RNASeq Xiaole Shirley Liu STAT 115215 BIOBST 282

RNA-Seq Xiaole Shirley Liu STAT 115/215, BIO/BST 282
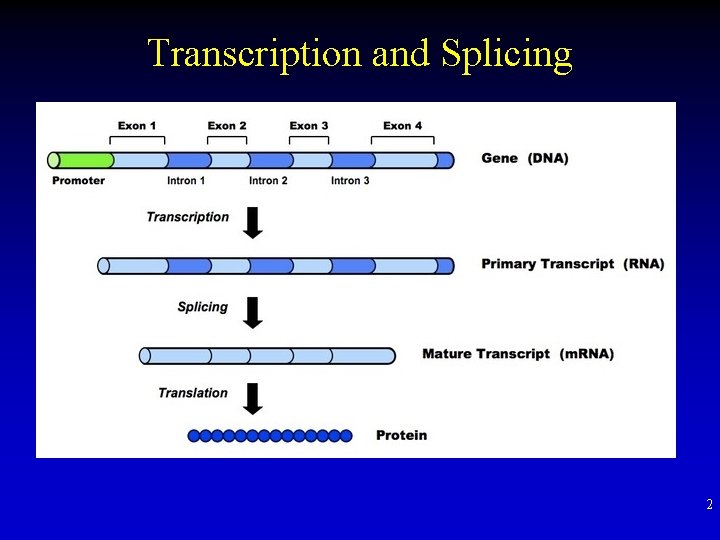
Transcription and Splicing 2
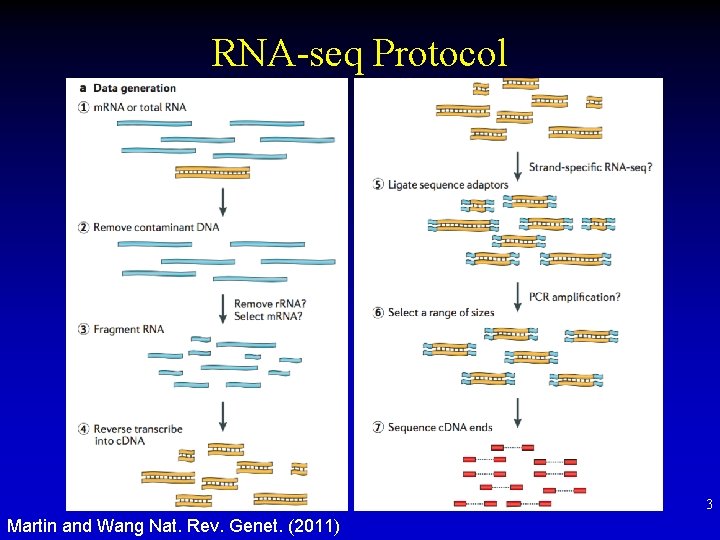
RNA-seq Protocol 3 Martin and Wang Nat. Rev. Genet. (2011)
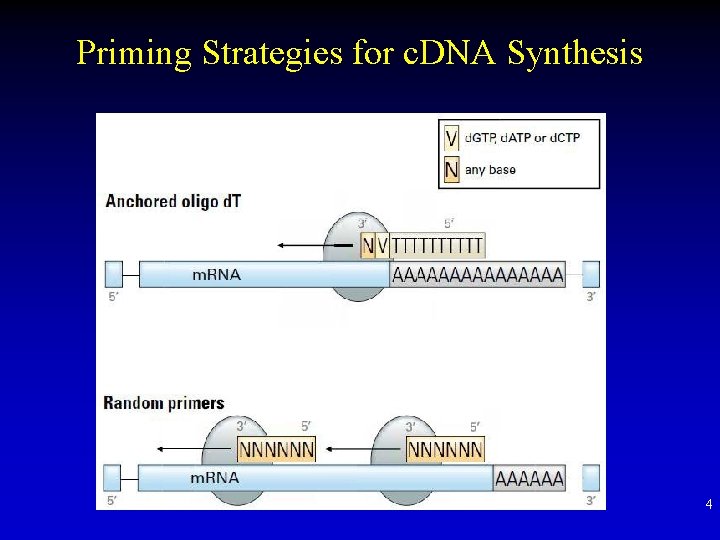
Priming Strategies for c. DNA Synthesis 4

