Protein Tertiary Structure Prediction Jingfen Zhang Digital Biology

Protein Tertiary Structure Prediction Jingfen Zhang Digital Biology Laboratory Computer Science Department University of Missouri 2012 -11 - 15

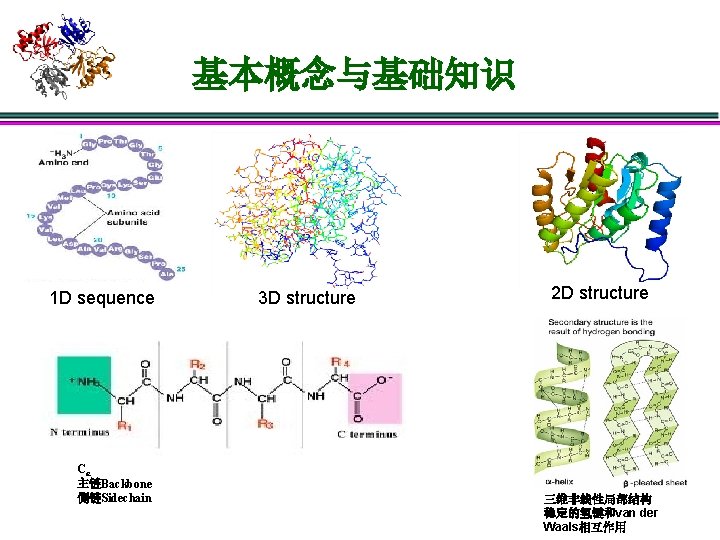
基本概念与基础知识 1 D sequence Cα 主链Backbone 侧链Sidechain 3 D structure 2 D structure 三维非线性局部结构 稳定的氢键和van der Waals相互作用

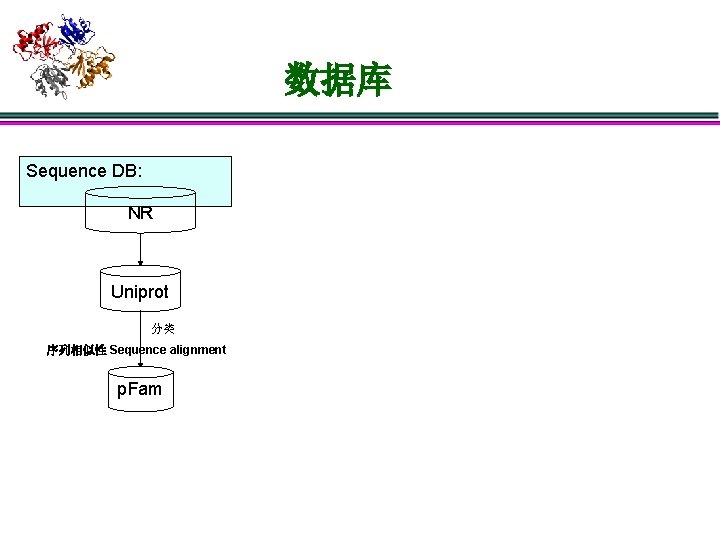
数据库 Sequence DB: NR Uniprot 分类 序列相似性 Sequence alignment p. Fam
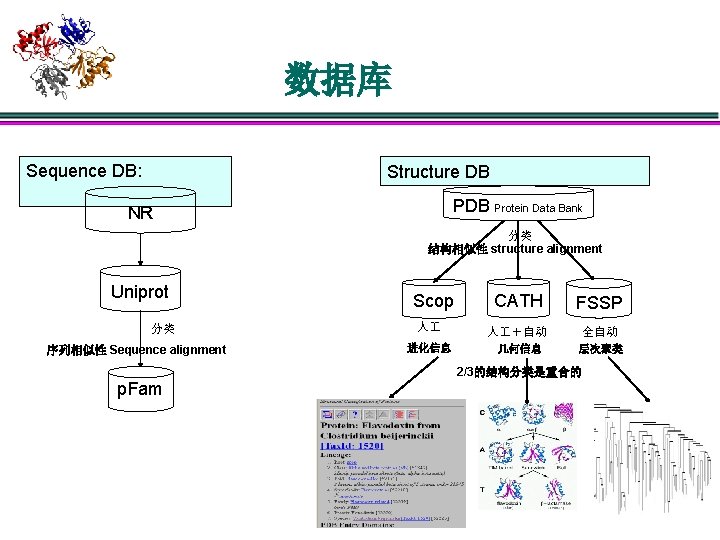
数据库 Sequence DB: Structure DB Protein Data Bank NR 分类 结构相似性 structure alignment Uniprot 分类 序列相似性 Sequence alignment CATH FSSP 人 人 +自动 全自动 进化信息 几何信息 层次聚类 Scop 2/3的结构分类是重合的 p. Fam
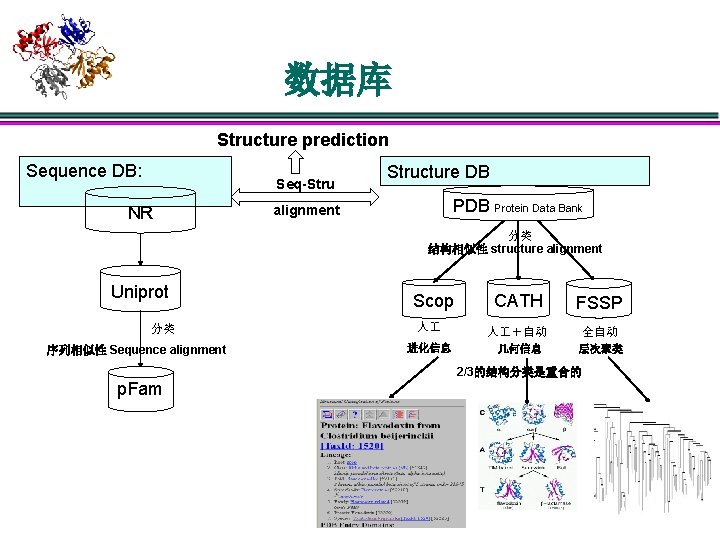
数据库 Structure prediction Sequence DB: Seq-Stru NR Structure DB Protein Data Bank alignment 分类 结构相似性 structure alignment Uniprot 分类 序列相似性 Sequence alignment CATH FSSP 人 人 +自动 全自动 进化信息 几何信息 层次聚类 Scop 2/3的结构分类是重合的 p. Fam
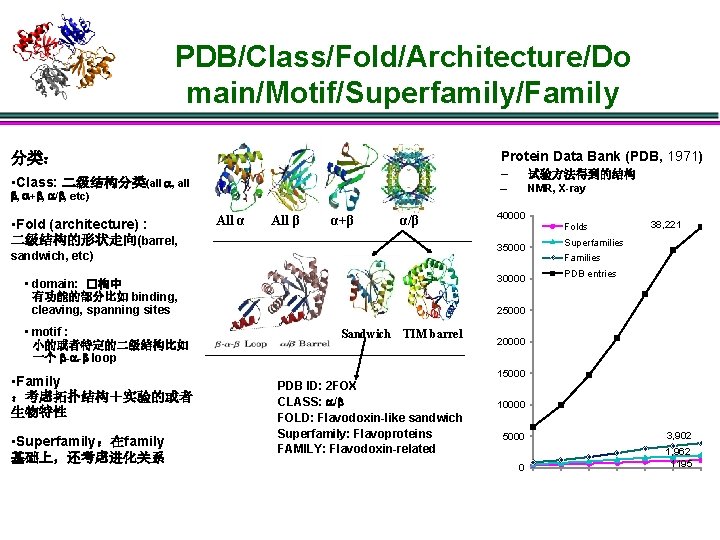
PDB/Class/Fold/Architecture/Do main/Motif/Superfamily/Family Protein Data Bank (PDB, 1971) 分类: • Class: 二级结构分类(all , + , / , etc) • Fold (architecture) : 二级结构的形状走向(barrel, All α All β α+β α/β • Superfamily:在family 基础上,还考虑进化关系 – NMR, X-ray 40000 30000 • domain: �构中 有功能的部分比如 binding, cleaving, spanning sites • Family :考虑拓扑结构+实验的或者 生物特性 试验方法得到的结构 35000 sandwich, etc) • motif : 小的或者特定的二级结构比如 一个 - - loop – Folds 38, 221 Superfamilies Families PDB entries 25000 Sandwich TIM barrel 20000 15000 PDB ID: 2 FOX CLASS: / FOLD: Flavodoxin-like sandwich Superfamily: Flavoproteins FAMILY: Flavodoxin-related 10000 5000 3, 902 0 1, 962 1195

