RNASeq Xiaole Shirley Liu STAT 115 STAT 215






























- Slides: 53

RNA-Seq Xiaole Shirley Liu STAT 115, STAT 215, BIO 298, BIST 520 Guest lecture by Wei Li

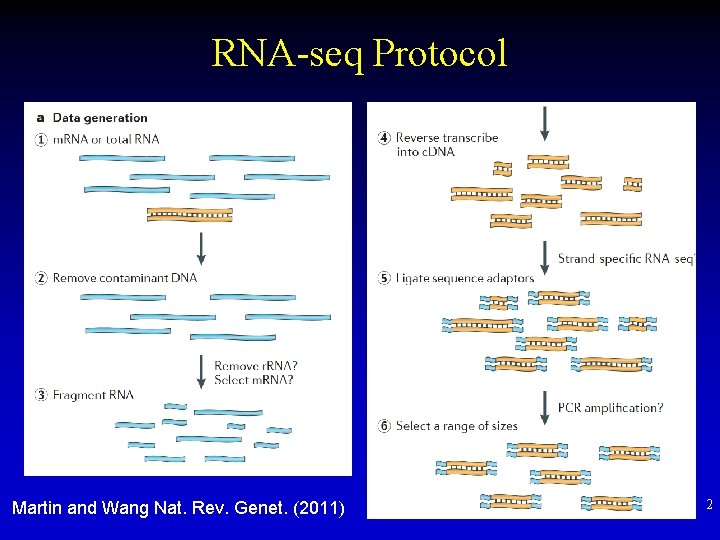
RNA-seq Protocol Martin and Wang Nat. Rev. Genet. (2011) 2

RNA-seq • https: //www. youtube. com/watch? v=V_4 n 8 n 5 Z 6 I 8 • (RNA-Seq using Ion Proton) 3

Why RNA-seq, not microarray? • No need to design microarray probes • Digital representation, higher detection range • Alternative splicing • Fusion • Mutations 4

RNA-seq Applications • Gene expression; differential expression 5

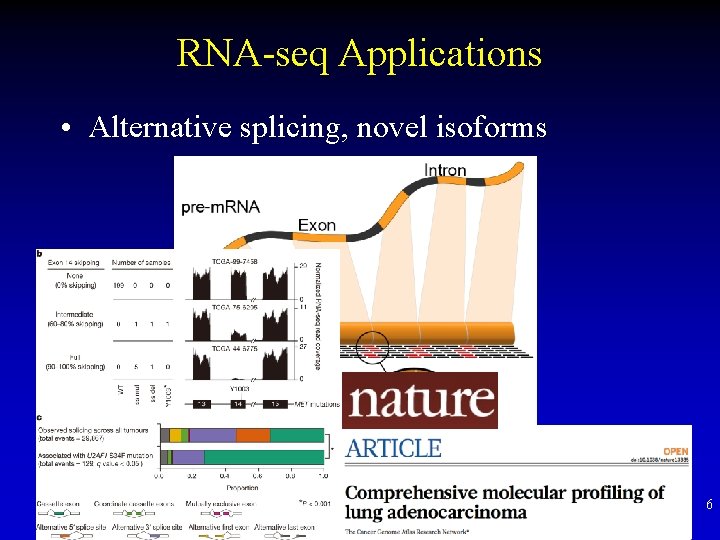
RNA-seq Applications • Alternative splicing, novel isoforms 6

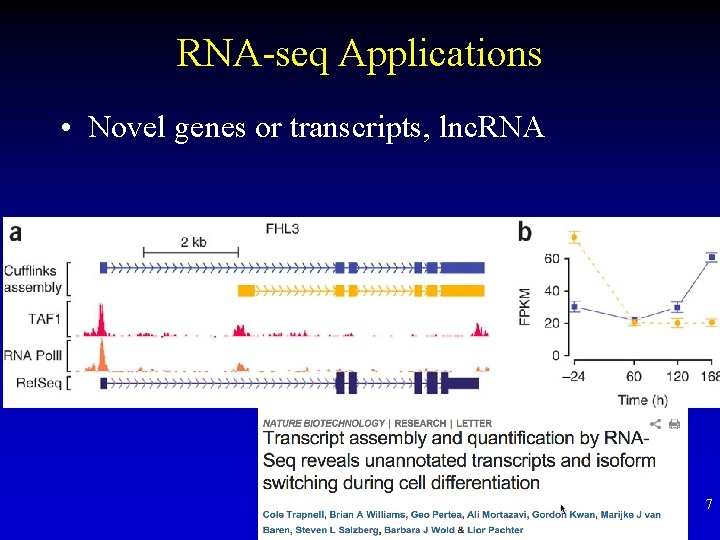
RNA-seq Applications • Novel genes or transcripts, lnc. RNA 7

