Regulation of Gene Expression David Shiuan Department of


























- Slides: 102

Regulation of Gene Expression David Shiuan Department of Life Science Institute of Biotechnology Interdisciplinary Program of Bioinformatics National Dong Hwa University

The fundamental problem of chemical physiology and of embryology --- is to understand why tissue cells do not all express, all the time, all the potentialities inherent in their genome. Francois Jacob and Jacques Monod J. Mol. Biol. 1961

• 1. Principle of gene regulation • 2. Regulation of gene expression in prokaryotes • 3. Regulation of gene expression in eukaryotes

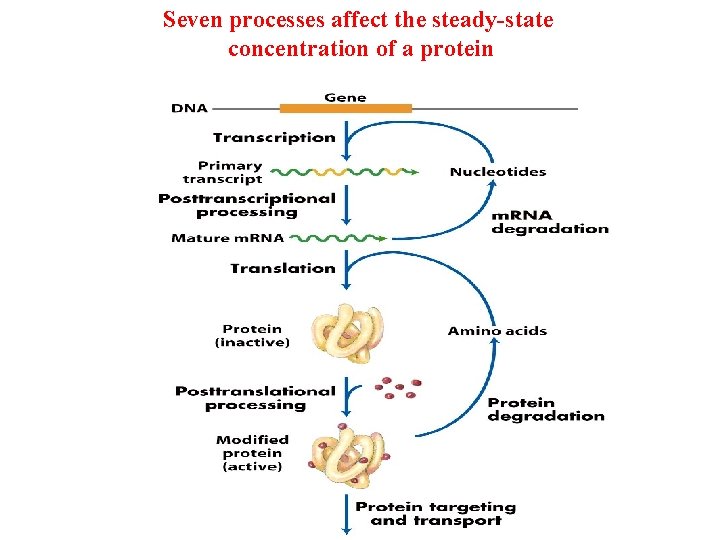
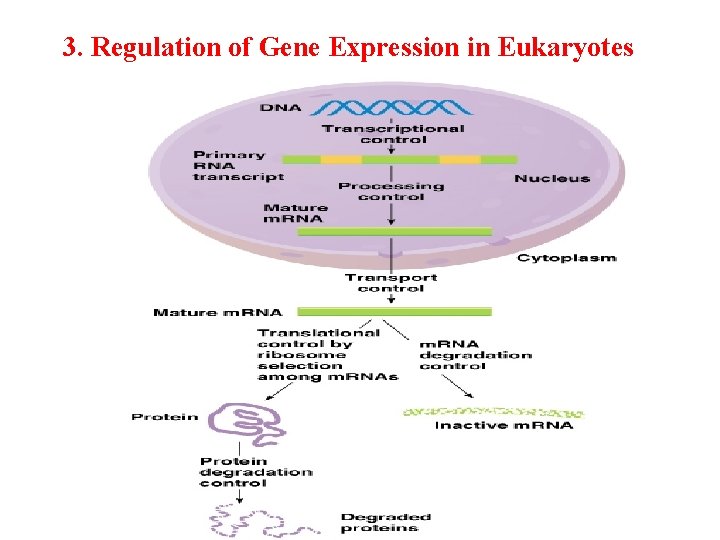
Seven processes affect the steady-state concentration of a protein

Potential Points of Regulation • Synthesis of primary RNA transcript (transcription) • Posttranscriptional modification of m. RNA • m. RNA degradation • Protein synthesis (translation) • Posttranslational modification of proteins • Protein targeting and transport • Protein degradation

