Protein Structure II Protein Methods Andy Howard Biochemistry

Protein Structure II; Protein Methods Andy Howard Biochemistry Lectures, Spring 2019 29 January 2019

Protein structure and methods n n We’ll complete our conversation about protein structure Then we’ll talk about methods of dealing with proteins 01/29/2019 Protein Structure & Methods P. 2 of 86

Topics for today n Protein structure Secondary structure n Tertiary Structure n Motifs & topology n Quaternary structure n n n Methods Purification n Characterization n Structure n Depicting structure 01/29/2019 Protein Structure & Methods Page 3 of 86

Other helices n n NH to C=O four residues earlier is not the only pattern found in proteins Other helical structures differ in that they have connections between amine N and C=O that is 3 or 5 residues earlier 01/29/2019 Protein Structure & Methods Page 4 of 86

310 and π helices n 310 helix is NH to C=O three residues earlier (Moran fig. 4. 15) n n More kinked; 3 residues per turn Often one H-bond of this kind at N-terminal end of an otherwise -helix § helix: even rarer: NH to C=O five residues earlier 01/29/2019 Protein Structure & Methods Page 5 of 86

Beta strands (fig. 4. 4, 4. 5) n n n Structures containing roughly extended polypeptide strands Extended conformation stabilized by interstrand main-chain hydrogen bonds No defined interval in sequence number between amino acids involved in H-bond 01/29/2019 Protein Structure & Methods Page 6 of 86
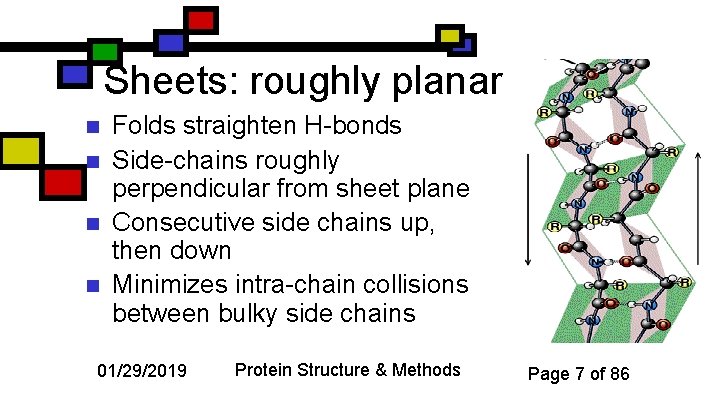
Sheets: roughly planar n n Folds straighten H-bonds Side-chains roughly perpendicular from sheet plane Consecutive side chains up, then down Minimizes intra-chain collisions between bulky side chains 01/29/2019 Protein Structure & Methods Page 7 of 86
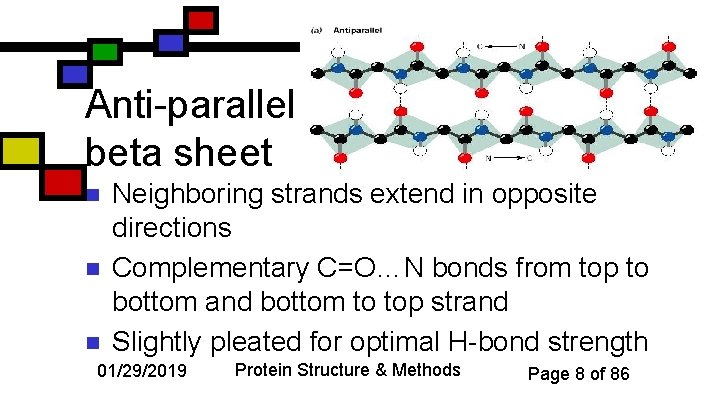
Anti-parallel beta sheet n n n Neighboring strands extend in opposite directions Complementary C=O…N bonds from top to bottom and bottom to top strand Slightly pleated for optimal H-bond strength 01/29/2019 Protein Structure & Methods Page 8 of 86

Parallel Beta Sheet n n N-to-C directions are the same for both strands You need to get from the C-end of one strand to the N-end of the other strand somehow H-bonds at more of an angle relative to the approximate strand directions Therefore: more pleated than anti-parallel sheet 01/29/2019 Protein Structure & Methods Page 9 of 86
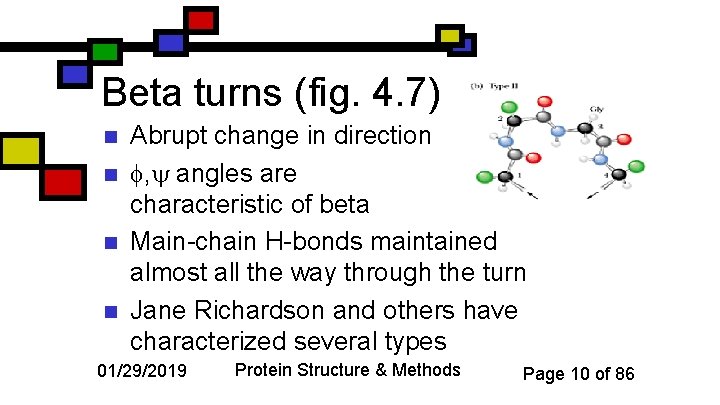
Beta turns (fig. 4. 7) n n Abrupt change in direction , angles are characteristic of beta Main-chain H-bonds maintained almost all the way through the turn Jane Richardson and others have characterized several types 01/29/2019 Protein Structure & Methods Page 10 of 86
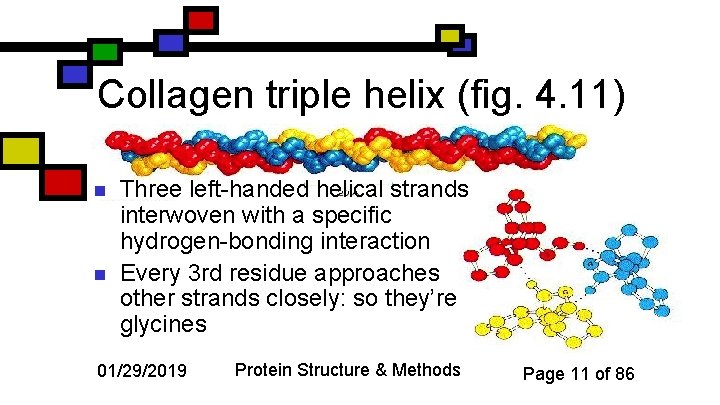
Collagen triple helix (fig. 4. 11) n n Three left-handed helical strands interwoven with a specific hydrogen-bonding interaction Every 3 rd residue approaches other strands closely: so they’re glycines 01/29/2019 Protein Structure & Methods Page 11 of 86

