Protein Structure Prediction by A Datalevel Parallel Proceedings

Protein Structure Prediction by A Data-level Parallel Proceedings of the 1989 ACM/IEEE conference on Supercomputing Speaker : Chuan-Cheng Lin Advisor : Prof. R. C. T. Lee CSIE National Chi Nan University 1

Outline l l l Concepts Introduction Approach Example Conclusions Reference 2
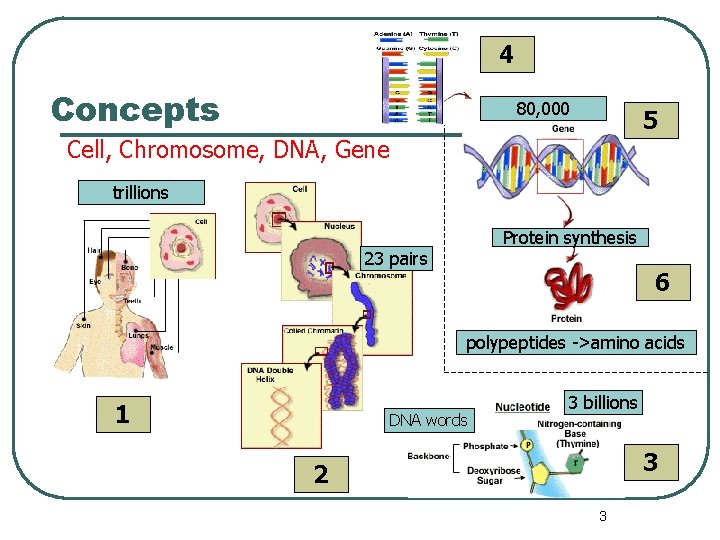
4 Concepts 80, 000 5 Cell, Chromosome, DNA, Gene trillions Protein synthesis 23 pairs 6 polypeptides ->amino acids 1 DNA words 3 billions 3 2 3