广泛应用的度量衡 å RMSD (root mean square deviation) å GDT_TS The largest set of ‘equivalent’ residues under some cutoff distances GDT_TS = (GDT_P 1Å + GDT_P 2Å + GDT_P 4 Å+ GDT_P 8Å)/4 å TM-score å DALI-score

广泛应用的度量衡 å RMSD (root mean square deviation) å GDT_TS The largest set of ‘equivalent’ residues under some cutoff distances GDT_TS = (GDT_P 1Å + GDT_P 2Å + GDT_P 4 Å+ GDT_P 8Å)/4 å TM-score å DALI-score å Dscores

广泛应用的度量衡 å RMSD (root mean square deviation) å GDT_TS The largest set of ‘equivalent’ residues under some cutoff distances GDT_TS = (GDT_P 1Å + GDT_P 2Å + GDT_P 4 Å+ GDT_P 8Å)/4 å TM-score å DALI-score å Dscores RMSD最小化差异性 TM-score, GDT_TS最大化相似性 RMSD, TM-score, GDT_TS: 涉及L*3的坐标转换,优化过程 DALI-score, Dscores直接比较L*L的距离矩阵 DALI-score, Dscore 2描述相似性, Dscore 1描述差异性

结构比对 - 方法 l 多目标优化: 尽可能多地匹配residues, 同时还优化比对 的距离/相似性 å CE (combinatorial extension structure alignment),优化RMSD å LGA (Local-Global Alignment)优化 GDT_TS X LCS (longest continuous segments) + GDT (global distance test) å TMAlign - 优化TM-score X 二级结构比对+ 最好的gapless比对+ 考虑gap-opening罚分的比对 X 启发式的迭代扩展比对,记录最好的比对 å DALI - 优化DALI-score X 比较6 -肽子结构,记录所有相似的片断 X Monte Carlo 拼接上述相似的子结构扩展成更长的比对

