PROTEIN SEONDARY SUPERSECONDARY STRUCTURE PREDICTION WITH HMM By

PROTEIN SEONDARY & SUPER-SECONDARY STRUCTURE PREDICTION WITH HMM By En-Shiun Annie Lee CS 882 Protein Folding Instructed by Professor Ming Li

0 OUTLINE. 1. 2. 3. 4. Introduction Problem Methods (4) HMM Examples (3) a. Segmentation HMM b. Profile HMM c. Conditional Random Field 5. Proposal

1 INTRODUCTION. 1. 2. 3. 4. Introduction * Problem Methods (4) HMM Examples (3) a. Segmentation HMM b. Profile HMM c. Conditional Random Field 5. Proposal

1 Genomics. • Achievements in Genomic – BLAST (Basic Local Alignment Search Tool) • most cited paper published in 1990 s • more than 15, 000 times – Human genome project • Completion April 2003

1 Proteomics. • Precedence to Proteomics – Protein Data Bank (PDB) • 40, 132 structures • cited more than 6, 000 times
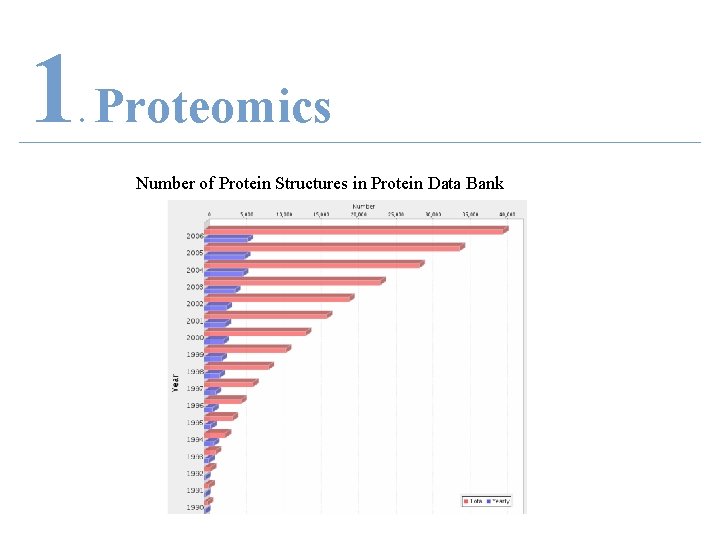
1 Proteomics. Number of Protein Structures in Protein Data Bank

