CS 5263 Bioinformatics Lecture 3 Dynamic Programming and

CS 5263 Bioinformatics Lecture 3: Dynamic Programming and Global Sequence Alignment
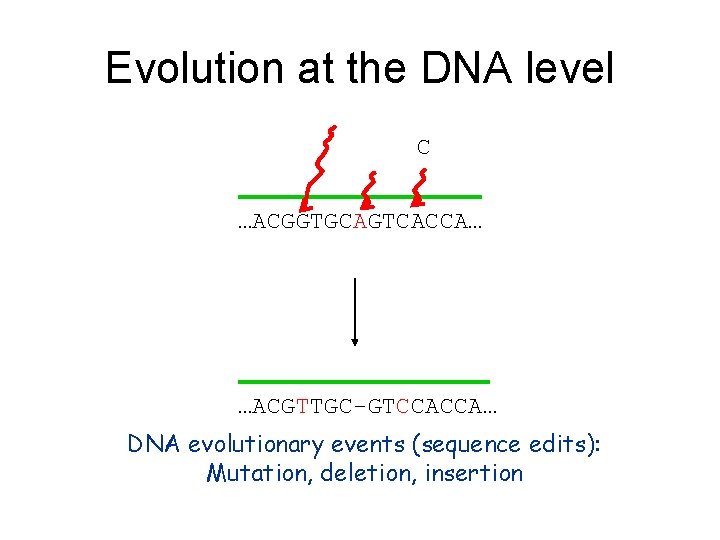
Evolution at the DNA level C …ACGGTGCAGTCACCA… …ACGTTGC-GTCCACCA… DNA evolutionary events (sequence edits): Mutation, deletion, insertion
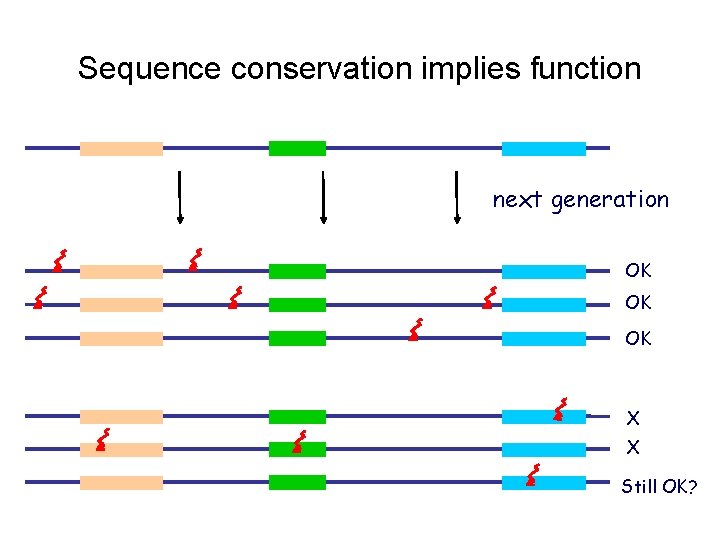
Sequence conservation implies function next generation OK OK OK X X Still OK?

Why sequence alignment? • Conserved regions are more likely to be functional – Can be used for finding genes, regulatory elements, etc. • Similar sequences often have similar origin and function – Can be used to predict functions for new genes / proteins • Sequence alignment is one of the most widely used computational tools in biology

Global Sequence Alignment S T S’ T’ AGGCTATCACCTGACCTCCAGGCCGATGCCC TAGCTATCACGACCGCGGTCGATTTGCCCGAC -AGGCTATCACCTGACCTCCAGGCCGA--TGCCC--TAG-CTATCAC--GACCGC--GGTCGATTTGCCCGAC Definition An alignment of two strings S, T is a pair of strings S’, T’ (with spaces) s. t. (1) |S’| = |T’|, and (|S| = “length of S”) (2) removing all spaces in S’, T’ leaves S, T

What is a good alignment? Alignment: The “best” way to match the letters of one sequence with those of the other How do we define “best”?

S’: -AGGCTATCACCTGACCTCCAGGCCGA--TGCCC--T’: TAG-CTATCAC--GACCGC--GGTCGATTTGCCCGAC • The score of aligning (characters or spaces) x & y is σ (x, y). • Score of an alignment: • An optimal alignment: one with max score

Scoring Function • Sequence edits: – Mutations – Insertions – Deletions Scoring Function: Match: +m Mismatch: -s Gap (indel): -d AGGCCTC AGGACTC AGGGCCTC AGG-CTC ~~~AAC~~~ ~~~A-A~~~

• Match = 2, mismatch = -1, gap = -1 • Score = 3 x 2 – 2 x 1 – 1 x 1 = 3

More complex scoring function • Substitution matrix – Similarity score of matching two letters a, b should reflect the probability of a, b derived from same ancestor – It is usually defined by log likelihood ratio (Durbin book) – Active research area. Especially for proteins. – Commonly used: PAM, BLOSUM
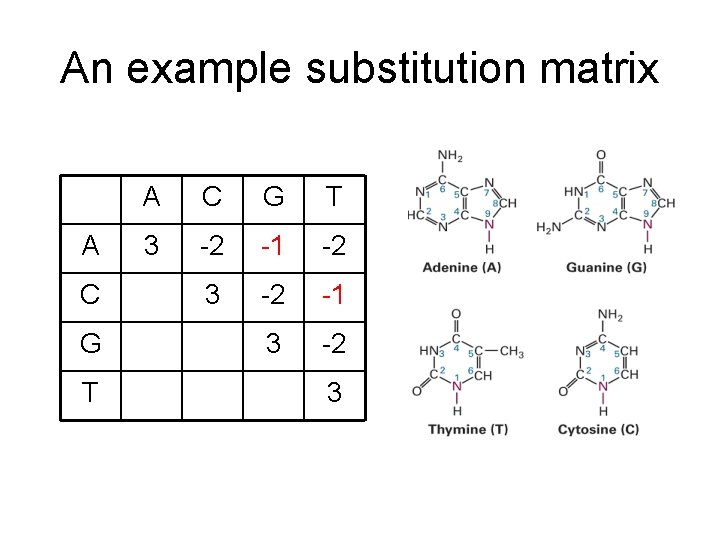
An example substitution matrix A C G T 3 -2 -1 -2 3 -2 -1 3 -2 3

How to find an optimal alignment? • A naïve algorithm: for all subseqs A of S, B of T s. t. |A| = |B| do align A[i] with B[i], 1 ≤i ≤|A| align all other chars to spaces compute its value S = abcd A = cd T = wxyz B = xz retain the max -abc-d a-bc-d end w--xyz -w-xyz output the retained alignment

Analysis • Assume |S| = |T| = n • Cost of evaluating one alignment: ≥n • How many alignments are there: – pick n chars of S, T together – say k of them are in S – match these k to the k unpicked chars of T • Total time: • E. g. , for n = 20, time is > 240 >1012 operations

Intro to Dynamic Programming

Dynamic programming • What is dynamic programming? – A method for solving problems exhibiting the properties of overlapping subproblems and optimal substructure – Key idea: tabulating sub-problem solutions rather than re-computing them repeatedly • Two simple examples: – Computing Fibonacci numbers – Find the special shortest path in a grid

