Histone Modifications Xiaole Shirley Liu STAT 115215 BIOBST

Histone Modifications Xiaole Shirley Liu STAT 115/215, BIO/BST 282
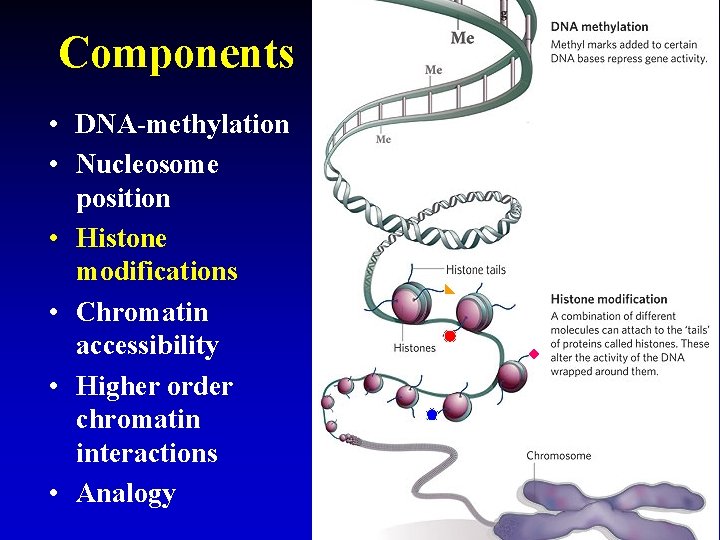
Components • DNA-methylation • Nucleosome position • Histone modifications • Chromatin accessibility • Higher order chromatin interactions • Analogy
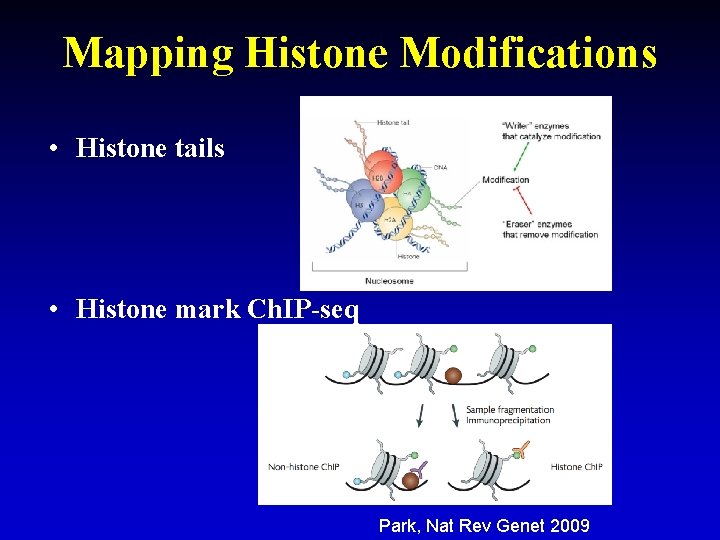
Mapping Histone Modifications • Histone tails • Histone mark Ch. IP-seq Park, Nat Rev Genet 2009

Histone Modifications • Different modifications at different locations by different enzymes
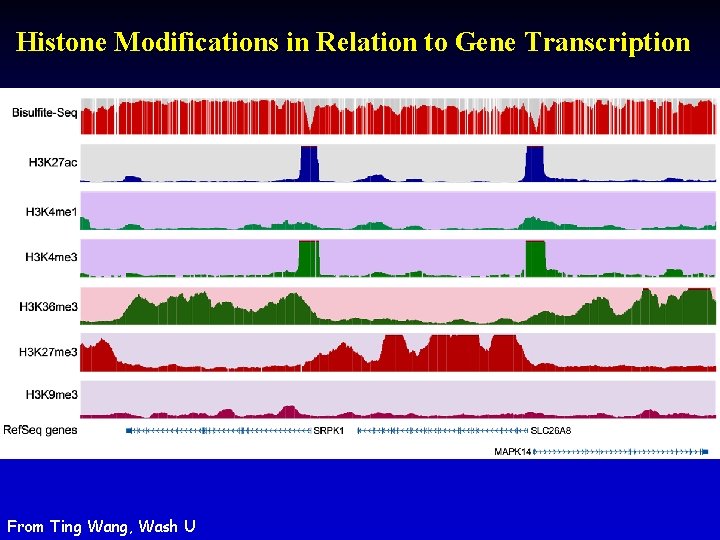
Histone Modifications in Relation to Gene Transcription From Ting Wang, Wash U

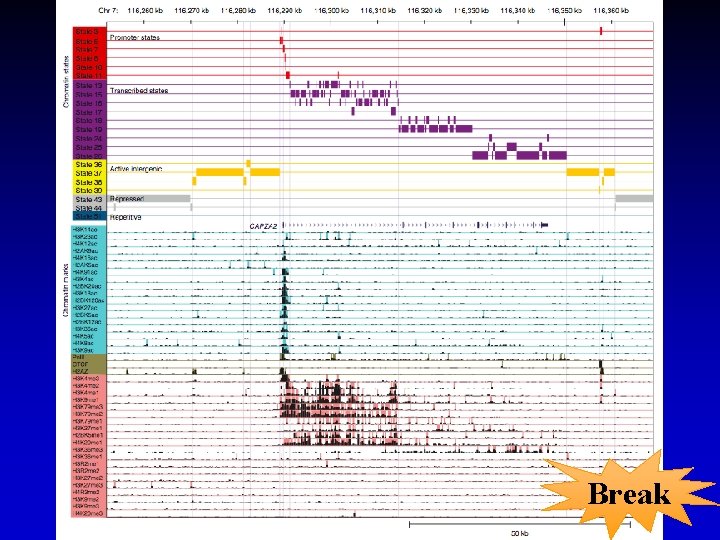
