CS 5263 Bioinformatics Lecture 13 Hidden Markov Models

CS 5263 Bioinformatics Lecture 13: Hidden Markov Models and applications

Project ideas • • • Implement an HMM Iterative refinement MSA Motif finder Implement an algorithm in a paper Write a survey paper

NAR Web Server Issue • Every December, Nucleic Acids Research has a special issue on web servers • Not necessary to invent original methods • Biologists appreciate web tools • You get a nice publication – And potentially many citations if your tool is useful (think about BLAST!) • Talk to me if you want to work on this project

Review of last lectures

Problems in HMM • Decoding – Predict the state of each symbol – Viterbi algorithm – Forward-backward algorithm • Evaluation – The probability that a sequence is generated by a model – Forward-backward algorithm • Learning – “unsupervised” – Baum-Welch – Viterbi
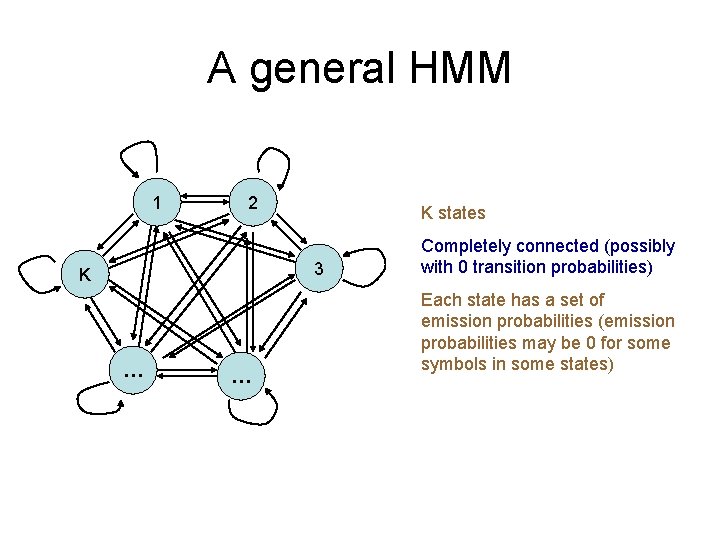
A general HMM 1 2 K states 3 K … … Completely connected (possibly with 0 transition probabilities) Each state has a set of emission probabilities (emission probabilities may be 0 for some symbols in some states)

The Viterbi algorithm V(j, i+1) = max V(1, i) + w(1, j) + r(j, xi+1), V(2, i) + w(2, j) + r(j, xi+1), V(3, i) + w(3, j) + r(j, xi+1), …… V(k, i) + w(k, j) + r(j, xi+1) Or simply: V(j, i+1) = Maxl {V(l, i) + w(l, j) + r(j, xi+1)}

• Viterbi finds the single best path – In many cases it is also interesting to know what are the other possible paths • Viterbi assigns a state to each symbol – In many cases it is also interesting to know the probability that the assignment is correct • Posterior decoding using Forwardbackward algorithm
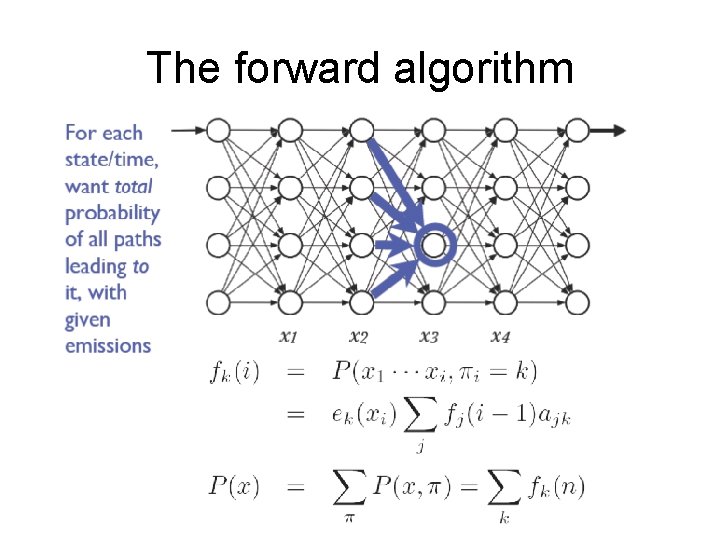
The forward algorithm
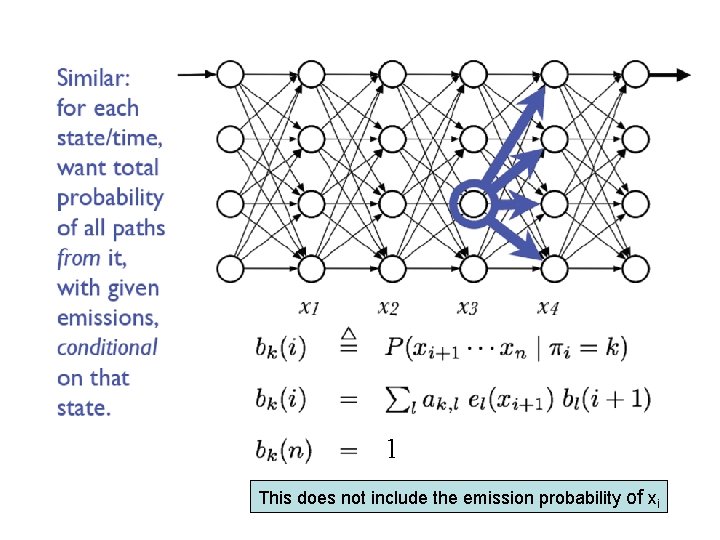
1 This does not include the emission probability of xi
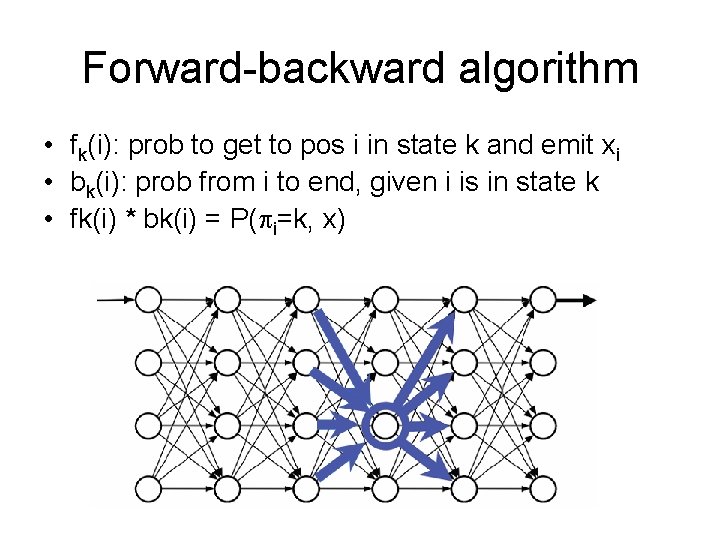
Forward-backward algorithm • fk(i): prob to get to pos i in state k and emit xi • bk(i): prob from i to end, given i is in state k • fk(i) * bk(i) = P( i=k, x)
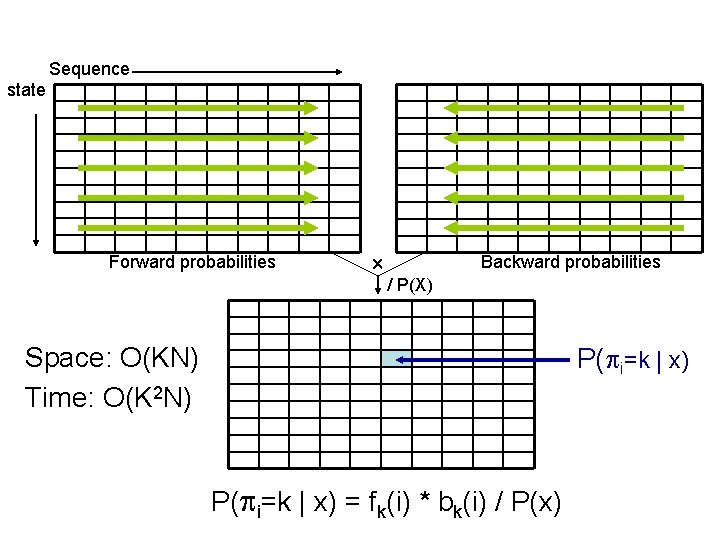
Sequence state Forward probabilities Backward probabilities / P(X) Space: O(KN) Time: O(K 2 N) P( i=k | x) = fk(i) * bk(i) / P(x)