n Hydrogen bonds, revisited Within proteins, H-bonds almost always are: n n n between carbonyl oxygen and hydroxyl: (C=O • • • H-O-) between carbonyl O and amine or amide: (C=O • • • H-N-) On the other hand, H-bonds between amino acid components and water molecules are common too 01/29/2019 Protein Structure & Methods Page 12 of 86

Hydrogen bonds provide enthalpic stabilization n These are stabilizing structures n n Any stabilization is (on its own) entropically disfavored; Sufficient enthalpic optimization overcomes that! In general the optimization is ~ 4 - 11 k. J/mol The short (low-barrier) H-bonds are rare but important; H < -9 k. J/mol 01/29/2019 Protein Structure & Methods Page 13 of 86

Secondary structures in structural proteins n n n Structural proteins often have uniform secondary structures Seeing instances of secondary structure provides a path toward understanding them in globular proteins Examples: n n Alpha-keratin (hair, wool, nails, …): -helical Silk fibroin (guess) is -sheet 01/29/2019 Protein Structure & Methods Page 14 of 86
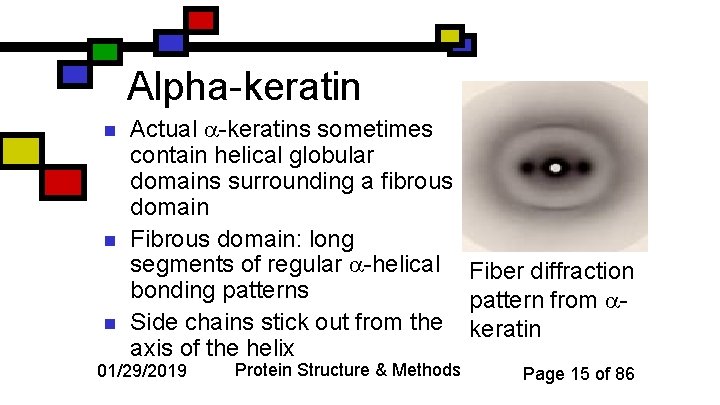
Alpha-keratin n Actual -keratins sometimes contain helical globular domains surrounding a fibrous domain Fibrous domain: long segments of regular -helical Fiber diffraction bonding patterns pattern from Side chains stick out from the keratin axis of the helix 01/29/2019 Protein Structure & Methods Page 15 of 86
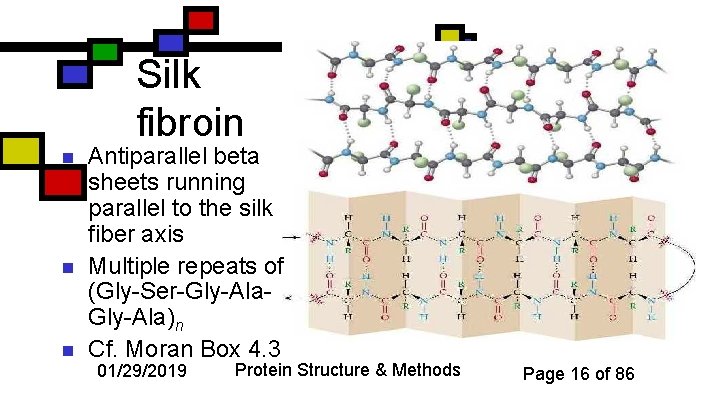
Silk fibroin n Antiparallel beta sheets running parallel to the silk fiber axis Multiple repeats of (Gly-Ser-Gly-Ala)n Cf. Moran Box 4. 3 01/29/2019 Protein Structure & Methods Page 16 of 86

Secondary structure in globular proteins n n n Segments with secondary structure are usually short: 2 -30 residues Some globular proteins are almost all helical, but even there are bends between short helices Other proteins: mostly beta Others: regular alternation of , Still others: irregular , , “coil” 01/29/2019 Protein Structure & Methods Page 17 of 86

Globular proteins with predominant structure types n n Globins, cytochromes, transmembrane proteins, ion channels: predominantly α-helical Immunoglobulins, viral coat proteins: predominantly beta-sheets Certain enzymes: alternating helices and strands (TIM barrels; see later slide) Many proteins have stretches that are neither helical nor. Protein strand-like (“coil”) Structure & Methods 01/29/2019 Page 18 of 86