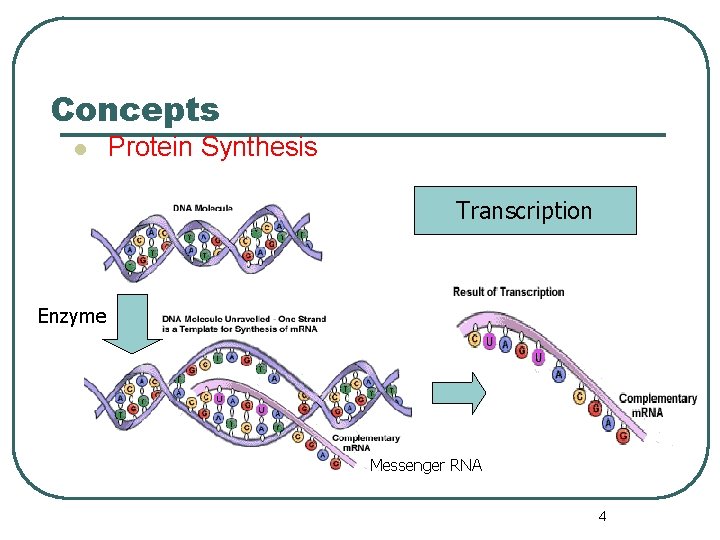
Concepts l Protein Synthesis Transcription Enzyme Messenger RNA 4
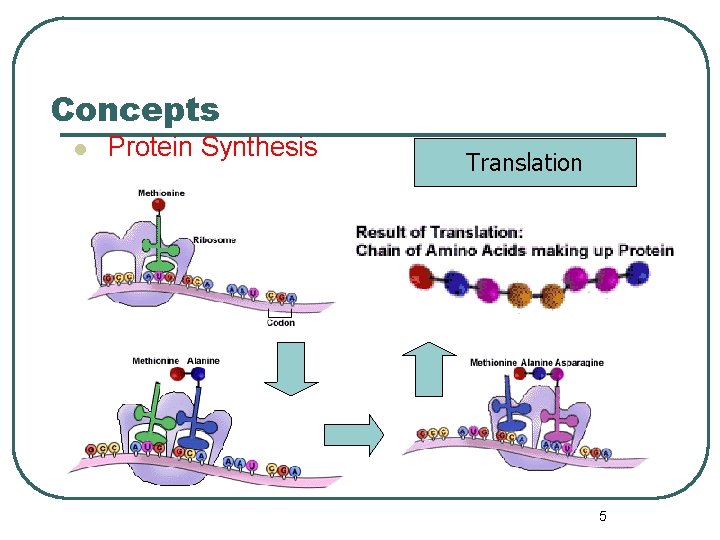
Concepts l Protein Synthesis Translation 5
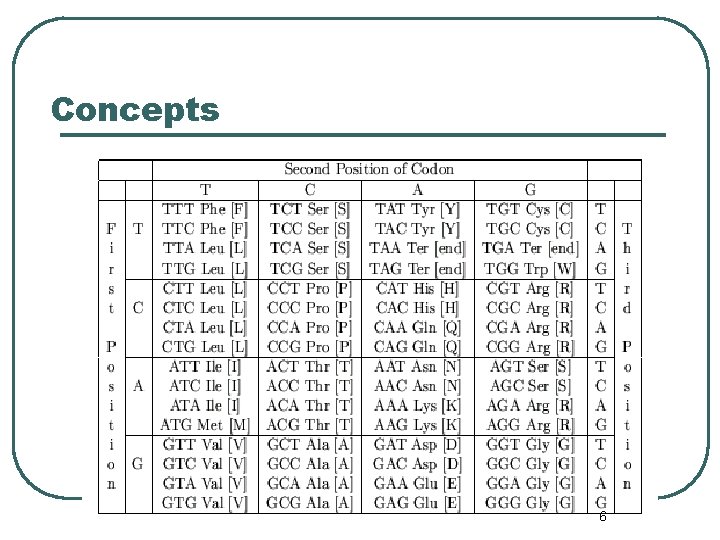
Concepts 6
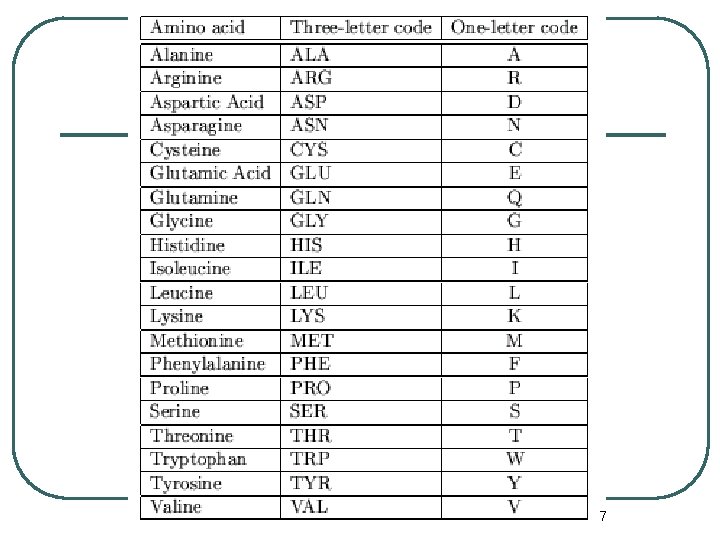
7
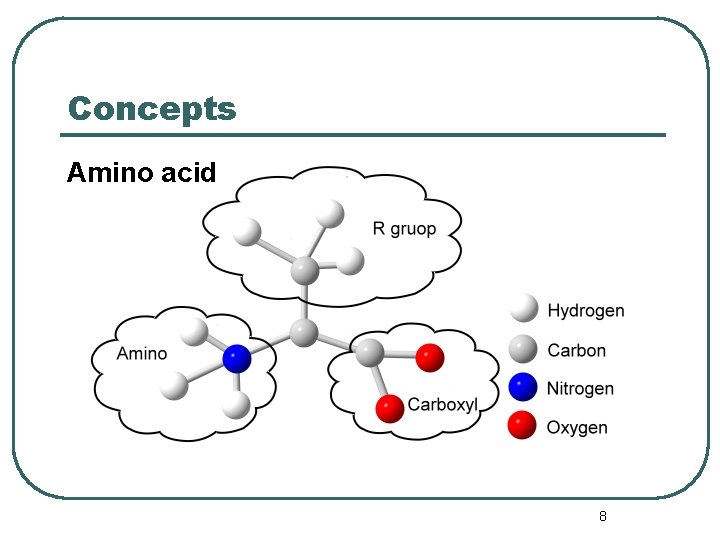
Concepts Amino acid 8
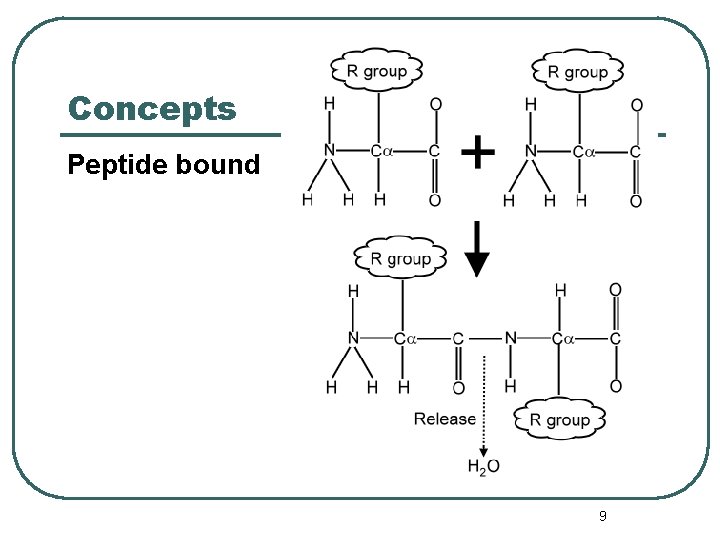
Concepts Peptide bound 9
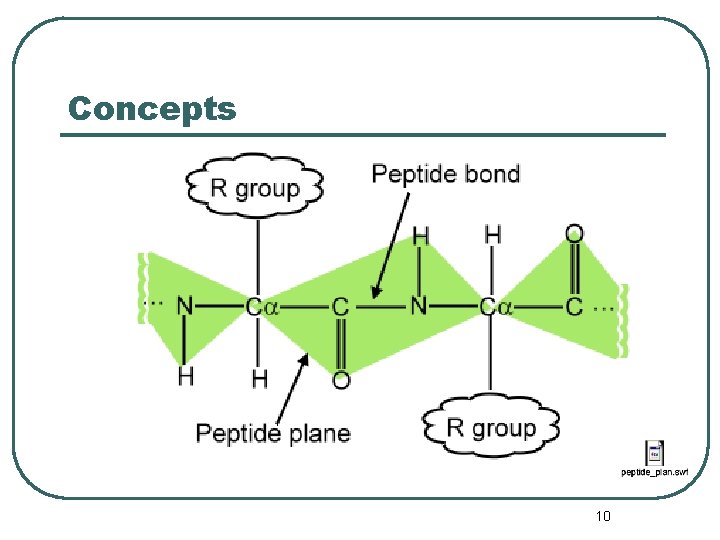
Concepts 10