Learning • When the states are known – “supervised” learning • When the states are unknown – Estimate parameters from unlabeled data – “unsupervised” learning How to find Cp. G islands in the porcupine genome?
Basic idea 1. Estimate our “best guess” on the model parameters θ 2. Use θ to predict the unknown labels 3. Re-estimate a new set of θ 4. Repeat 2 & 3 until converge Two ways

Viterbi training 1. Estimate our “best guess” on the model parameters θ 2. Find the Viterbi path using current θ 3. Re-estimate a new set of θ based on the Viterbi path 4. Repeat 2 & 3 until converge

Baum-Welch training 1. Estimate our “best guess” on the model parameters θ 2. Find P( i=k | x, θ) using forward- E-step backward algorithm 3. Re-estimate a new set of θ based on all M-step possible paths 4. Repeat 2 & 3 until converge

Viterbi vs Baum-Welch training • Viterbi training – – Returns a single path Each position labeled with a fixed state Each transition counts one Each emission also counts one • Baum-Welch training – Does not return a single path – Considers the probability that each transition is used – and the probability that a symbol is generated by a certain state – They only contribute partial counts
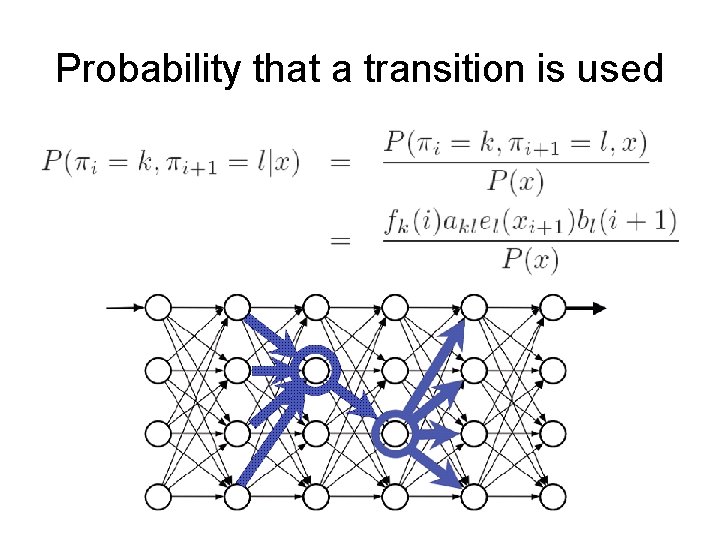
Probability that a transition is used

Viterbi vs Baum-Welch training • Both guaranteed to converges • Baum-Welch improves the likelihood of the data in each iteration – True EM (expectation-maximization) • Viterbi improves the probability of the most probable path in each iteration – EM-like

Today • Some practical issues in HMM modeling • HMMs for sequence alignment

Duration modeling • For any sub-path, the probability consists of two components – The product of emission probabilities • Depend on symbols and state path – The product of transition probabilities • Depend on state path