Tertiary Structure (CF&M § 4. 4) n n The overall 3 -D arrangement of atoms in a single polypeptide chain Made up of secondary-structure elements & locally unstructured strands Described in terms of sequence, topology, overall fold, domains Stabilized by van der Waals interactions, hydrogen bonds, disulfides, . . . 01/29/2019 Protein Structure & Methods Page 19 of 86
Motifs n n n Combinations of secondary structure elements that recur in many proteins Helix-loop-helix, coiled-coil, helix bundle Beta-dominated motifs: , hairpin, -meander, Greek key, -sandwich 01/29/2019 Protein Structure & Methods Page 20 of 86
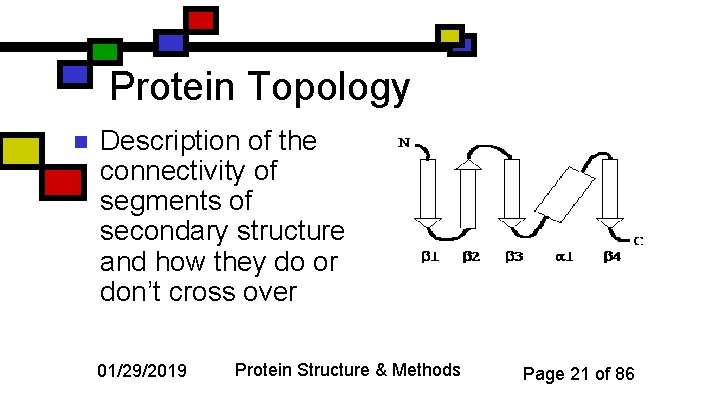
Protein Topology n Description of the connectivity of segments of secondary structure and how they do or don’t cross over 01/29/2019 Protein Structure & Methods Page 21 of 86
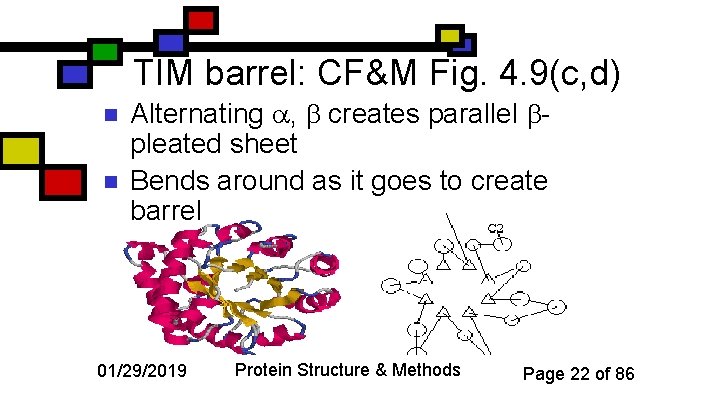
TIM barrel: CF&M Fig. 4. 9(c, d) n n Alternating , creates parallel pleated sheet Bends around as it goes to create barrel 01/29/2019 Protein Structure & Methods Page 22 of 86

Other tertiary structures n n Spend some time contemplating figures 4. 9, 4. 14, 4. 15: they show various tertiary structures Some researchers speculate that there are only N folds (N=2000? ) and all structures are combinations of those 01/29/2019 Protein Structure & Methods Page 23 of 86
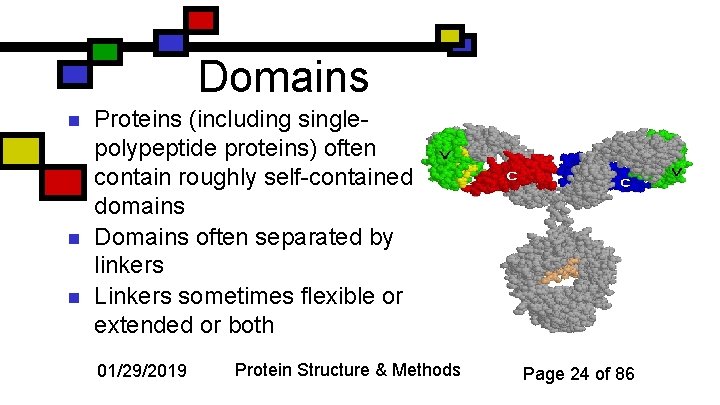
Domains n n n Proteins (including singlepolypeptide proteins) often contain roughly self-contained domains Domains often separated by linkers Linkers sometimes flexible or extended or both 01/29/2019 Protein Structure & Methods Page 24 of 86

Quaternary structure (CF&M § 4. 5) n n Arrangement of individual polypeptide chains to form a complete oligomeric, functional protein Individual chains can be identical or different n n If they’re the same, they can be coded for by the same gene If they’re different, you need more than one gene 01/29/2019 Protein Structure & Methods Page 25 of 86

Quaternary structures often involve symmetries n n Unsurprising: two identical objects tend to align themselves in symmetric ways 38% of all E. coli proteins are dimers Can be homooligomers or heterooligomers or combinations of those Note figs. Protein 4. 20, Structure 4. 23 & Methods 01/29/2019 Page 26 of 86

i. Clicker quiz, question 1 1. Which is the best way to describe the effect of a proline residue on helix? n (a) Prolines disrupt helices n (b) Prolines encourage helix formation n (c) Prolines disrupt -helices n (d) Proliines encourage -helix formation 01/29/2019 Protein Structure & Methods P. 27 of 86
i. Clicker quiz, question 2 n 2. Bovine trypsin (above) is a single polypeptide containing 223 amino acids. Which type(s) of structure does it lack? 01/29/2019 n n n (a) Primary (b) Secondary (c) Tertiary (d) Quaternary (e) Tertiary and quaternary Protein Structure & Methods P. 28 of 86
i. Clicker quiz, question #3 3. Mammalian hemoglobins consist of two identical chains and two identical chains. How many genes would be required to specify hemoglobin? . 01/29/2019 n n (a) One (b) Two (c) Four (d) None of the above Protein Structure & Methods P. 29 of 86
How do we visualize protein structures? n n n It’s often as important to decide what to omit as it is to decide what to include Any segment larger than about 10Å (1 nm) needs to be simplified if you want to understand it What you omit depends on what you want to emphasize 01/29/2019 Protein Structure & Methods P. 30 of 86
Styles of protein depiction n n All atoms All non-H atoms Main-chain (backbone) only One dot per residue (typically at C ) Ribbon diagrams: n n n Helical ribbon for helix Flat ribbon for strand Thin string for coil 01/29/2019 Protein Structure & Methods P. 31 of 86