Concepts l Protein • Primary structure • Secondary structure • Tertiary structure • Quaternary structure 11

Introduction l What is protein? Why do we prediction protein structure? l How? l • X-ray • NMR • Known 19006 protein structure(22 -Oct-2002 ) 12

Introduction l Method of protein structure prediction • AI • Neural Network • PHI-PSI • Potential Energy • Statistical method • … 13

Introduction To determine the native folded state of a protein given only the primary sequence of amino acids is referred to as the protein folding problem. 14

Introduction The protein folding problem is, given an amino acid sequence, to find its correctly folded 3 D protein structure. Protein Folding in the Hydrophobic. Hydrophilic(HP) Model is NP-Complete [BL 98]. 15

Given a test protein sequence, we want to compare every part of it against every part of every protein in the database, then to select some similar parts of proteins in the database. 16

The Basic Algorithm Step 1: Specify the initial parameters, such as the initial windows size W, the window weight pattern P, and N, the number of best matches to keep. 17

Window size: 1. Large or small 2. The five and seven are good choices for the initial window size. 3. A smaller windows is used in finding the best matches for prediction of the next larger window. 18
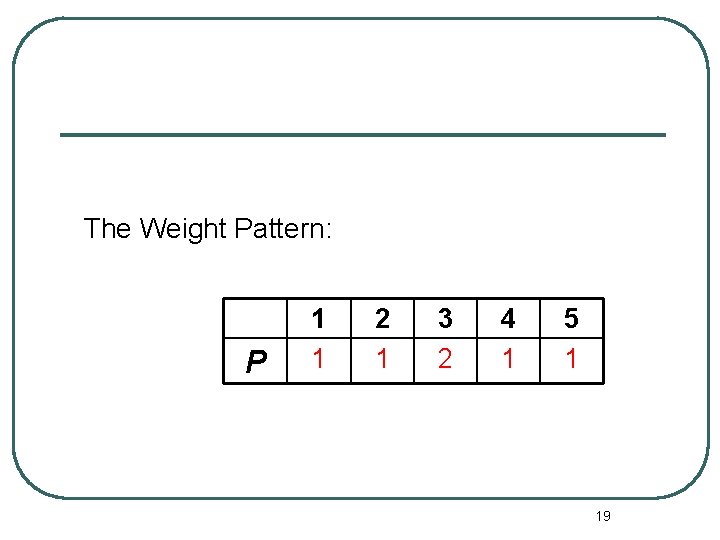
The Weight Pattern: P 1 1 2 1 3 2 4 1 5 1 19