Example 1: Fibonacci numbers • 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89, … F(0) = 1; F(1) = 1; F(n) = F(n-1) + f(n-2) • How to compute F(n)?
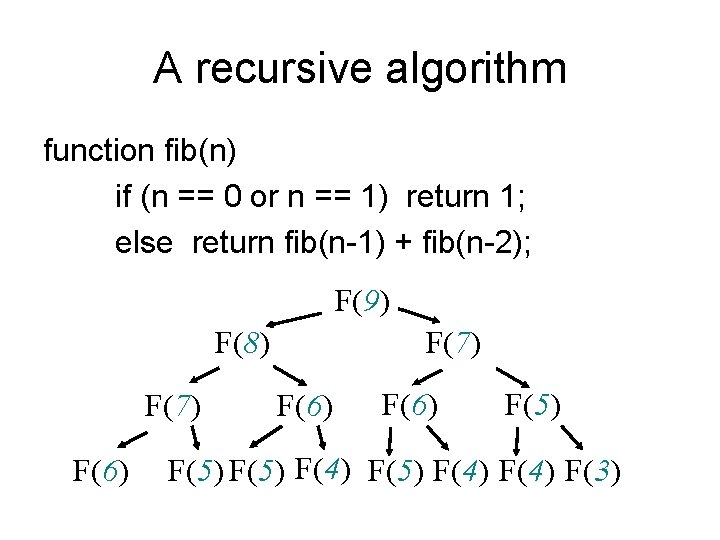
A recursive algorithm function fib(n) if (n == 0 or n == 1) return 1; else return fib(n-1) + fib(n-2); F(9) F(8) F(7) F(6) F(5) F(4) F(3)
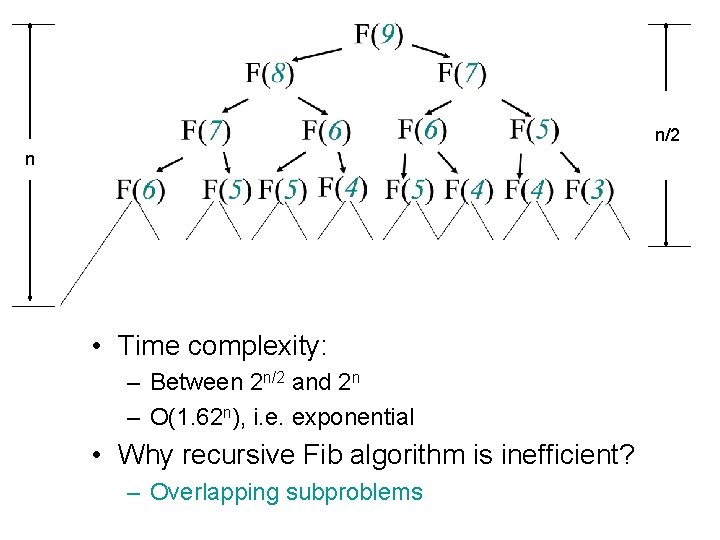
n/2 n • Time complexity: – Between 2 n/2 and 2 n – O(1. 62 n), i. e. exponential • Why recursive Fib algorithm is inefficient? – Overlapping subproblems
![An iterative algorithm function fib(n) F[0] = 1; F[1] = 1; for i = An iterative algorithm function fib(n) F[0] = 1; F[1] = 1; for i =](http://slidetodoc.com/presentation_image_h/147c4abcbd6a718eba46a31cb604a321/image-19.jpg)
An iterative algorithm function fib(n) F[0] = 1; F[1] = 1; for i = 2 to n F[i] = F[i-1] + F[i-2]; Return F[n]; Time complexity: Time: O(n), space: O(n)
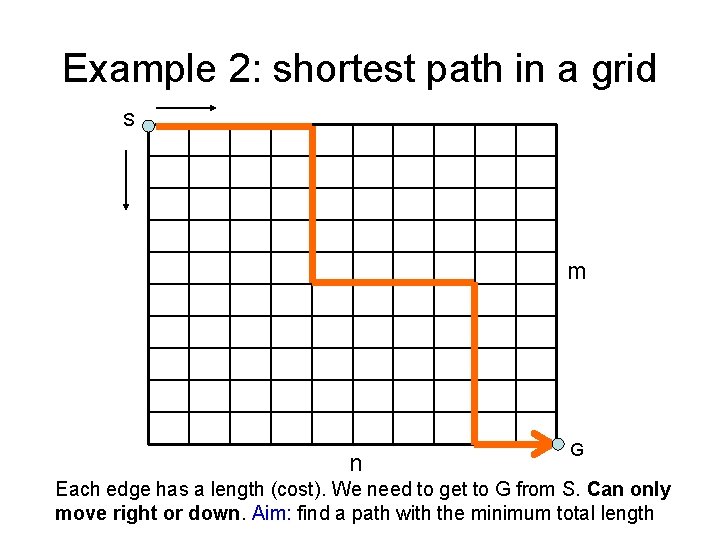
Example 2: shortest path in a grid S m n G Each edge has a length (cost). We need to get to G from S. Can only move right or down. Aim: find a path with the minimum total length