Why RNA-seq, not Microarray? • No need to know the genome sequence or predict genes • No need to design microarray probes • Digital representation • Higher detection range • New genes • Alternative splicing • Mutations and gene fusion 5

Experimental Design • Poly. A (m. RNA after splicing, enrich for exons) • Ribo-minus (remove too abundant house keeping genes) • Strand specific (directionality of lnc. RNA) • Sequencing: what does $500 / sample mean? – SE or PE: PE for splicing, novel transcripts – Depth: 30 -50 M differential expression, deeper transcript assembly – Read length: longer for transcript assembly, mutation calls 6

Experimental Design • Assessing biological variation requires biological replicates (no need for technical replicates) • 3 preferred, 2 OK, 1 only for exploratory assays (not good for publications) • Batch effects still exist, try to be consistent or process all samples at the same time • Better technology never eliminate the needs for good experimental design 7
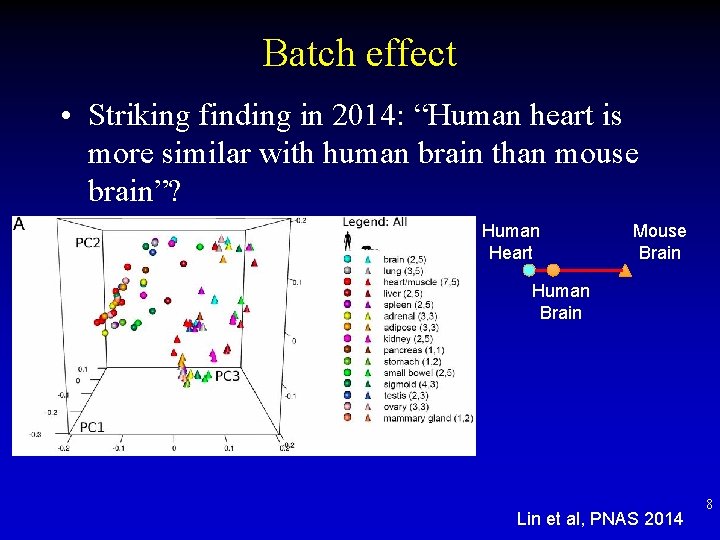
Batch effect • Striking finding in 2014: “Human heart is more similar with human brain than mouse brain”? Human Heart Mouse Brain Human Brain Lin et al, PNAS 2014 8

Batch Effect • • • 1 st batch: human tissues 2 nd batch: human tissues 3 rd batch: mouse tissues 4 th batch: mouse tissues 5 th batch: human/mouse tissues • After batch removal, tissues cluster Break 9

