Let M 1 be the class of all scoring matrices that contain a sub-matrix isomorphic to a generic matrix M with z > v. T. • Let M be the class of all scoring matrices that contain a submatrix isomorphic to a generic matrix M with y > u and z > v. T. Theorem 1: (a) The gap-0 -1 multiple alignment problem is NP-hard for every scoring matrix M in M 2. (b) The gap-0 multiple alignment problem is NP-hard for every M in M 1 (c) The multiple alignment problem is NP-hard for every M in M Note that M is quite broad and covers most scoring schemes used in biological applications. My T. Thai mythai@cise. ufl. edu 28

Reduction Ø Reduce the MAX-CUT-B: q Given G=(V, E) where k=|V| and each vertex has a degree at most B q Find a partition of V into two disjoint sets such that to maximize the number of edges crossing these two sets Ø Given a graph G=(V, E) with k vertices v 0, …, vk -1 and l edges e 0, …, el-1. We will construct a set of k 2 sequences t 0, …, tk -1 as follows: 2 My T. Thai mythai@cise. ufl. edu 29

Reduction Ø For each vertex vi, construct a sequence ti such that q for each edge em={vh, vi} incident at vi, h < i, n < k 5, set where ti, j represents the character at the jth position in ti. q For other j, let ti, j = T Ø For i ≥ k, set ti = T T T … T with length k 12 l My T. Thai mythai@cise. ufl. edu 30

An Example My T. Thai mythai@cise. ufl. edu 31

Proof of Theorem 1(a) Ø We will show that a gap-0 -1 alignment will partition V into two disjoint subsets V 0 and V 1: q V 0: all vertices vi such that ti remains in place (a space appends at the end) q V 1: all vertices vi such that ti shifts to the right Ø Thus, based on the alignment, we can find the cut. And vice versa, based on the cut, we can find the alignment Ø The left part is: prove that if k is sufficiently large, the optimal gap-0 -1 alignment yields a partion of V with maximum edge cut. My T. Thai mythai@cise. ufl. edu 32

Proof of Theorem 1(a) Ø Let c denote the cut based on the alignment A Ø Consider all the sequences ti after that alignment A: q The total indel penalties is of order O(k 4) (appears at the first and last column in the SP score matrix) q The total number of mismatches before the alignment is 3 k 5 l(k 2 -1) q To maximally reduce this number: § § § 1 A-A match reduces 2 A-T mismatches For each edge (vh, vi), if there are in different subsets (of the partition), then a total of k 5 A-A matches between sequences th and ti are created No other A-T mismatches can be elimiated q Thus the SP-score: q k 12 lv. Tk 2(k 2 -1)2+3 k 5 l(u-v. T)(k 2 -1)-ck 5(2 u-v. A-v. T)+O(k 4) My T. Thai mythai@cise. ufl. edu 33

Theorem 2 Consider the following scoring matrix M 0 for the alphabet ∑ 0 = {A, T, C}. (a) The gap-0 -1 MSA problem is MAX-SNPhard (b) The gap-0 MSA problem in MAX-SNPhard My T. Thai (c) The MSA problem in MAX-SNP-hard mythai@cise. ufl. edu 34

MAX-SNP-hard Proof Ø To prove problem A’ is MAX-SNP-hard, we need to L-reduce problem A, which is MAXSNP-hard to A’ Ø L-reduce: q There are two polynomial-time algorithms f, g and constants a, b > 0 such that for each instance I of A: § f produces an instance I’ = f(I) of A’ such that OPT(I’) ≤ a. OPT(I) § Given any solution of I’ with cost c’, g produces a solution of I with cost c such that |c-OPT(I)| ≤ b|c’OPT(I’)| My T. Thai mythai@cise. ufl. edu 35

Proof of Theorem 2 Ø To prove MSA (with M 0 and the scoring matrix mentioned before) MAX-SNP-hard: q L-reduce the MAX-CUT-B to another optimization problem, called A’, which is L-reduce to a scaled version of MSA Ø Problem A’: q Given a graph G=(V, E) with bounded degree B. For every partition P={V 0, V 1}, let cp be the size of cut determined by P. q Find the partition P of V that minimizes dp = 3|E|2 cp My T. Thai mythai@cise. ufl. edu 36

Show A’ is MAX-SNP-hard Ø Let f and g be an identity function Ø Set a = 3 B and b = 2, we can easily prove the two properties of the L-reduction since: q cp ≥|E|/B and dp = 3|E| - 2 cp ≤ 3 |E| q Any increase of cp by 1 = decrease dp by 2 My T. Thai mythai@cise. ufl. edu 37