Optimal substructures • Naïve algorithm: enumerate all possible paths and compare costs – Exponential number of paths • Key observation: – If a path P(S, G) is the shortest from S to G, any of its sub-path P(S, x), where x is on P(S, G), is the shortest from S to x
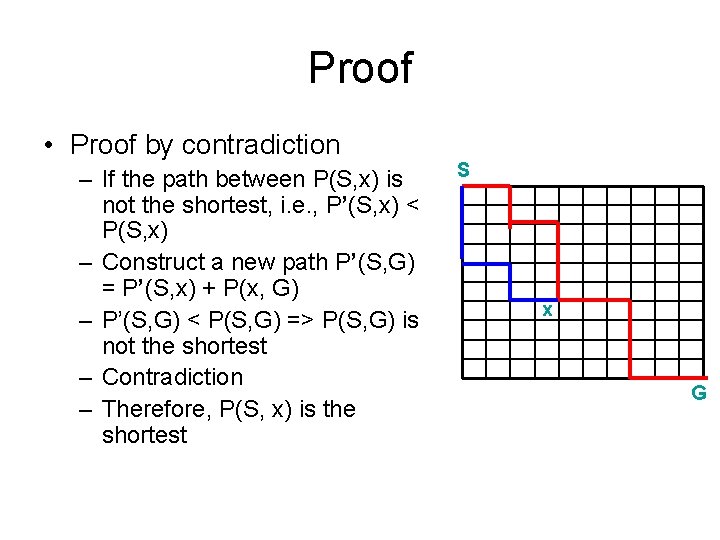
Proof • Proof by contradiction – If the path between P(S, x) is not the shortest, i. e. , P’(S, x) < P(S, x) – Construct a new path P’(S, G) = P’(S, x) + P(x, G) – P’(S, G) < P(S, G) => P(S, G) is not the shortest – Contradiction – Therefore, P(S, x) is the shortest S x G
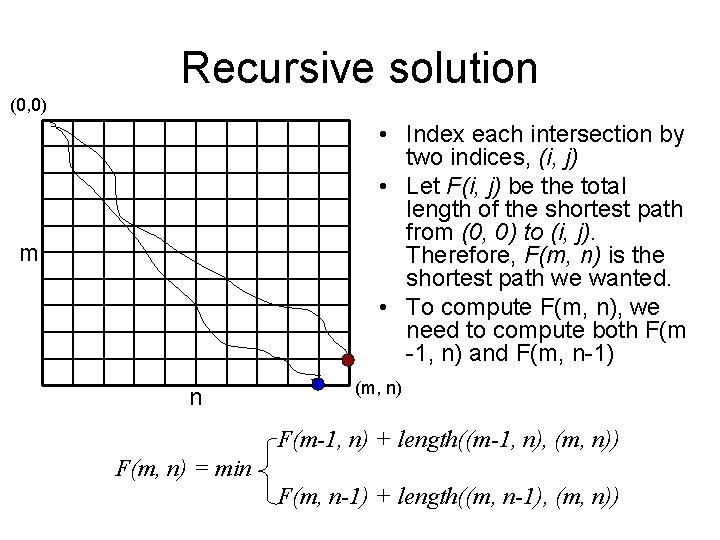
Recursive solution (0, 0) • Index each intersection by two indices, (i, j) • Let F(i, j) be the total length of the shortest path from (0, 0) to (i, j). Therefore, F(m, n) is the shortest path we wanted. • To compute F(m, n), we need to compute both F(m -1, n) and F(m, n-1) m n (m, n) F(m-1, n) + length((m-1, n), (m, n)) F(m, n) = min F(m, n-1) + length((m, n-1), (m, n))
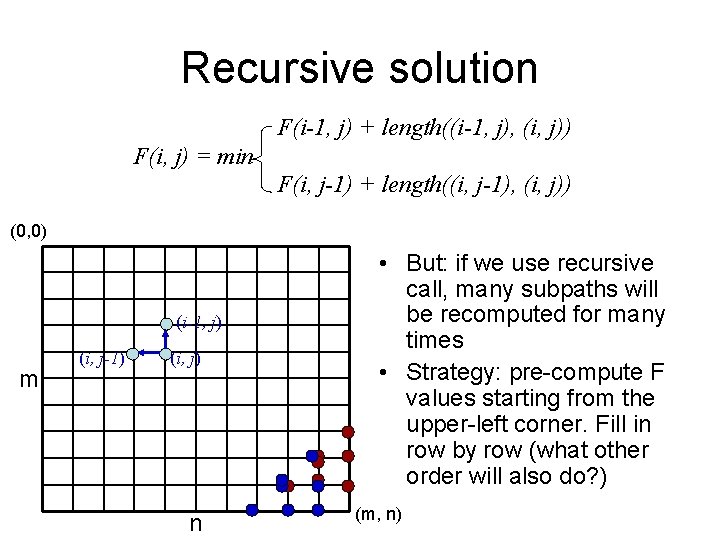
Recursive solution F(i-1, j) + length((i-1, j), (i, j)) F(i, j) = min F(i, j-1) + length((i, j-1), (i, j)) (0, 0) (i-1, j) m (i, j-1) (i, j) n • But: if we use recursive call, many subpaths will be recomputed for many times • Strategy: pre-compute F values starting from the upper-left corner. Fill in row by row (what other order will also do? ) (m, n)
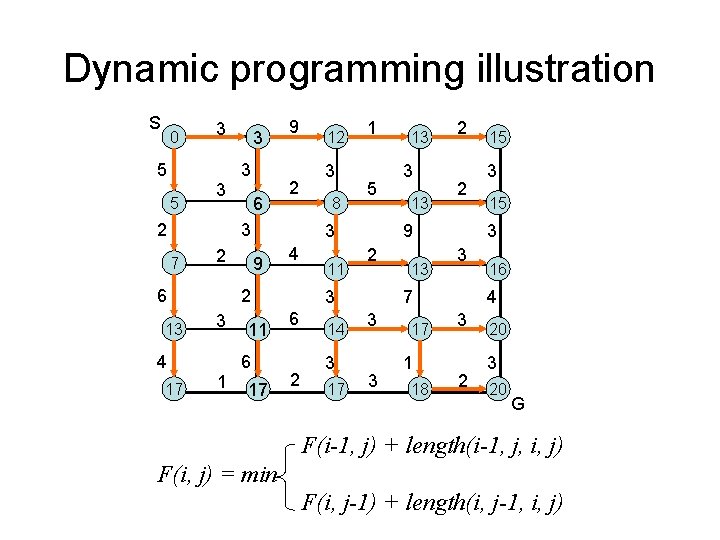
Dynamic programming illustration S 0 5 5 3 3 2 3 3 6 9 2 3 7 2 6 13 4 17 1 3 8 1 5 3 9 4 2 3 12 11 6 17 11 2 14 3 17 3 13 2 2 9 2 3 6 13 13 3 17 1 18 3 15 3 3 7 3 15 16 4 3 2 20 3 20 G F(i-1, j) + length(i-1, j, i, j) F(i, j) = min F(i, j-1) + length(i, j-1, i, j)
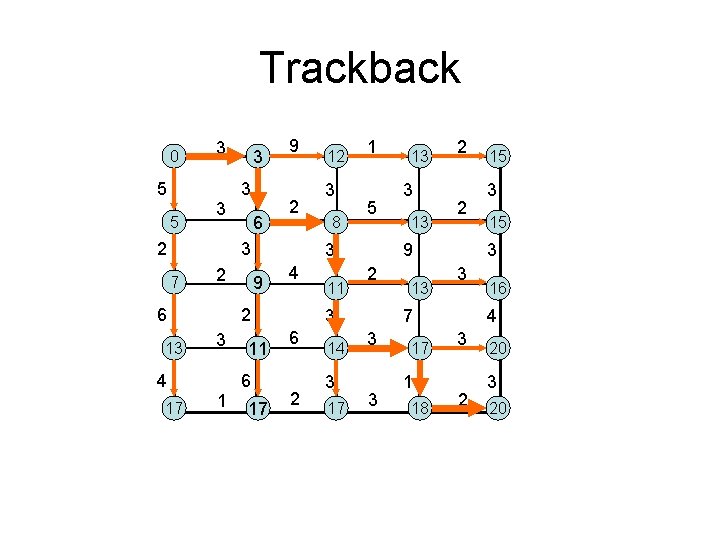
Trackback 0 5 5 3 3 2 3 3 6 9 2 3 7 2 6 13 4 17 1 3 8 1 5 3 9 4 2 3 12 11 6 17 11 2 14 3 17 3 13 2 2 9 2 3 6 13 13 3 17 1 18 3 15 3 3 7 3 15 16 4 3 2 20 3 20

Elements of dynamic programming • Optimal sub-structures – Optimal solutions to the original problem contains optimal solutions to sub-problems • Overlapping sub-problems – Some sub-problems appear in many solutions • Memorization and reuse – Carefully choose the order that sub-problems are solved

Dynamic Programming for sequence alignment Suppose we wish to align x 1……x. M y 1……y. N Let F(i, j) = optimal score of aligning x 1……xi y 1……yj Scoring Function: Match: +m Mismatch: -s Gap (indel): -d
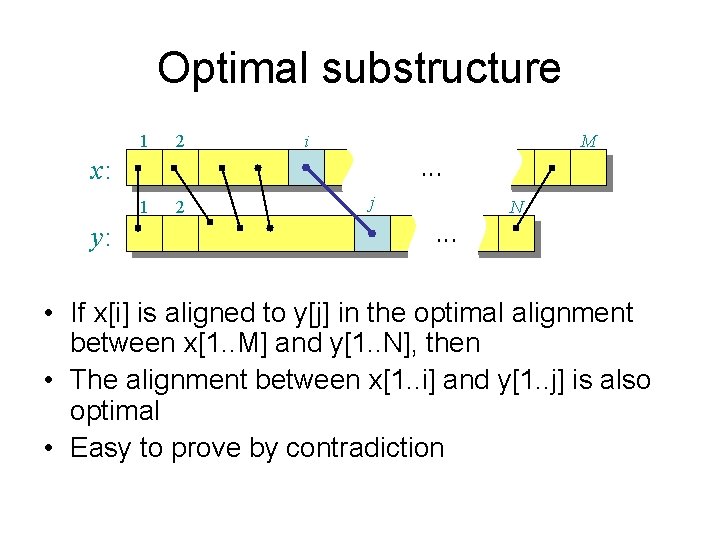
Optimal substructure 1 2 i M . . . x: 1 y: 2 j N . . . • If x[i] is aligned to y[j] in the optimal alignment between x[1. . M] and y[1. . N], then • The alignment between x[1. . i] and y[1. . j] is also optimal • Easy to prove by contradiction

Recursive formula Notice three possible cases: 1. x. M aligns to y. N ~~~~~~~ x. M ~~~~~~~ y. N 2. F(M, N) = F(M-1, N-1) + -s, if not x. M aligns to a gap ~~~~~~~ x. M ~~~~~~~ 3. m, if x. M = y. N F(M, N) = F(M-1, N) - d y. N aligns to a gap ~~~~~~~ y. N F(M, N) = F(M, N-1) - d