RNA-seq Analysis

Alignment • Prefer splice-aware aligners • Top. Hat, BWA, STAR… • Sometimes need to trim the beginning bases 11
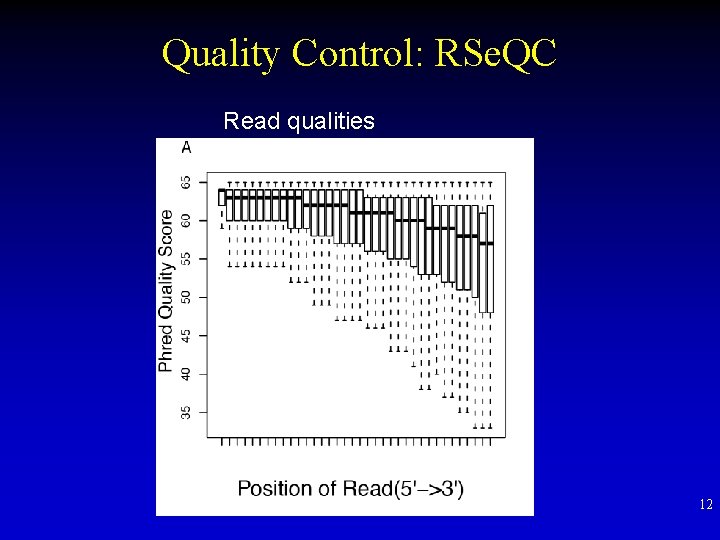
Quality Control: RSe. QC Read qualities 12
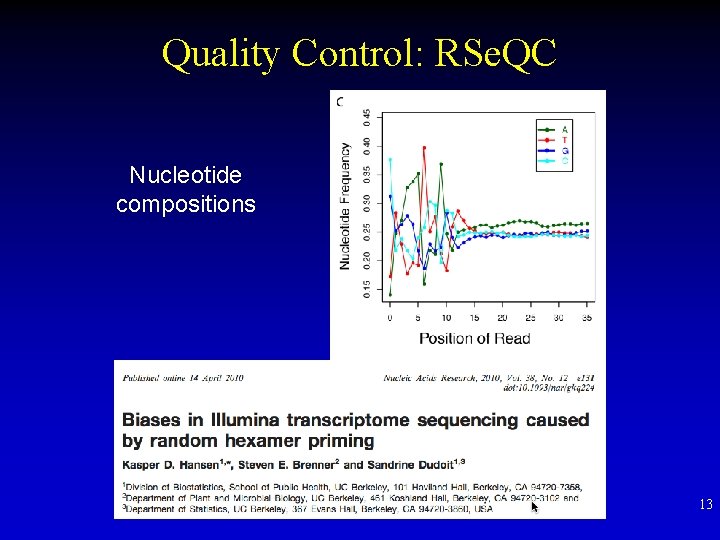
Quality Control: RSe. QC Nucleotide compositions 13
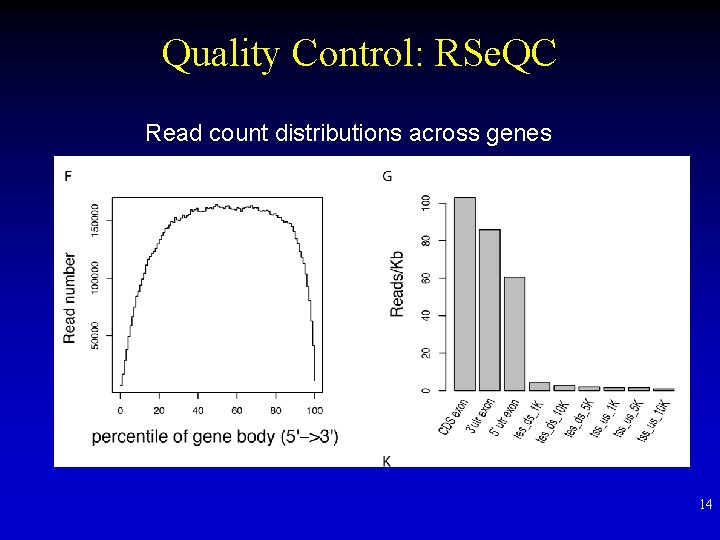
Quality Control: RSe. QC Read count distributions across genes 14
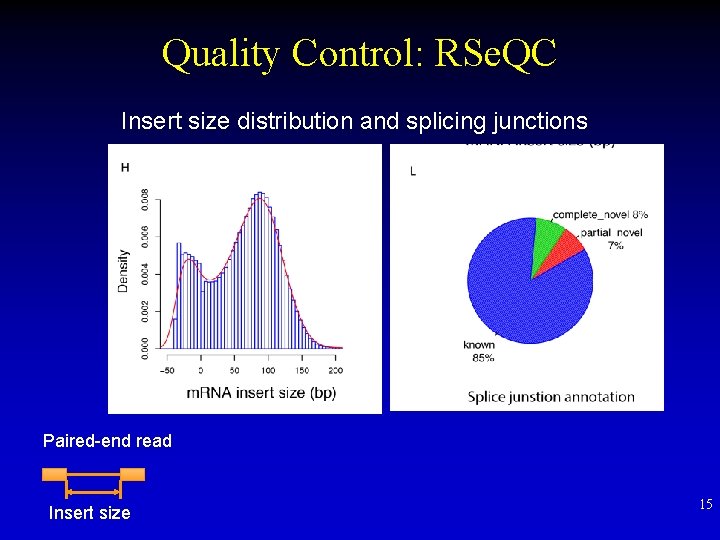
Quality Control: RSe. QC Insert size distribution and splicing junctions Paired-end read Insert size 15