The Basic Algorithm Step 2: Move the window over the test protein sequence, And at each position, extract an amino acid segment S of length W, and do: 20

The Basic Algorithm 2 -1. set the window size in every processor to be of length W; 2 -2. send S to every processor; 2 -3. match S against all si , i=1, 2, . . , m in all the processors, and compute a score for each si using a scoring function; 2 -4. select the N segments from {s 1, …, sm} which have the highest N scores. 21

Compute a score: 22

Why do we bother to use the top N matches rather than just the one with the highest score? Among the top N matchers, the majority have a similar structure, then the input will at least have the tendency to form that structure as well. 23

The Basic Algorithm Step 3: If the recursive mode is chosen, adjust the parameters (e. g. the window size) and repeat Step 2 unless the end conditions are met or PHI-PSI has gone though a pre-specified number Recursive levels. 24

Example: Step 1: Initial parameters W: 5 N: 2 Recursive level=1 Sr=0 25
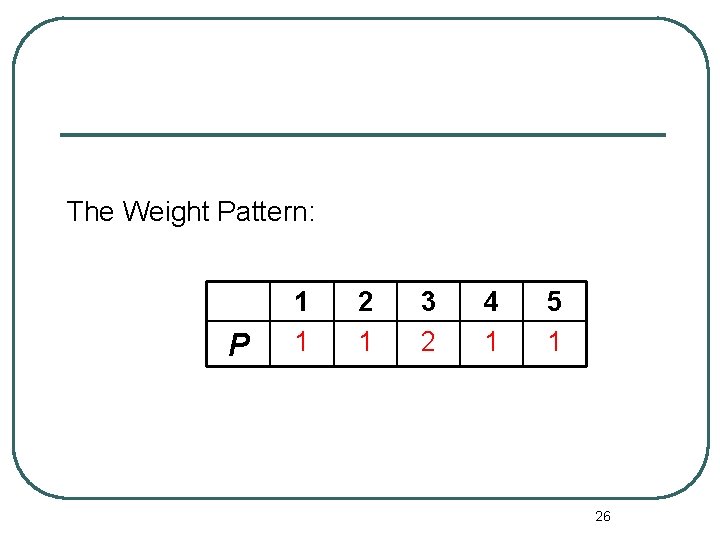
The Weight Pattern: P 1 1 2 1 3 2 4 1 5 1 26
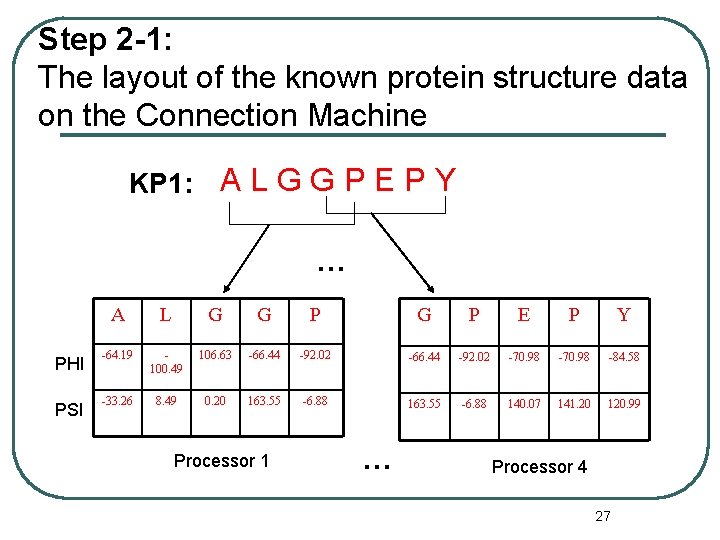
Step 2 -1: The layout of the known protein structure data on the Connection Machine KP 1: A L G G P E P Y … PHI PSI A L G G P E P Y -64. 19 100. 49 106. 63 -66. 44 -92. 02 -70. 98 -84. 58 -33. 26 8. 49 0. 20 163. 55 -6. 88 140. 07 141. 20 120. 99 Processor 1 … Processor 4 27
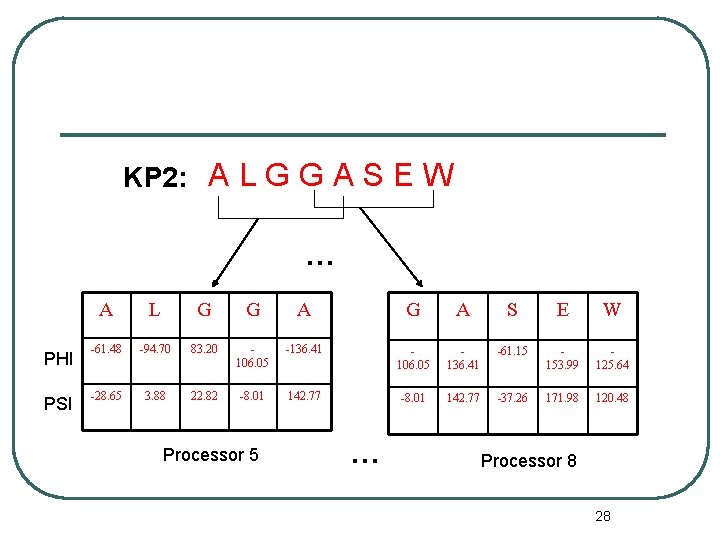
KP 2: A L G G A S E W … PHI PSI A L G G A S E W -61. 48 -94. 70 83. 20 106. 05 -136. 41 106. 05 136. 41 -61. 15 153. 99 125. 64 -28. 65 3. 88 22. 82 -8. 01 142. 77 -37. 26 171. 98 120. 48 Processor 5 … Processor 8 28