Histone Modifications • Gene body mark: H 3 K 36 me 3, H 3 K 79 me 3 • Active promoter (TSS) mark: H 3 K 4 me 3 • Active enhancer (TF binding) mark: H 3 K 4 me 1, H 3 K 27 ac • Both enhancers and promoters: H 3 K 4 me 2, H 3/H 4 ac, H 2 AZ • Repressive mark: H 3 K 27 me 3, H 3 K 9 me 3 Annotate / segment the genome based on histone marks
7
Break 8
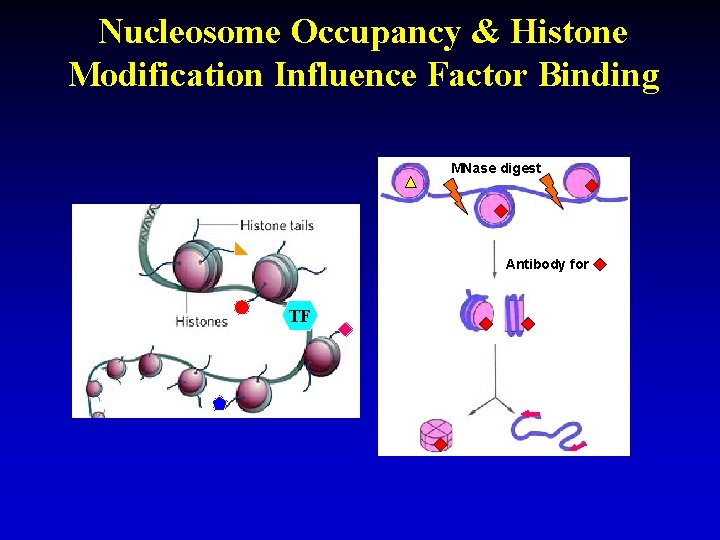
Nucleosome Occupancy & Histone Modification Influence Factor Binding MNase digest Antibody for TF

Combine Tags From All Ch. IP-Seq
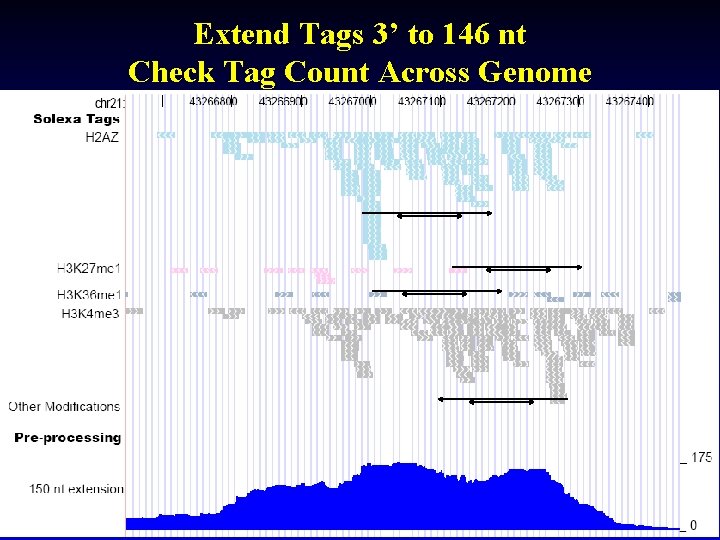
Extend Tags 3’ to 146 nt Check Tag Count Across Genome
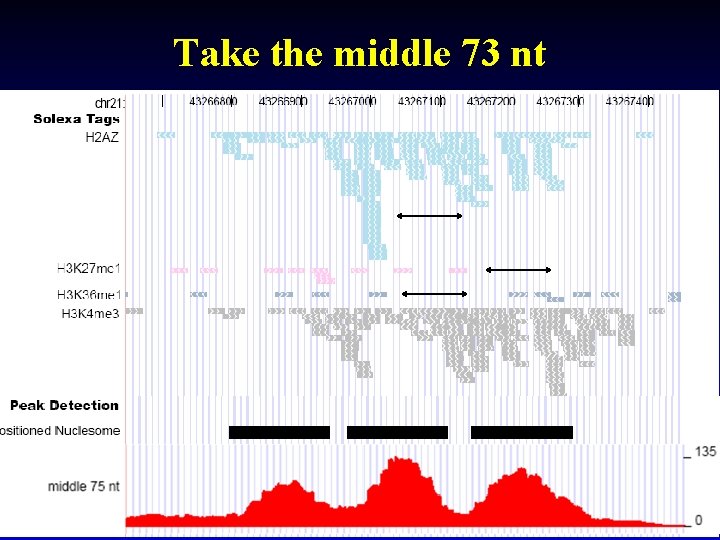
Take the middle 73 nt
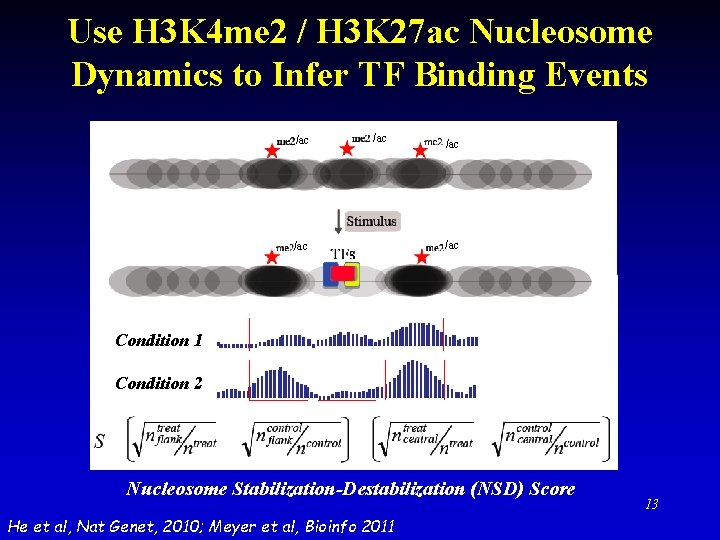
Use H 3 K 4 me 2 / H 3 K 27 ac Nucleosome Dynamics to Infer TF Binding Events /ac /ac /ac Condition 1 Condition 2 Nucleosome Stabilization-Destabilization (NSD) Score He et al, Nat Genet, 2010; Meyer et al, Bioinfo 2011 13