Expression Index • RPKM (Reads per kilobase of transcript per million reads of library) – Total reads / 1 M, divide by gene length in KB – Corrects for coverage, gene length – Top. Hat / Cufflinks • FPKM (Fragments), PE libraries, ~RPKM/2 • TPM (transcripts per million) – Divide read count by gene length in KB (RPK), divide by scaling factor (sum of RKP across all genes / 1 M) – Proportion of reads mapped to a gene in each sample is comparable – RSEM 16
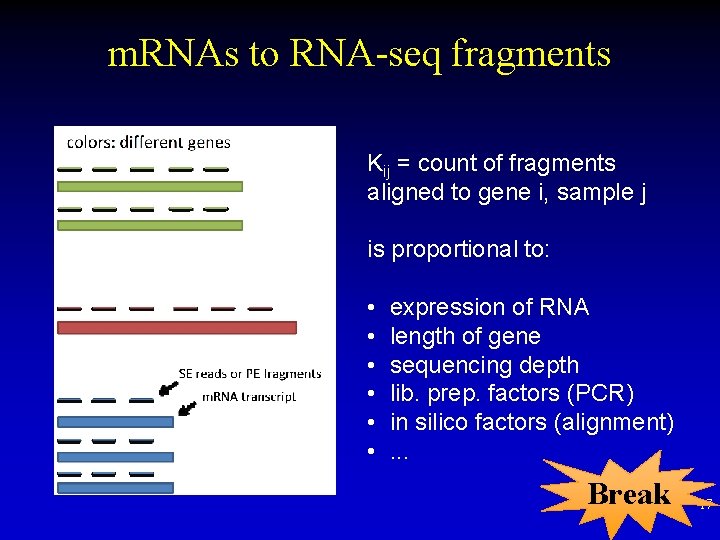
m. RNAs to RNA-seq fragments Kij = count of fragments aligned to gene i, sample j is proportional to: • • • expression of RNA length of gene sequencing depth lib. prep. factors (PCR) in silico factors (alignment). . . Break 17

Differential Expression

Statistical Power of Detecting Differential Expression • Range of count – Expression, Sequencing depth, Gene length Gene A (1 kb) Gene B (8 kb) • • • Sample size Dispersion True fold change Cufflinks: RPKM/FPKM LIMMA-VOOM and DESeq: Read count 19
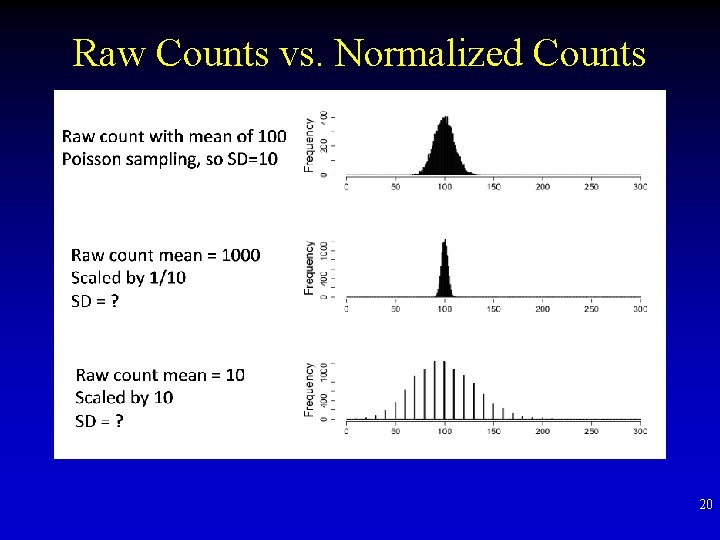
Raw Counts vs. Normalized Counts 20