RNA-seq Applications • Detect gene fusions • Mutations, RNA editing 8

RNA-seq Experimental Design and Analysis

Experimental Design • Assessing biological variation requires biological replicates (no need for technical replicates) • 3 preferred, 2 OK, 1 only for exploratory assays (not good for publications) 10

Experimental Design • For differential expression, don’t pool RNA from multiple biological replicates • Batch effects still exist, try to be consistent or process all samples at the same time 11

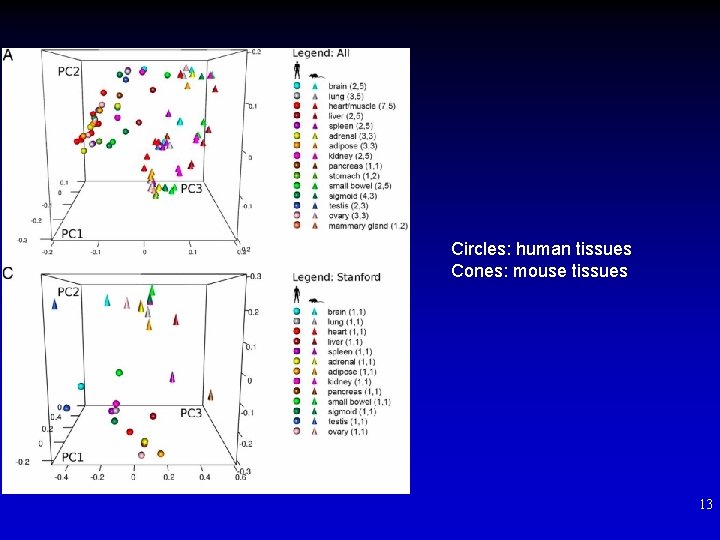
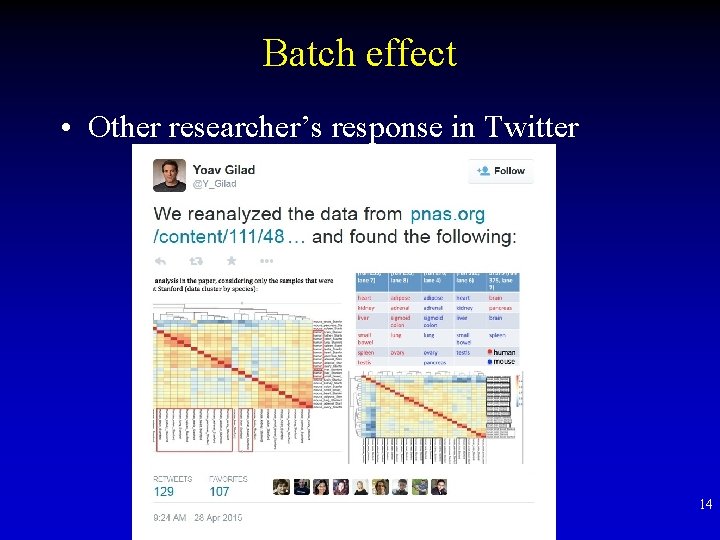
Batch effect • A research group’s striking finding in 2014 • “Human heart is more similar with human brain than mouse brain” Human Heart Mouse Brain Human Brain 12

Circles: human tissues Cones: mouse tissues 13

Batch effect • Other researcher’s response in Twitter 14

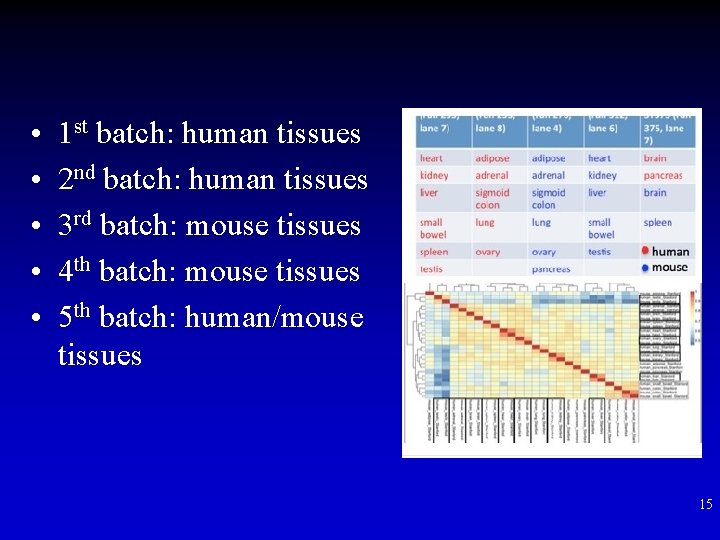
• • • 1 st batch: human tissues 2 nd batch: human tissues 3 rd batch: mouse tissues 4 th batch: mouse tissues 5 th batch: human/mouse tissues 15