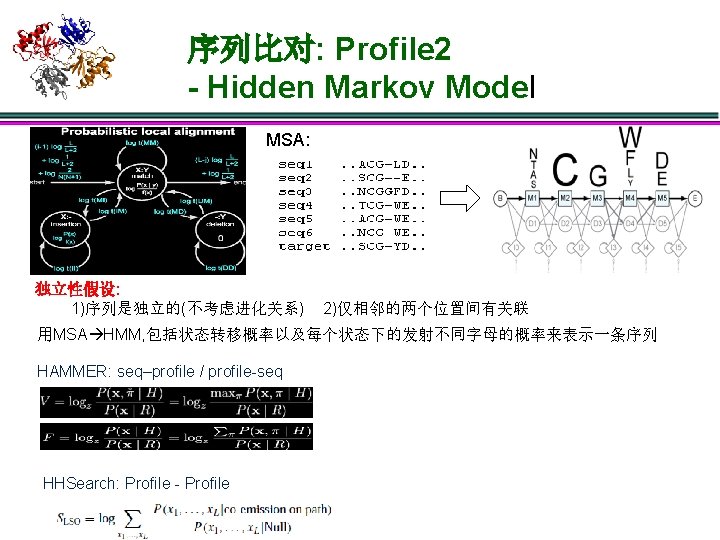
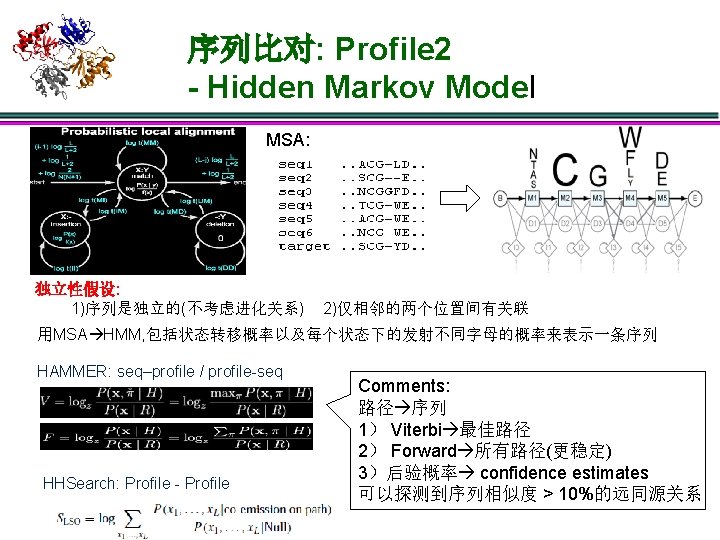
序列比对: Sequence – Sequence seq 1 . . . NCGE… target . . . SCGD… 1)独立性假设: 一个sequence里的residue之间是独立的 2)score = ∑sa, b, sa, b single score for residues a, b: sa, b = log P(a | b) f(a) 从b的同源性导出 a 随机出现的频率 3)P(a|b): 固定进化距离可以统计得到 20 x 20的score matrix(也称为替代 矩阵),比如BLOSUM, PAM等;给定已知的数据库则可以估计f(a) 4)gap open cost α,gap extend cost β Smith–Waterman algorithm: DP optimal BLAST: (Basic Local Alignment Search Tool) seeding + heuristic extension FASTA: (FAST-All), seeding + banded Smith-Waterman

序列比对: Sequence – Sequence seq 1 . . . NCGE… target . . . SCGD… 1)独立性假设: 一个sequence里的residue之间是独立的 2)score = ∑sa, b, sa, b single score for residues a, b: sa, b = log P(a | b) f(a) 从b的同源性导出 a 随机出现的频率 3)P(a|b): 固定进化距离可以统计得到 20 x 20的score matrix(也称为替代 矩阵),比如BLOSUM, PAM等;给定已知的数据库则可以估计f(a) 4)gap open cost α,gap extend cost β Smith–Waterman algorithm: DP optimal BLAST: (Basic Local Alignment Search Tool) seeding + heuristic extension FASTA: (FAST-All), seeding + banded Smith-Waterman Comments: 1)序列的比较实际上是对residues匹 配与失配进行评分 2)序列相似度足够高时性能最好