Recursive formula • Generalize: F(i, j) = max F(i-1, j-1) + (Xi, Yj) F(i-1, j) – d F(i, j-1) – d (Xi, Yj) = m if Xi = Yj, and –s otherwise • Boundary conditions: – F(0, 0) = 0. -jd: y[1. . j] aligned to gaps. – F(0, j) = ? – F(i, 0) = ? -id: x[1. . i] aligned to gaps.
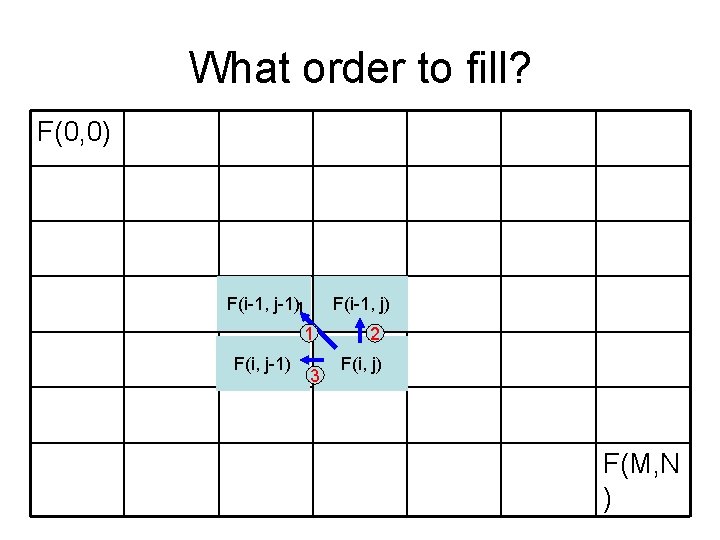
What order to fill? F(0, 0) F(i-1, j-1)1 F(i-1, j) 1 F(i, j-1) 3 2 F(i, j) F(M, N )
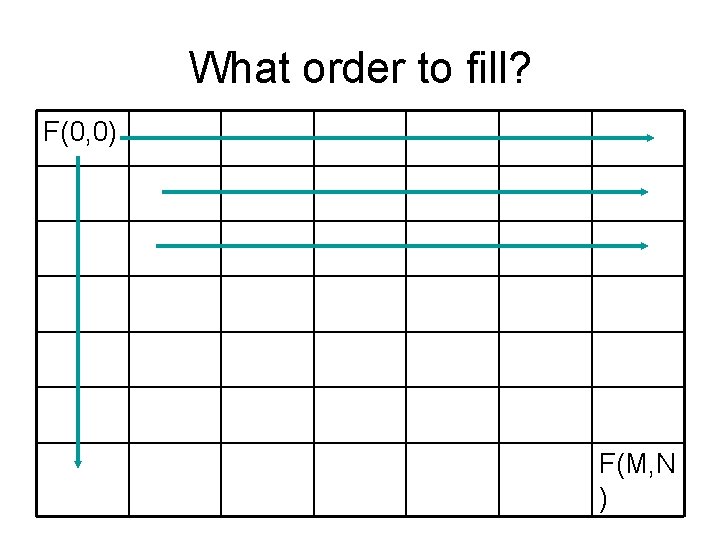
What order to fill? F(0, 0) F(M, N )
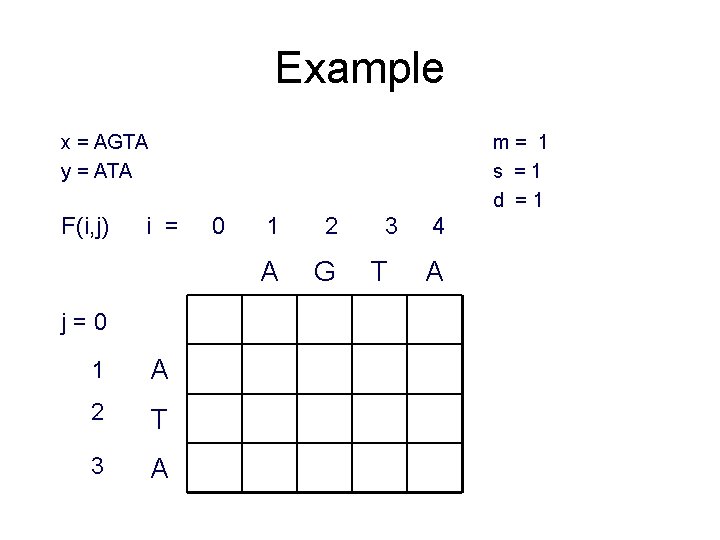
Example x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 1 A 2 T 3 A 0 1 2 A G 3 T 4 A
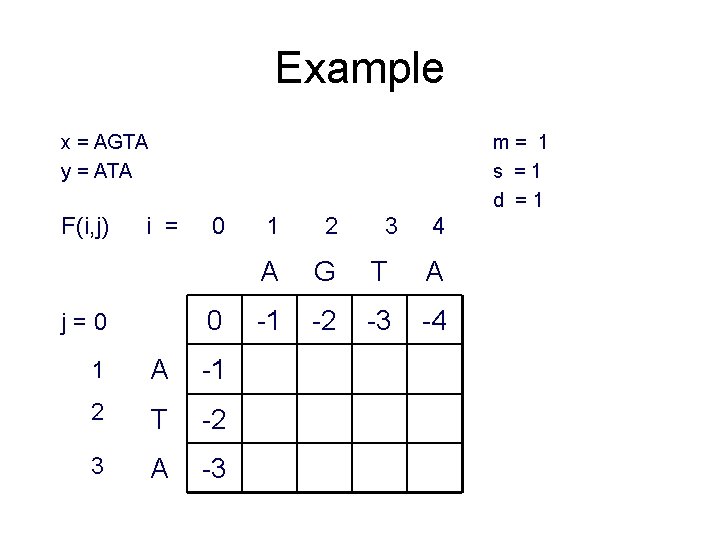
Example x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = 0 0 j=0 1 A -1 2 T -2 3 A -3 1 2 3 4 A G T A -1 -2 -3 -4
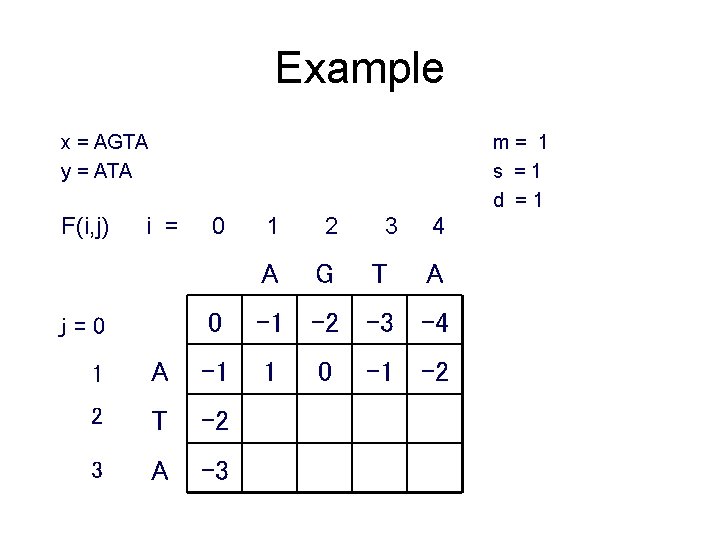
Example x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 A G 3 T 4 A 0 -1 -2 -3 -4 1 1 A -1 2 T -2 3 A -3 0 -1 -2
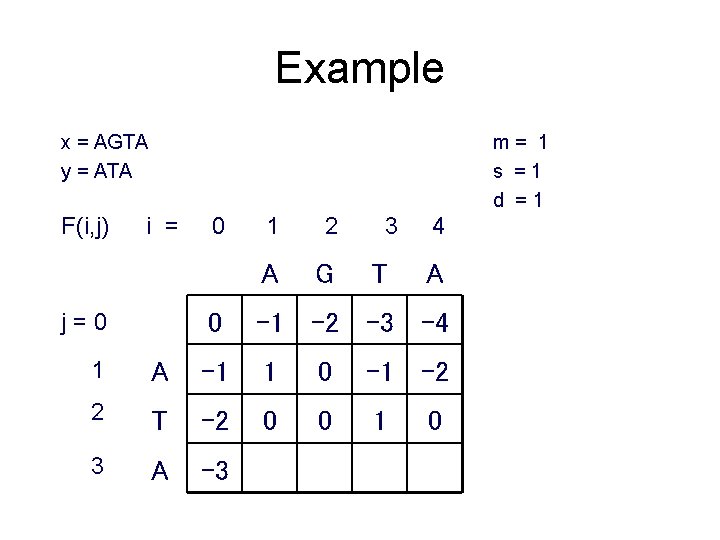
Example x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 A G 3 T 4 A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 3 A -3 0
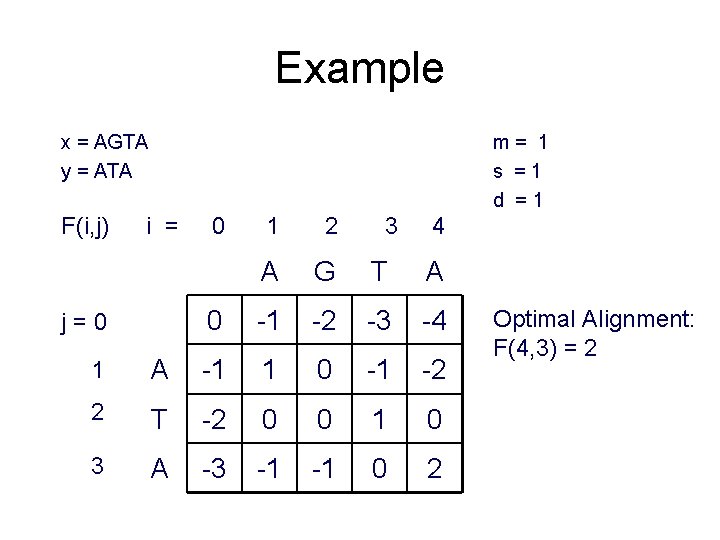
Example x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2 Optimal Alignment: F(4, 3) = 2
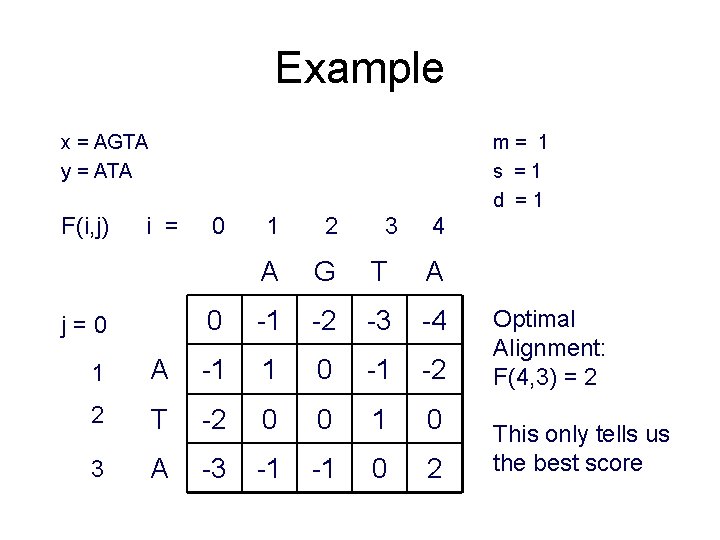
Example x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2 Optimal Alignment: F(4, 3) = 2 This only tells us the best score
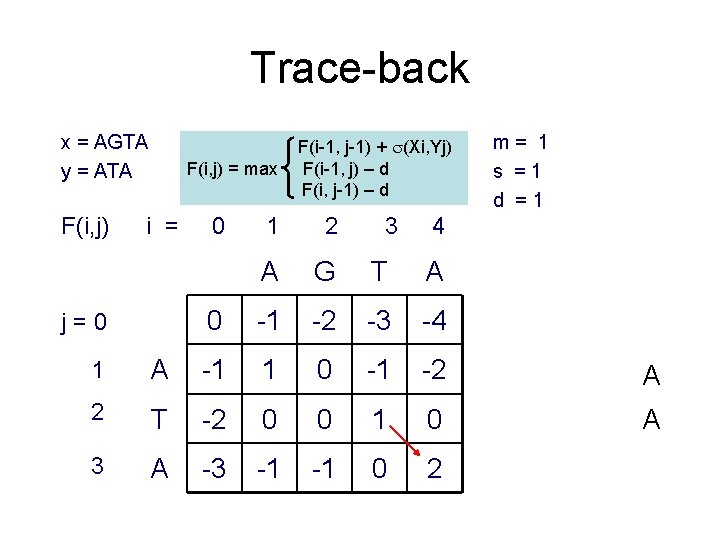
Trace-back x = AGTA y = ATA F(i, j) = max i = j=0 0 F(i-1, j-1) + (Xi, Yj) F(i-1, j) – d F(i, j-1) – d 1 2 3 A G T A 0 -1 -2 -3 -4 m= 1 s =1 d =1 4 1 A -1 1 0 -1 -2 A 2 T -2 0 0 1 0 A 3 A -3 -1 -1 0 2
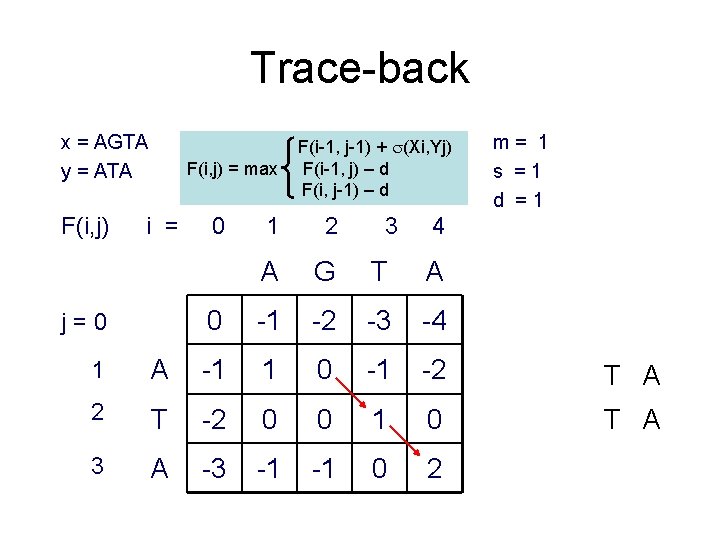
Trace-back x = AGTA y = ATA F(i, j) = max i = j=0 0 F(i-1, j-1) + (Xi, Yj) F(i-1, j) – d F(i, j-1) – d 1 2 3 A G T A 0 -1 -2 -3 -4 m= 1 s =1 d =1 4 1 A -1 1 0 -1 -2 T A 2 T -2 0 0 1 0 T A 3 A -3 -1 -1 0 2