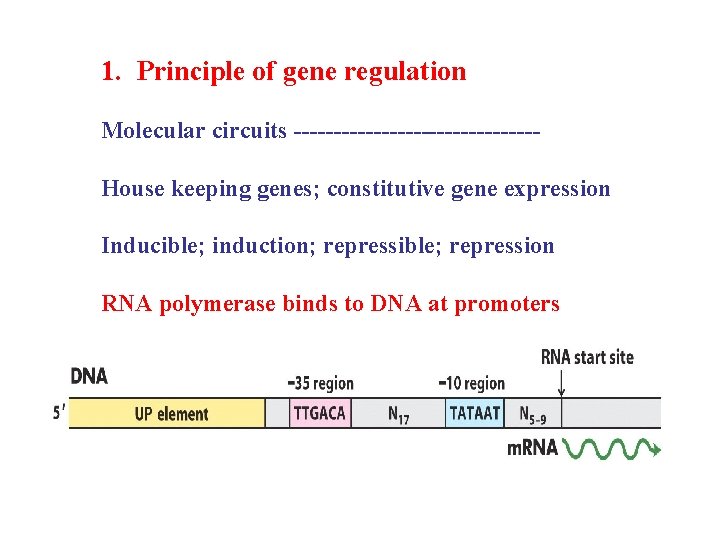
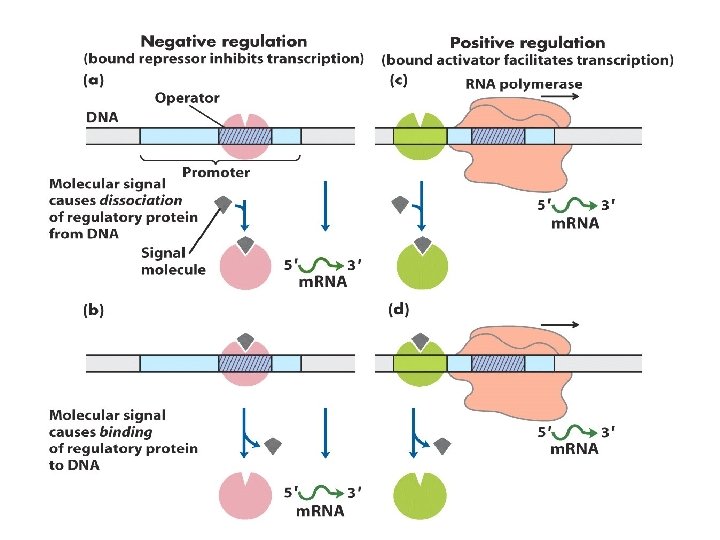
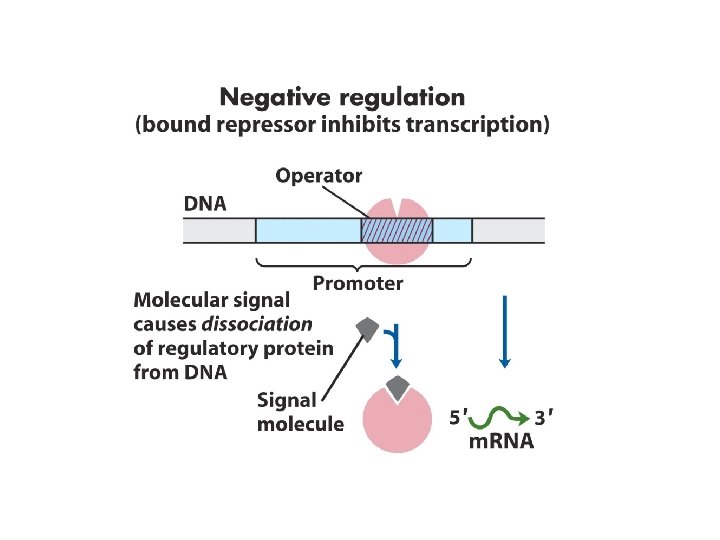
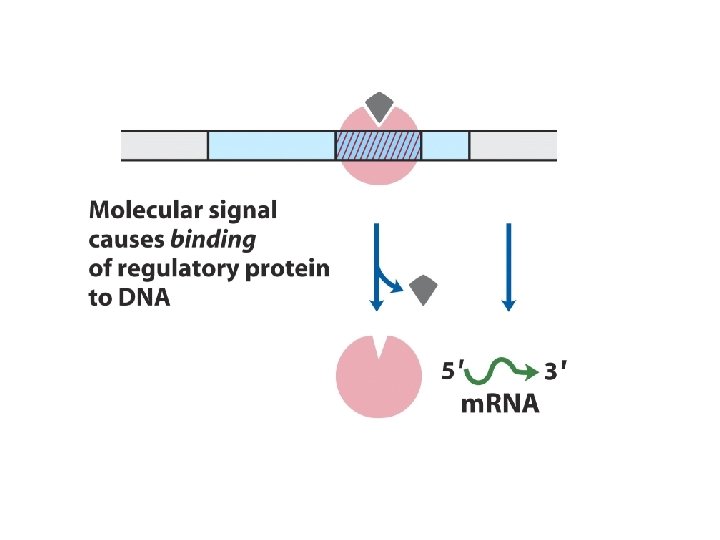
1. Principle of gene regulation Molecular circuits ---------------House keeping genes; constitutive gene expression Inducible; induction; repressible; repression RNA polymerase binds to DNA at promoters