Condition-Specific Binding, Epigenetics and Gene Expression C 1 C 2 • Condition-specific TF bindings are associated with epigenetic signatures • Can we use the epigenetic profile and TF motif analysis to simultaneous guess the binding of many TFs together? Genes TF 1 TF 2 TF 3 Epigenetics 14
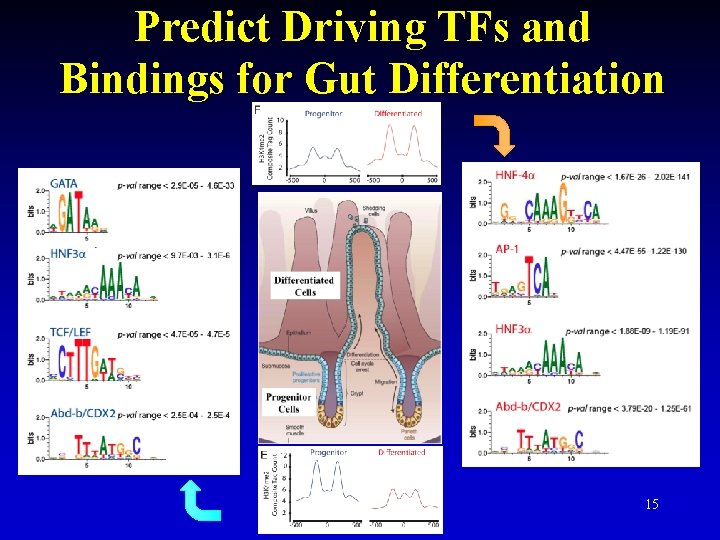
Predict Driving TFs and Bindings for Gut Differentiation 15
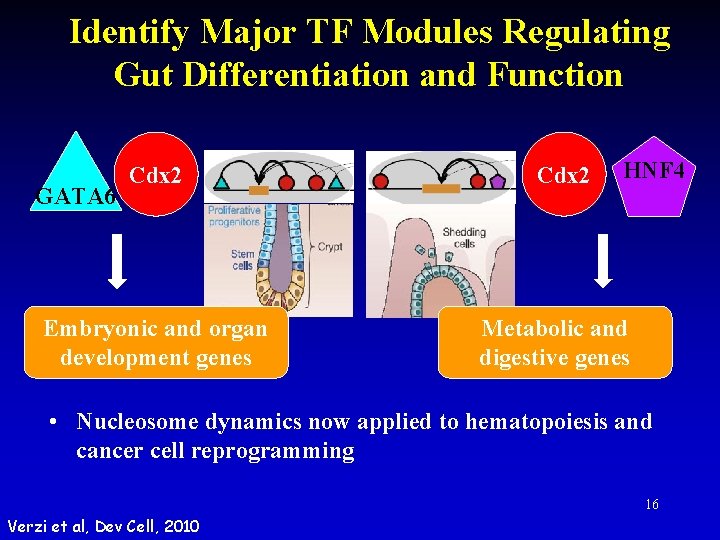
Identify Major TF Modules Regulating Gut Differentiation and Function GATA 6 Cdx 2 Embryonic and organ development genes Cdx 2 HNF 4 Metabolic and digestive genes • Nucleosome dynamics now applied to hematopoiesis and cancer cell reprogramming 16 Verzi et al, Dev Cell, 2010
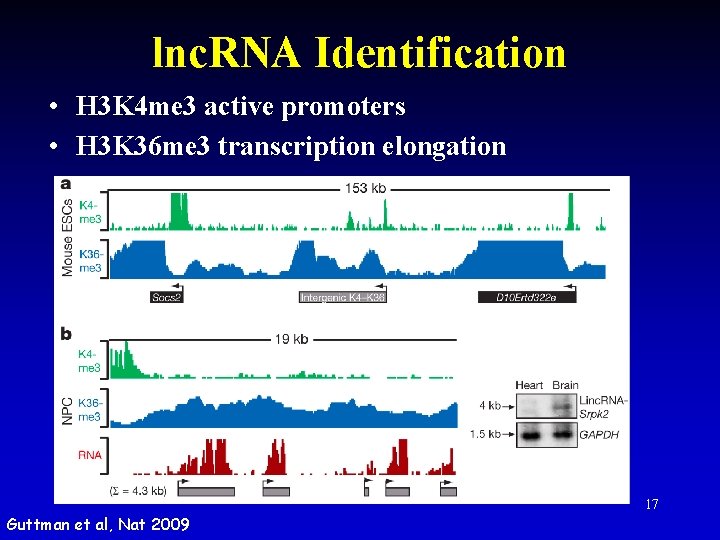