Batch effect 16

Batch effect • Before experiments: careful design • After experiments: batch effect removal (combat) 17

Experimental Design • • Ribo-minus (remove too abundant genes) Poly. A (m. RNA, enrich for exons) Strand specific (anti-sense lnc. RNA) Sequencing: – PE (resolve redundancy) or SE: expression – PE for splicing, novel transcripts – Depth: 30 -50 M differential expression, deeper transcript assembly – Read length: longer for transcript assembly 18

Alignment • Prefer splice-aware aligners • Top. Hat, BWA, STAR (not DNASTAR) • Sometimes need to trim the beginning bases 19

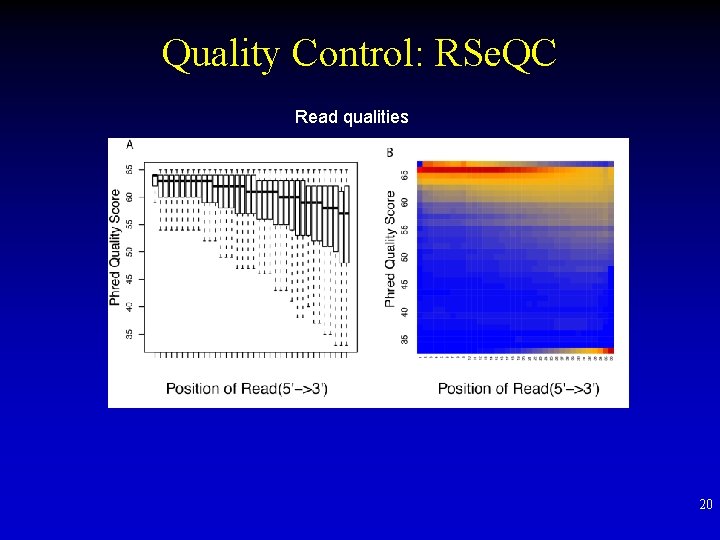
Quality Control: RSe. QC Read qualities 20

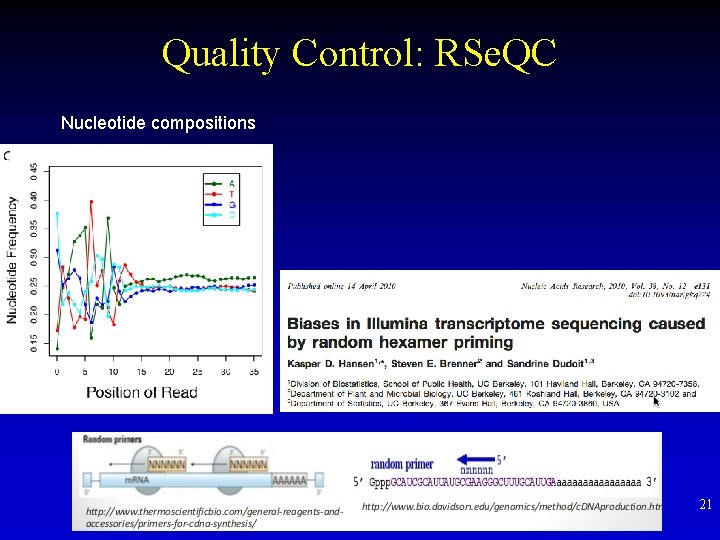
Quality Control: RSe. QC Nucleotide compositions 21

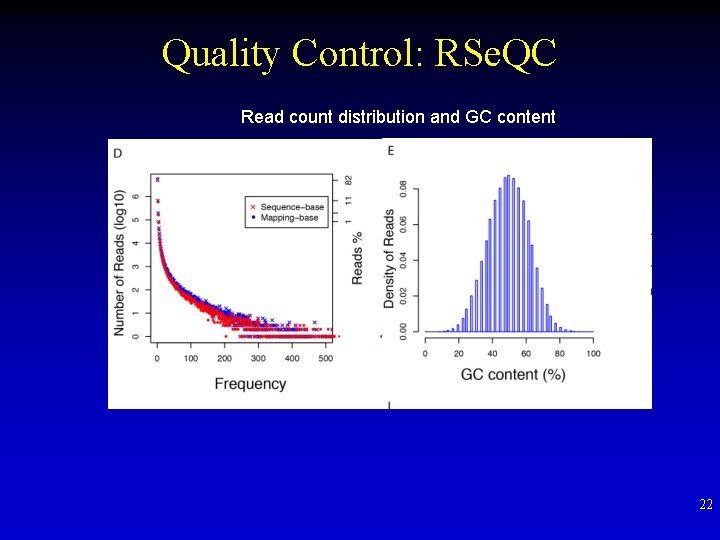
Quality Control: RSe. QC Read count distribution and GC content 22