1 Secondary Structure. • Importance – The known secondary structure may be used as an input for the tertiary structure predictions.
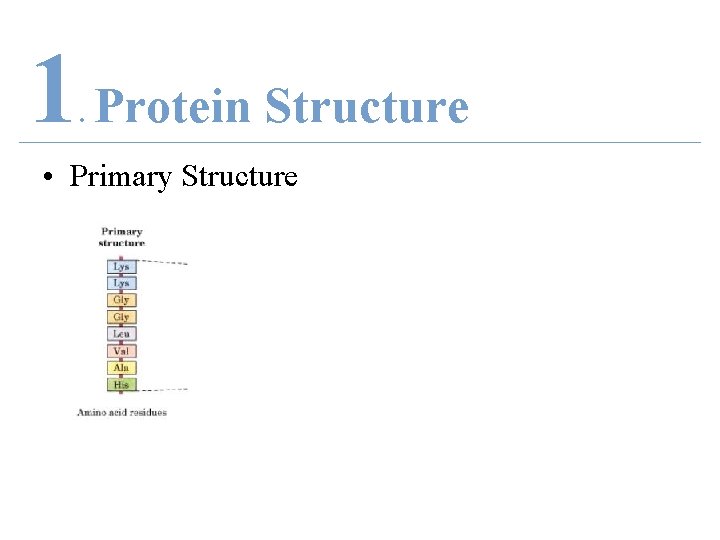
1 Protein Structure. • Primary Structure
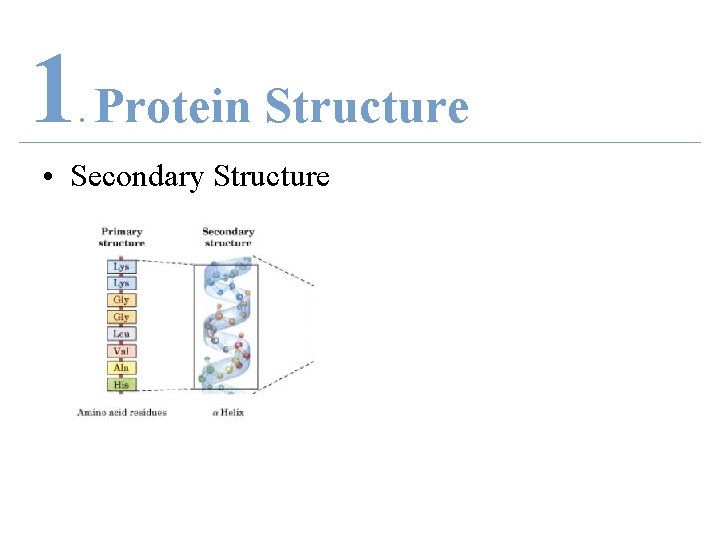
1 Protein Structure. • Secondary Structure
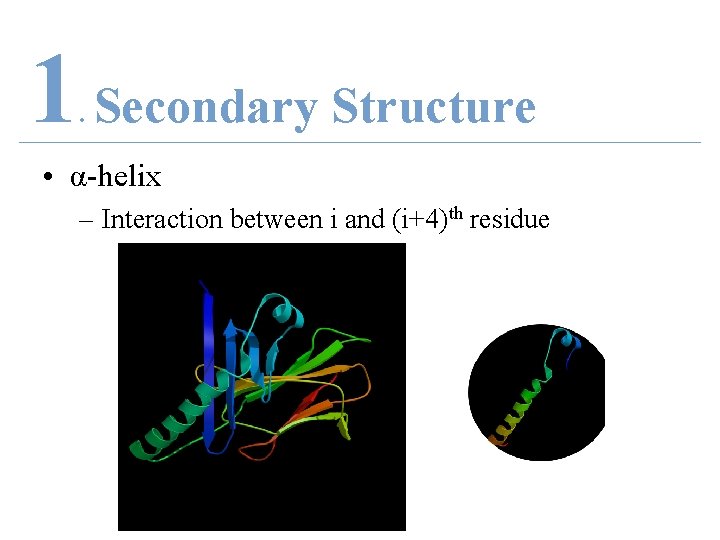
1 Secondary Structure. • α-helix – Interaction between i and (i+4)th residue
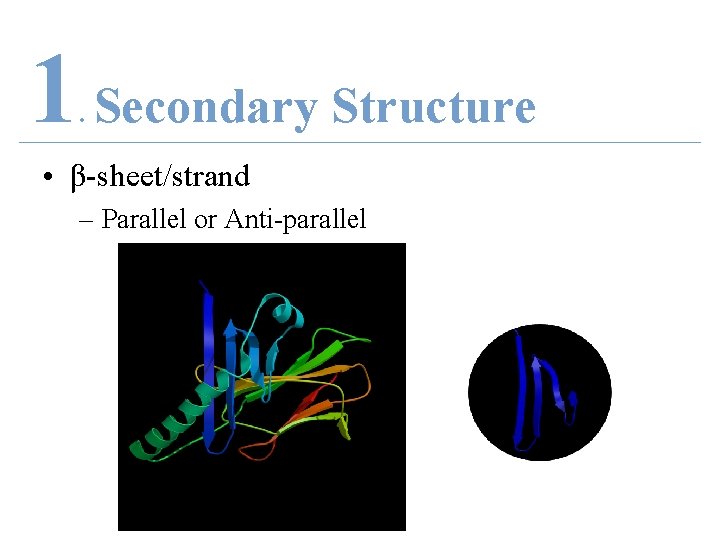
1 Secondary Structure. • β-sheet/strand – Parallel or Anti-parallel
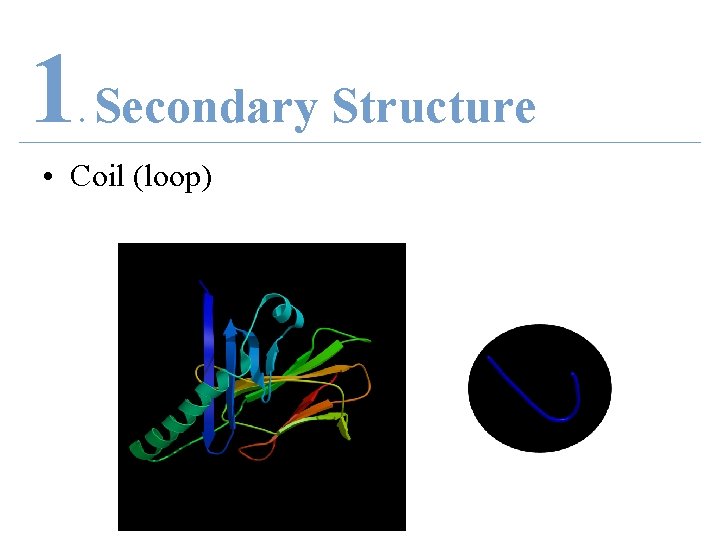
1 Secondary Structure. • Coil (loop)
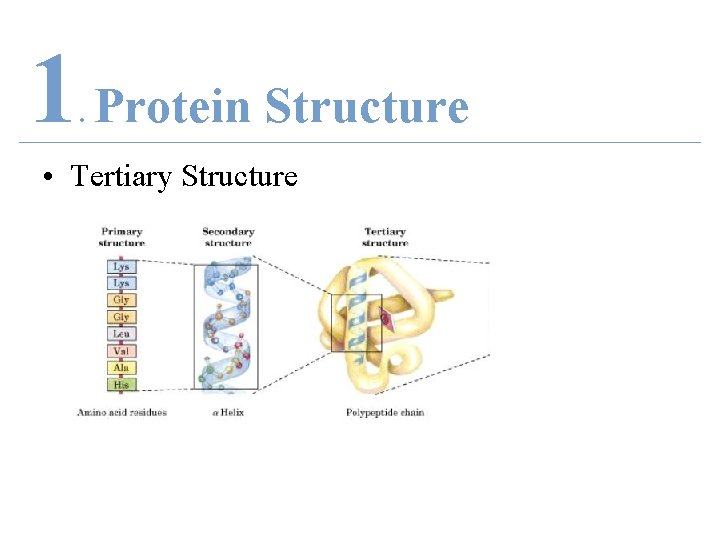
1 Protein Structure. • Tertiary Structure
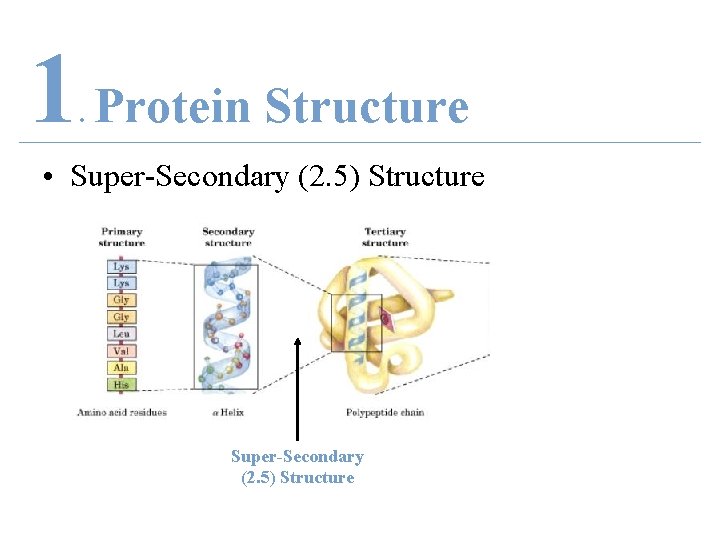
1 Protein Structure. • Super-Secondary (2. 5) Structure
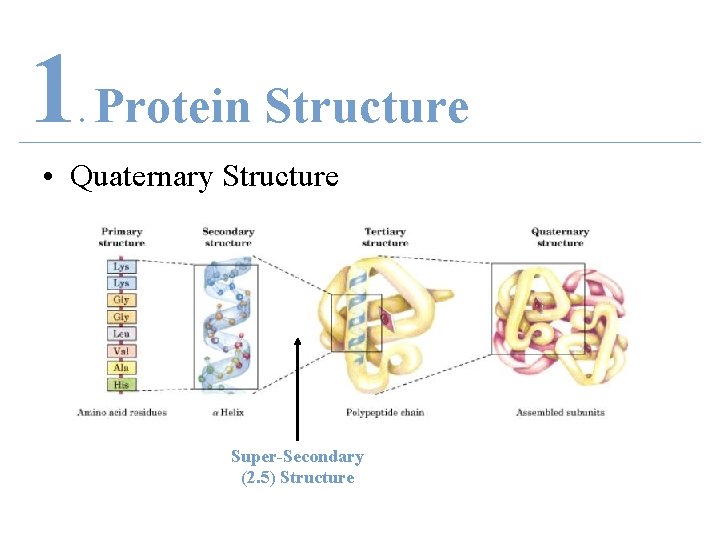
1 Protein Structure. • Quaternary Structure Super-Secondary (2. 5) Structure

2 PROBLEM. 1. 2. 3. 4. Introduction Problem * Methods (4) HMM Examples (3) a. Segmentation HMM b. Profile HMM c. Conditional Random Field 5. Proposal

2 Secondary Structure. • Problem – Given: • A primary sequence of amino acids – a 1 a 2…an – Find: • Secondary structure of each ai as – α-helix = H – β-strand = E * – coil = C

2 Secondary Structure. • Example – Given: • Primary Sequence – GHWIATRGQLIREAYEDYRHFSSECPFIP – Find: • Secondary Structure Element – CEEEEECHHHHHHCCCHHCCCCCC – Note: segments

2 Prediction Quality. • Three-state prediction accuracy – Q 3 = # of correctly predicted residues total # of number of residues – Q , Qβ , Qc – Q 3 for random prediction is 33% – Theoretical limit Q 3=90%.

2 Prediction Quality. • Segment Overlap (SOV) – Higher penalties for core segment regions • Matthews Correlation Coefficients (MCC) – Prediction errors made for each state

2 True Structures. • Three dimensional PDB data – DSSP (Dictionary of Secondary Structure of Proteins) • 8 states – – – – H = alpha helix G = 310 - helix I = 5 helix (pi helix) E = extended strand (beta ladder) E B = residue in isolated beta-bridge E T = hydrogen bonded turn S = bend C = coil – STRIDE H H H C C C