Duration modeling • Model a stretch of DNA for which the distribution does not change for a certain length • The simplest model implies that P(length = L) = p. L-1(1 -p) • i. e. , length follows geometric distribution – Not always appropriate P Duration: the number of steps that a state is used consecutively without visiting other states p s 1 -p L
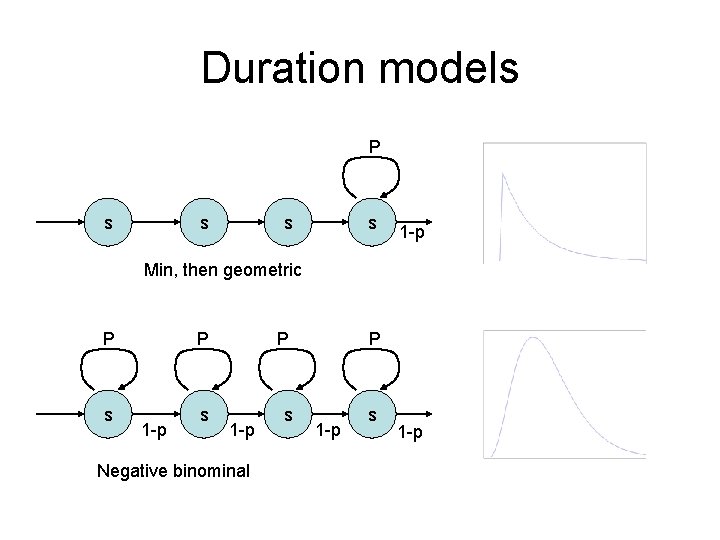
Duration models P s s 1 -p Min, then geometric P P s s 1 -p Negative binominal 1 -p
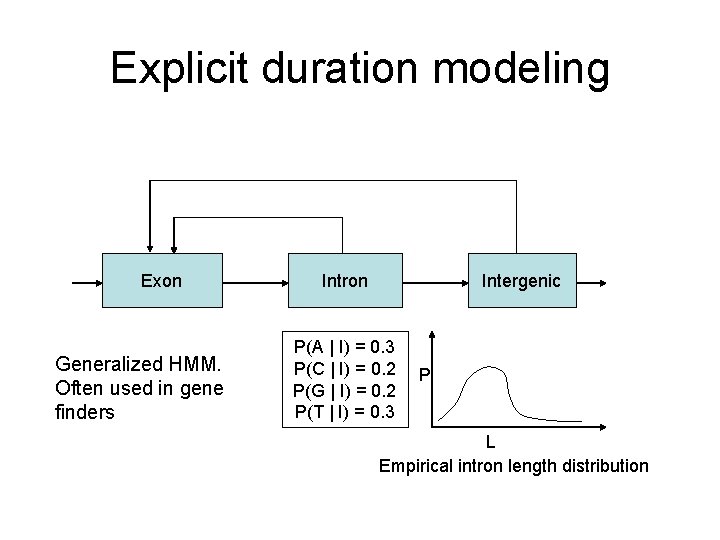
Explicit duration modeling Exon Generalized HMM. Often used in gene finders Intron Intergenic P(A | I) = 0. 3 P(C | I) = 0. 2 P(G | I) = 0. 2 P(T | I) = 0. 3 P L Empirical intron length distribution

Silent states • Silent states are states that do not emit symbols (e. g. , the state 0 in our previous examples) • Silent states can be introduced in HMMs to reduce the number of transitions
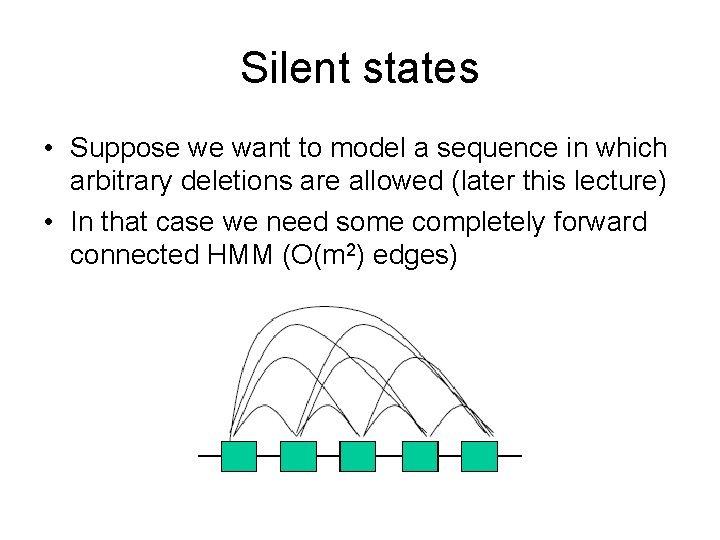
Silent states • Suppose we want to model a sequence in which arbitrary deletions are allowed (later this lecture) • In that case we need some completely forward connected HMM (O(m 2) edges)
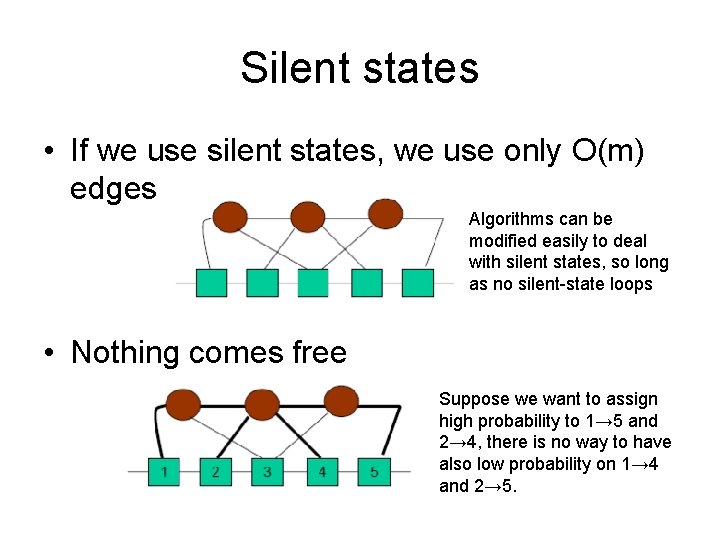
Silent states • If we use silent states, we use only O(m) edges Algorithms can be modified easily to deal with silent states, so long as no silent-state loops • Nothing comes free Suppose we want to assign high probability to 1→ 5 and 2→ 4, there is no way to have also low probability on 1→ 4 and 2→ 5.