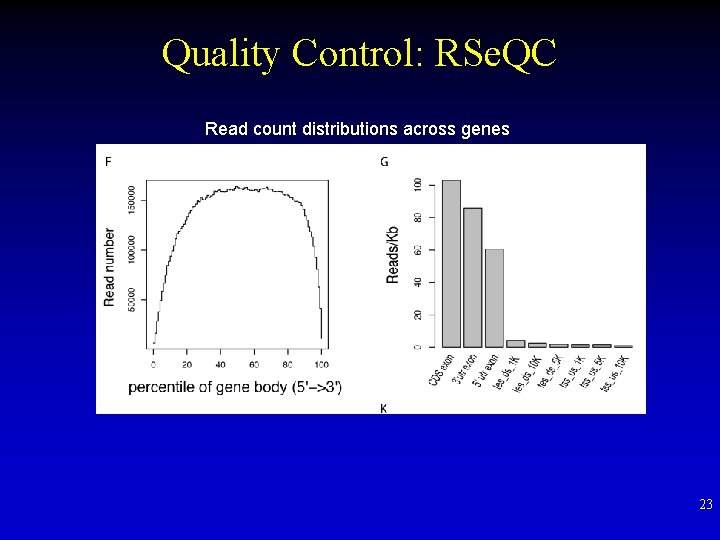
Quality Control: RSe. QC Read count distributions across genes 23

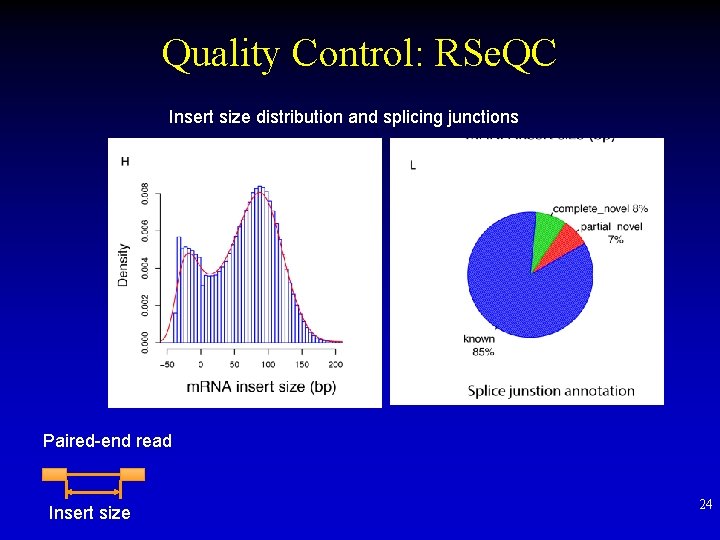
Quality Control: RSe. QC Insert size distribution and splicing junctions Paired-end read Insert size 24

Differential Expression

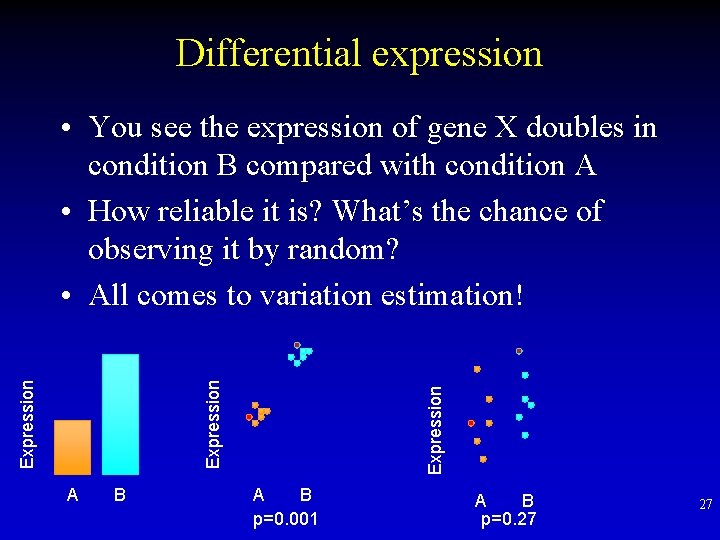
Differential expression A B Expression • You see the expression of gene X doubles in condition B compared with condition A • How reliable it is? What’s the chance of observing it by random? • All comes to variation estimation! A B p=0. 001 A B p=0. 27 27