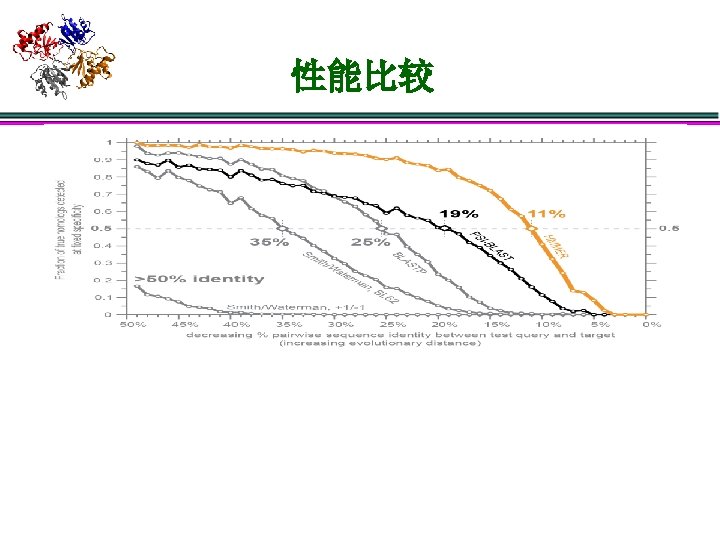

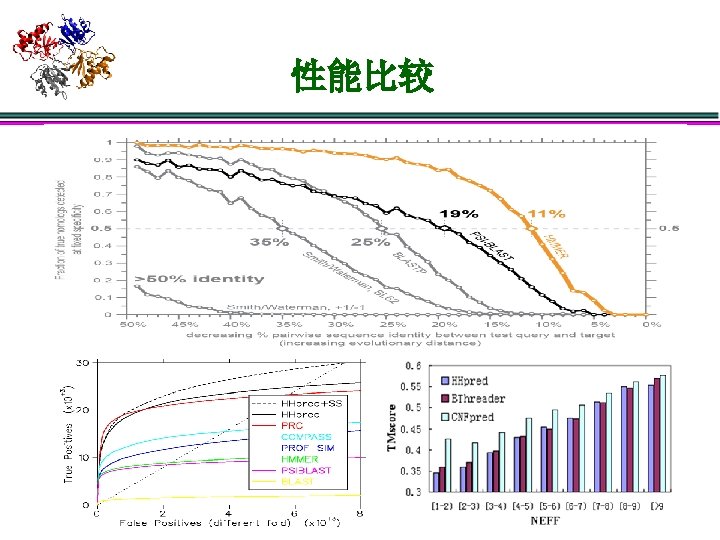

Potential future work l 松弛独立性假设 序列不独立 l l l New Model l l 研究进化关系 l 比较sequence. DB规范的进化关系与structure. DB规范的进化关系 l 新的组织结构 统一序列结构的进化 建立含进化关系的Profile HMM, CRF, CNF, … ? New search Method l l Sensitivity, 尤其是local alignment Accuracy: 假定fold或者template正确了,如何提高比对的正确性