Modular design of HMM • HMM can be designed modularly • Each modular has own begin / end states (silent) • Each module communicates with other modules only through begin/end states
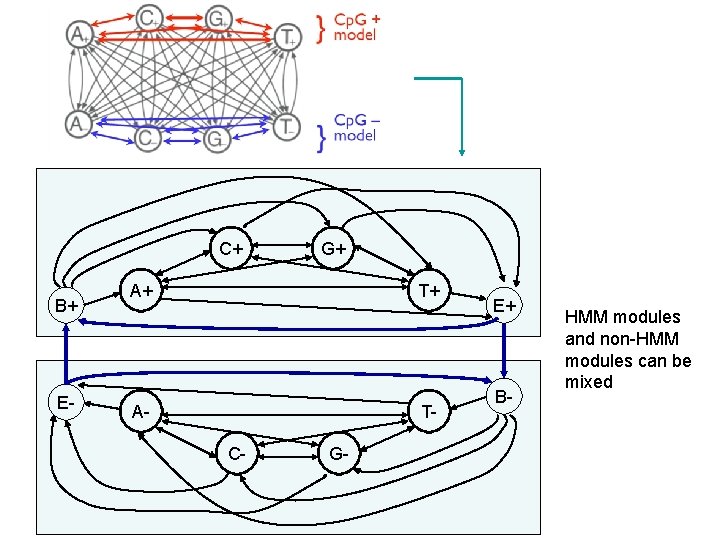
C+ B+ E- G+ A+ T+ A- TC- G- E+ B- HMM modules and non-HMM modules can be mixed
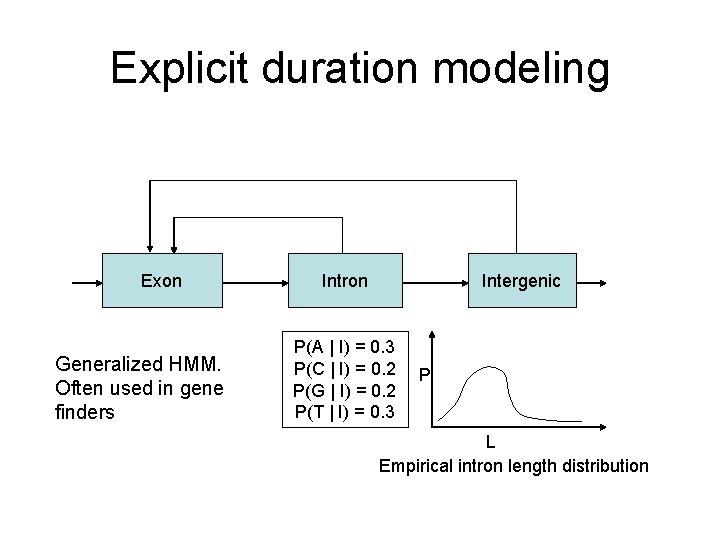
Explicit duration modeling Exon Generalized HMM. Often used in gene finders Intron Intergenic P(A | I) = 0. 3 P(C | I) = 0. 2 P(G | I) = 0. 2 P(T | I) = 0. 3 P L Empirical intron length distribution

HMM applications • • Pair-wise sequence alignment Multiple sequence alignment Gene finding Speech recognition: a good tutorial on course website • Machine translation • Many others
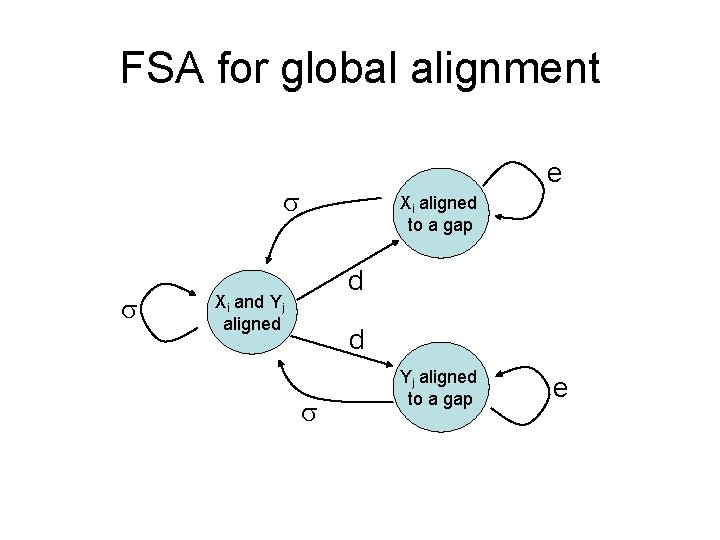
FSA for global alignment e Xi aligned to a gap d Xi and Yj aligned d Yj aligned to a gap e
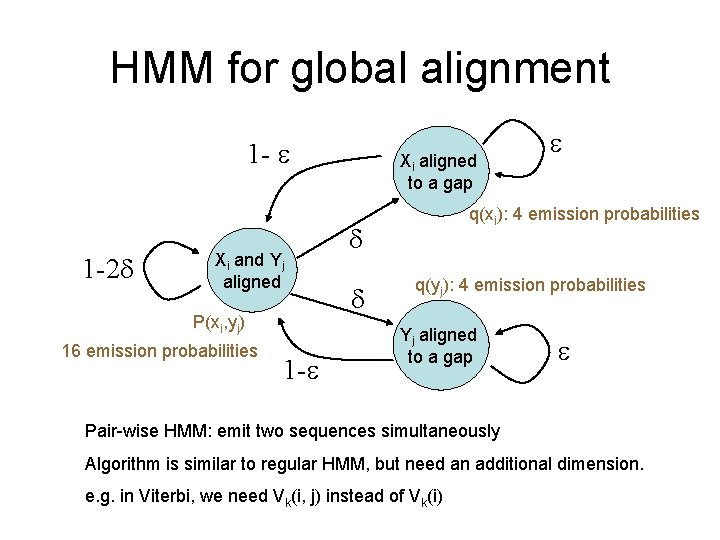
HMM for global alignment 1 - 1 -2 Xi and Yj aligned P(xi, yj) 16 emission probabilities 1 - Xi aligned to a gap q(xi): 4 emission probabilities q(yj): 4 emission probabilities Yj aligned to a gap Pair-wise HMM: emit two sequences simultaneously Algorithm is similar to regular HMM, but need an additional dimension. e. g. in Viterbi, we need Vk(i, j) instead of Vk(i)

HMM and FSA for alignment • After proper transformation between the probabilities and substitution scores, the two are identical Ø (a, b) log [p(a, b) / (q(a) q(b))] Ø d log Ø e log • Details in Durbin book chap 4 • Finding an optimal FSA alignment is equivalent to finding the most probable path with Viterbi

HMM for pair-wise alignment • Theoretical advantages: – Full probabilistic interpretation of alignment scores – Probability of all alignments instead of the best alignment (forward-backward alignment) – Posterior probability that Ai is aligned to Bj – Sampling sub-optimal alignments • Not commonly used in practice – Needleman-Wunsch and Smith-Waterman algorithms work pretty well, and more intuitive to biologists – Other reasons?