lnc. RNA Identification • H 3 K 4 me 3 active promoters • H 3 K 36 me 3 transcription elongation Guttman et al, Nat 2009 17
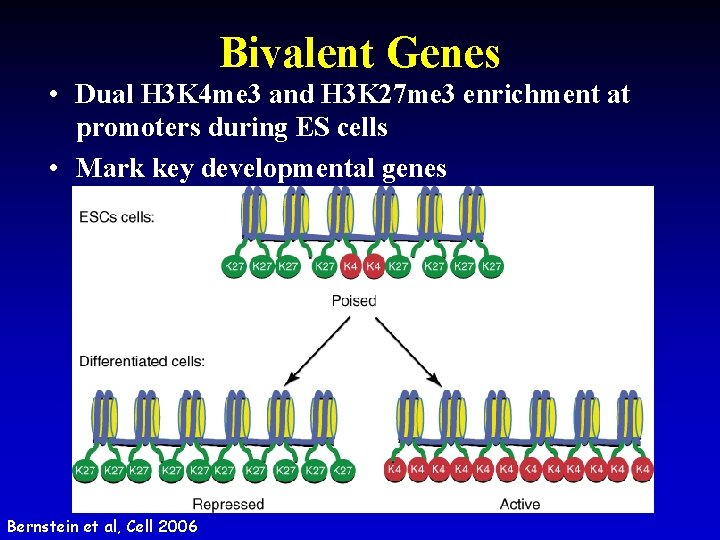
Bivalent Genes • Dual H 3 K 4 me 3 and H 3 K 27 me 3 enrichment at promoters during ES cells • Mark key developmental genes Bernstein et al, Cell 2006
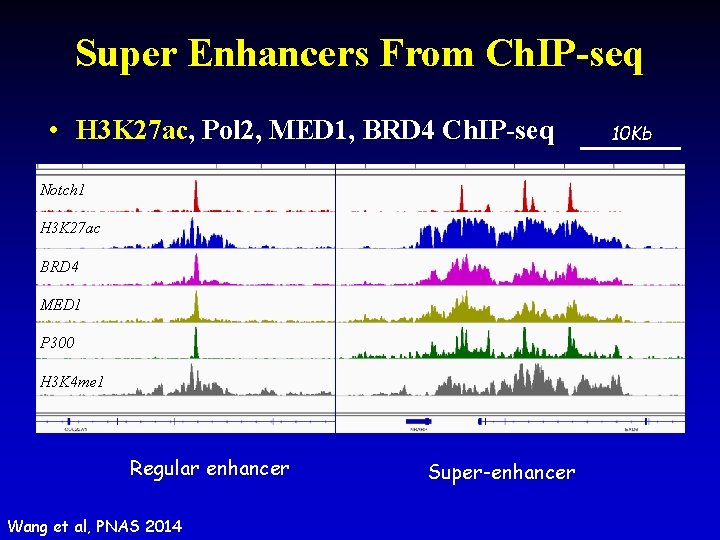
Super Enhancers From Ch. IP-seq • H 3 K 27 ac, Pol 2, MED 1, BRD 4 Ch. IP-seq Notch 1 H 3 K 27 ac BRD 4 MED 1 P 300 H 3 K 4 me 1 Regular enhancer Wang et al, PNAS 2014 Super-enhancer 10 Kb
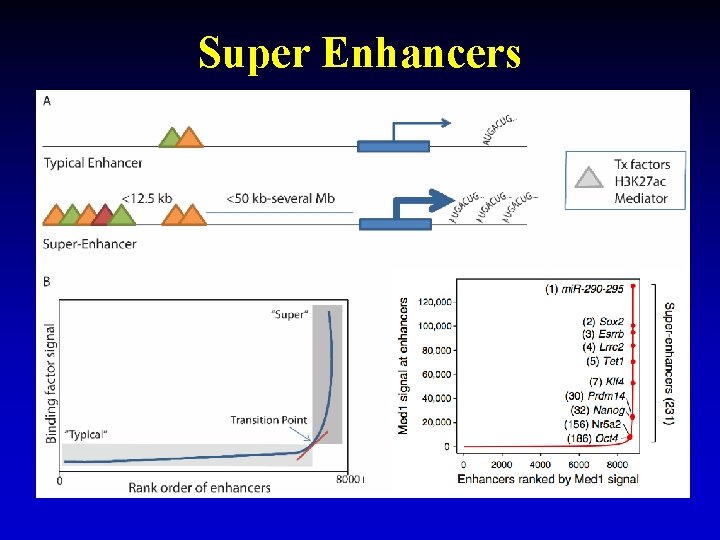
Super Enhancers

Super Enhancers at Key Genes 21

BET Inhibitor JQ 1 Targets Super-Enhancer Target Genes 22
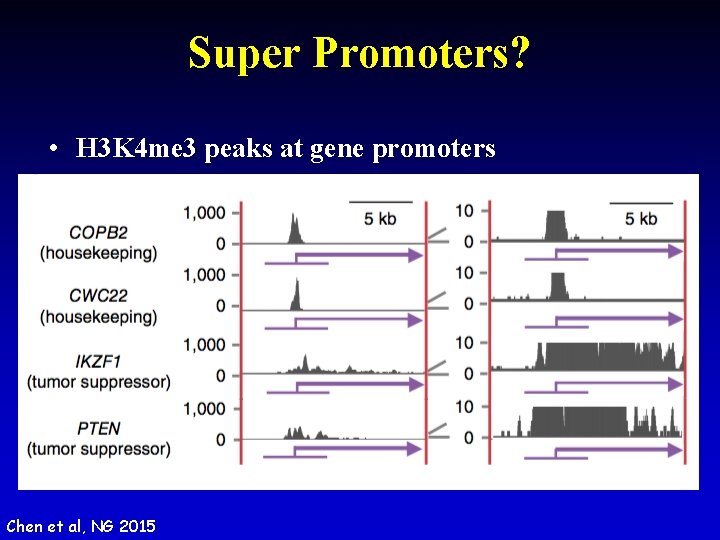
Super Promoters? • H 3 K 4 me 3 peaks at gene promoters Chen et al, NG 2015