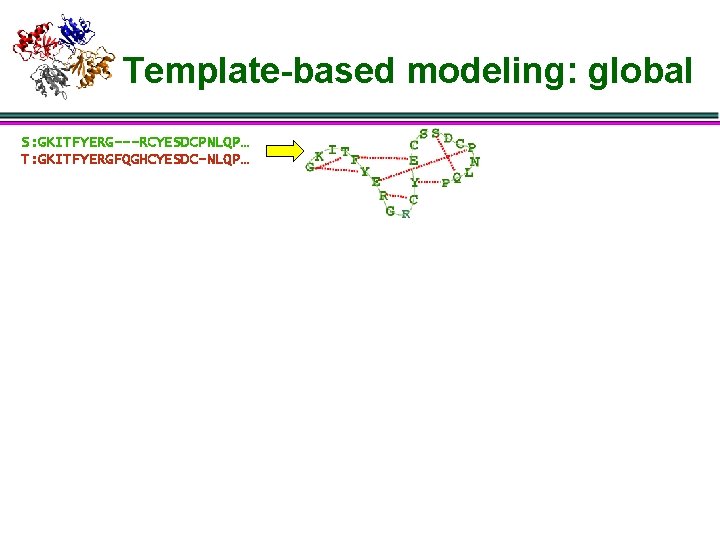
Template-based modeling: global S: GKITFYERG---RCYESDCPNLQP… T: GKITFYERGFQGHCYESDC-NLQP…
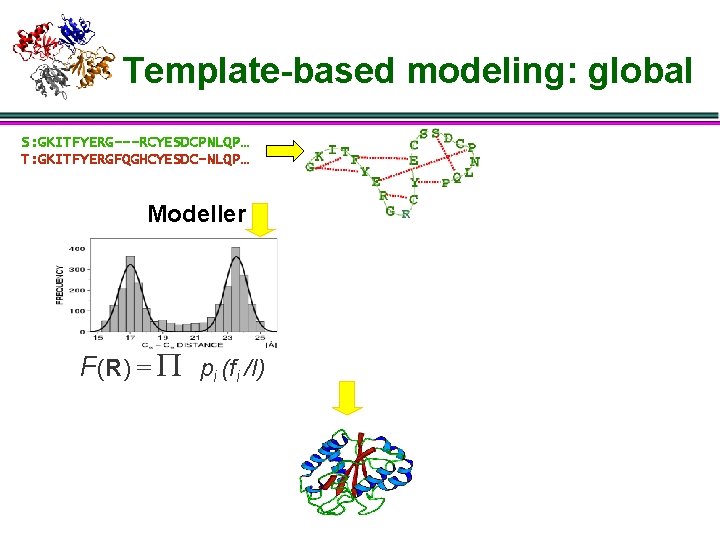
Template-based modeling: global S: GKITFYERG---RCYESDCPNLQP… T: GKITFYERGFQGHCYESDC-NLQP… Modeller F(R) = P pi (fi /I)
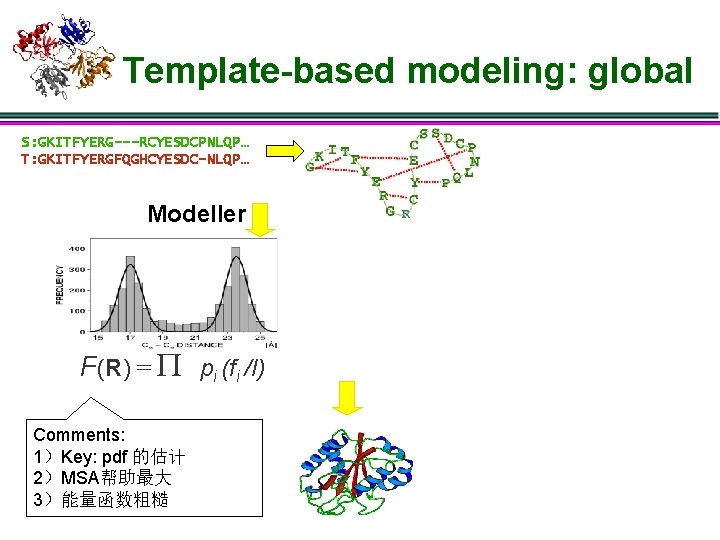
Template-based modeling: global S: GKITFYERG---RCYESDCPNLQP… T: GKITFYERGFQGHCYESDC-NLQP… Modeller F(R) = P pi (fi /I) Comments: 1)Key: pdf 的估计 2)MSA帮助最大 3)能量函数粗糙
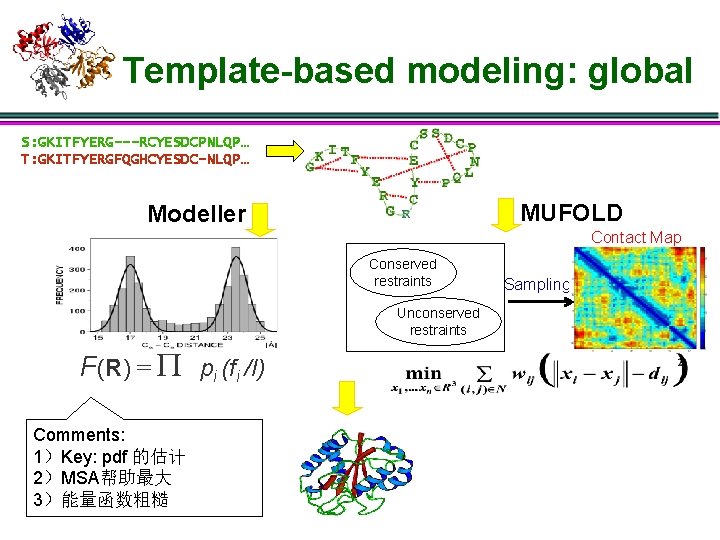
Template-based modeling: global S: GKITFYERG---RCYESDCPNLQP… T: GKITFYERGFQGHCYESDC-NLQP… MUFOLD Modeller Contact Map Conserved restraints Unconserved restraints F(R) = P pi (fi /I) Comments: 1)Key: pdf 的估计 2)MSA帮助最大 3)能量函数粗糙 Sampling
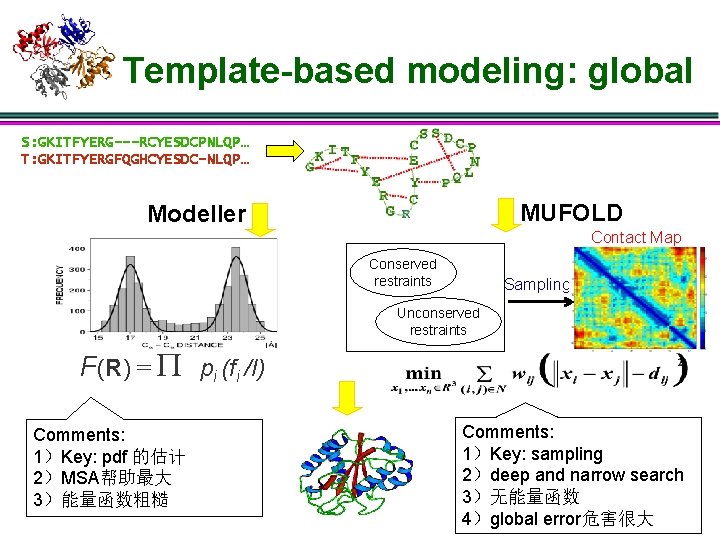
Template-based modeling: global S: GKITFYERG---RCYESDCPNLQP… T: GKITFYERGFQGHCYESDC-NLQP… MUFOLD Modeller Contact Map Conserved restraints Sampling Unconserved restraints F(R) = P pi (fi /I) Comments: 1)Key: pdf 的估计 2)MSA帮助最大 3)能量函数粗糙 Comments: 1)Key: sampling 2)deep and narrow search 3)无能量函数 4)global error危害很大
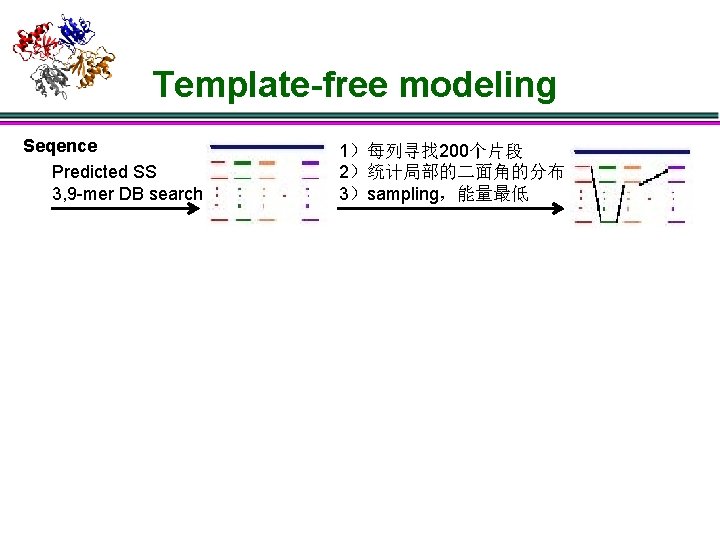