Show A’ L-reduce to scaled MSA Similar to the above construction, we have: My T. Thai mythai@cise. ufl. edu 38

Ø Similar to the proof of Theorem 1, we have the optimal SP-score: where Ø If the SP-score is scaled by a factor of k-5/2 for a MSA of k sequences, then A’ L-reduce to MSA. My T. Thai mythai@cise. ufl. edu 39

GENETIC ALGORITHMS

How do GAs work? Ø Create a population of random solutions Ø Use natural selection: § crossover and mutation to improve the solutions Ø Stop the operation if satisfying some certain criteria such as: § No improvement on fitness function § The improvement is less than some certain threshold § The number of iteration is more than some certain threhold

Terms and Definitions Ø Chromosomes q Potential solutions Ø Population q Collection of chromosomes Ø Generations q Successive populations

Terms and Definitions Ø Crossover q Exchange of genes between two chromosomes Ø Mutation q Random change of one or more genes in a chromosome Ø Elitism q Copy the best solutions without doing crossover or mutation.

Terms and Definitions Ø Offspring § New chromosome created by crossover between two parent chromosomes Ø Fitness function § Measures how “good” a chromosome is. Ø Encoding scheme § How do we represent every chromosome/gene? Binary, combination, syntax trees.

Why are GAs attractive? Ø No need for a particular algorithm to solve the given problem. Only the fitness function is required to evaluate the quality of the solutions. Ø Implicitly a parallel technique and can be implement efficiently on powerful parallel computers for demanding large scale problems.

Basic Outline of a GA Ø Initial population composed of random chromosomes, called first generation Ø Evaluate the fitness of each chromosome in the population Ø Create a new population: q Select two parent chromosomes from a population according to their fitness q Crossover (with some probability) to form a new offspring q Mutation (with some probability) to mutate new offspring q Place new offspring in a new population Ø Process is repeated until a satisfactory solution evolves

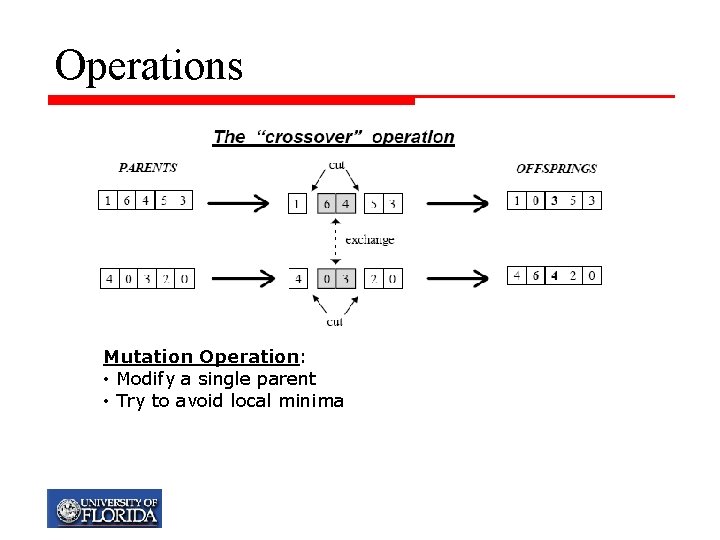
Operations Mutation Operation: • Modify a single parent • Try to avoid local minima

Let's see some running examples Ø Minimum of a function: q http: //cs. felk. cvut. cz/~xobitko/ga/example_f. html Ø Elitism: q http: //cs. felk. cvut. cz/~xobitko/ga/params. html Ø The travelling salesman problem: q http: //cs. felk. cvut. cz/~xobitko/ga/tspexample. htm l

Multiple Sequence Alignment Ø Fitness function is used to compare the different alignments q Based on the number of matching symbols and the number and size of gaps q Also called the cost function § Different weights for different types of matches § Gap costs q can be simple and count the total matching symbols q can be complicated and consider the type of matching symbols, location in the sequence, neighboring symbols etc.

Approximation Algorithms My T. Thai mythai@cise. ufl. edu 51

Scoring method Ø Score zero for a match or for two opposing spaces Ø Score one for a mismatch or for a character opposite a space

Assumptions: Ø Assume that two opposing spaces have a zero value Ø Assume other values satisfies triangle inequality q s(x, z) ≤ s(x, y) + s(y, z) q s(x, z) – cost of transforming character x into character z

Objective Functions Ø Two objective functions q SP § The sum of the values of pairwise alignments induced by an alignment A q TA § Using the topology of the tree, map the strings to the nodes of the tree § The sum of the selected pairwise alignments is called tree alignment