Other applications • HMM for multiple alignment – Widely used • HMM for gene finding – Foundation for most gene finders – Include many knowledge-based fine-tunes and GHMM extensions – We’ll only discuss basic ideas

HMM for multiple seq alignment • Proteins form families both across and within species – Ex: Globins, Zinc finger – Descended from a common ancestor – Typically have similar three-dimensional structures, functions, and significant sequence similarity • Identifying families is very useful: suggest functions • So: search and alignment are both useful • Multiple alignment is hard • One very useful approach: profile-HMM

Say we already have a database of reliable multiple alignment of protein families When a new protein comes, how do we align it to the existing alignments and find the family that the protein may belong to?

Solution 1 • Use regular expression to represent the consensus sequences – Implemented in the Prosite database – for example C - x(2) - P - F - x - [FYWIV] - x(7) - C x(8, 10) - W - C - x(4) - [DNSR] - [FYW] x(3, 5) - [FYW] - x - [FYWI] - C

Multi-alignments represented by consensus • Consensus sequences are very intuitive • Gaps can be represented by Do-not-cares • Aligning sequences with regular expressions can be done easily with DP • Possible to allow mismatches in searching • Problems – Limited power in expressing more divergent positions • E. g. among 100 seqs, 60 have Alanine, 20 have Glycine, 20 others – Specify boundaries of indel not so easy • unaligned regions have more flexibilities to evolve – May have to change the regular expression when a new member of a protein family is identified
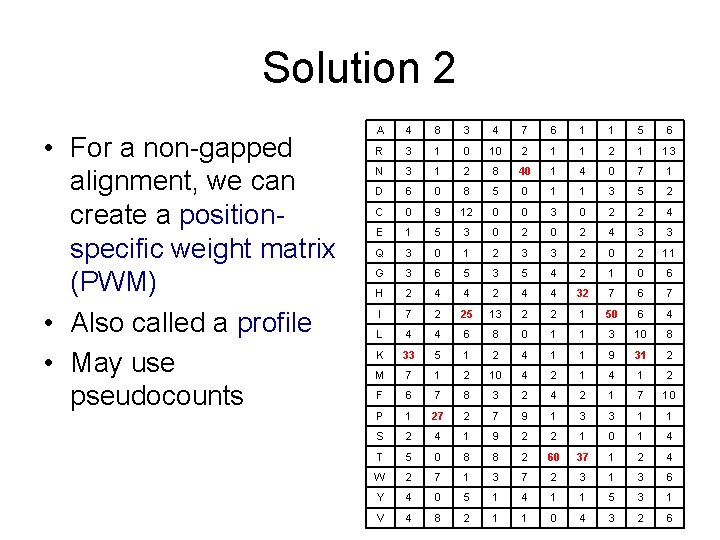
Solution 2 • For a non-gapped alignment, we can create a positionspecific weight matrix (PWM) • Also called a profile • May use pseudocounts A 4 8 3 4 7 6 1 1 5 6 R 3 1 0 10 2 1 13 N 3 1 2 8 40 1 4 0 7 1 D 6 0 8 5 0 1 1 3 5 2 C 0 9 12 0 0 3 0 2 2 4 E 1 5 3 0 2 4 3 3 Q 3 0 1 2 3 3 2 0 2 11 G 3 6 5 3 5 4 2 1 0 6 H 2 4 4 32 7 6 7 I 7 2 25 13 2 2 1 50 6 4 L 4 4 6 8 0 1 1 3 10 8 K 33 5 1 2 4 1 1 9 31 2 M 7 1 2 10 4 2 1 4 1 2 F 6 7 8 3 2 4 2 1 7 10 P 1 27 2 7 9 1 3 3 1 1 S 2 4 1 9 2 2 1 0 1 4 T 5 0 8 8 2 60 37 1 2 4 W 2 7 1 3 7 2 3 1 3 6 Y 4 0 5 1 4 1 1 5 3 1 V 4 8 2 1 1 0 4 3 2 6

Scoring by PWMs A 4 8 3 4 7 6 1 1 5 6 R 3 1 0 10 2 1 13 N 3 1 2 8 40 1 4 0 7 1 D 6 0 8 5 0 1 1 3 5 2 C 0 9 12 0 0 3 0 2 2 4 E 1 5 3 0 2 4 3 3 Q 3 0 1 2 3 3 2 0 2 11 G 3 6 5 3 5 4 2 1 0 6 H 2 4 4 32 7 6 7 I 7 2 25 13 2 2 1 50 6 4 L 4 4 6 8 0 1 1 3 10 8 K 33 5 1 2 4 1 1 9 31 2 M 7 1 2 10 4 2 1 4 1 2 F 6 7 8 3 2 4 2 1 7 10 P 1 27 2 7 9 1 3 3 1 1 S 2 4 1 9 2 2 1 0 1 4 T 5 0 8 8 2 60 37 1 2 4 W 2 7 1 3 7 2 3 1 3 6 Y 4 0 5 1 4 1 1 5 3 1 V 4 8 2 1 1 0 4 3 2 6 x = KCIDNTHIKR P(x | M) = i ei(xi) Random model: each amino acid xi can be generated with probability q(xi) P(x | random) = i q(xi) Log-odds ratio: S = log P(X|M) / P(X|random) = i log ei(xi) / q(xi)