Template-free modeling Seqence Predicted SS 3, 9 -mer DB search 1)每列寻找 200个片段 2)统计局部的二面角的分布 3)sampling,能量最低
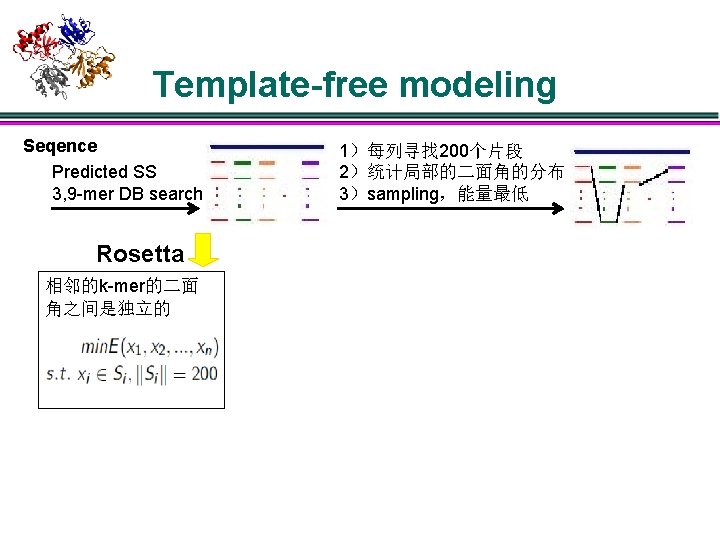
Template-free modeling Seqence Predicted SS 3, 9 -mer DB search Rosetta 相邻的k-mer的二面 角之间是独立的 1)每列寻找 200个片段 2)统计局部的二面角的分布 3)sampling,能量最低
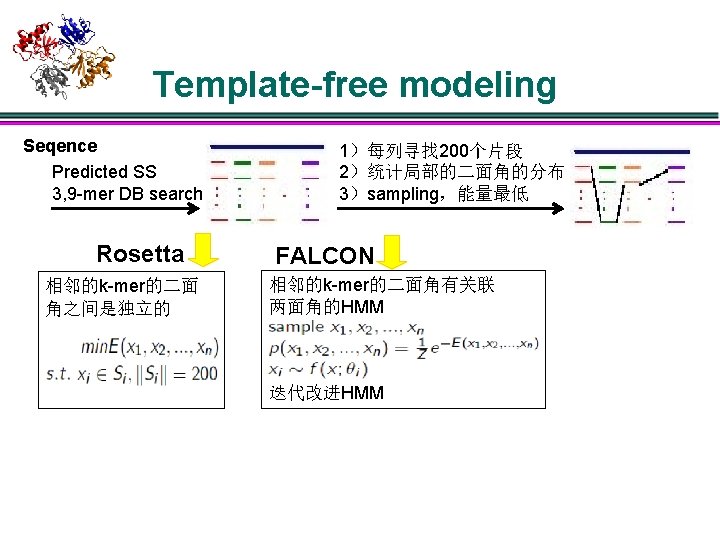
Template-free modeling Seqence Predicted SS 3, 9 -mer DB search Rosetta 相邻的k-mer的二面 角之间是独立的 1)每列寻找 200个片段 2)统计局部的二面角的分布 3)sampling,能量最低 FALCON 相邻的k-mer的二面角有关联 两面角的HMM 迭代改进HMM
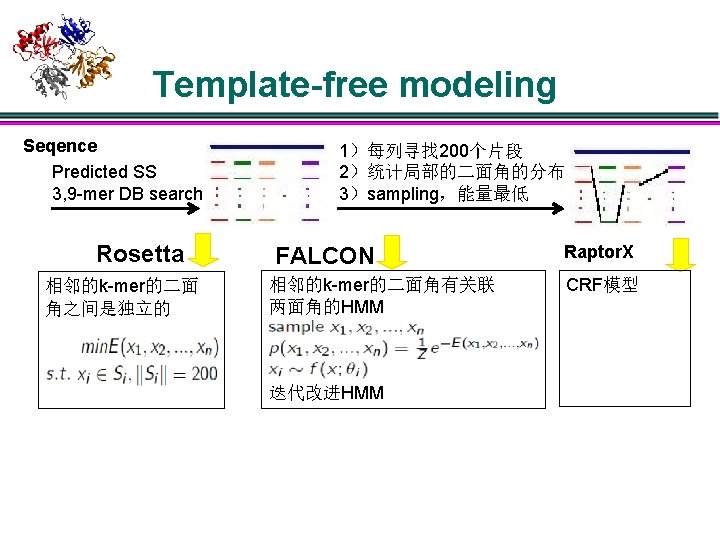
Template-free modeling Seqence Predicted SS 3, 9 -mer DB search Rosetta 相邻的k-mer的二面 角之间是独立的 1)每列寻找 200个片段 2)统计局部的二面角的分布 3)sampling,能量最低 FALCON 相邻的k-mer的二面角有关联 两面角的HMM 迭代改进HMM Raptor. X CRF模型
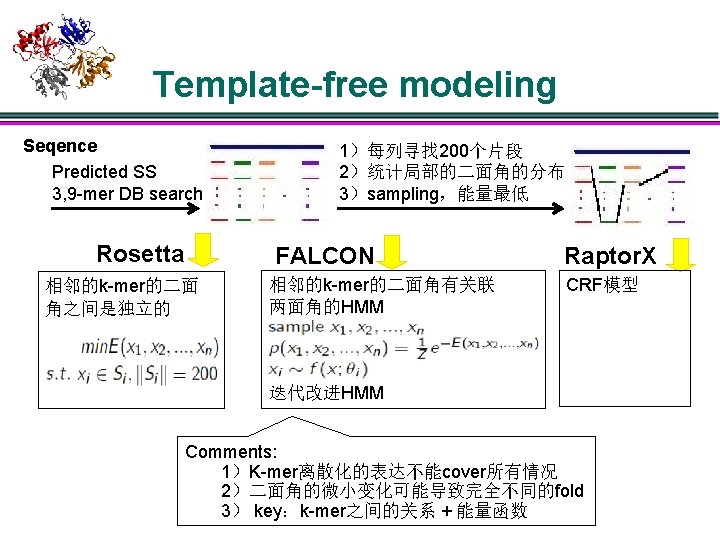
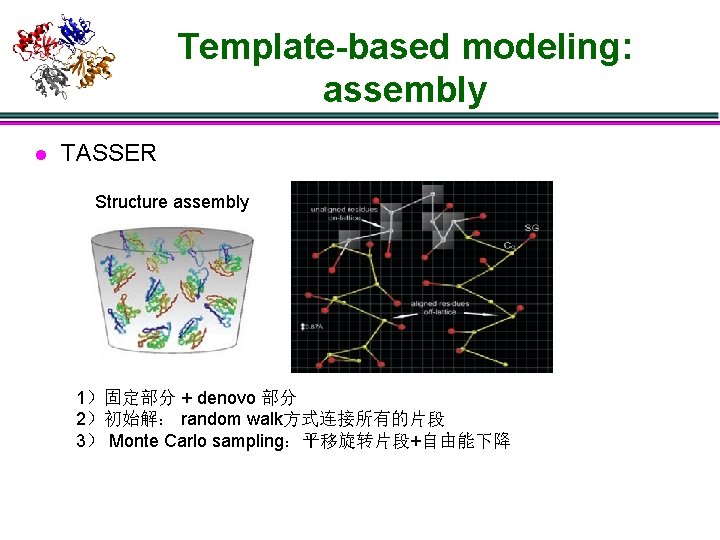
Template-based modeling: assembly l TASSER Structure assembly 1)固定部分 + denovo 部分 2)初始解: random walk方式连接所有的片段 3) Monte Carlo sampling:平移旋转片段+自由能下降
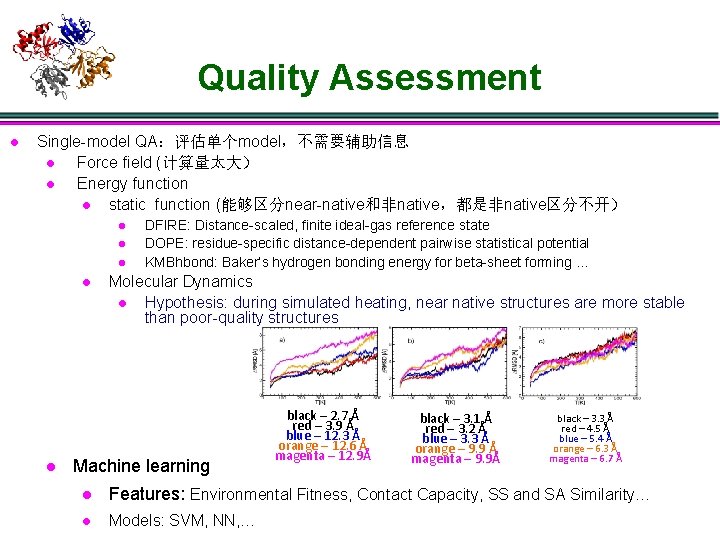
Quality Assessment l Single-model QA:评估单个model,不需要辅助信息 l Force field (计算量太大) l Energy function l static function (能够区分near-native和非native,都是非native区分不开) l l l DFIRE: Distance-scaled, finite ideal-gas reference state DOPE: residue-specific distance-dependent