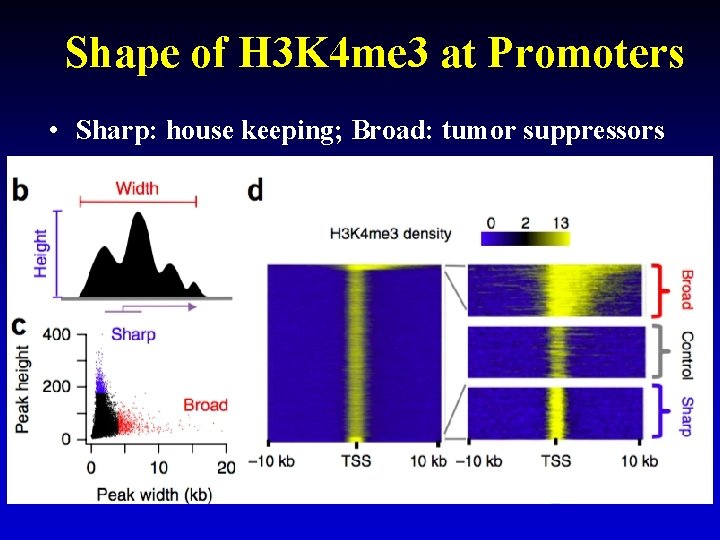
Shape of H 3 K 4 me 3 at Promoters • Sharp: house keeping; Broad: tumor suppressors
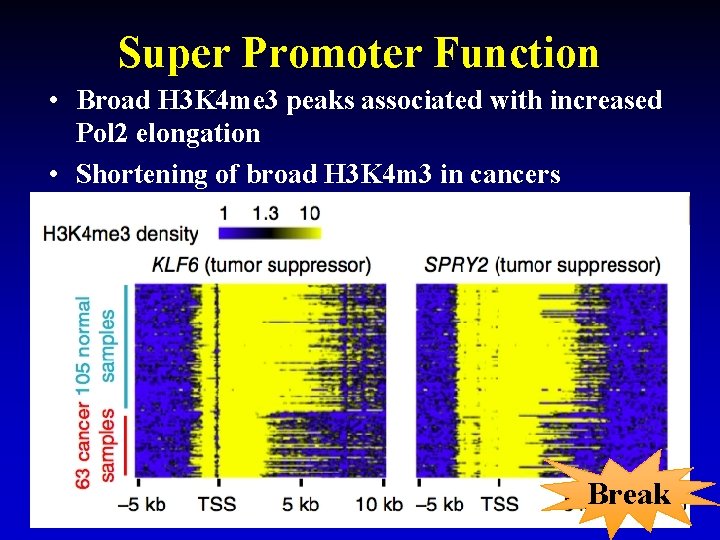
Super Promoter Function • Broad H 3 K 4 me 3 peaks associated with increased Pol 2 elongation • Shortening of broad H 3 K 4 m 3 in cancers Break
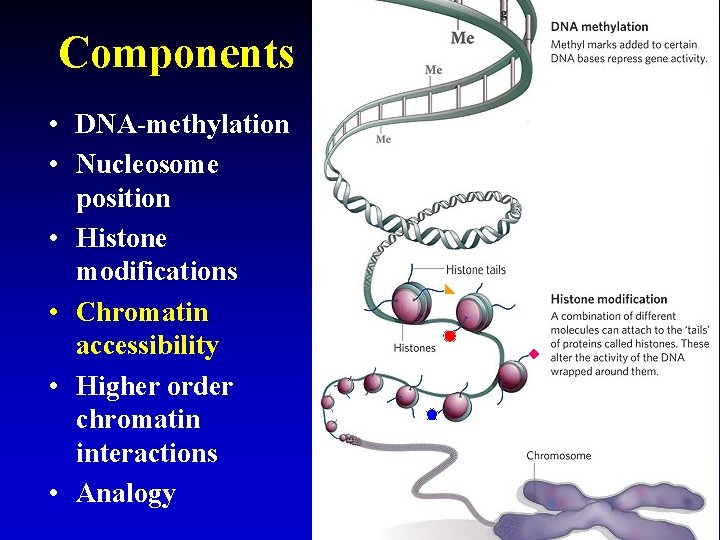
Components • DNA-methylation • Nucleosome position • Histone modifications • Chromatin accessibility • Higher order chromatin interactions • Analogy
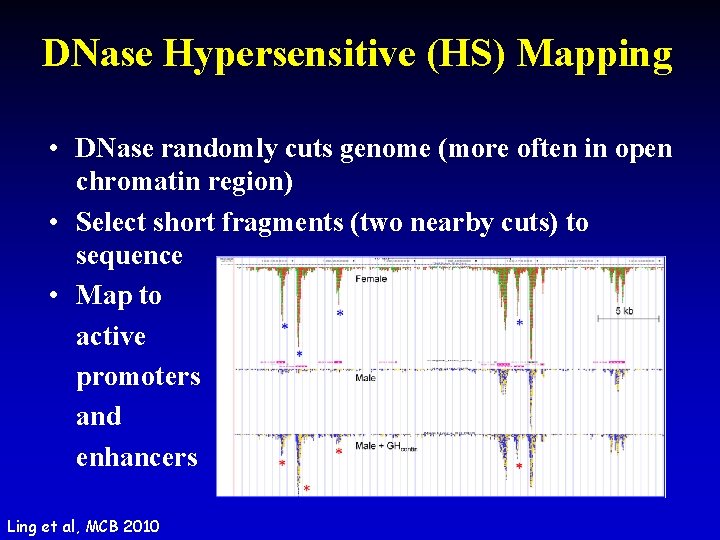
DNase Hypersensitive (HS) Mapping • DNase randomly cuts genome (more often in open chromatin region) • Select short fragments (two nearby cuts) to sequence • Map to active promoters and enhancers Ling et al, MCB 2010
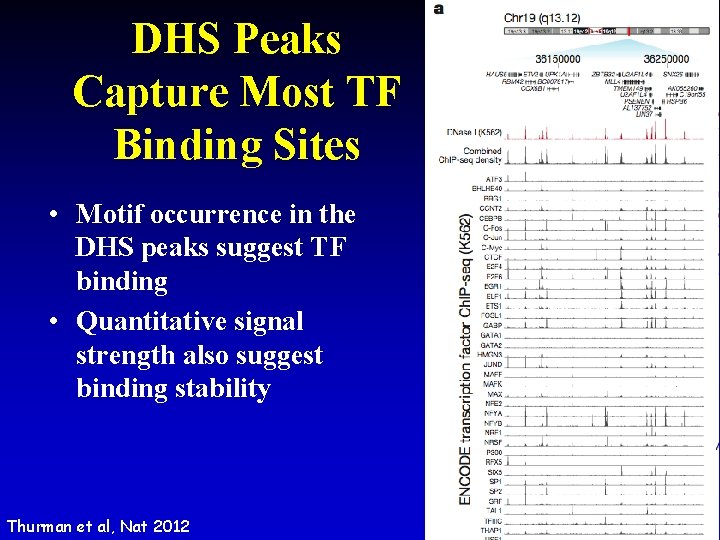