PWMs • Advantage: – Can be used to identify both strong and weak homologies – Easy to implement and use – Probabilistic interpretation • PWMs are used in PSI-BLAST – – Given a sequence, get k similar seqs by BLAST Construct a PWM with these sequences Search the database for seqs matching the PWM Improved sensitivity • Problem: – Not intuitive to deal with gaps
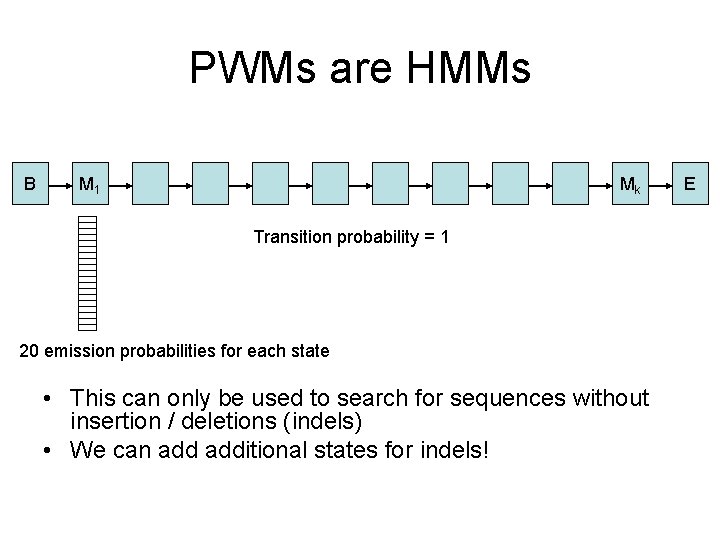
PWMs are HMMs B M 1 Mk Transition probability = 1 20 emission probabilities for each state • This can only be used to search for sequences without insertion / deletions (indels) • We can additional states for indels! E
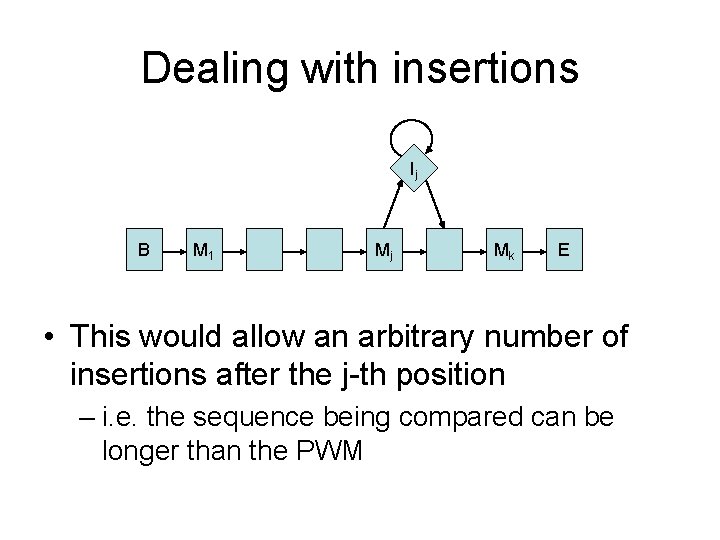
Dealing with insertions Ij B M 1 Mj Mk E • This would allow an arbitrary number of insertions after the j-th position – i. e. the sequence being compared can be longer than the PWM
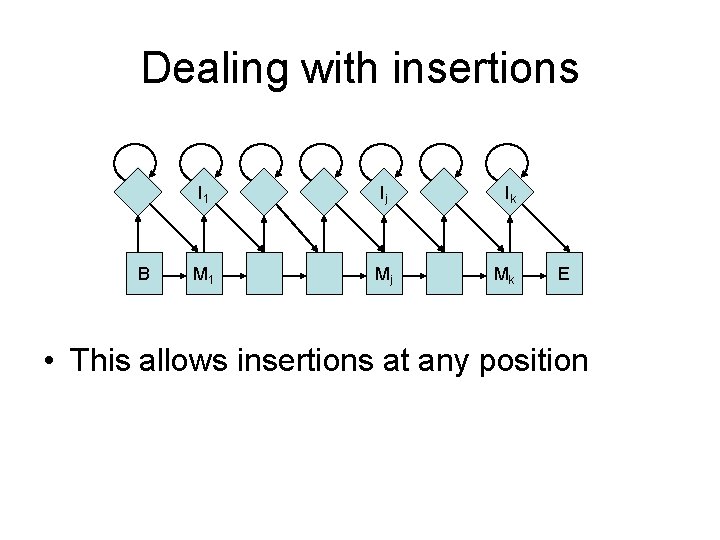
Dealing with insertions B I 1 Ij Ik M 1 Mj Mk E • This allows insertions at any position
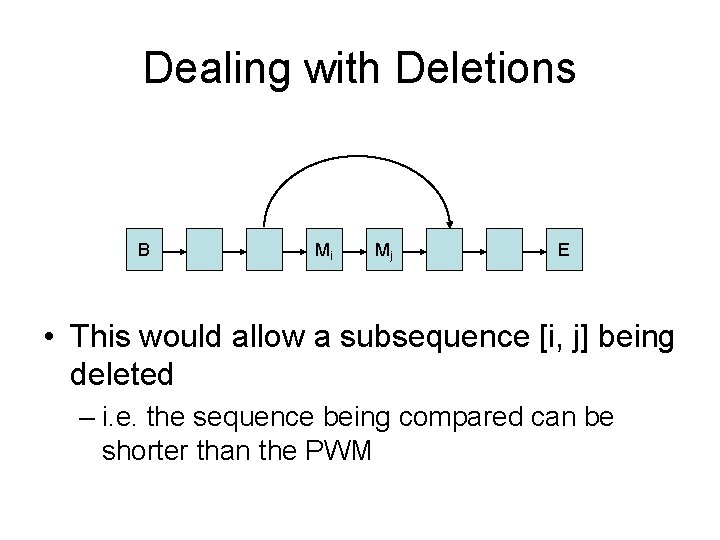
Dealing with Deletions B Mi Mj E • This would allow a subsequence [i, j] being deleted – i. e. the sequence being compared can be shorter than the PWM
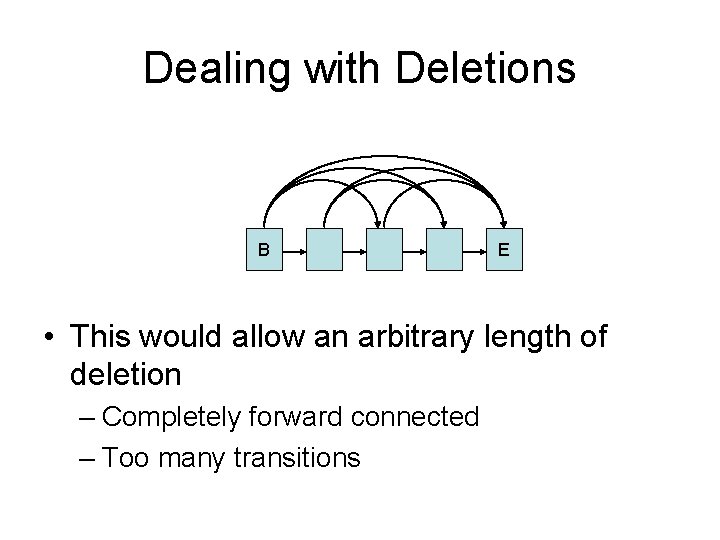
Dealing with Deletions B E • This would allow an arbitrary length of deletion – Completely forward connected – Too many transitions
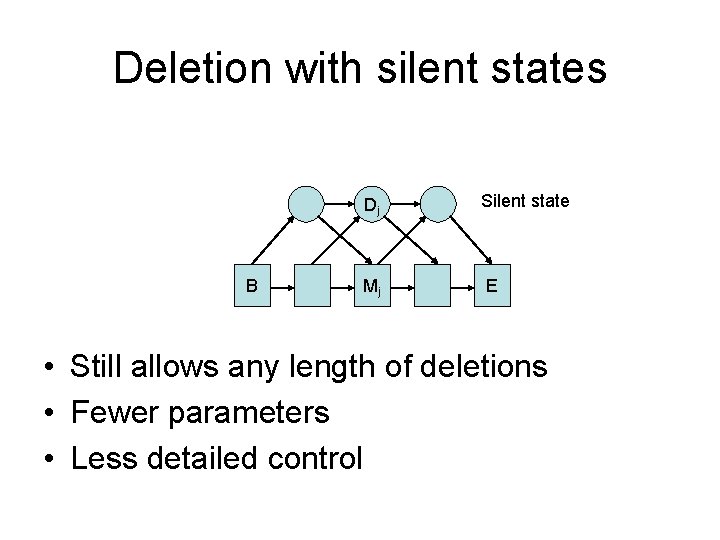
Deletion with silent states B Dj Silent state Mj E • Still allows any length of deletions • Fewer parameters • Less detailed control
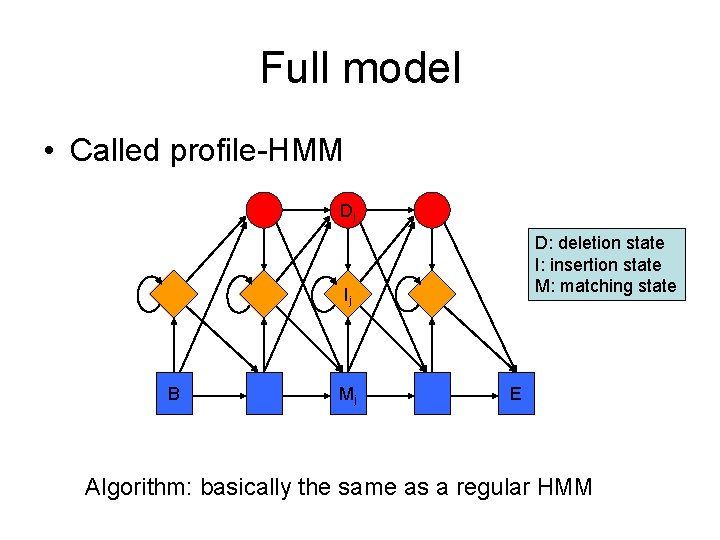
Full model • Called profile-HMM Dj D: deletion state I: insertion state M: matching state Ij B Mj E Algorithm: basically the same as a regular HMM