pairwise statistical potential KMBhbond: Baker’s hydrogen bonding energy for beta-sheet forming … Molecular Dynamics l Hypothesis: during simulated heating, near native structures are more stable than poor-quality structures Machine learning black – 2. 7 Å red – 3. 9 Å blue – 12. 3 Å orange – 12. 6 Å magenta – 12. 9Å black – 3. 1 Å red – 3. 2 Å blue – 3. 3 Å orange – 9. 9 Å magenta – 9. 9Å black – 3. 3 Å red – 4. 5 Å blue – 5. 4 Å orange – 6. 3 Å magenta – 6. 7 Å l Features: Environmental Fitness, Contact Capacity, SS and SA Similarity… l Models: SVM, NN, …
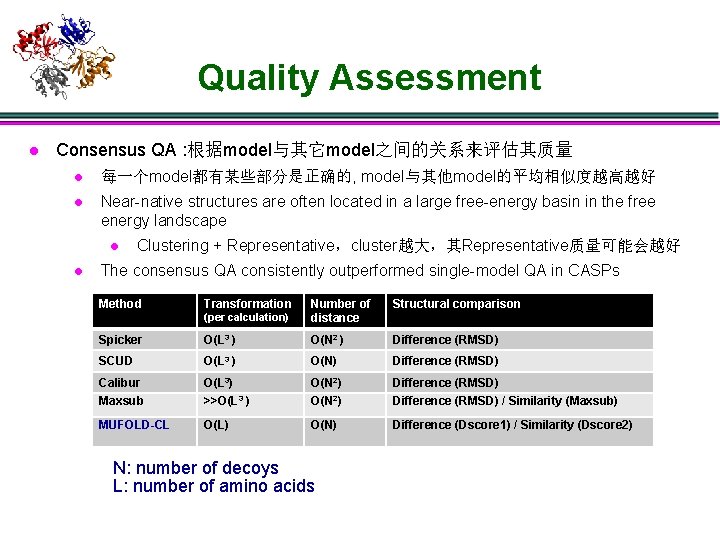
Quality Assessment l Consensus QA : 根据model与其它model之间的关系来评估其质量 l 每一个model都有某些部分是正确的, model与其他model的平均相似度越高越好 l Near-native structures are often located in a large free-energy basin in the free energy landscape l l Clustering + Representative,cluster越大,其Representative质量可能会越好 The consensus QA consistently outperformed single-model QA in CASPs Method Transformation Structural comparison (per calculation) Number of distance Spicker O(L 3 ) O(N 2 ) Difference (RMSD) SCUD O(L 3 ) O(N) Difference (RMSD) Calibur Maxsub O(L 3) >>O(L 3 ) O(N 2) Difference (RMSD) / Similarity (Maxsub) MUFOLD-CL O(L) O(N) Difference (Dscore 1) / Similarity (Dscore 2) N: number of decoys L: number of amino acids