Differential expression • Variation can be estimated if you have many biological replicates • But in practice, only 2 -3 replicates are available • What to do next? – Proper statistical models 28

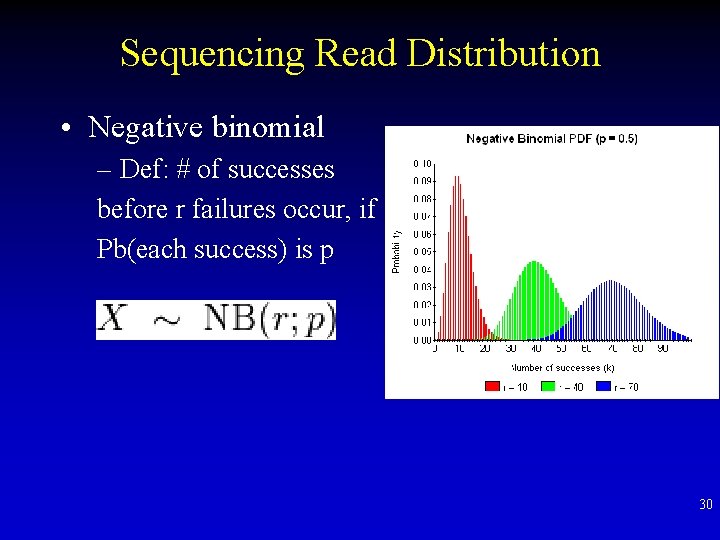
Sequencing Read Distribution • Poisson distribution: – # events within an interval – Mean = Variance • But: sequencing data is over-dispersed (Mean<Variance) 29

Sequencing Read Distribution • Negative binomial – Def: # of successes before r failures occur, if Pb(each success) is p 30

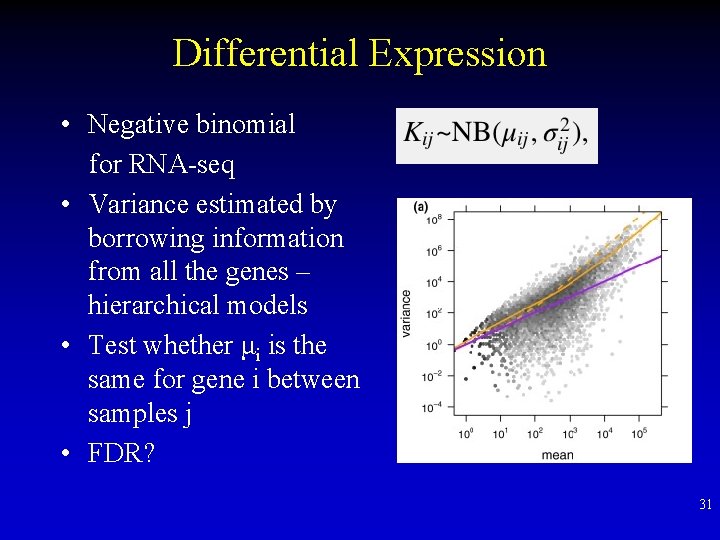
Differential Expression • Negative binomial for RNA-seq • Variance estimated by borrowing information from all the genes – hierarchical models • Test whether μi is the same for gene i between samples j • FDR? 31

Differential expression • Edge. R • DESeq/DESeq 2 32

Expression Index • RPKM (Reads per kilobase of transcript per million reads of library) – Corrects for coverage, gene length – 1 RPKM ~ 0. 3 -1 transcript / cell – Comparable between different genes within the same dataset – Top. Hat / Cufflinks • FPKM (Fragments), PE libraries, RPKM/2 • TPM (transcripts per million) – Normalizes to transcript copies instead of reads – Longer transcripts have more reads – RSEM, HTSeq 33

Differential Expression • Should we do differential expression on RPKM/FPKM or TPM? Gene A (1 kb) Gene B (8 kb) • • Cufflinks: RPKM/FPKM LIMMA-VOOM and DESeq: TPM Power to detect DE is proportional to length Continued development and updates 34