Using profile HMM • Alignment – Align a sequence to a profile HMM – Viterbi • Searching – Given a sequence and HMMs for different protein families, which family the sequence may belong to? – Given a HMM for a protein family and many proteins, which protein may belong to the family? – Viterbi or forward – How to score? • Training / Learning – Given a multiple alignment, how to construct a HMM? – Given an unaligned protein family, how to construct a HMM?

Pfam • A database of protein families – Developed by Sean Eddy and colleagues while working in Durbin’s lab • • Hand-curated “seed” multiple alignment Train HMM from seed alignment Hand-chosen score thresholds Automatic classification / classification of all other protein sequences • 7973 families in Rfam 18. 0, 8/2005 (covers ~75% of proteins)
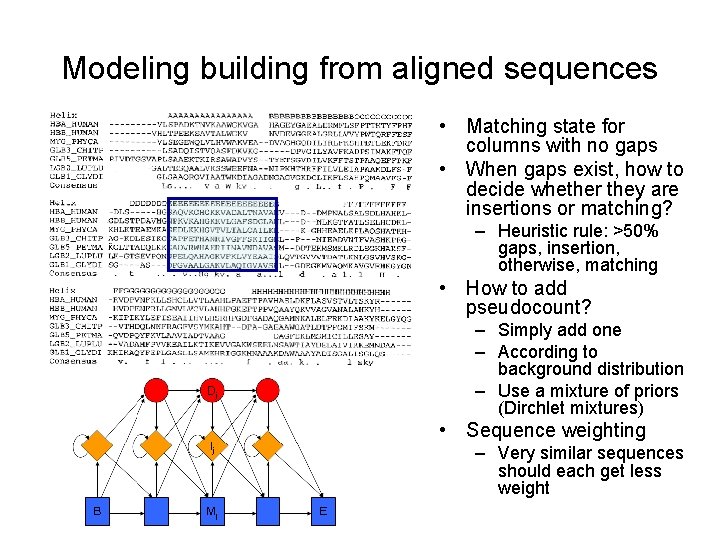
Modeling building from aligned sequences • Matching state for columns with no gaps • When gaps exist, how to decide whether they are insertions or matching? – Heuristic rule: >50% gaps, insertion, otherwise, matching • How to add pseudocount? – Simply add one – According to background distribution – Use a mixture of priors (Dirchlet mixtures) • Sequence weighting – Very similar sequences should each get less weight