P 1: A L G G P P 2: L G G P E P 3: G G P E P P 4: G P E P Y P 5: A L G G A P 6: L G G A S P 7: G G A S E P 8: G A S E W 29

Step 2 -2: Testing protein sequence : ALGGPNAWTG S : ALGGP Send S to P 1~P 8 30
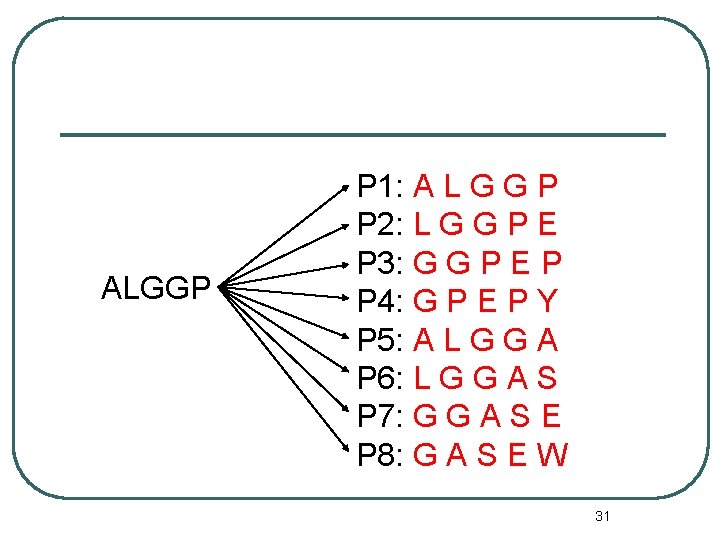
ALGGP P 1: A L G G P P 2: L G G P E P 3: G G P E P P 4: G P E P Y P 5: A L G G A P 6: L G G A S P 7: G G A S E P 8: G A S E W 31