Alternative Splicing • Assign reads to splice isoforms (Top. Hat) 35

Alternative Splicing • Different AS events 36

Alternative Splicing • MATS: Multivariate Analysis of Transcript Splicing 37

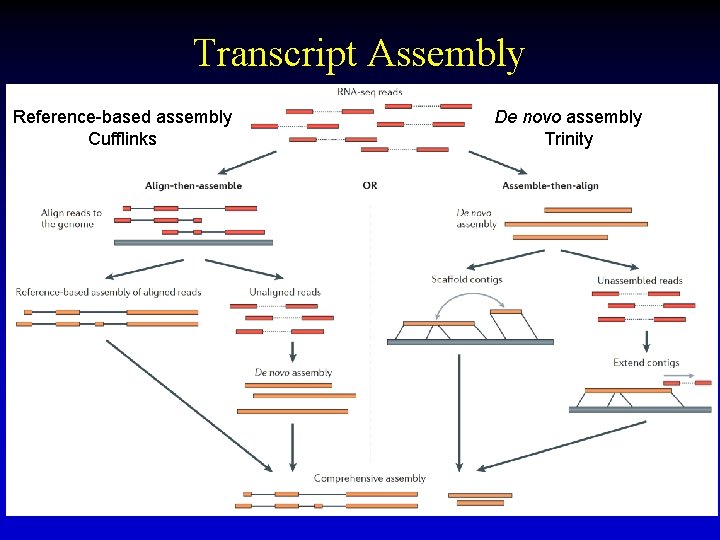
Transcript Assembly Reference-based assembly Cufflinks De novo assembly Trinity 38

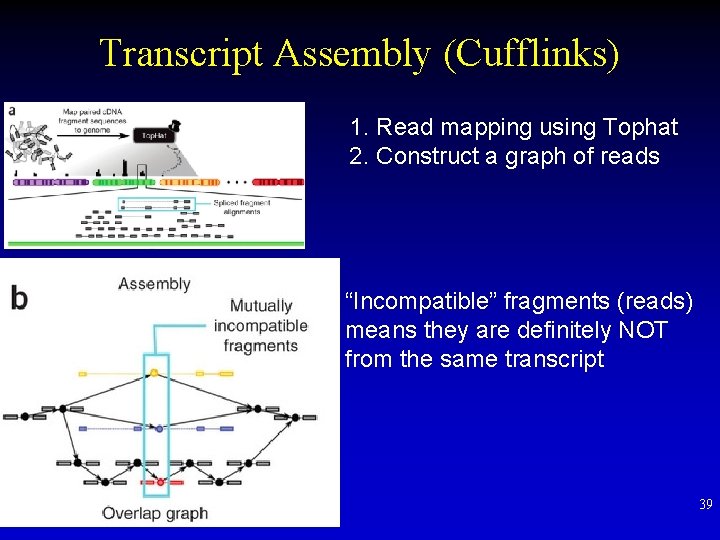
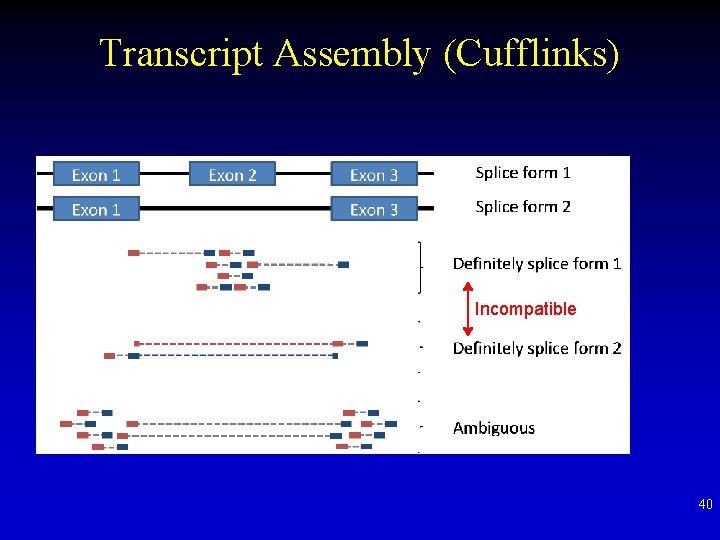
Transcript Assembly (Cufflinks) 1. Read mapping using Tophat 2. Construct a graph of reads “Incompatible” fragments (reads) means they are definitely NOT from the same transcript 39

Transcript Assembly (Cufflinks) Incompatible 40

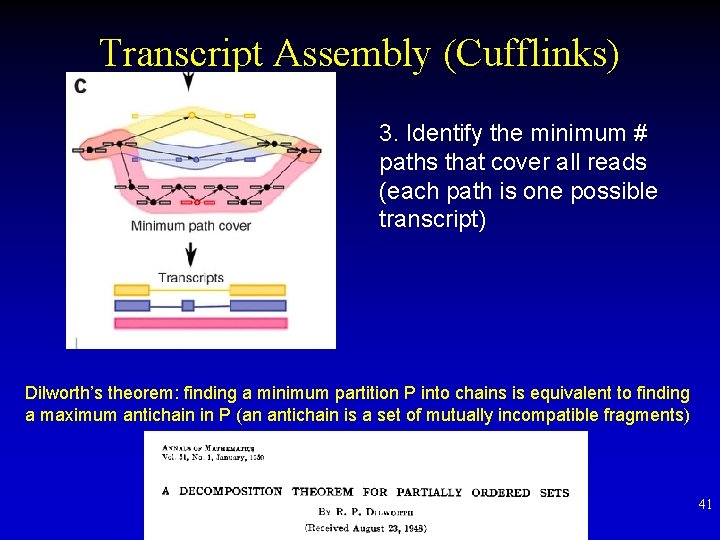
Transcript Assembly (Cufflinks) 3. Identify the minimum # paths that cover all reads (each path is one possible transcript) Dilworth’s theorem: finding a minimum partition P into chains is equivalent to finding a maximum antichain in P (an antichain is a set of mutually incompatible fragments) 41