Modeling building from unaligned sequences • Choose a model length and initial parameters – Commonly use average seq length as model length • Baum-Welch or Viterbi training – Usually necessary to use multiple starting points or other heuristics to escape from local optima • Align all sequences to the final model using Viterbi
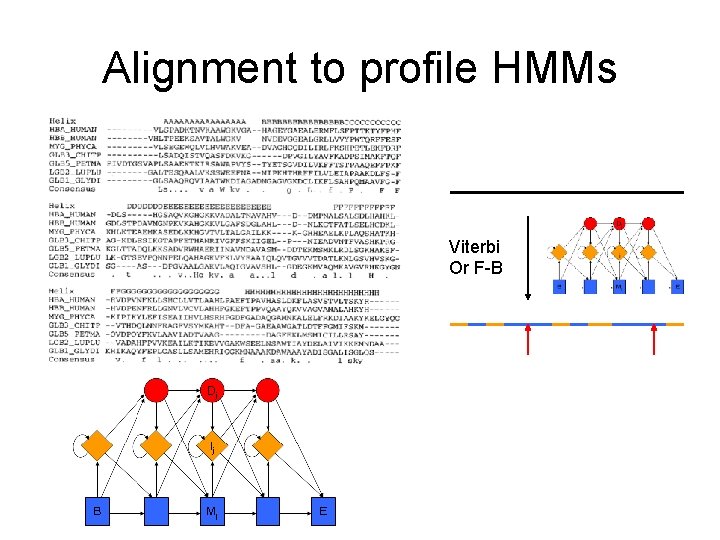
Alignment to profile HMMs Viterbi Or F-B
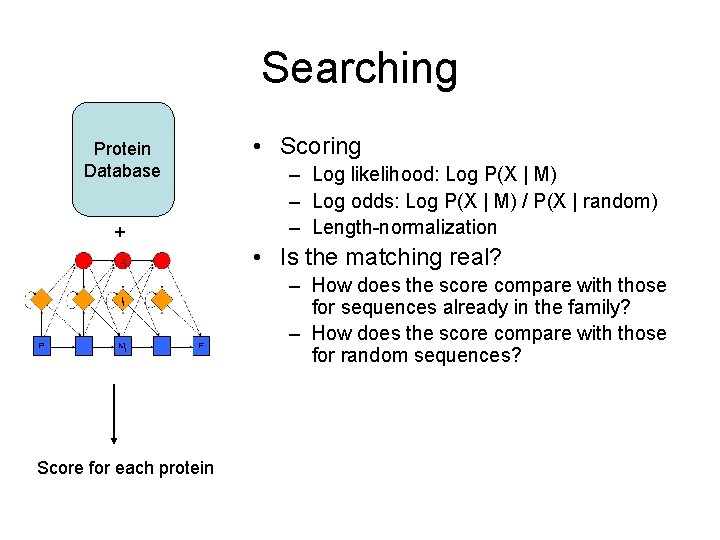
Searching Protein Database + • Scoring – Log likelihood: Log P(X | M) – Log odds: Log P(X | M) / P(X | random) – Length-normalization • Is the matching real? – How does the score compare with those for sequences already in the family? – How does the score compare with those for random sequences? Score for each protein

Example: modeling and searching for globins • 300 random picked globin sequence • Build a profile HMM from scratch (without pre-alignment) • Align 60, 000 proteins to the HMM
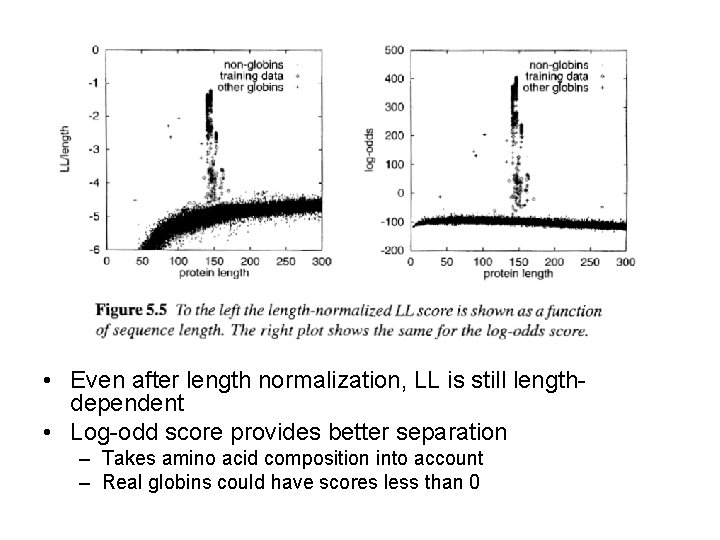
• Even after length normalization, LL is still lengthdependent • Log-odd score provides better separation – Takes amino acid composition into account – Real globins could have scores less than 0
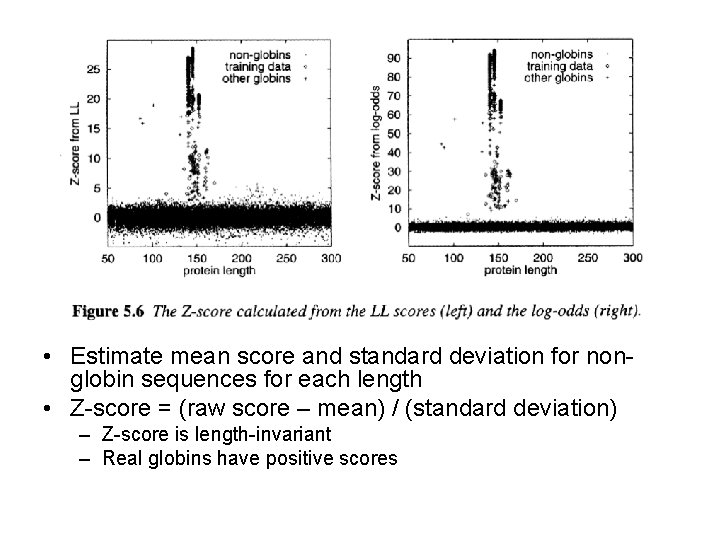
• Estimate mean score and standard deviation for nonglobin sequences for each length • Z-score = (raw score – mean) / (standard deviation) – Z-score is length-invariant – Real globins have positive scores

Next lecture • Gene finding • HMM wrap up
- Slides: 60