3 METHODS. 1. 2. 3. 4. Introduction Problem Methods (4) * HMM Examples (3) a. Segmentation HMM b. Profile HMM c. Conditional Random Field 5. Proposal
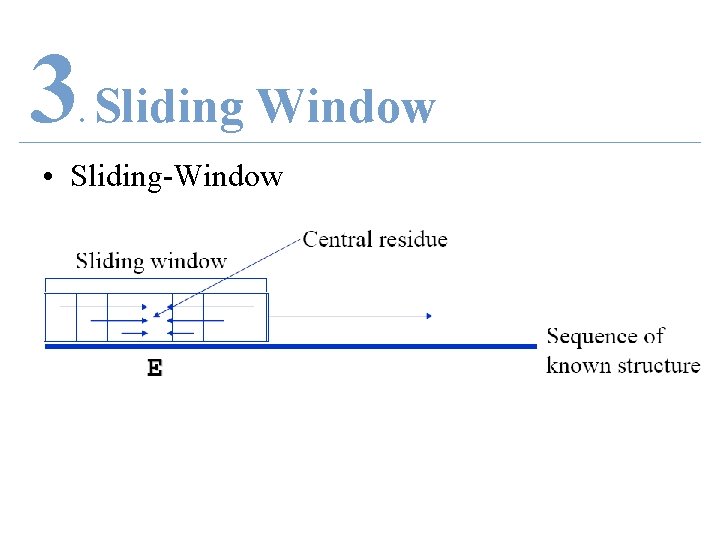
3 Sliding Window. • Sliding-Window
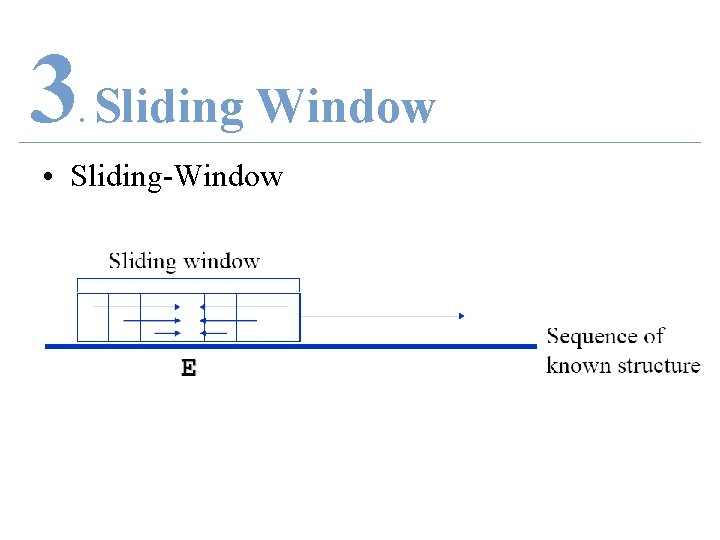
3 Sliding Window. • Sliding-Window
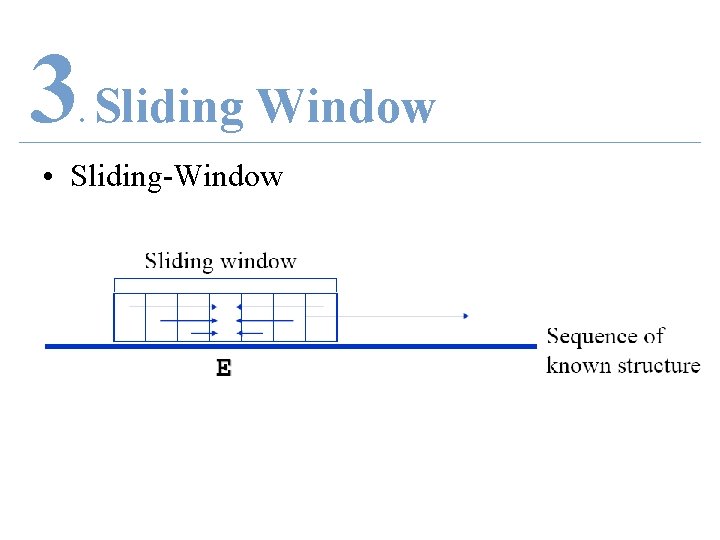
3 Sliding Window. • Sliding-Window
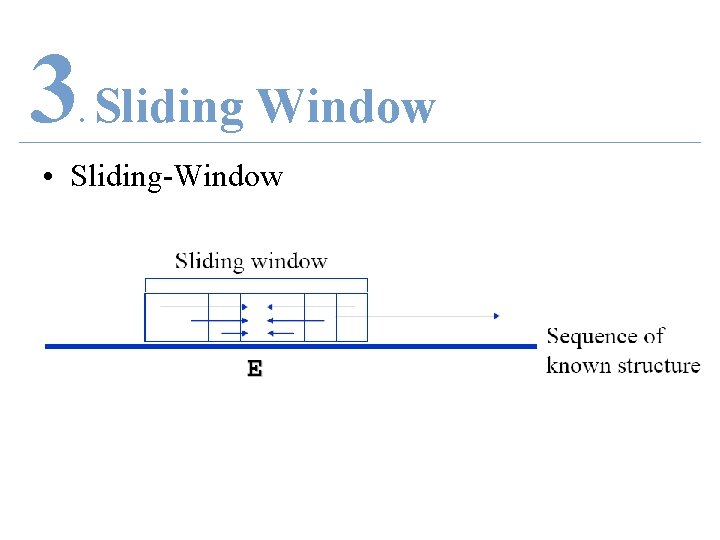
3 Sliding Window. • Sliding-Window