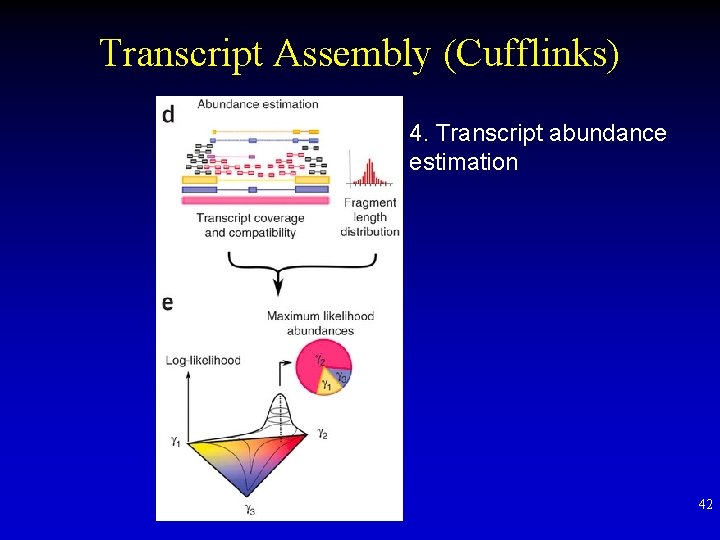
Transcript Assembly (Cufflinks) 4. Transcript abundance estimation 42

Isoform Inference • If given known set of isoforms • Estimate x to maximize the likelihood of observing n 43

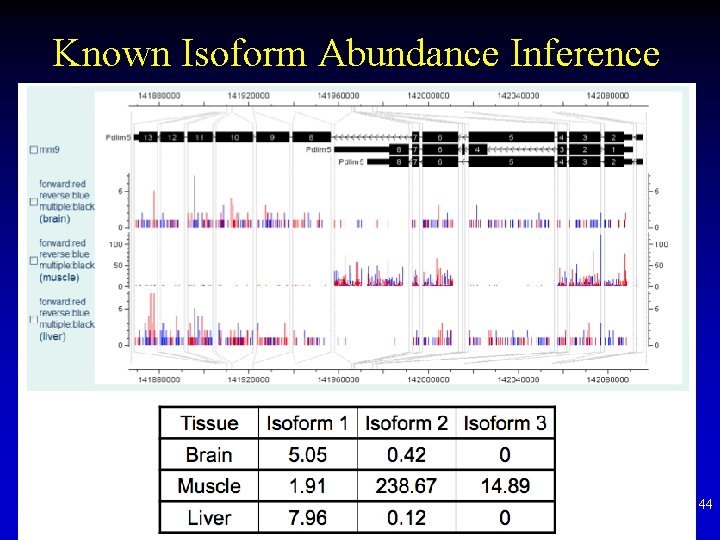
Known Isoform Abundance Inference 44

Isoform Inference • With known isoform set, sometimes the gene-level expression level inference is great, although isoform abundances have big uncertainty (e. g. known set incomplete) • De novo isoform inference is a nonidentifiable problem if RNA-seq reads are short and gene is long with too many exons • Algorithm: Trinity 45