DHS Peaks Capture Most TF Binding Sites • Motif occurrence in the DHS peaks suggest TF binding • Quantitative signal strength also suggest binding stability Thurman et al, Nat 2012

TF Network from DNase Footprint 29
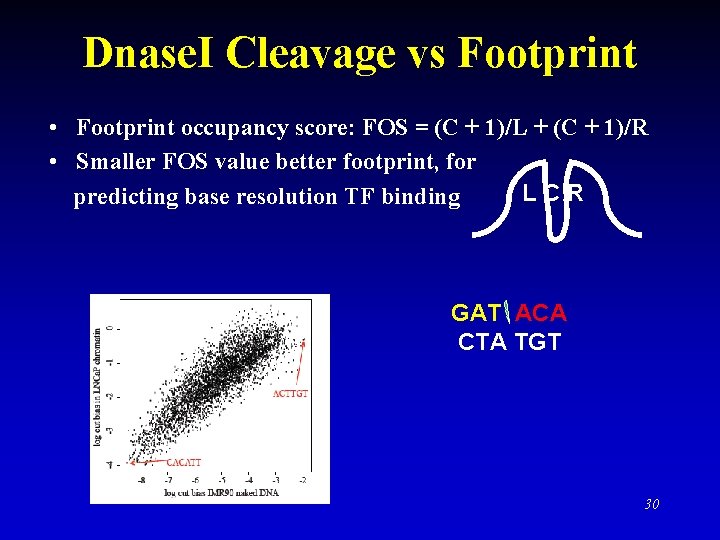
Dnase. I Cleavage vs Footprint • Footprint occupancy score: FOS = (C + 1)/L + (C + 1)/R • Smaller FOS value better footprint, for LCR predicting base resolution TF binding GAT ACA CTA TGT 30
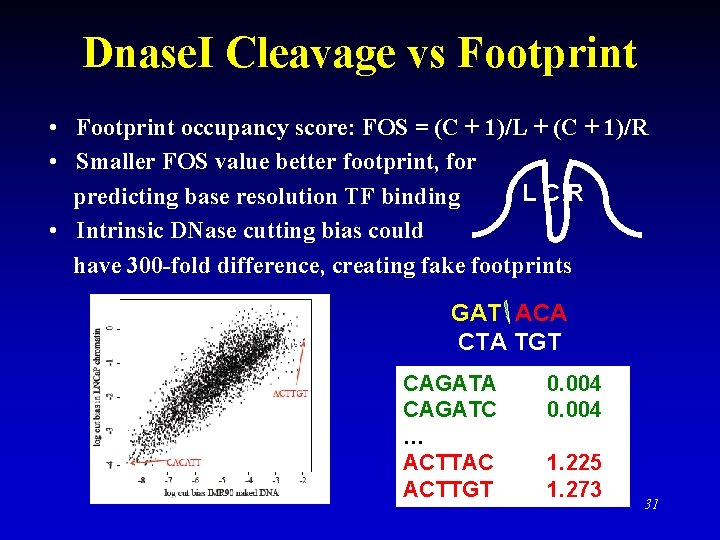
Dnase. I Cleavage vs Footprint • Footprint occupancy score: FOS = (C + 1)/L + (C + 1)/R • Smaller FOS value better footprint, for LCR predicting base resolution TF binding • Intrinsic DNase cutting bias could have 300 -fold difference, creating fake footprints GAT ACA CTA TGT CAGATA CAGATC … ACTTAC ACTTGT 0. 004 1. 225 1. 273 31