Sequencing Read Distribution • The number of patients arriving in an emergency room between 10 and 11 pm • # Reads mapped to a gene of 1 KB long • Poisson dist – λ average events per interval – K # events in an interval – Var = mean = λ 21
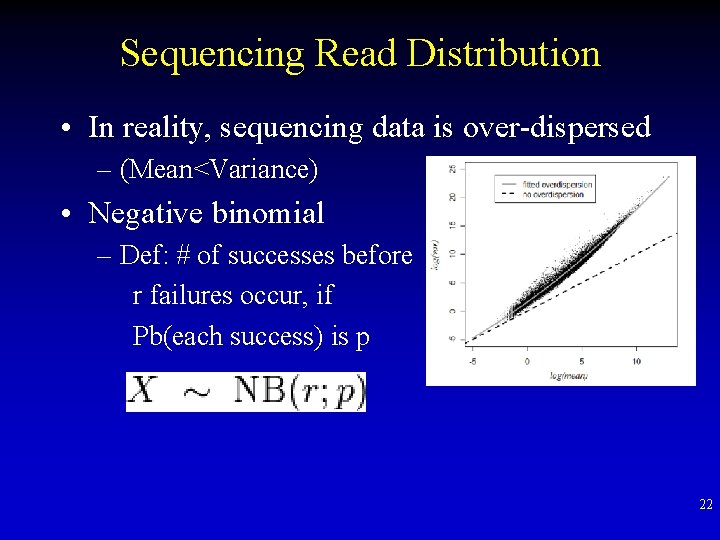
Sequencing Read Distribution • In reality, sequencing data is over-dispersed – (Mean<Variance) • Negative binomial – Def: # of successes before r failures occur, if Pb(each success) is p 22
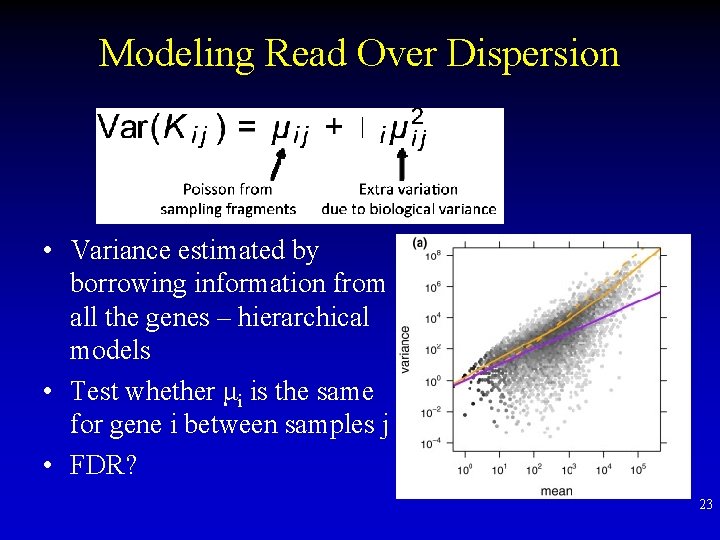
Modeling Read Over Dispersion • Variance estimated by borrowing information from all the genes – hierarchical models • Test whether μi is the same for gene i between samples j • FDR? 23
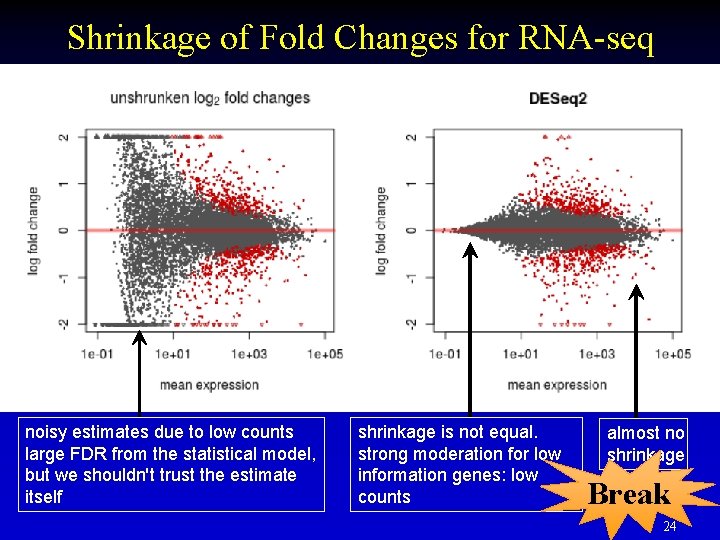
Shrinkage of Fold Changes for RNA-seq noisy estimates due to low counts large FDR from the statistical model, but we shouldn't trust the estimate itself shrinkage is not equal. strong moderation for low information genes: low counts almost no shrinkage Break 24