Center Star Method Ø For a set of k strings X q Choose a center string Xc of X which minimizes Σj≠c. D(Xc, Xj) q Let M = min Σj≠c. D(Xc, Xj) q Center star is a star tree of k nodes with the center node labeled Xc and each of the k-1 remaining nodes labeled by a distinct string in X {Xc} q If Xi and Xj are strings labeling adjacent nodes of tree T, then alignment of Xi and Xj induced by A(T) has value D(Xi, Xj)

Center Star Method – Alg Ac Ø Do an optimal alignment for each pair (Xc, Xj) for all j ≠ c Ø s 0 = max number of spaces placed before the first char of Xc Ø sf = max number of spaces placed after the last char of Xc Ø si = max number of spaces placed between Xc(i) and Xc(i+1)

Center Star Method – Alg Ac Ø For Xc, insert s 0, si, and sf spaces at the beginning, between, and the end of Xc respectively. Call X’c Ø Then for each Xj, do the optimal alignment without modifying X’c My T. Thai mythai@cise. ufl. edu 57

Analysis Ø d(Xi, Xj) ≥ D(Xi, Xj) Ø V(Ac) = Σi<jd(Xi, Xj) Ø V(Ac) is at most twice the value of the optimal multiple alignment of X My T. Thai mythai@cise. ufl. edu 58

Analysis Ø Lemma 3. 1: For any 2 strings Xi, Xj, we have: d(Xi, Xj) ≤ d(Xi, Xc) + d(Xc, Xj) = D(Xi, Xc) + D(Xc, Xj) q triangle inequality

Analysis Ø A* be the optimal multiple alignment of k strings X Ø Define: V(A*) = Σi<jd*(Xi, Xj)

Analysis Ø Theorem 3. 1 V(Ac) / V(A*) ≤ 2(k-1)/ k < 2 Ø Proof:

Disadvantages Ø Requires all pairwise alignments Ø Computationally expensive Ø Faster, Randomized alignments q q Randomly select string Xi Build multiple alignment with star centered at Xi Select best multiple alignment A from p such stars At most (k-1)p pairwise alignments need to be computed

Randomized Alignments Ø Theorem 3. 2 For any r >1, let e(r) be the expected number of stars needed to be chosen at random before the value of best resulting alignment is within a factor of 2+1/(r-1) of the optimal alignment. Then e(r) ≤ r. Ø e(r) is independent of k and the length of the strings.

Proof of Theorem 3. 2 Ø For r = 2, for each string Xi define M(i) = Σj. D(Xi, Xj) then M(c) = M From Theorem 3. 1, Σ(i, j)D(Xi, Xj) = Σj. M(i) ≤ 2(k-1)M so the Avg value of M(i) < 2 M Ø Since min M(i) = M, then Median M(i) < 3 M Number of centers selected before a selected M(i) is less than the median = 2

Proof Ø Suppose median is ∂M for 1 ≤ ∂ ≤ 3 Then Σ(i, j)D(Xi, Xj)≥ k. M/2 + k ∂ M/2 Ø Value of the alignment obtained from any below median star ≤ 2(k-1) ∂ M Therefore, error ratio for this star ≤ = 2 ∂ / (1/2 + ∂ /2) Ø When ∂ = 3, error ratio = 3. Ø So we have e(2) ≤ 2

Proof Ø Now generalize this proof for r > 2 Ø At least k/r stars have M(i) less than or equal to (2 r-1)M/(r-1) q Minimum M(i) is M q Mean < 2 M Ø expected number of stars to pick with M(i) < ∂ M is r for 1 ≤ ∂ ≤ (2 r-1)/(r-1) Ø error ratio = 2 ∂ /[1/r + (r-1) ∂ /r] Ø (2 r-1)/(r-1)=2 + 1/(r-1)

Theorem 3. 3 Ø Picking p stars at random, the best resulting alignment will have value within a factor of 2 + 1/(r-1) of the optimal with probability at least 1 – [(r-1)/r]p

Center Star Method Ø Proof Ø From theorem 3. 2, if Median value was actually 3 M q For half the stars M(i) = M and M(i) = 3 M for the other half Ø Σ(i, j)D(Xi, Xj)=2 k. M Ø optimal SP alignment can be obtained from any center string Xiwith M(i) = M q Probability of selecting such a string is one-half

Tree Alignment Method Ø Typical approach: q first find multiple alignment and then build a tree showing the evolutionary derivations Ø Another approach (called tree alignment): q first choose the typology of the tree and then map the strings to the nodes of the tree q Alignment is the pairwise alignments of the strings at the ends of the edges of the tree

Formal Definitions Ø Let K be an input set of k strings Ø K’ K be a set of strings containing K Ø Evolutionary tree TK’ for K is a tree: q with at least k nodes q each string in K’ labels exactly one node & each node gets exactly one label in K’ Ø The value of TK’ : V(TK’) = ΣD(X, Y) Ø the problem is to find a set of strings K’ and T(K’) for K which minimizes V (TK’)

Ø The alignment value D(X, Y ) is interpreted as the minimum “cost" to transform string X to string Y Ø The sum of the alignment values of the edges gives the evolutionary cost implied by the tree.