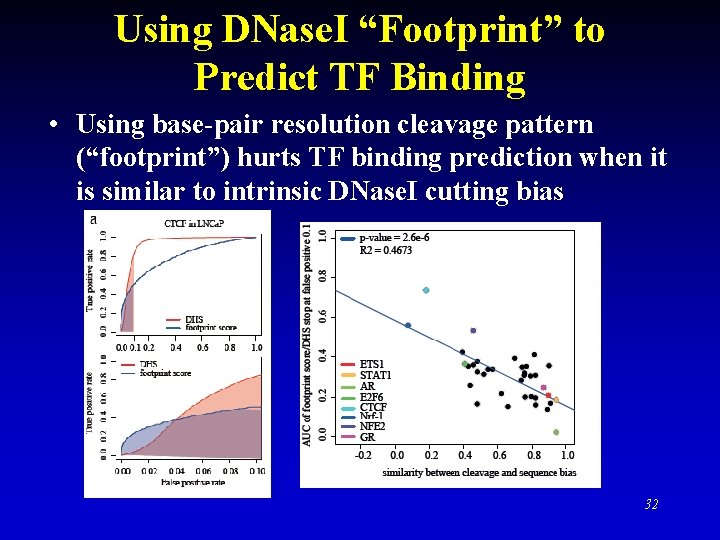
Using DNase. I “Footprint” to Predict TF Binding • Using base-pair resolution cleavage pattern (“footprint”) hurts TF binding prediction when it is similar to intrinsic DNase. I cutting bias 32
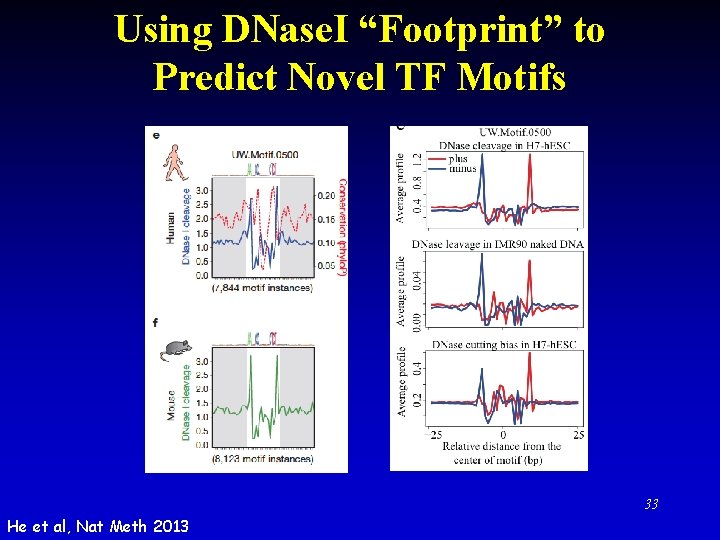
Using DNase. I “Footprint” to Predict Novel TF Motifs 33 He et al, Nat Meth 2013
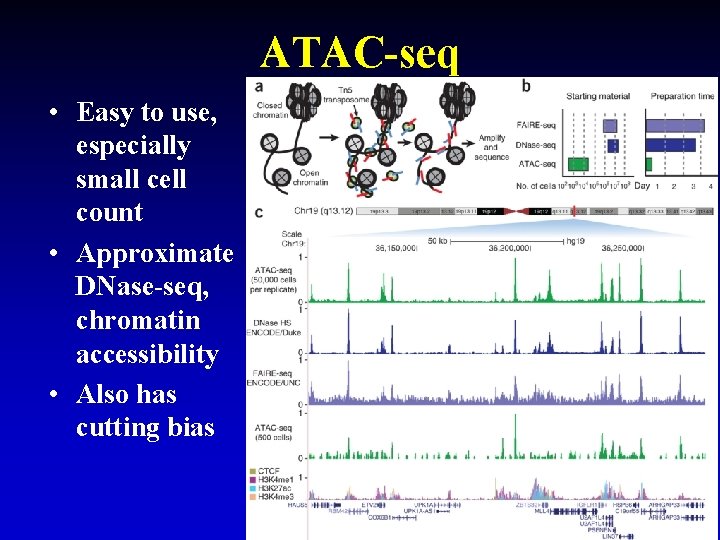
ATAC-seq • Easy to use, especially small cell count • Approximate DNase-seq, chromatin accessibility • Also has cutting bias
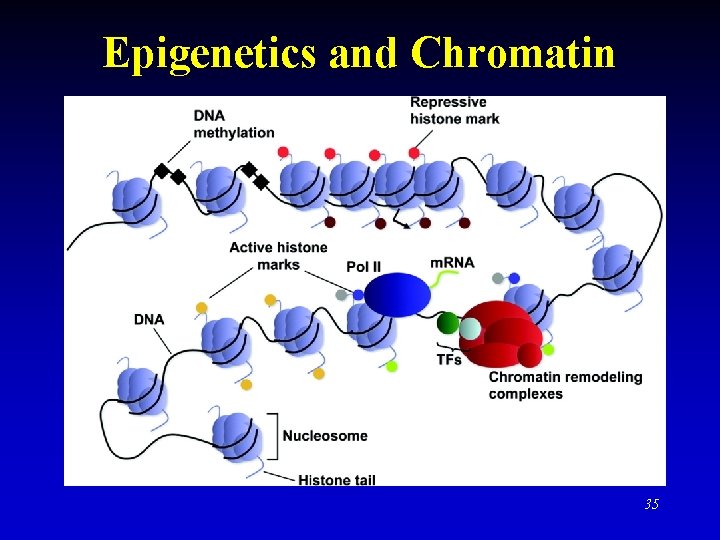
Epigenetics and Chromatin 35

Transcription and Epigenetic Regulation • Stem cell differentiation • Aging brain • Cancer

Histone Marks and Chromatin Accessibility • Epigenetic beacons in the genome • Histone marks – – Annotate functional elements in the genome