3 Four Methods. a. b. c. d. Statistical Method Neural Network Support Vector Machine Hidden Markov Model
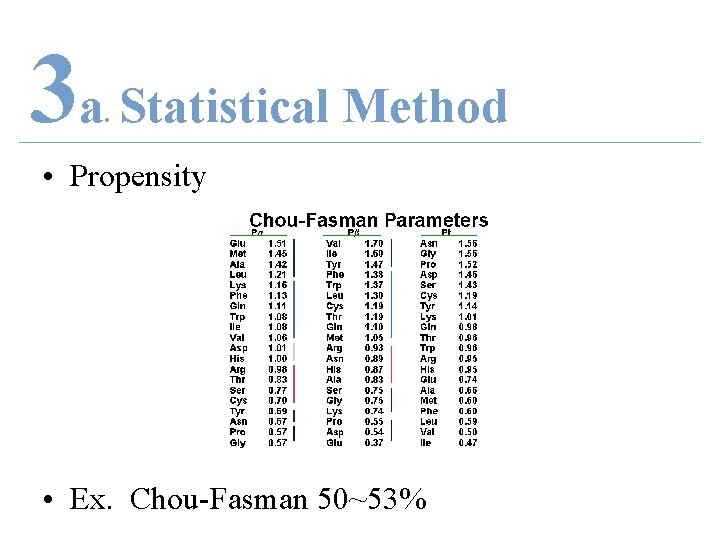
3 a Statistical Method. • Propensity • Ex. Chou-Fasman 50~53%
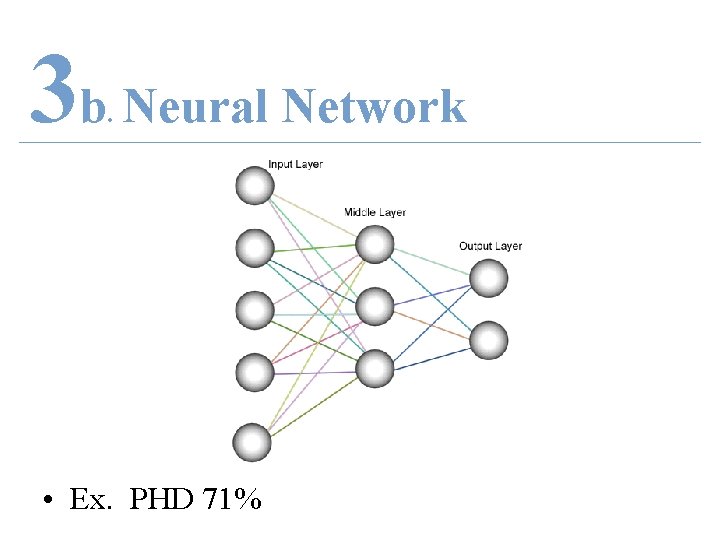
3 b Neural Network. • Ex. PHD 71%
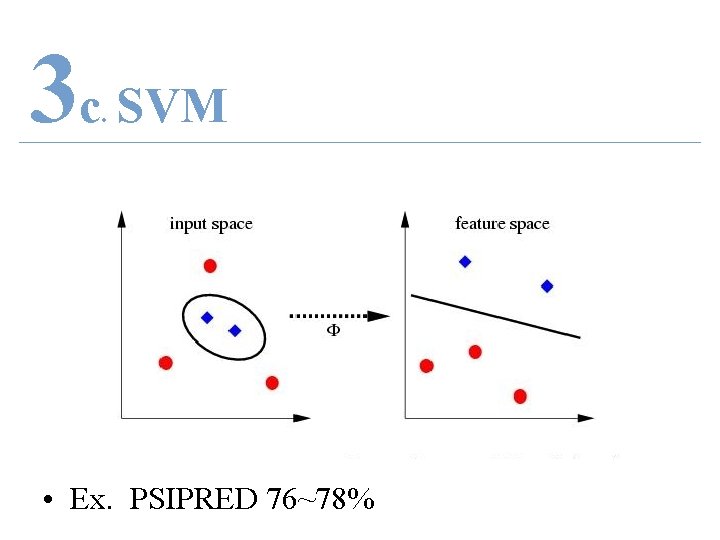
3 c SVM. • Ex. PSIPRED 76~78%
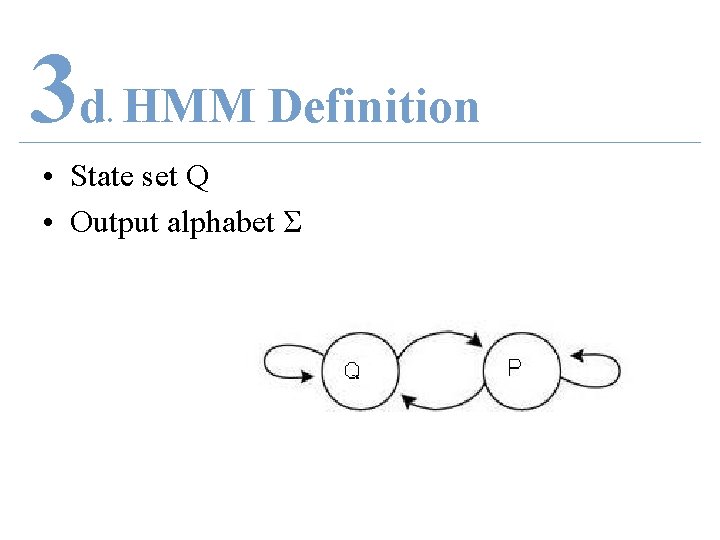
3 d HMM Definition. • State set Q • Output alphabet Σ
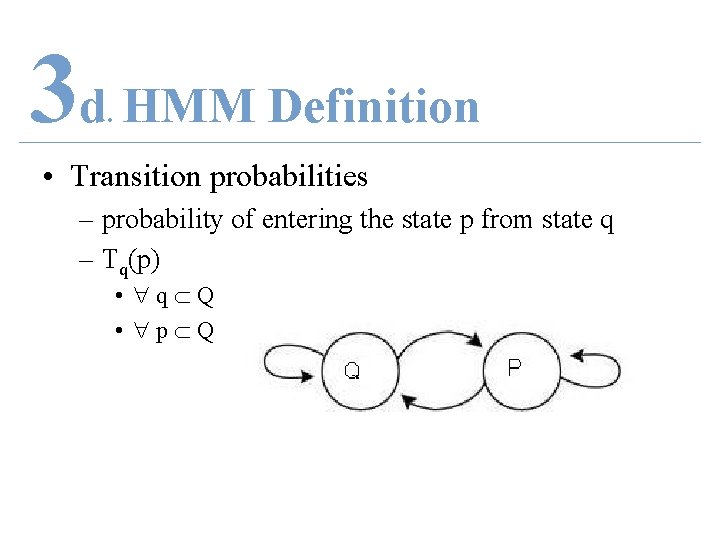
3 d HMM Definition. • Transition probabilities – probability of entering the state p from state q – Tq(p) • q Q • p Q

3 d HMM Definition. • Emission probabilities – probability emits each letter of Σ from state q – Eq(ai) • ai Σ • q Q

3 d HMM Decoding. • Problem – Given: • HMM = (Q, Σ, E, T) and • Sequence S – Where S = S 1, S 2, …, Sn – Find: • Most probable path of state gone through to get S – Where X = X 1, X 2, …, Xn = state sequence
![4 HMM Decoding. • Optimize – Pr [ S , X ] • X 4 HMM Decoding. • Optimize – Pr [ S , X ] • X](http://slidetodoc.com/presentation_image_h/12edc0bfe37edf93c476c4d9ecd2a4dd/image-35.jpg)
4 HMM Decoding. • Optimize – Pr [ S , X ] • X = X 1, X 2, …, Xn = state sequence • S = S 1, S 2, …, Sn – Pr [ S | X ]
![4 HMM Decoding. • Dynamic programming – Memoryless – Pr [Xn|Sn] = Pr [Xn-1|Sn-1] 4 HMM Decoding. • Dynamic programming – Memoryless – Pr [Xn|Sn] = Pr [Xn-1|Sn-1]](http://slidetodoc.com/presentation_image_h/12edc0bfe37edf93c476c4d9ecd2a4dd/image-36.jpg)
4 HMM Decoding. • Dynamic programming – Memoryless – Pr [Xn|Sn] = Pr [Xn-1|Sn-1] Tn-1[Xn] EXn [Sn]

4 HMM EXAMPLES. 1. 2. 3. 4. Introduction Problem Methods (4) HMM Examples (3) * a. Segmentation HMM b. Profile HMM c. Conditional Random Field 5. Proposal

4 a SEMI-HMM. 1. 2. 3. 4. Introduction Problem Methods (4) HMM Examples (3) a. Semi-HMM * b. Profile HMM c. Conditional Random Field 5. Proposal
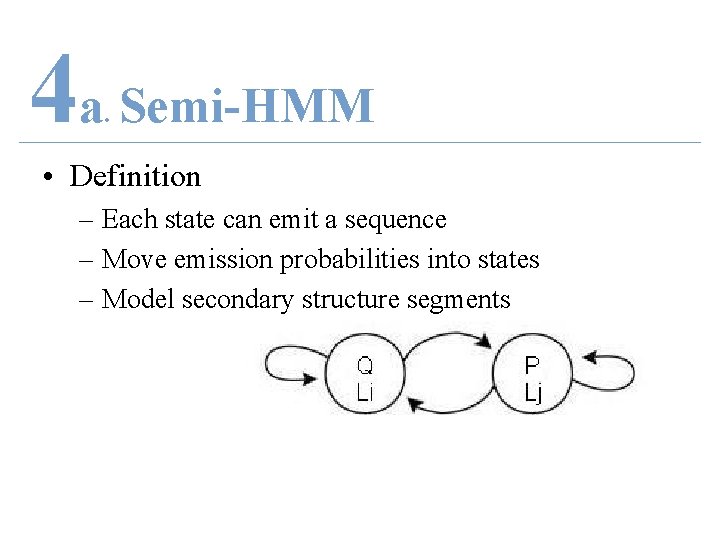
4 a Semi-HMM. • Definition – Each state can emit a sequence – Move emission probabilities into states – Model secondary structure segments