Step 2 -3 S: ALGGP P 1: ALGGP 32

33

S: ALGGP P 2: LGGPE 34

S: ALGGP P 3: GGPEP 35

S: ALGGP P 4: GPEPY 36

S: ALGGP P 5: ALGGA 37

Step 2 -4: Score 1=9 Score 2=3 Score 3=1. 5 Score 4=0 Score 5=7. 5 Score 6=3 Score 7=0 Score 8=0 38
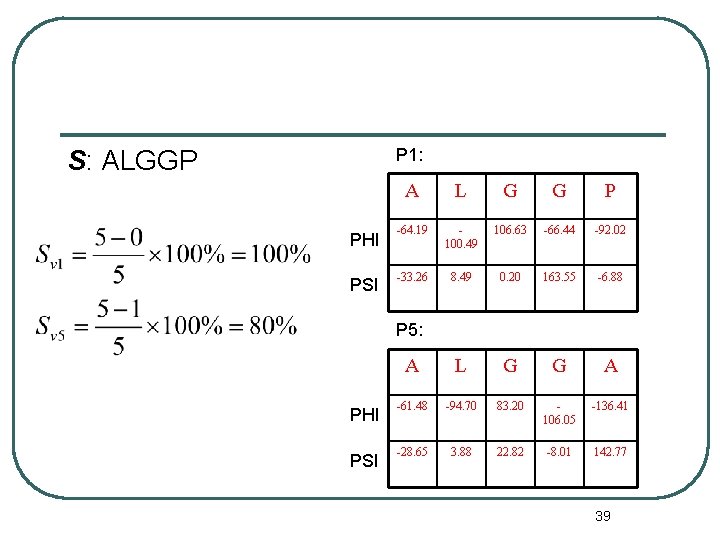
P 1: S: ALGGP PHI PSI A L G G P -64. 19 100. 49 106. 63 -66. 44 -92. 02 -33. 26 8. 49 0. 20 163. 55 -6. 88 A L G G A -61. 48 -94. 70 83. 20 106. 05 -136. 41 -28. 65 3. 88 22. 82 -8. 01 142. 77 P 5: PHI PSI 39
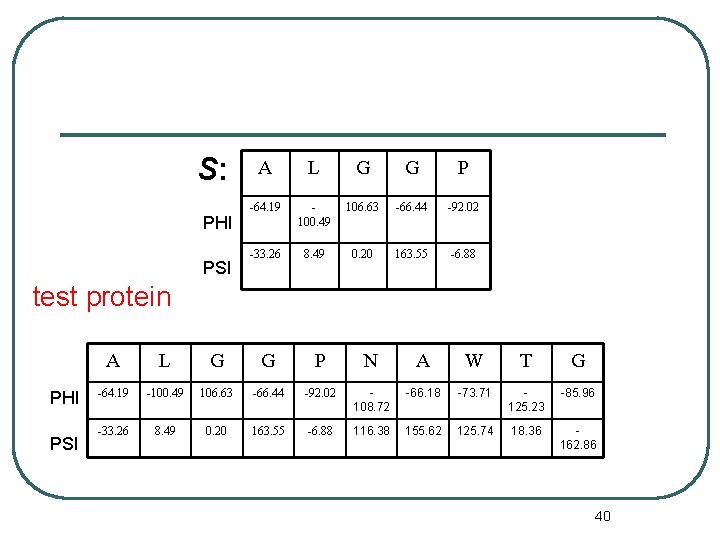
S: PHI PSI A L G G P -64. 19 100. 49 106. 63 -66. 44 -92. 02 -33. 26 8. 49 0. 20 163. 55 -6. 88 test protein PHI PSI A L G G P N A W T G -64. 19 -100. 49 106. 63 -66. 44 -92. 02 108. 72 -66. 18 -73. 71 125. 23 -85. 96 -33. 26 8. 49 0. 20 163. 55 -6. 88 116. 38 155. 62 125. 74 18. 36 162. 86 40