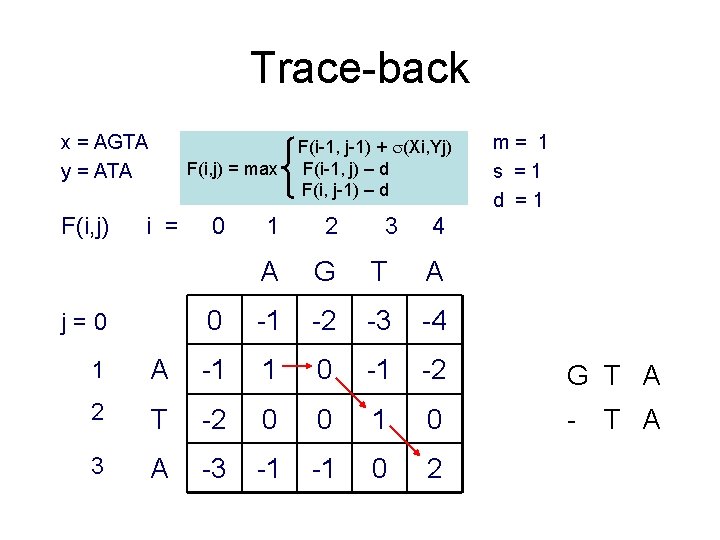
Trace-back x = AGTA y = ATA F(i, j) = max i = j=0 0 F(i-1, j-1) + (Xi, Yj) F(i-1, j) – d F(i, j-1) – d 1 2 3 A G T A 0 -1 -2 -3 -4 m= 1 s =1 d =1 4 1 A -1 1 0 -1 -2 G T A 2 T -2 0 0 1 0 - 3 A -3 -1 -1 0 2 T A
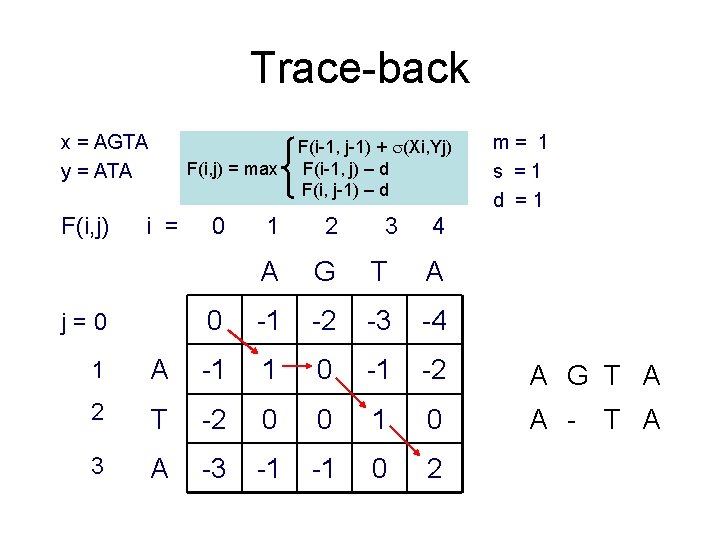
Trace-back x = AGTA y = ATA F(i, j) = max i = j=0 0 F(i-1, j-1) + (Xi, Yj) F(i-1, j) – d F(i, j-1) – d 1 2 3 A G T A 0 -1 -2 -3 -4 m= 1 s =1 d =1 4 1 A -1 1 0 -1 -2 A G T A 2 T -2 0 0 1 0 A - 3 A -3 -1 -1 0 2 T A
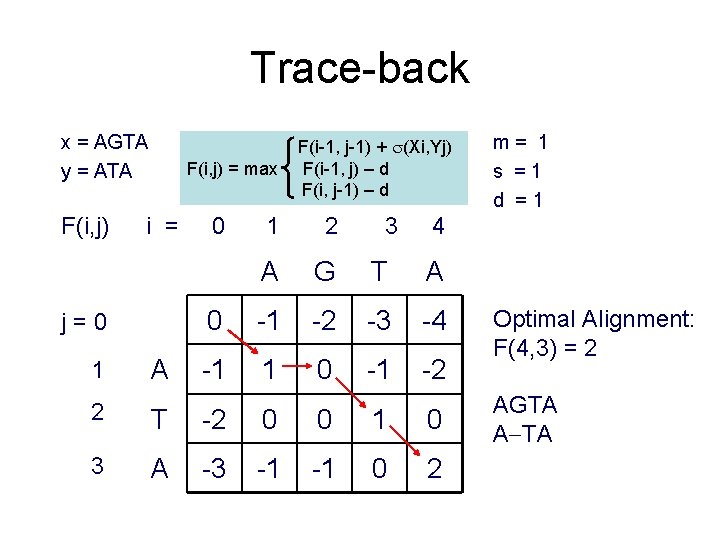
Trace-back x = AGTA y = ATA F(i, j) = max i = j=0 1 A 0 F(i-1, j-1) + (Xi, Yj) F(i-1, j) – d F(i, j-1) – d 1 2 3 A G T A 0 -1 -2 -3 -4 -1 1 0 -1 -2 m= 1 s =1 d =1 4 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2 Optimal Alignment: F(4, 3) = 2 AGTA A TA
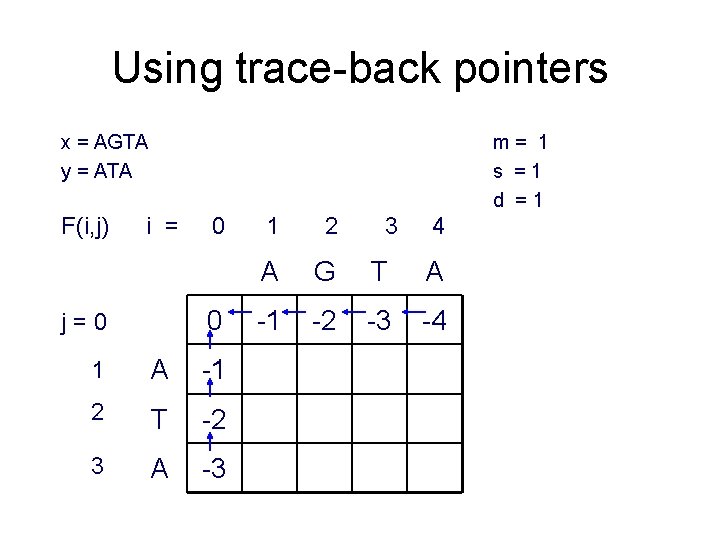
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = 0 0 j=0 1 A -1 2 T -2 3 A -3 1 2 3 4 A G T A -1 -2 -3 -4
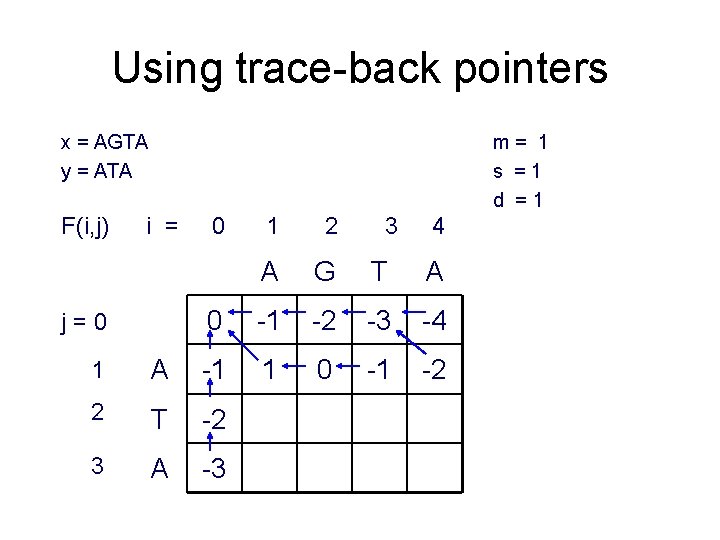
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 A G T A 0 -1 -2 -3 -4 1 0 -1 -2 1 A -1 2 T -2 3 A -3 3 4
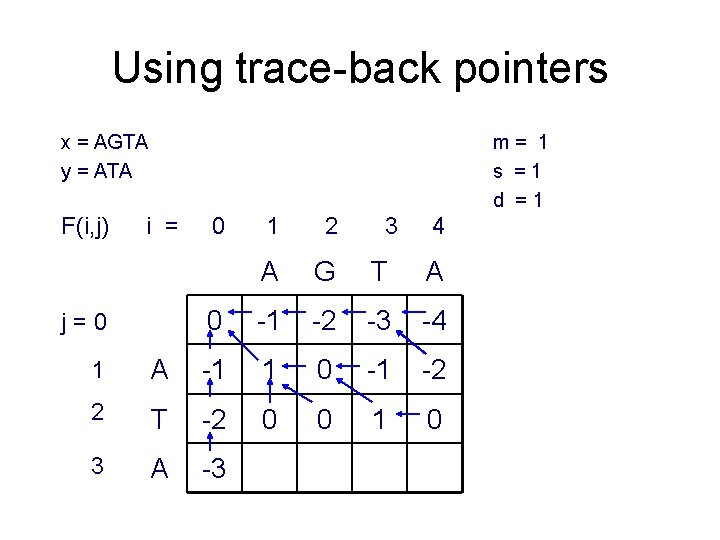
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3
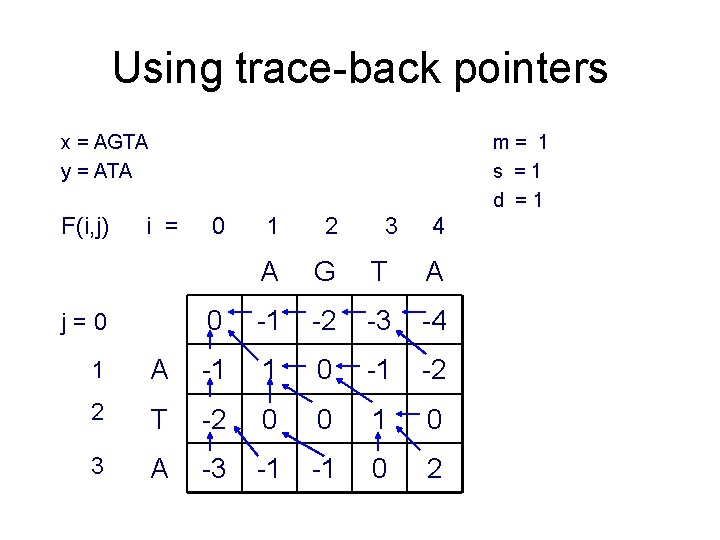
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2
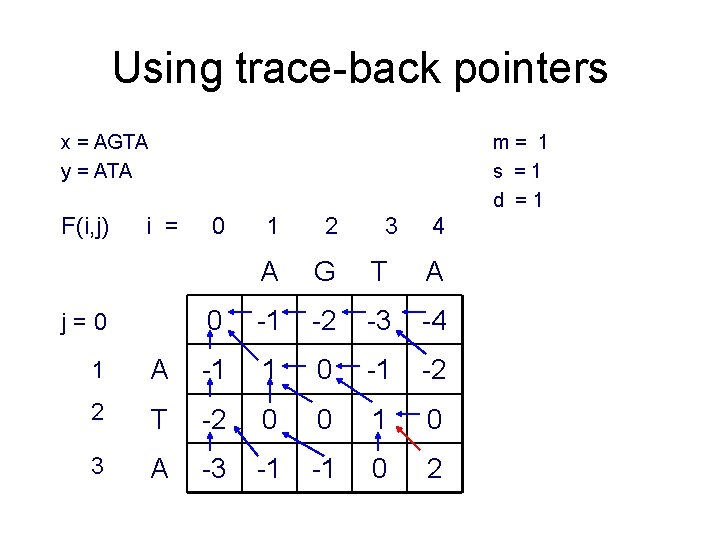
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2
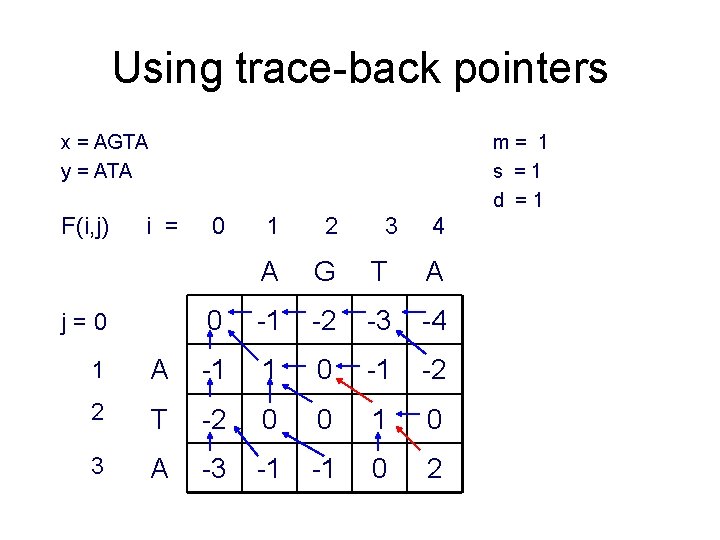
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2
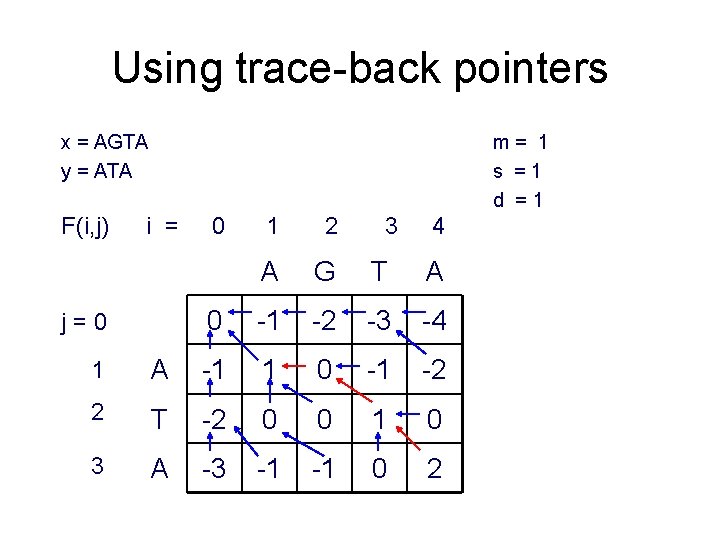
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 1 A -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2
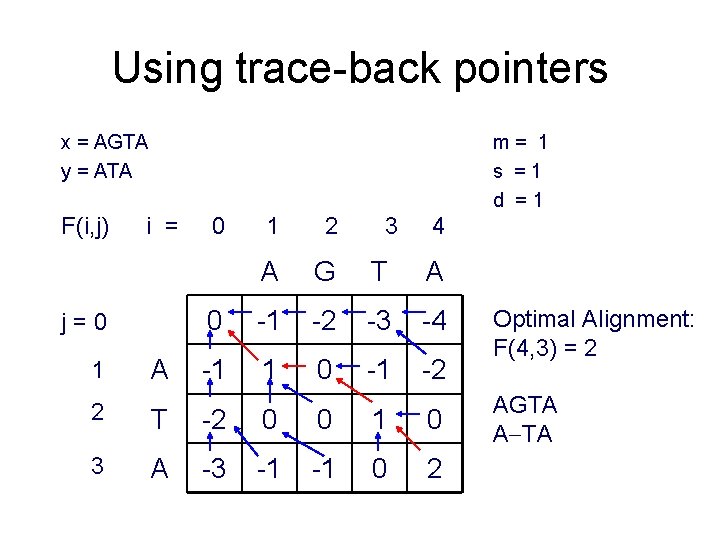
Using trace-back pointers x = AGTA y = ATA F(i, j) m= 1 s =1 d =1 i = j=0 1 A 0 1 2 3 4 A G T A 0 -1 -2 -3 -4 -1 1 0 -1 -2 2 T -2 0 0 1 0 3 A -3 -1 -1 0 2 Optimal Alignment: F(4, 3) = 2 AGTA A TA