Nucleosome positions and chromatin dynamics lnc. RNAs, bivalent genes Super enhancer and super promoter targets • Chromatin accessibility – Collection of TF binding sites – Caution of modeling “footprint”

Clarifications During Office Hour • DNA: naked, double helix • Chromatin: DNA in its compaction structure in the cells • Nucleosome: basic unit of chromatin, 146 bp DNA wrapped around 8 histones proteins • Regular TF Ch. IP-seq: sonication, antibody against TF • Histone mark Ch. IP-seq: sonication or MNase, antibody against the histone modification
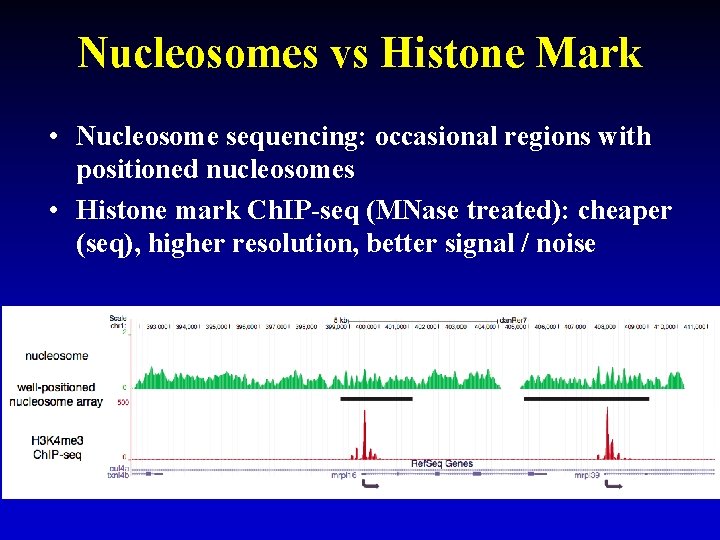
Nucleosomes vs Histone Mark • Nucleosome sequencing: occasional regions with positioned nucleosomes • Histone mark Ch. IP-seq (MNase treated): cheaper (seq), higher resolution, better signal / noise
- Slides: 39