Step 3: if Sr<=recursive level then W=W+2 Sr++ go to Step 2 else end 41
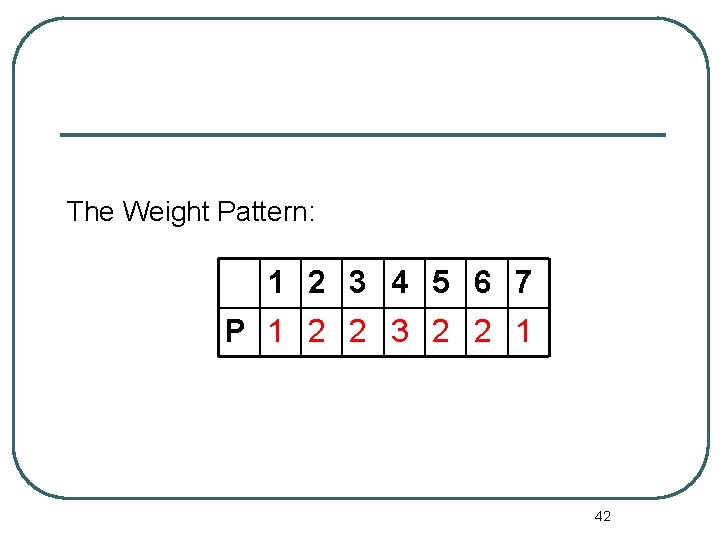
The Weight Pattern: 1 2 3 4 5 6 7 P 1 2 2 3 2 2 1 42
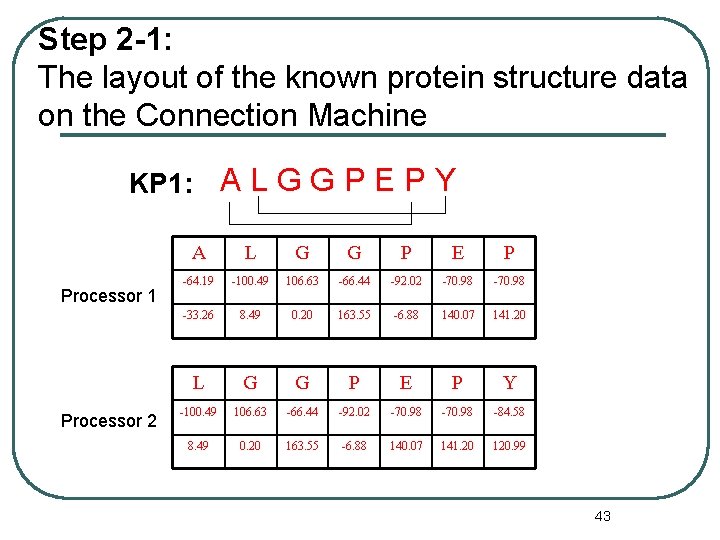
Step 2 -1: The layout of the known protein structure data on the Connection Machine KP 1: A L G G P E P Y Processor 1 Processor 2 A L G G P E P -64. 19 -100. 49 106. 63 -66. 44 -92. 02 -70. 98 -33. 26 8. 49 0. 20 163. 55 -6. 88 140. 07 141. 20 L G G P E P Y -100. 49 106. 63 -66. 44 -92. 02 -70. 98 -84. 58 8. 49 0. 20 163. 55 -6. 88 140. 07 141. 20 120. 99 43
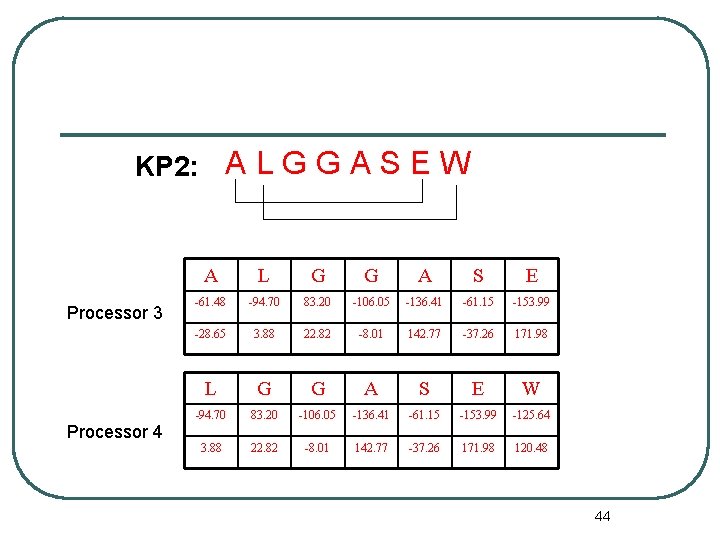
KP 2: A L G G A S E W Processor 3 Processor 4 A L G G A S E -61. 48 -94. 70 83. 20 -106. 05 -136. 41 -61. 15 -153. 99 -28. 65 3. 88 22. 82 -8. 01 142. 77 -37. 26 171. 98 L G G A S E W -94. 70 83. 20 -106. 05 -136. 41 -61. 15 -153. 99 -125. 64 3. 88 22. 82 -8. 01 142. 77 -37. 26 171. 98 120. 48 44

Step 2 -2: Testing protein sequence : AALGGPNA… ALGGPNA S : ALGGPNA Send S to P 1~P 4 45
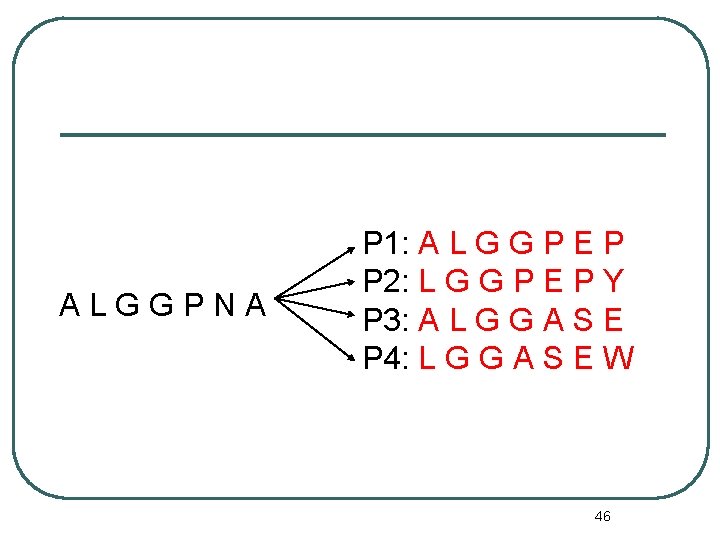
ALGGPNA P 1: A L G G P E P P 2: L G G P E P Y P 3: A L G G A S E P 4: L G G A S E W 46