The Needleman-Wunsch Algorithm 1. Initialization. a. b. c. 2. F(0, 0) F(0, j) F(i, 0) = 0 =-j d =-i d Main Iteration. Filling in scores a. For each i = 1……M For each j = 1……N F(i, j) Ptr(i, j) 3. = max = F(i-1, j) – d F(i, j-1) – d F(i-1, j-1) + σ(xi, yj) UP, LEFT DIAG [case 1] [case 2] [case 3] if [case 1] if [case 2] if [case 3] Termination. F(M, N) is the optimal score, and from Ptr(M, N) can trace back optimal alignment

Performance • Time: O(NM) • Space: O(NM) • Later we will cover more efficient methods
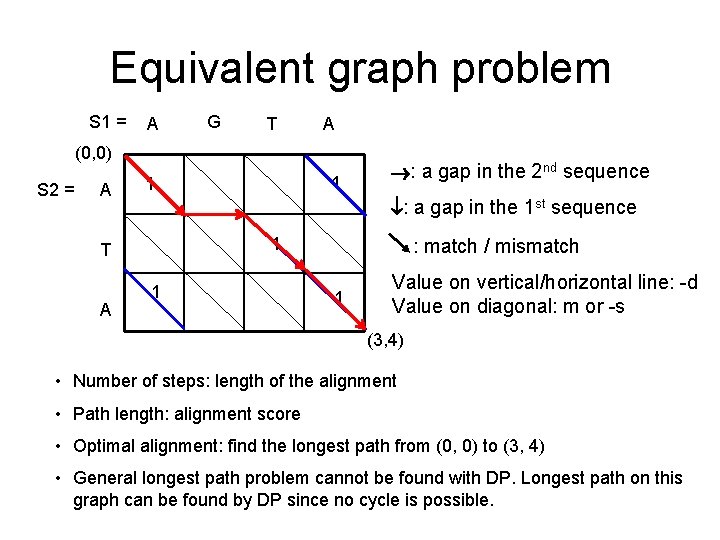
Equivalent graph problem S 1 = A G T A (0, 0) S 2 = A : a gap in the 1 st sequence 1 T A 1 1 : a gap in the 2 nd sequence 1 : match / mismatch 1 Value on vertical/horizontal line: -d Value on diagonal: m or -s (3, 4) • Number of steps: length of the alignment • Path length: alignment score • Optimal alignment: find the longest path from (0, 0) to (3, 4) • General longest path problem cannot be found with DP. Longest path on this graph can be found by DP since no cycle is possible.