How do we show 3 -D? You need to be aware that several forms of protein depiction suffer from the basic difficulty of representing three dimensions on a 2 -D image. So… n Stereo pairs n Dynamics: rotation of flat image n Perspective (hooray, Renaissance) 01/29/2019 Protein Structure & Methods P. 32 of 86

Stereo pairs 01/29/2019 Protein Structure & Methods P. 33 of 86
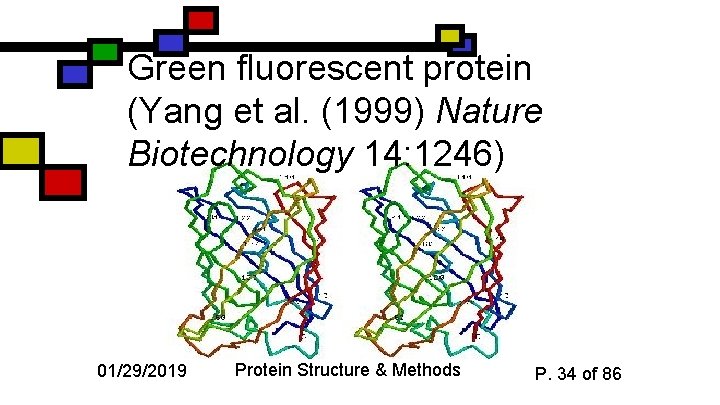
Green fluorescent protein (Yang et al. (1999) Nature Biotechnology 14: 1246) 01/29/2019 Protein Structure & Methods P. 34 of 86
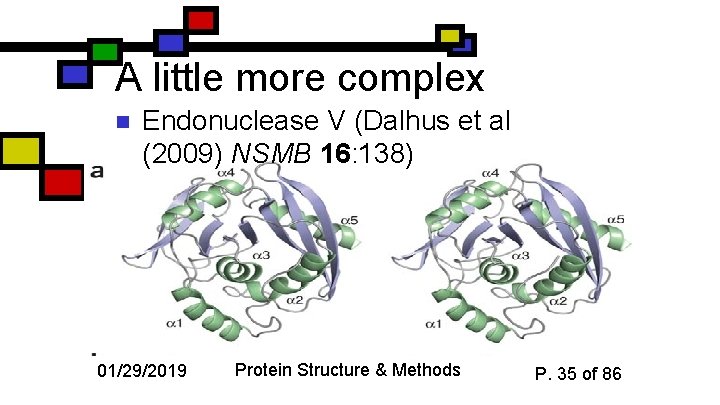
A little more complex n Endonuclease V (Dalhus et al (2009) NSMB 16: 138) 01/29/2019 Protein Structure & Methods P. 35 of 86
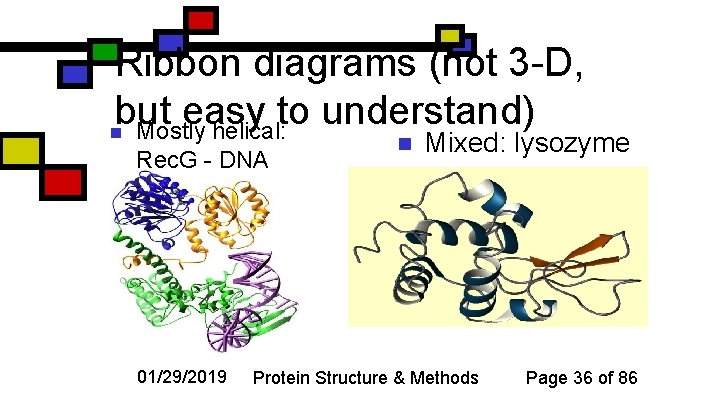
Ribbon diagrams (not 3 -D, but easy to understand) n Mostly helical: Rec. G - DNA 01/29/2019 n Mixed: lysozyme Protein Structure & Methods Page 36 of 86

The Protein Data Bank n n n http: //www. rcsb. org/ This is an electronic repository for threedimensional structural information of polypeptides and polynucleotides 126060 structures as of this month; 40092 distinct sequences 01/29/2019 Protein Structure & Methods P. 37 of 86

How were they determined? n n n Most are determined by X-ray crystallography Smaller number are high-field NMR or cryo. EM structures A few calculated structures, most of which are either close relatives of experimental structures or else they’re small, all-alphahelical proteins 01/29/2019 Protein Structure & Methods P. 38 of 86

What you can do with the PDB n n Display structures Look up specific coordinates Run clever software that compares and synthesizes the knowledge contained there Use it as a source for determining additional structures 01/29/2019 Protein Structure & Methods P. 39 of 86