Method Ø Let G be a graph with k nodes labeled with a distinct string in K Ø Each edge (X, Y) has a weight D(X, Y) Ø Find the MST of G. This MST is an evolutionary tree for K

Analysis Ø T* denote the optimal evolutionary tree for K. q Prove: V(MST)/V(T*) < 2 OPT Ø Let C be a traversal of edges of T* which traverses everyy edge exactly once in each direction Ø Let C 1, …, Ck be the order that C encounters Ø Let V(C) = D(Ck, C 1) + Σi<k. D(Ci, Ci+1)

Analysis My T. Thai mythai@cise. ufl. edu 74

Analysis Ø Corollary 4. 1: V(C) ≤ 2 V(T*), Ø Let D(Ci*, Ci*+1) be the largest distance of any adjacent strings in C traversal Ø Lemma(4. 2) V(MST) ≤ V(C) – D(Ci*, Ci*+1) ≤ V(C) – V(C)/K

Analysis Ø Theorem 4. 1 For any set K of k strings, we have: V(MST)/ V(T*k) ≤ 2(k-1)/k < 2 Ø Theorem 4. 2 V(MST) / V(T*k) ≤ (k-1)/k V(C)/V(T*k) ≤ 2 (k-1)/k Ø Corollary 4. 2 V(T*k) > k. V(MST)/2(k-1)

Constrained MSA

Motivation General SP MSA problem: Ø NP-completeness has already been established Ø Appromixation algorithms have been developed Ø Heuristics are also avaliable Constrained MSA: Ø Biologists often have additional knowledge of data (e. g. active site residues) Ø Additional knowledge can specify matches at certain locations Ø Models allow users to provide additional constraints

Definition of CMSA Problem Ø Suppose that P = p 1 p 2. . . pα is a common subsequence of S 1, S 2, . . . , SK Ø The constrained multiple sequence alignment of S with respect to P is: q an MSA A with the constraints that there are α columns in A, c 1, c 2, . . . , cα with c 1 < c 2 < …< cα, such that the characters of column ci, 1 ≤ i ≤ α, are all equal to pi.

Optimal CPSA

Dynamic Algorithm My T. Thai mythai@cise. ufl. edu 81

Time and Space Complexities My T. Thai mythai@cise. ufl. edu 82

CMSA The improvement of CPSA in turn improves the time & space complexity of Progressive CMSA from O(αkn 4) and O(αn 4) to O(αk 2 n 2) and O(αn 2). Optimal CMSA This Optimal CMSA algorithm involves the creation of a matrix with k+1 dimensions. (Assume δ(x, y) is the distance function and satisfies the triangle inequality. ) Ø Let D(i 1, . . . , ik; γ) be the optimal CMSA score matrix for {S 1[1. . i 1], . . . , Sk[1. . ik]} where P[1. . γ] is aligned in γ columns. Ø Then optimal alignment score is D(n 1, . . . , nk; α), where ni =|Si|. Computing D: Ø D({0}k; 0) = 0 Ø Let εj = 0 or 1 with εj. Sj[ij] where j = 0 represents a space, and δ(x 1, . . . , xk) = Σ 1≤i<j≤kδ(xi, xj). D(i 1, i 2, . . . , ik; γ) is the minimum of: q if S 1[i 1] =. . . = Sk[ik] = P[γ], § q D(i 1 − 1, . . . , ik − 1; γ − 1) + δ(S 1[i 1], . . . , Sk[ik]) minε∈{0, 1}k (D(i 1 − ε 1, . . . , ik − εk; γ) + δ(ε 1 S 1[i 1], . . . , εk. Sk[ik])). These values can be computed using dynamic programming.

CMSA (Center Star) The Center-Star method proposed for the general MSA problem can be modified to apply to the CMSA problem. Ø Consider each sequence as the center, Sc. Consider each list position that Sc is aligned with P. Ø Find the minimum star-sum score Sc. Ø Create a constrained alignment matrix by merging the constrained pairwise sequence alignments between Sc & S j.

CMSA (Center Star) The recurrence of Thm. 3. 1 is only slightly modified:

Example My T. Thai mythai@cise. ufl. edu 86
- Slides: 86