Splicing Transcripts • Assign reads to splice isoforms (Top. Hat) 25
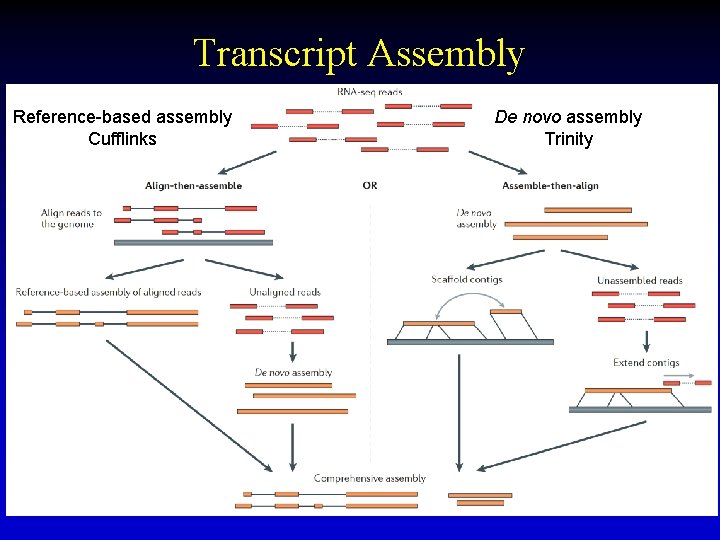
Transcript Assembly Reference-based assembly Cufflinks De novo assembly Trinity 26

Isoform Inference • If given known set of isoforms • Estimate x to maximize the likelihood of observing n 27
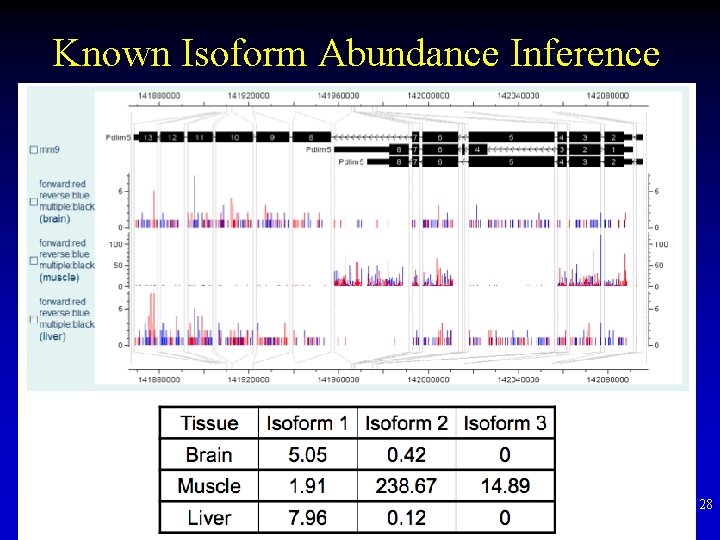
Known Isoform Abundance Inference 28

Identification of Differential Splicing Between RNA-seq Samples • Most differential splicing detection algorithms call differentially expressed exons, not whole transcripts, esp for novel splicing 29

Splicing Isoform Inference • With known isoform set, sometimes the gene-level expression level inference is great, although isoform abundances might have uncertainty (e. g. known set incomplete) • De novo method are usually better at detecting differential exon splicing, but not whole transcripts • De novo isoform inference is a non-identifiable problem if RNA-seq reads are short and gene is long with too many exons • Experimental validation of quantitative differential splicing is still quite hard 30

Active Field • HISAT 2 for fast alignment – Hierarchical index – https: //ccb. jhu. edu/software/hisat 2/index. shtml • Kallisto and Sleuth – Kallisto TPM, Sleuth differential expression – Known genes and transcripts – https: //scilifelab. github. io/courses/rnaseq/labs/k allisto 31

Summary • • RNA-seq design considerations Read mapping: BWA, STAR Quality control: RSe. QC Expression index: R/FPKM and TPM Differential expression: LIMMA-VOOM and DESeq Transcriptome assembly: Cufflinks, Trinity Alternative splicing: r/MATs New developments: HISAT 2, Kallisto and Sleuth 32

Acknowledgement • • • Wei Li Michael Love Alisha Holloway Simon Andrews Radhika Khetani 33
- Slides: 33