Protein methods n n n In a rapid survey course like this, it’s tempting to ignore problems of methodology Some texts tend to simply present results without any discussion of how they were obtained The next section of this lecture is an attempt to redress this in the context of proteins 01/29/2019 Protein Structure & Methods P. 40 of 86

How will we study protein methods? n n n How to purify proteins How to characterize them How to determine their structures 01/29/2019 Protein Structure & Methods P. 41 of 86

Protein Purification (Moran § 3. 6) n Why do we purify proteins? n n To get a basic idea of function we need to see a protein in isolation from its environment That necessitates purification An instance of reductionist science Full characterization requires a knowledge of the protein’s action in context 01/29/2019 Protein Structure & Methods P. 42 of 86
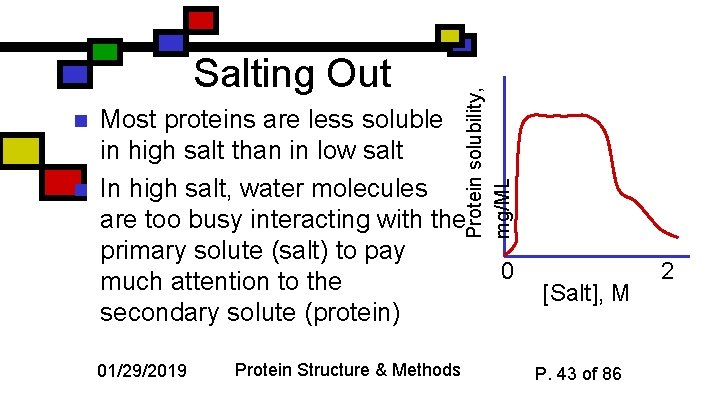
n n Protein solubility, mg/ML Salting Out Most proteins are less soluble in high salt than in low salt In high salt, water molecules are too busy interacting with the primary solute (salt) to pay much attention to the secondary solute (protein) 01/29/2019 Protein Structure & Methods 0 [Salt], M P. 43 of 86 2

How do we use that for purification? n n Various proteins differ in the degree to which their solubility disappears as [salt] goes up We can separate proteins by their differential solubility in high salt. 01/29/2019 Protein Structure & Methods P. 44 of 86

How to do it n n n Dissolve protein mixture in highly soluble salt like Li 2 SO 4, (NH 4)2 SO 4, Na. Cl Increase [salt] until some proteins precipitate and others don’t You may be able to recover both: n n n The supernatant (get rid of salt; move on) The pellet (redissolve, desalt, move on) Typical salt concentrations > 1 M 01/29/2019 Protein Structure & Methods P. 45 of 86

Dialysis n n n Some plastics allow molecules to pass through if and only if MW < Cutoff Protein will stay inside bag, smaller proteins will leave Non-protein impurities may leave too. 01/29/2019 Protein Structure & Methods P. 46 of 86

Gel-filtration chromatography n n n Pass a protein solution through a beadcontaining medium at low pressure Beads retard small molecules Beads don’t retard bigger molecules Can be used to separate proteins of significantly different sizes Suitable for preparative work Cf. Moran fig. 3. 11 Protein Structure & Methods 01/29/2019 P. 47 of 86

Ion-exchange chromatography n n Charged species affixed to column Phosphonates (-) retard (+)charged proteins: Cation exchange Quaternary ammonium salts (+) retard ( -)charged proteins: Anion exchange Separations facilitated by adjusting p. H 01/29/2019 Protein Structure & Methods P. 48 of 86

Affinity chromatography n n n Stationary phase contains a species that has specific favorable interaction with the protein we want DNA-binding protein specific to AGCATGCT: bind AGCATGCT to a column, and the protein we want will stick; every other protein falls through Often used to purify antibodies by binding the antigen to the column 01/29/2019 Protein Structure & Methods P. 49 of 86

Metal-ion affinity chromatography n n Immobilize a metal ion, e. g. Ni 2+ or Co 2+, to the column material Proteins with affinity to that metal will stick 01/29/2019 Protein Structure & Methods P. 50 of 86

How do you complete that? n n Wash them off afterward with a ligand with an even higher affinity, e. g imidazole We can engineer proteins to contain the affinity tag: poly-histidine at N- or C-terminus 01/29/2019 Protein Structure & Methods P. 51 of 86

High-performance liquid chromatography n n Many LC separations can happen faster and more effectively under high pressure Works for small molecules Protein application is routine too, both for analysis and purification FPLC is a trademark, but it’s used generically 01/29/2019 Protein Structure & Methods P. 52 of 86