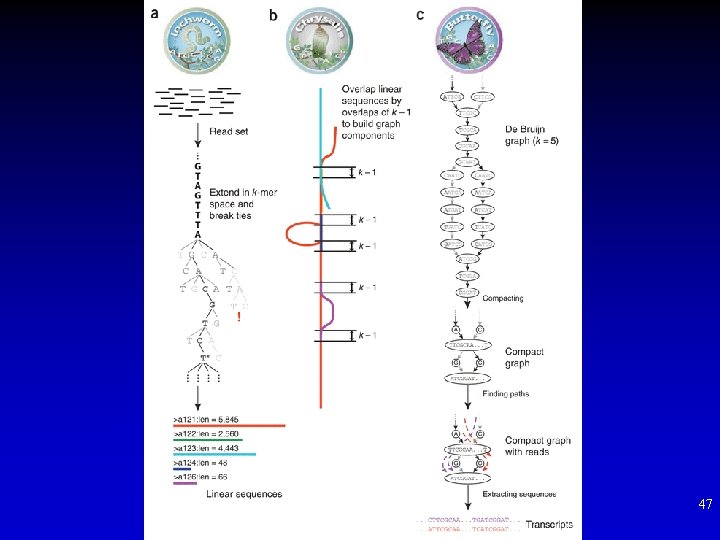
De-novo transcriptome assembly 46

47

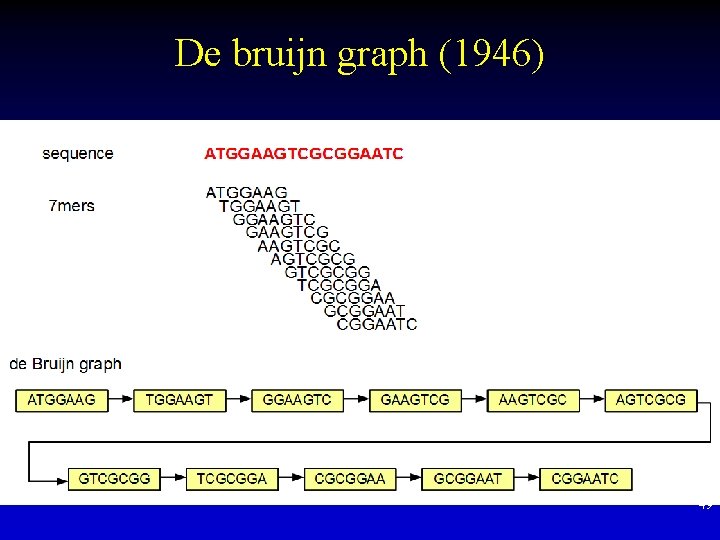
De bruijn graph (1946) • Used in the earliest human genome assemblies • Standard algorithm for genome assembly • A sequence of length k can be represented as an edge between two sequences (length k -1) 48

De bruijn graph (1946) 49

De bruijn graph • How to do genome assembly? • Sequences as nodes -> traverse all nodes in a graph -> Hamilton path problem -> NP complete problem! • De bruijn graph: Sequences as edges -> traverse all edges in a graph -> Euler graph > Polynomial algorithm! 50

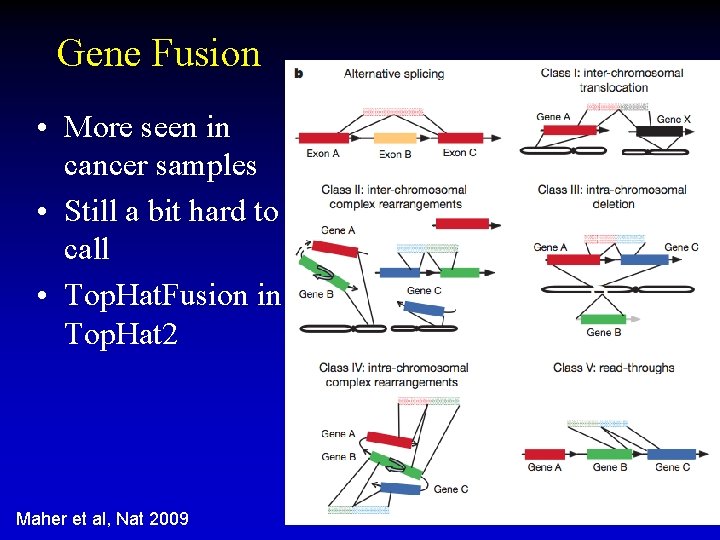
Gene Fusion • More seen in cancer samples • Still a bit hard to call • Top. Hat. Fusion in Top. Hat 2 Maher et al, Nat 2009 51

Other Applications • RNA editing – Change on RNA sequence after transcription – Most frequent: A to I (behaves like G), C to U – Evolves from mononucleotide deaminases, might be involved in RNA degradation • Circular RNA – Mostly arise from splicing – Varying length, abundance, and stability – Possible function: sponge for RBP or mi. RNA 52

Summary • • • RNA-seq design considerations Read mapping: Top. Hat, BWA, STAR De novo transcriptome assembly: TRINITY Quality control: RSe. QC Expression index: FPKM and TPM Differential expression – Cufflinks: versatile – LIMMA-VOOM and DESeq: better variance estimates • Alternative splicing: MATS • Gene fusion, genome editing, circular RNA 53

Acknowledgement • Alisha Holloway • Simon Andrews • Radhika Khetani 54