4 a Segmentation. • Sequence Segments
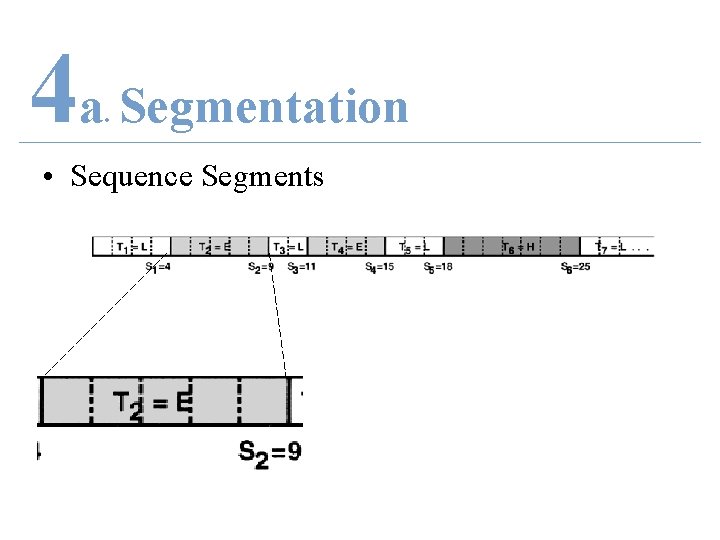
4 a Segmentation. • Sequence Segments
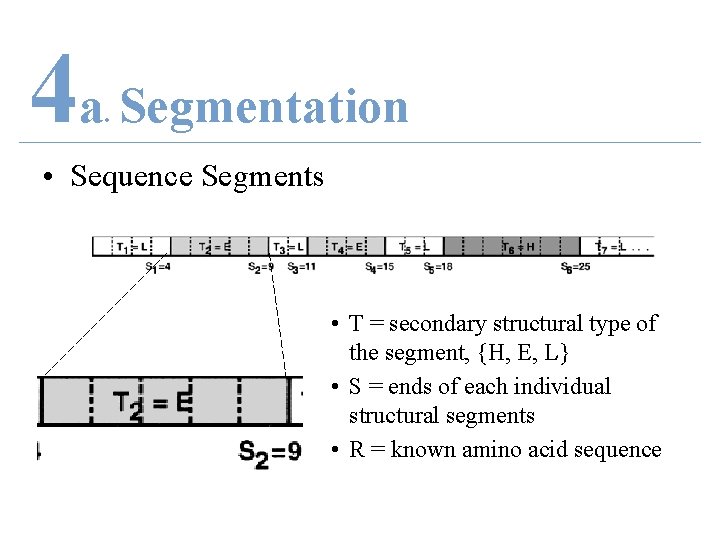
4 a Segmentation. • Sequence Segments • T = secondary structural type of the segment, {H, E, L} • S = ends of each individual structural segments • R = known amino acid sequence
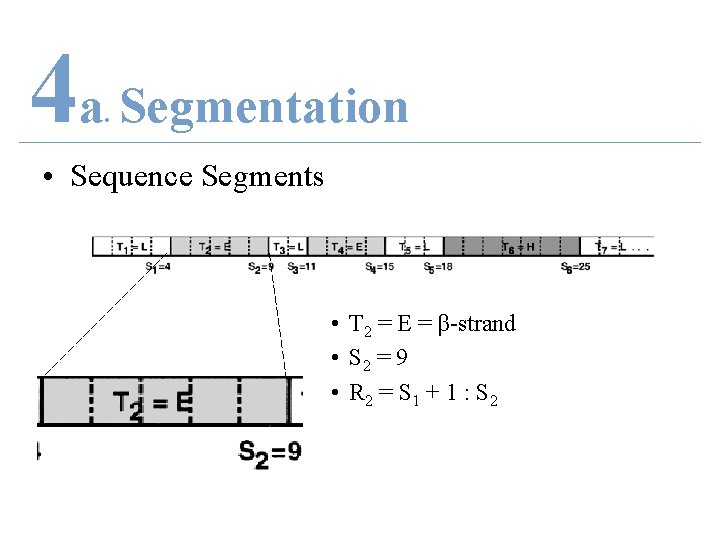
4 a Segmentation. • Sequence Segments • T 2 = E = β-strand • S 2 = 9 • R 2 = S 1 + 1 : S 2

4 a Bayesian. • Bayesian Formulation • R = Sequence of ALL amino acid residues • S = End of the segments • T = Secondary structural type of the segments – {H, E, L}

4 a Bayesian. • Bayesian Formulation 1. Likelihood 2. Priori Probability 3. Constant (S, T) dropped

4 a Bayesian. • Likelihood • m = Total number of segments • Sj = End of the jth segments • Tj = Secondary structural type of the jth segments

4 a Bayesian. • Likelihood

4 a Bayesian. • Likelihood

4 a Bayesian. • Likelihood N-terminus Internal C-terminus
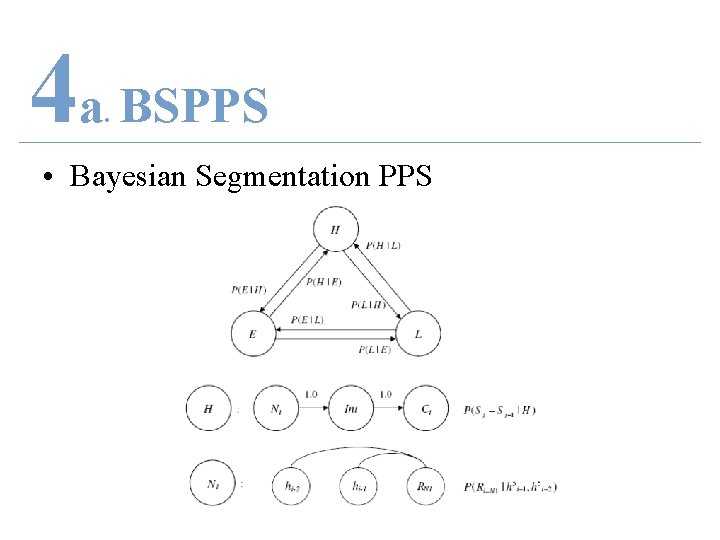
4 a BSPPS. • Bayesian Segmentation PPS
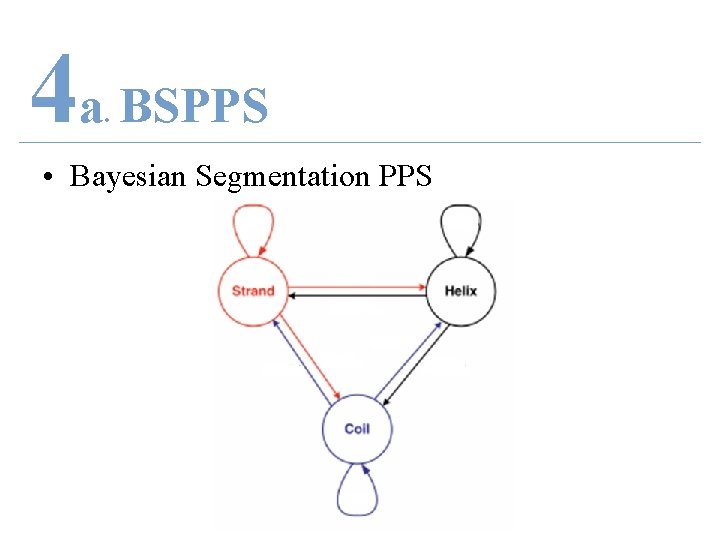
4 a BSPPS. • Bayesian Segmentation PPS