Question • If we change the scoring scheme, will the optimal alignment be changed? – Old: Match = 1, mismatch = gap = -1 – New: match = 2, mismatch = gap = 0 – New: Match = 2, mismatch = gap = -2?
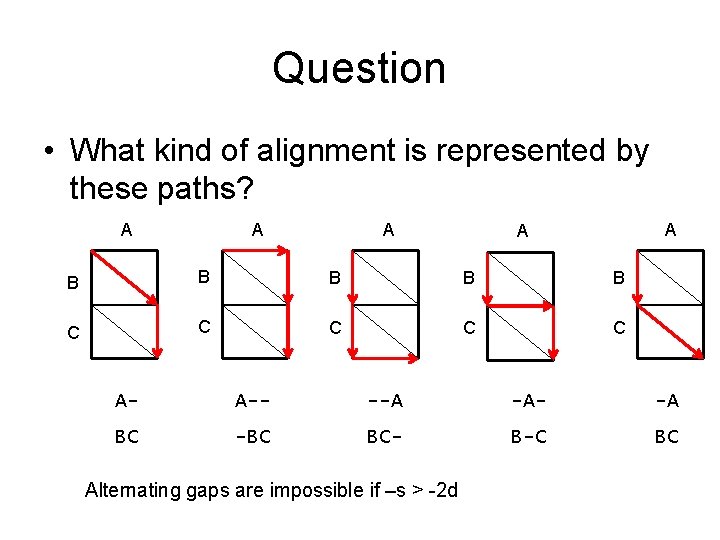
Question • What kind of alignment is represented by these paths? A A A B B B C C C A- A-- --A -A- -A BC -BC BC- B-C BC Alternating gaps are impossible if –s > -2 d