Electrophoresis (CF&M§ 5. 3) n n n Separating analytes by charge by subjecting a mixture to a strong electric field Gel electrophoresis: field applied to a semisolid matrix Can be used for charge (directly) or size (indirectly) 01/29/2019 Protein Structure & Methods P. 53 of 86
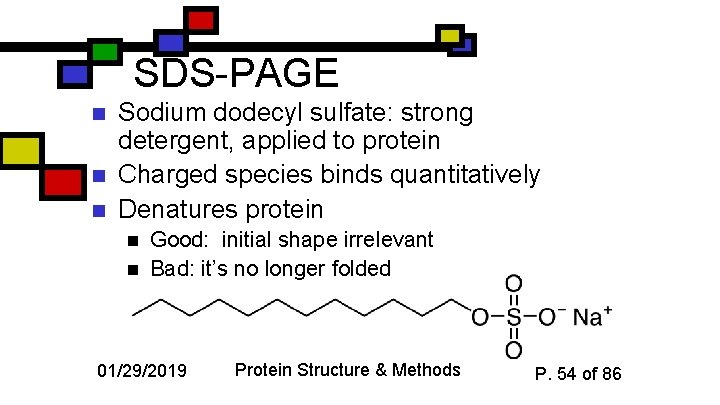
SDS-PAGE n n n Sodium dodecyl sulfate: strong detergent, applied to protein Charged species binds quantitatively Denatures protein n n Good: initial shape irrelevant Bad: it’s no longer folded 01/29/2019 Protein Structure & Methods P. 54 of 86
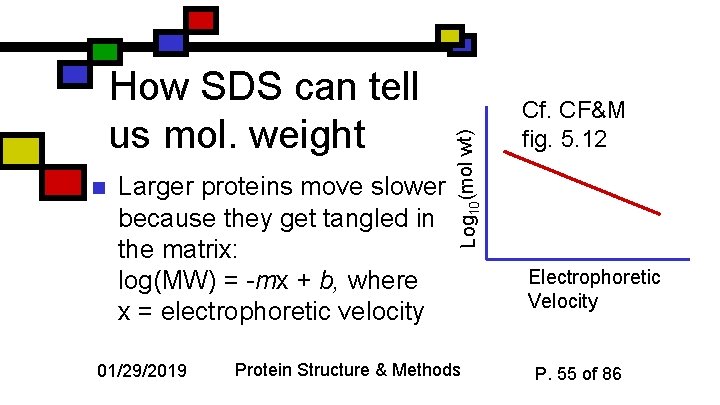
n Larger proteins move slower because they get tangled in the matrix: log(MW) = -mx + b, where x = electrophoretic velocity 01/29/2019 Log 10(mol wt) How SDS can tell us mol. weight Protein Structure & Methods Cf. CF&M fig. 5. 12 Electrophoretic Velocity P. 55 of 86
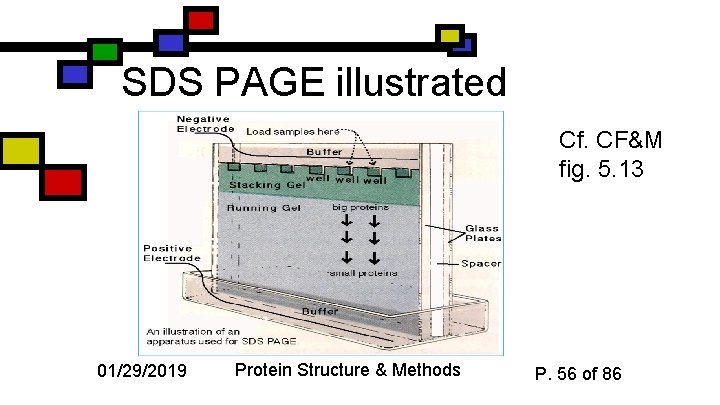
SDS PAGE illustrated Cf. CF&M fig. 5. 13 01/29/2019 Protein Structure & Methods P. 56 of 86
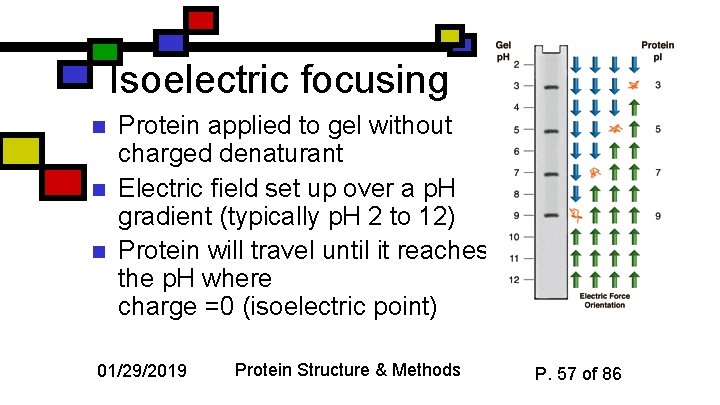
Isoelectric focusing n n n Protein applied to gel without charged denaturant Electric field set up over a p. H gradient (typically p. H 2 to 12) Protein will travel until it reaches the p. H where charge =0 (isoelectric point) 01/29/2019 Protein Structure & Methods P. 57 of 86

Using Isoelectric Focusing n n Sensitive to single changes in charge (e. g. asp asn) Readily used preparatively with samples that are already semi-pure 01/29/2019 Protein Structure & Methods P. 58 of 86

Applying this method n n Spectroscopy is more relevant for identification of moieties than for structure determination Quenching of fluorescence sometimes provides structural information 01/31/2019 Protein Methods & Functions P. 59 of 86

Mass spectrometry as an analytical tool n n n Mass spectrometry separates molecular species according to their mass/charge value It’s been used in chemistry for a century but couldn’t be applied to proteins until two techniques where developed in the 1980’s that preserved their properties: Electrospray and MALDI; Cf. CF&M 5 A 01/31/2019 Protein Methods & Functions P. 60 of 86

i. Clicker quiz question 4 4. A protein has p. I = 4. At p. H=7 its charge will be n (a) positive n (b) negative n (c) neutral n (d) insufficient information provided. 01/29/2019 Protein Structure & Methods P. 61 of 86

i. Clicker question 5 5. Which of the following techniques does not separate proteins by size? n (a) SDS-PAGE n (b) Size-exclusion n (c) Isoelectric focusing n (d) Mass spectrometry n (e) All four of these separate by size. 01/29/2019 Protein Structure & Methods P. 62 of 86

Structure Methods!. . . Warning: Specialty Content! n n n I determine protein structures (and develop methods for determining protein structures) as my own research focus So it’s hard for me to avoid putting a lot of emphasis on this material But today I’m allowed to do that, because it’s one of the stated topics of the day. 01/31/2019 Protein Methods & Functions P. 63 of 86