4 a Results. • Better than PSIPRED – (w/o homology information)
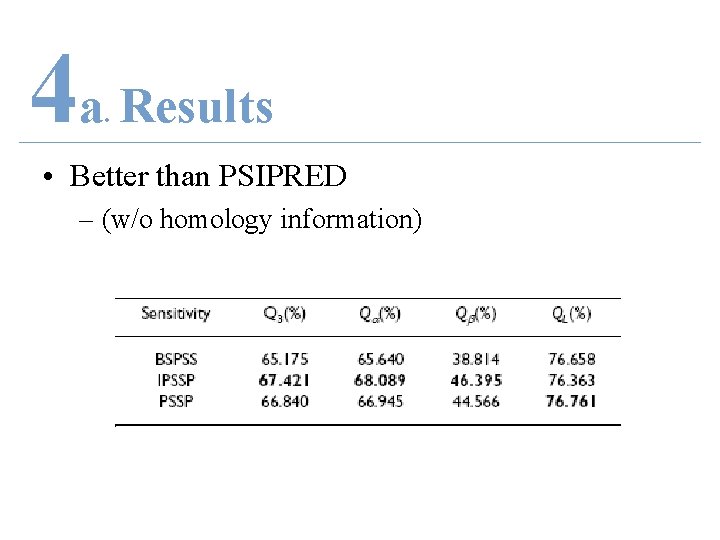
4 a Results. • Better than PSIPRED – (w/o homology information)

4 b PROFILE-HMM. 1. 2. 3. 4. Introduction Problem Methods (4) HMM Examples (3) a. Semi-HMM b. Profile HMM * c. Conditional Random Field 5. Proposal
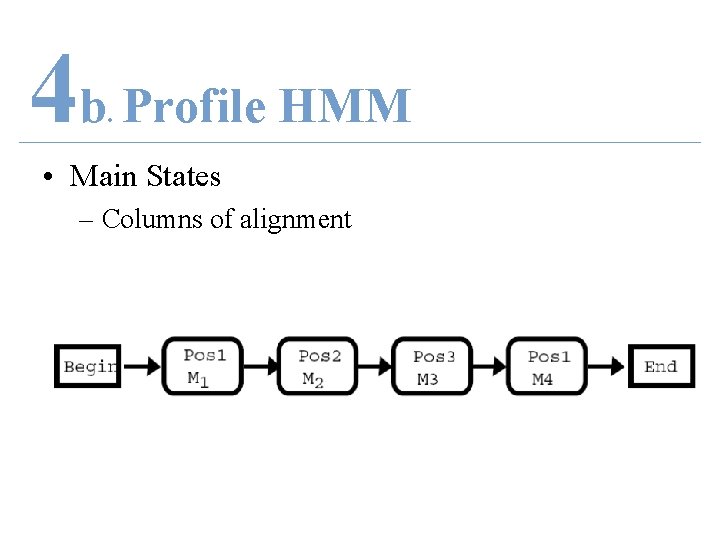
4 b Profile HMM. • Main States – Columns of alignment
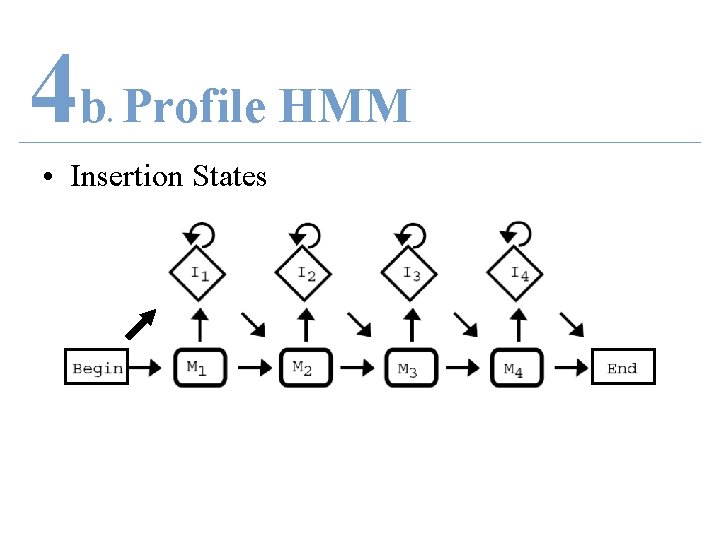
4 b Profile HMM. • Insertion States
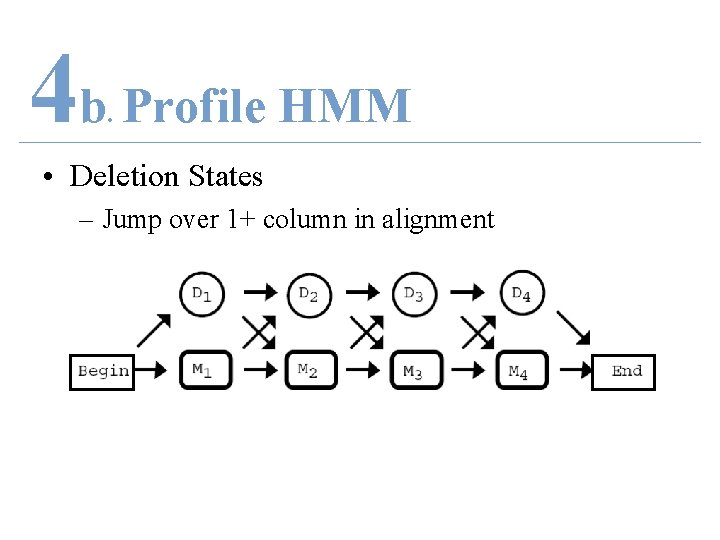
4 b Profile HMM. • Deletion States – Jump over 1+ column in alignment

4 b Profile HMM. • Combined

4 b HMMSTR. • HMM for local protein STRucture

4 b HMMSTR. • HMM for local protein STRucture • Pronounced “hamster”

4 b I-Site Library. • I-sites Library – Motif = short basic structural fragments • 3~19 residues • 262 motifs • Highly predictable – Non-redundant PDB data (<25% similarity) – Fold uniquely across protein family – Exhaustive motif clustering

4 b Build HMM. • States – Amino acid sequence and – Structural attribute • Transition from state – Adjacent positions in motif – No gap or insertion states
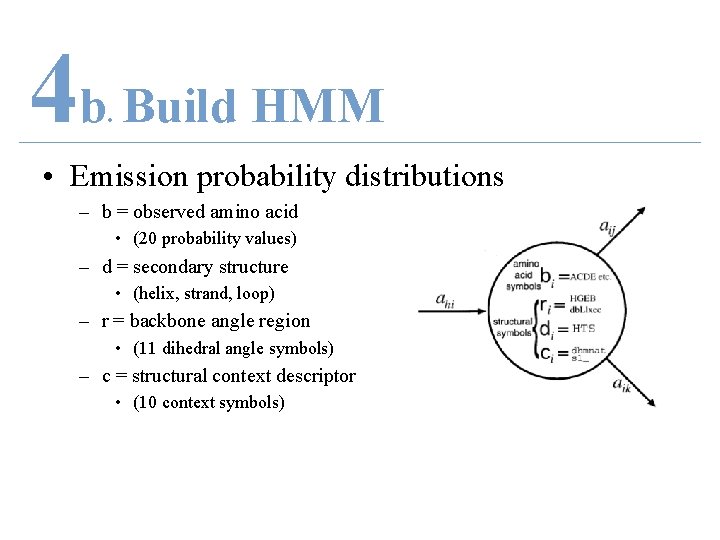
4 b Build HMM. • Emission probability distributions – b = observed amino acid • (20 probability values) – d = secondary structure • (helix, strand, loop) – r = backbone angle region • (11 dihedral angle symbols) – c = structural context descriptor • (10 context symbols)