Step 2 -3 S: ALGGPNA P 1: ALGGPEP 47

48

S: ALGGPNA P 2: LGGPEPY 49

50

S: ALGGPNA P 3: ALGGASE 51

52

S: ALGGPNA P 4: LGGASEW 53

54

Step 2 -4: Score 1=-74. 9 Score 2=-970. 63 Score 3=-1592. 74 Score 4=-860. 25 55
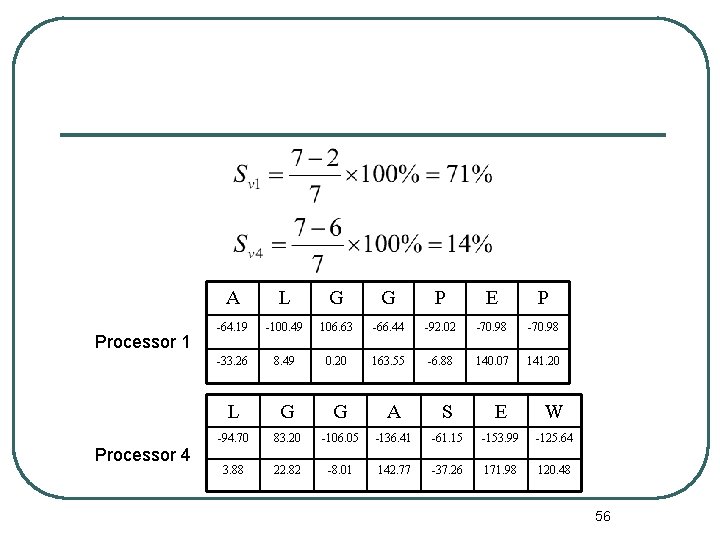
Processor 1 Processor 4 A L G G P E P -64. 19 -100. 49 106. 63 -66. 44 -92. 02 -70. 98 -33. 26 8. 49 0. 20 163. 55 -6. 88 140. 07 141. 20 L G G A S E W -94. 70 83. 20 -106. 05 -136. 41 -61. 15 -153. 99 -125. 64 3. 88 22. 82 -8. 01 142. 77 -37. 26 171. 98 120. 48 56
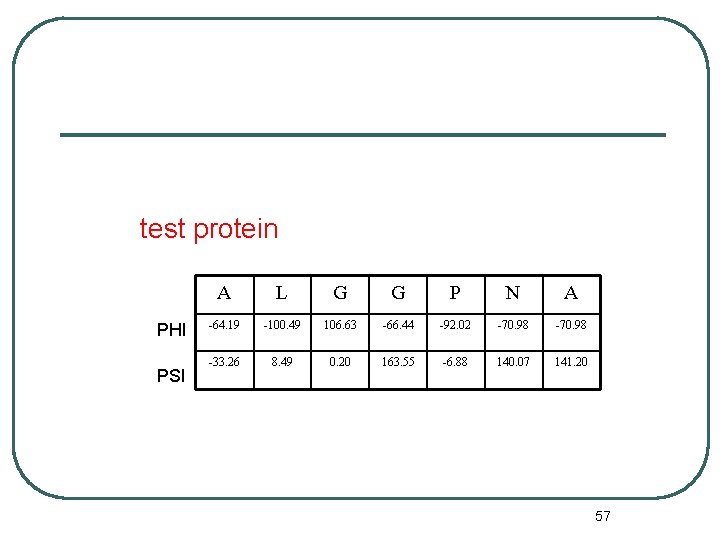
test protein PHI PSI A L G G P N A -64. 19 -100. 49 106. 63 -66. 44 -92. 02 -70. 98 -33. 26 8. 49 0. 20 163. 55 -6. 88 140. 07 141. 20 57

Prediction errors The prediction errors are measured in terms of PHI and PSI angles. There are several ways to measure the errors, such as: 1. Residue error 2. Overall errors 58

Residue errors – the difference between the real angle values computed from the 3 D coordinates and the values predicted by the algorithm for a particular residue in a protein. Overall errors – the average of the residue errors of all the proteins in the database. 59

Conclusions Secondary Structure Prediction 60

61
![Reference l [BL 98] Protein Folding in the Hydrophobic. Hydrophilic(HP) Model is NP-Complete, Berger, Reference l [BL 98] Protein Folding in the Hydrophobic. Hydrophilic(HP) Model is NP-Complete, Berger,](http://slidetodoc.com/presentation_image_h2/3521b56fa1ec3b64fa5e5ef715e69163/image-62.jpg)
Reference l [BL 98] Protein Folding in the Hydrophobic. Hydrophilic(HP) Model is NP-Complete, Berger, B. and Leighton, T. , Journal of Computational Biology, Vol. 5, No. 1, 1998, pp. 27 -40. l Protein Structure Prediction • http: //cmgm. stanford. edu/WWW/www_predict. html l PDB (Protein Data Bank) • http: //www. rcsb. org/pdb/ 62

Thank you 63
- Slides: 63