How do we determine structure? (CF&M § 4. 4) n n We can distinguish between methods that require little prior knowledge (crystallography, NMR, Cryo. EM) and methods that answer specific questions (XAFS, fiber, …) This distinction isn’t entirely clear-cut 01/31/2019 Protein Methods & Functions P. 64 of 86

Crystallography: overview n n Crystals are translationally ordered 3 -D arrays of molecules Conventional solids are usually crystals Proteins have to be coerced into crystallizing … but once they’re crystals, they behave like other crystals, mostly 01/31/2019 Protein Methods & Functions P. 65 of 86

How are protein crystals unusual? n n Aqueous interactions required for crystal integrity: they disintegrate if dried Bigger unit cells (~10 nm, not 1 nm) Intermolecular forces are weak ionic forces Small # of unit cells and static disorder means they don’t scatter terribly well Determining 3 D structures is feasible but difficult n 01/31/2019 Protein Methods & Functions P. 66 of 86

Crystal structures: Fourier transforms of diffraction results n n n Position of spots tells you how big the unit cell is Intensity tells you what the contents are We’re using electromagnetic radiation, which behaves like a wave, exp(2 ik • x) = cos 2 k • x + isin 2 k • x 01/31/2019 Protein Methods & Functions P. 67 of 86

Relating ρ(r) to intensities n n n Therefore intensity Ihkl = C*|Fhkl|2 Fhkl is a complex coefficient in the Fourier transform of the electron density in the unit cell: (r) = (1/V) hkl Fhkl exp(-2 ih • r) Inverse of that: Fhkl = �V (r) exp(2 ih • r) 01/31/2019 Protein Methods & Functions P. 68 of 86
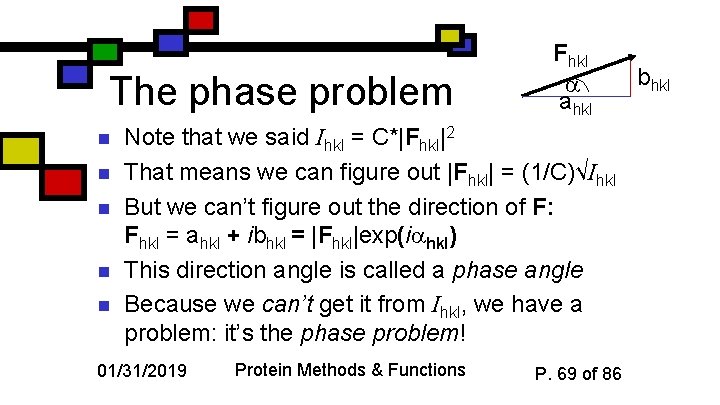
The phase problem n n n Fhkl ahkl Note that we said Ihkl = C*|Fhkl|2 That means we can figure out |Fhkl| = (1/C)√Ihkl But we can’t figure out the direction of F: Fhkl = ahkl + ibhkl = |Fhkl|exp(i hkl) This direction angle is called a phase angle Because we can’t get it from Ihkl, we have a problem: it’s the phase problem! 01/31/2019 Protein Methods & Functions P. 69 of 86 bhkl
What can we learn? n n n Electron density map + sequence we can determine the positions of all the non-H atoms in the protein—maybe! Best resolution possible: Dmin = / 2 Realistic resolution usually poorer than that 01/31/2019 Protein Methods & Functions P. 70 of 86

What else can we learn? n n Hydrogen positions can be inferred, especially if you are able to get highresolution data (see next slide) Atomic mobility can estimated for intermediate to high resolution data 01/31/2019 Protein Methods & Functions P. 71 of 86

Limitations of resolution n Often the crystal doesn’t diffract that well, so Dmin is larger— 1. 5Å, 2. 5Å, or worse Dmin ~ 2. 5Å tells us where backbone and most side-chain atoms are Dmin ~ 1. 2Å: all protein non-H atoms, most solvent, some disordered atoms; some H’s 01/31/2019 Protein Methods & Functions P. 72 of 86

What does this look like? n n Takes some experience to interpret Automated fitting programs work pretty well with Dmin < 2. 1Å 01/31/2019 ATP binding to a protein of unknown function: S. H. Kim Protein Methods & Functions P. 73 of 86

Macromolecular NMR n n n NMR is a mature field Depends on resonant interaction between EM fields and unpaired nucleons (1 H, 15 N, 31 S) Raw data yield interatomic distances Conventional spectra of proteins are too muddy to interpret Multi-dimensional (2 -4 D) techniques: initial resonances coupled with others 01/31/2019 Protein Methods & Functions P. 74 of 86
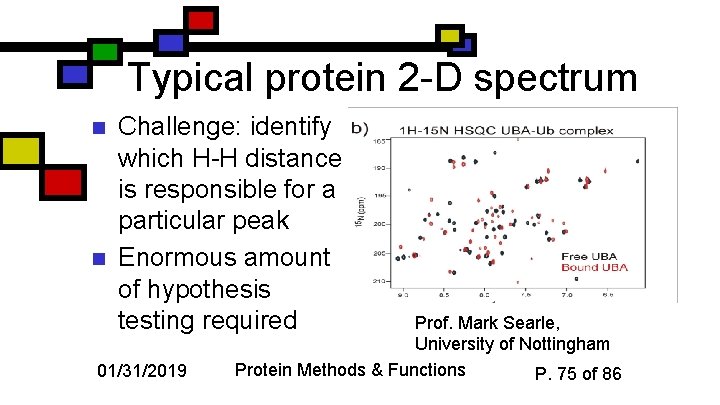
Typical protein 2 -D spectrum n n Challenge: identify which H-H distance is responsible for a particular peak Enormous amount of hypothesis testing required 01/31/2019 Prof. Mark Searle, University of Nottingham Protein Methods & Functions P. 75 of 86