4 b Build HMM. • Model I-site Library – Each 262 motif is a chain in HMM – Merge states base on similarity of • Sequence • Structure
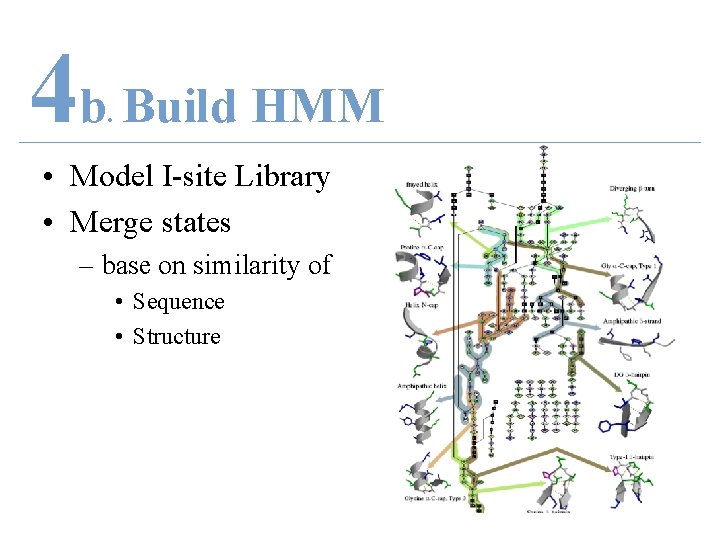
4 b Build HMM. • Model I-site Library • Merge states – base on similarity of • Sequence • Structure
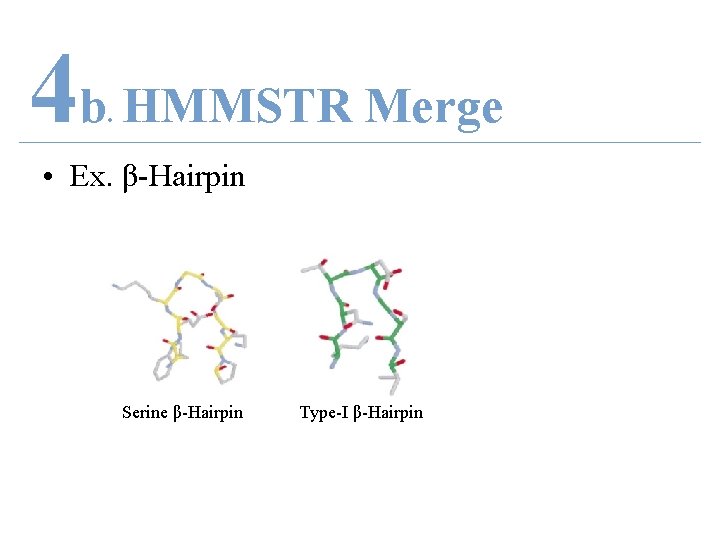
4 b HMMSTR Merge. • Ex. β-Hairpin Serine β-Hairpin Type-I β-Hairpin
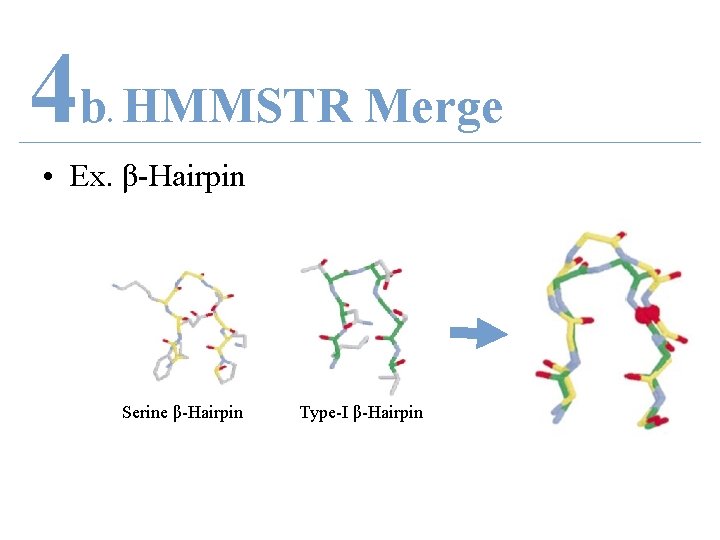
4 b HMMSTR Merge. • Ex. β-Hairpin Serine β-Hairpin Type-I β-Hairpin

4 b HMMSTR Merge. • Ex. β-Hairpin

4 b HMMSTR Merge. • Ex. β-Hairpin

4 b HMMSTR Training. • Input: PDB proteins • Find – best state sequence for sequence – probability distribution of one amino acid • Integrate 3 data set – Aligned probability distribution – Amino acid and context information – Contact map
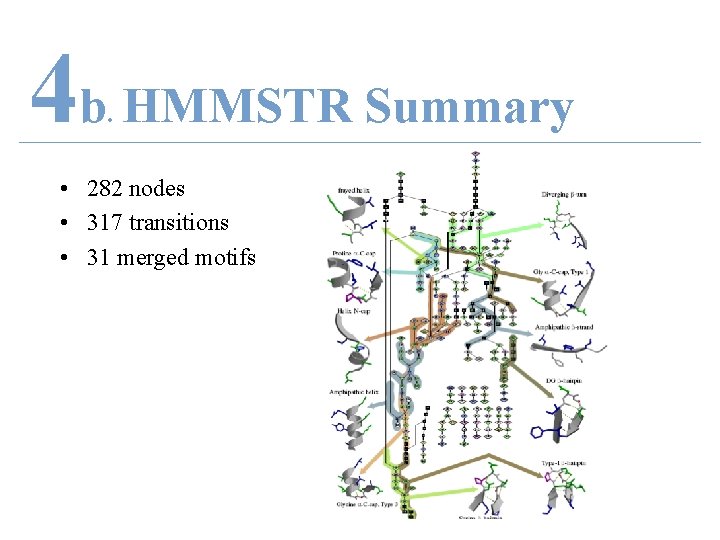
4 b HMMSTR Summary. • 282 nodes • 317 transitions • 31 merged motifs

4 b HMMSTR Summary. • Introduce structural context on level of supersecondary structure • Predict higher-order 3 D tertiary structure – Side-result = predict 1 D secondary structure

4 b PROFILE-HMM. 1. 2. 3. 4. Introduction Problem Methods (4) HMM Examples (3) a. Semi-HMM b. Profile HMM c. Conditional Random Field * 5. Proposal

4 c HMM Disadvantages. • Does not model – Multiple interacting features – Long-range dependencies • Strict independence assumptions

4 c Conditional Model. • Allow – Arbitrary features – Non-independent features • Transition probability – With respect to past and future observations
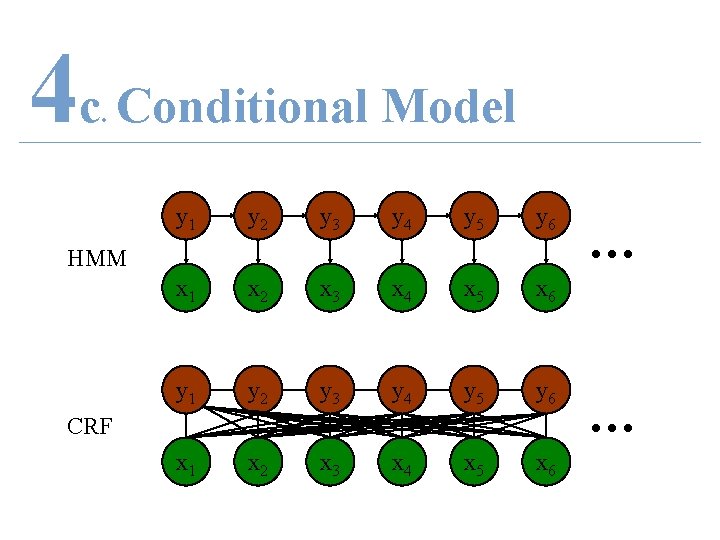
4 c Conditional Model. y 1 y 2 y 3 y 4 y 5 y 6 x 1 x 2 x 3 x 4 x 5 x 6 HMM CRF … …