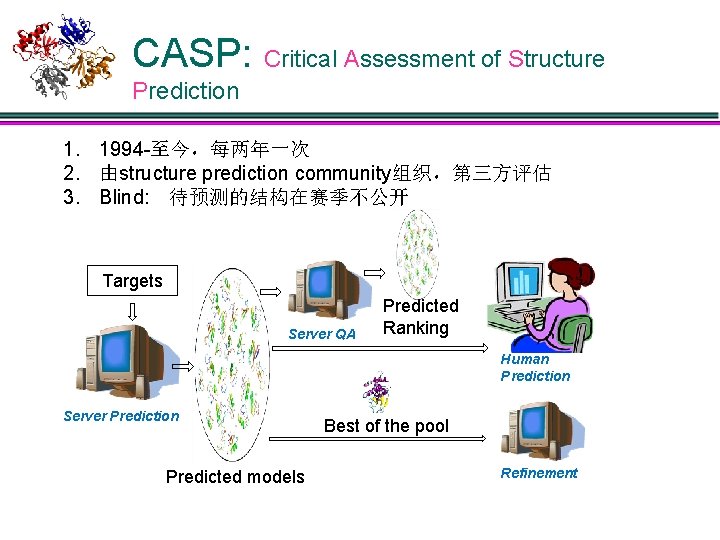
CASP: Critical Assessment of Structure Prediction 1. 1994 -至今,每两年一次 2. 由structure prediction community组织,第三方评估 3. Blind: 待预测的结构在赛季不公开 Targets Server QA Predicted Ranking Human Prediction Server Prediction Predicted models Best of the pool Refinement

Our CASP History CASP 8: invited to give a talk at the meeting Server Prediction: #1 in Free modeling category QA: No significant result CASP 9: invited to give a talk at the meeting and publish a paper in the special issue Server Prediction : #10 in Template-based modeling category (#1 in trivial cases) #7 in Free modeling category Human Prediction: (We did by our server not by human in fact) #1 in Human prediction QA: #1 in correlation #2 in top-1 selection CASP 10: Should have better performance than CASP 9

Acknowledgements l Team (MUFOLD development) å Dong Xu (Computer Science) X Jingfen Zhang X Zhiquan He å Yi Shang (Computer Science) X Qingguo Wang å Ioan Kosztin (Physics) X l Jiong Zhang Funding Sources å University of Missouri Research Board Grant å NIH: R 21/R 33 GM 078601

Thanks for your patience!
- Slides: 47