Results n n Often there’s a family of structures that satisfy the NMR data equally well Can be portrayed as a series of threads tied down at unambiguous assignments They portray the protein’s structure in solution Ambiguities partly represent real molecular diversity; but they also represent atoms that area in truth well-defined, but the NMR data don’t provide the unambiguous assignment 01/31/2019 Protein Methods & Functions P. 76 of 86

Comparing NMR to X-ray n n n NMR family of structures often reflects real conformational heterogeneity Nonetheless, it’s hard to visualize what’s happening at the active site at any instant Hydrogens sometimes well-located in NMR; they’re often the least defined atoms in an Xray structure 01/31/2019 Protein Methods & Functions P. 77 of 86
NMR vs. X-ray, continued n n The NMR structure is obtained in solution! Hard to make NMR work if MW > 55 k. Da, and even when you can, it takes a lot of computer time 01/31/2019 Protein Methods & Functions P. 78 of 86

What does it mean when NMR and X-ray structures differ? n n Lattice forces may have tied down or moved surface amino acids in X-ray structure NMR may have errors in it X-ray may have errors in it (measurable) X-ray structure often closer to true atomic resolution X-ray structure has built-in reliability checks n 01/31/2019 Protein Methods & Functions P. 79 of 86

n n n Cryoelectron microscopy Like X-ray crystallography, EM damages the samples Samples analyzed < 100 K survive better 2 -D arrays of molecules n Spatial averaging to improve resolution n Discerning details ≥ 4Å resolution Can be used with crystallography n 01/31/2019 Protein Methods & Functions P. 80 of 86

Solution scattering n n n Proteins in solution scatter X-rays in characteristic ways Low-resolution structural information available Does not require crystals Until ~ 2000 you needed high [protein] Thanks to Bio. CAT, SAXS on dilute proteins is becoming more feasible Hypothesis-based analysis n 01/31/2019 Protein Methods & Functions P. 81 of 86

Fiber Diffraction Some proteins, like many DNA molecules, possess approximate fibrous order (2 -D ordering) n Produce characteristic fiber diffraction patterns n Collagen, muscle proteins, filamentous. Protein viruses Methods & Functions 01/31/2019 n P. 82 of 86
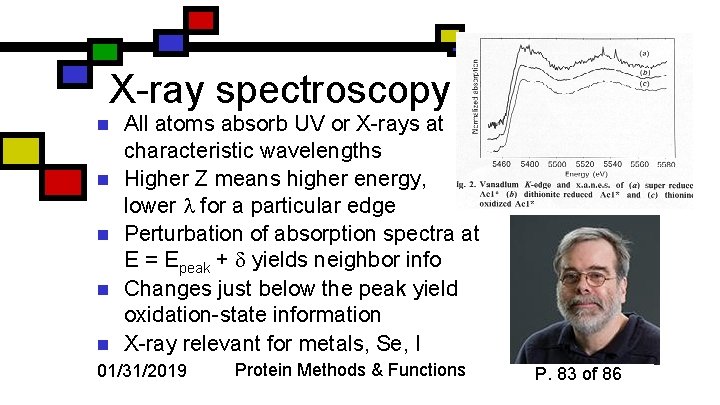
X-ray spectroscopy n n n All atoms absorb UV or X-rays at characteristic wavelengths Higher Z means higher energy, lower for a particular edge Perturbation of absorption spectra at E = Epeak + yields neighbor info Changes just below the peak yield oxidation-state information X-ray relevant for metals, Se, I 01/31/2019 Protein Methods & Functions P. 83 of 86

n n n Mass spectrometry as a structural tool MS tells you molecular weights Can give high precision in m/m Not inherently a way of determining structure Can distinguish oligomeric state Coupled with proteolytic digestion, it can be used to find fragmentation patterns 01/31/2019 Protein Methods & Functions P. 84 of 86

Circular dichroism n n n Proteins in solution can rotate polarized light Amount of rotation varies with Effect depends on interaction with secondary structure elements, esp. 01/31/2019 Protein Methods & Functions P. 85 of 86
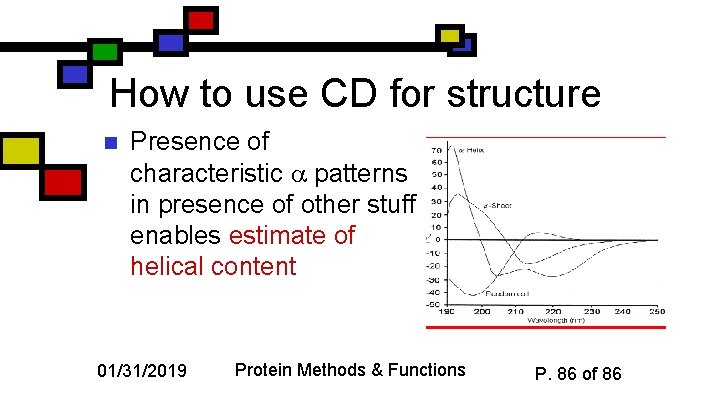
How to use CD for structure n Presence of characteristic patterns in presence of other stuff enables estimate of helical content 01/31/2019 Protein Methods & Functions P. 86 of 86
- Slides: 86