4 c Random Field. • Random Field (Undirected graphical model) – Let G = (Y, E) be a graph • Where each vertex Yv = a random variable – If P(Yv|all other Y)= P(Yv|neighbours of Yv) Then Y is a random field

4 c Random Field. • Example: – P(Y 5 | all other Y) = P(Y 5 | Y 4, Y 6)

4 c Conditional RF. • Conditional Random Field – Let X = r. v. data sequences to be labeled • observations – Let Y = r. v. corresponding label sequences • labels – Let G = (V, E) be a graph • S. t. Y = (Yv)v Y so Y is indexed by vertices of G – If P(Yv | X, Yw w≠v) = P(Yv | X, Yw, w~v) Then (X, Y) is a random field
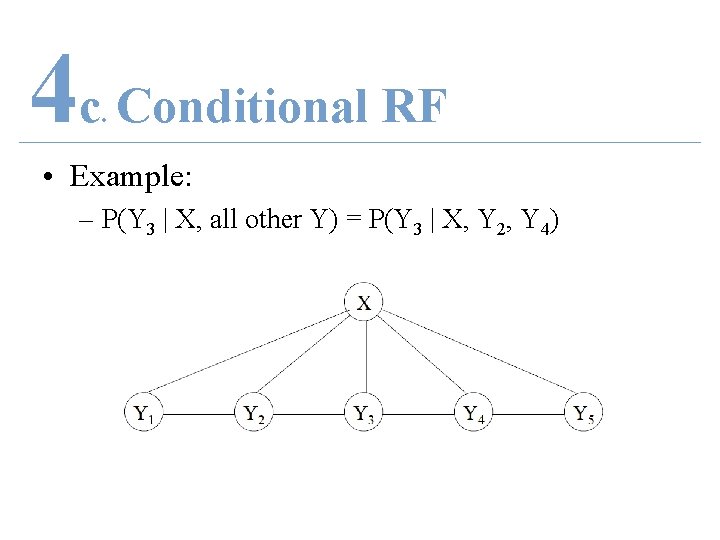
4 c Conditional RF. • Example: – P(Y 3 | X, all other Y) = P(Y 3 | X, Y 2, Y 4)

4 c HMM vs. CRF. • HMM: – Maximize P(x, y|θ)=P(y|x, θ)P(x|θ) – Transition and emission probabilities – Transition/emission base only one x • CRF: – Maximize P(y|x, θ) – Feature function f(i, j, k) – Feature function base on all x
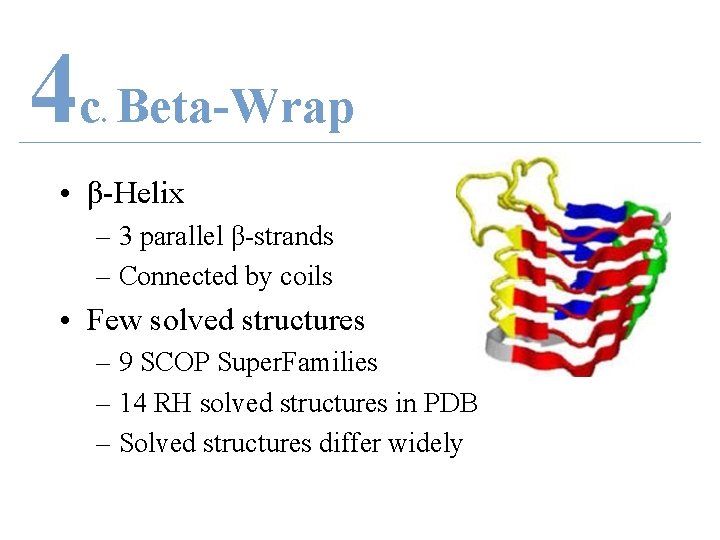
4 c Beta-Wrap. • β-Helix – 3 parallel β-strands – Connected by coils • Few solved structures – 9 SCOP Super. Families – 14 RH solved structures in PDB – Solved structures differ widely

4 c Graph Definition. • Let G = (V, E 1, E 2) be a graph – V = Nodes/States = Secondary structures – Edges = interactions • E 1 – Edges between adjacent neighbors – Implied in the model • E 2 – Edges for long-term interactions – Explicitly considered
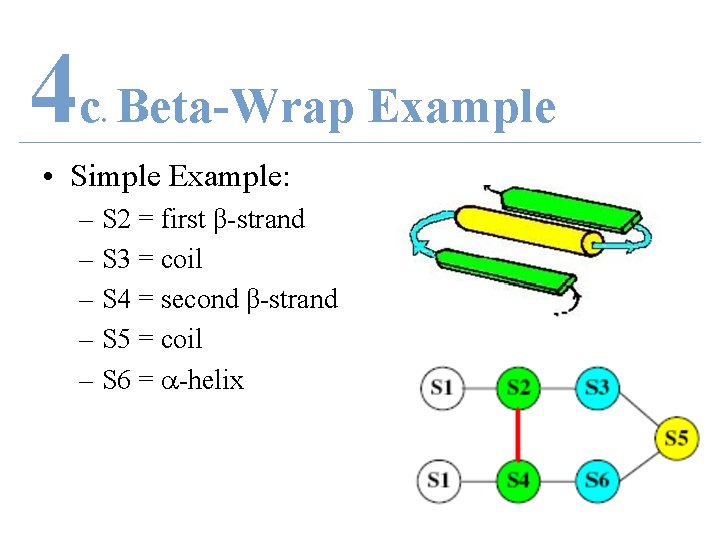
4 c Beta-Wrap Example. • Simple Example: – S 2 = first β-strand – S 3 = coil – S 4 = second β-strand – S 5 = coil – S 6 = -helix
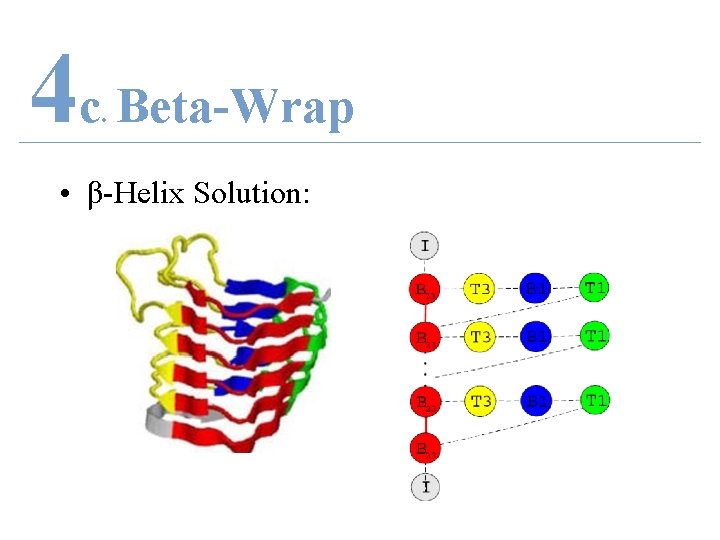
4 c Beta-Wrap. • β-Helix Solution:

5 PROPOSAL. 1. 2. 3. 4. Introduction Problem Methods (4) HMM Examples (3) a. Segmentation HMM b. Profile HMM c. Conditional Random Field 5. Proposal *

5 Difficulties. • Do not infer global interaction – i. e. Beta-sheet interactions • Protein structure definition constraint

5 Possible Future Work. • Novel methods of secondary structure prediction – Model as Integer Programming • Super-secondary structure prediction

5 Acknowledgement. • Professor Ming Li – Guidance in • knowledge and • expertise • Bioinformatics lab • Mentoring a “rookie” • Class • Attention and listening
- Slides: 89