A variant of the basic algorithm Scoring scheme: m = s = d: 1 Seq 1: CAGCA-CTTGGATTCTCGG || |: ||| Seq 2: ---CAGCGTGG-------Seq 1: CAGCACTTGGATTCTCGG |||| | | || Seq 2: CAGC-----G-T----GG Score = -7 Score = -2 The first alignment may be biologically more realistic

A variant of the basic algorithm • Maybe it is OK to have an unlimited # of gaps in the beginning and end: -----CTATCACCTGACCTCCAGGCCGATGCCCCTTCCGGC GCGAGTTCATCTATCAC--GACCGC--GGTCG------- • Then, we don’t want to penalize gaps in the ends
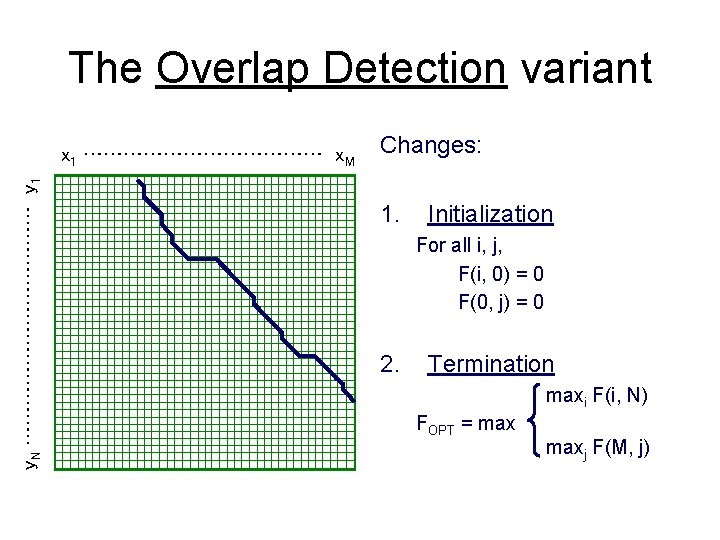
The Overlap Detection variant y. N ……………… y 1 x 1 ……………… x. M Changes: 1. Initialization For all i, j, F(i, 0) = 0 F(0, j) = 0 2. Termination maxi F(i, N) FOPT = maxj F(M, j)
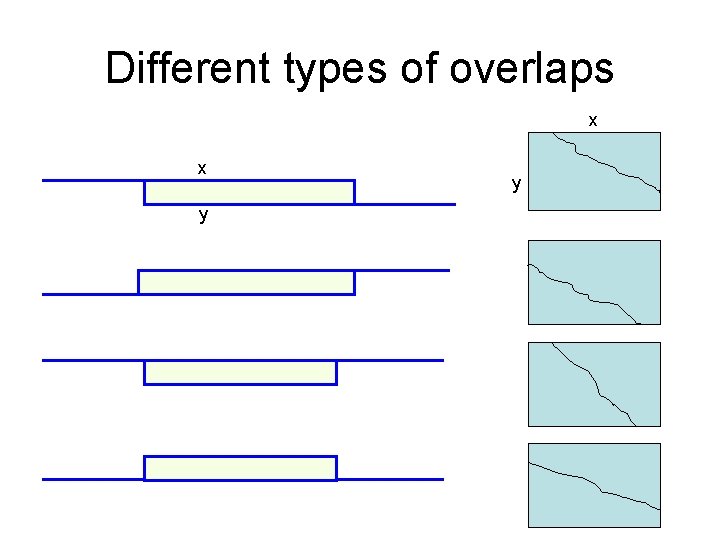
Different types of overlaps x x y y

A non-bio variant • Shell command diff: Compare two text files – Given file 1 and file 2 – Find the difference between file 1 and file 2 – Similar to sequence alignment – How to score? • • Longest common subsequence (LCS) Match has score 1 No mismatch penalty No gap penalty
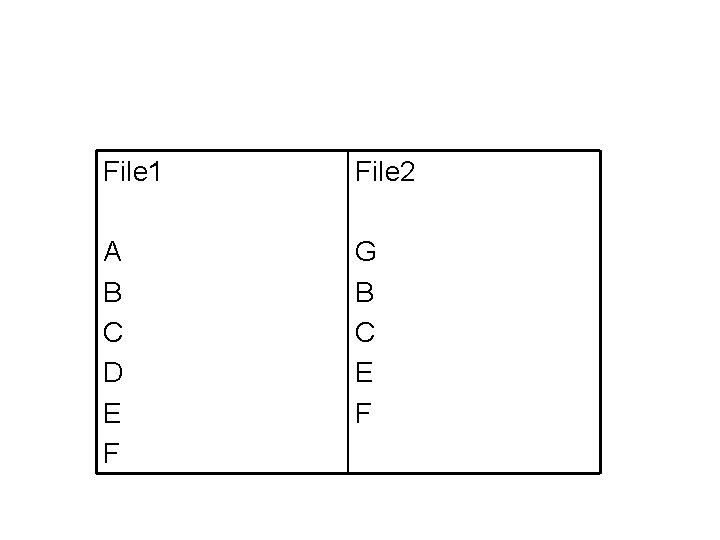
File 1 File 2 A B C D E F G B C E F
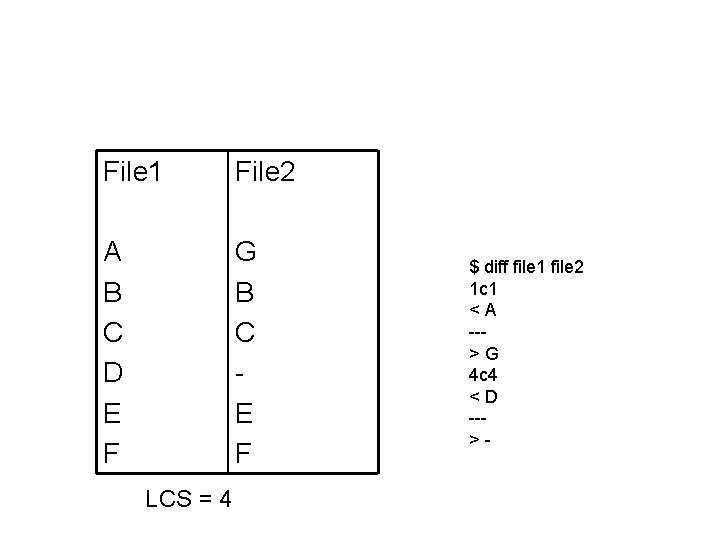
File 1 File 2 A B C D E F G B C E F LCS = 4 $ diff file 1 file 2 1 c 1 <A -->G 4 c 4 <D -->-

The LCS variant Changes: 1. Initialization For all i, j, F(i, 0) = F(0, j) = 0 2. Filling in table F(i-1, j) F(i, j) = max F(i, j-1) F(i-1, j-1) + σ(xi, yj) where σ(xi, yj) = 1 if xi = yj and 0 otherwise. 3. Termination FOPT = maxi F(i, N) maxj F(M, j)

More efficient algorithms • What happens if you have 1 million lines of text in each file? • O(mn) algorithm is too inefficient • Memory inefficient – 1 TB memory to store the matrix • Bounded DP – maybe the majority of the two files are the same? (e. g. , two versions of a software) • Linear-space algorithm – same time complexity
- Slides: 66