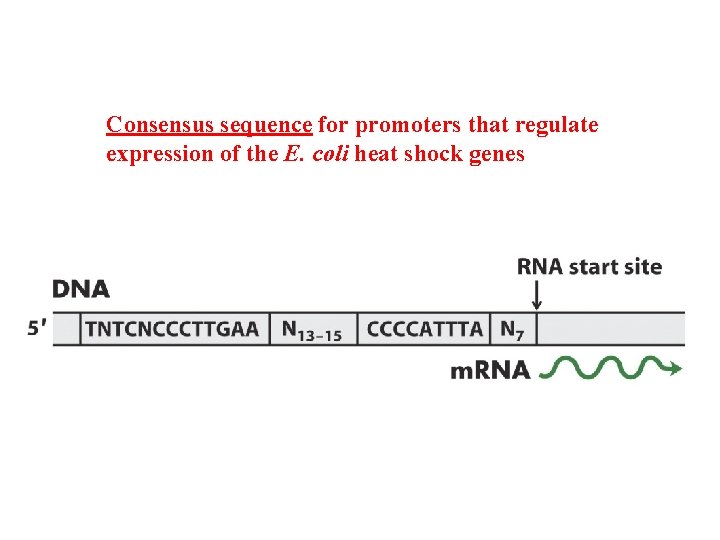
Consensus sequence for promoters that regulate expression of the E. coli heat shock genes






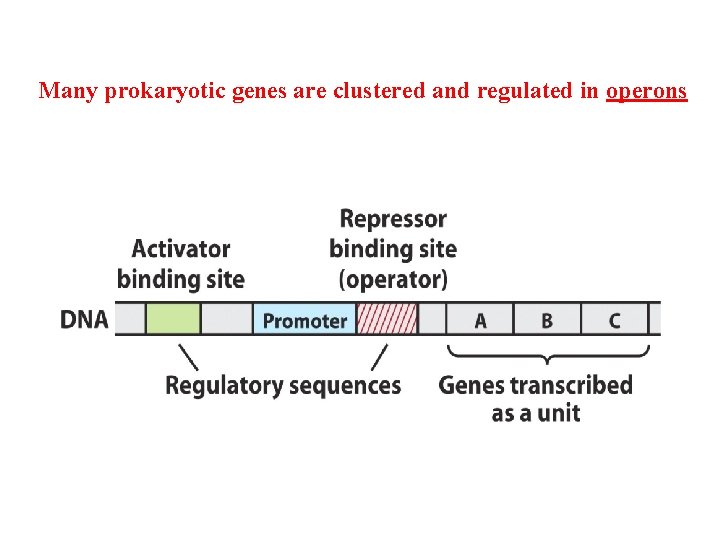
Many prokaryotic genes are clustered and regulated in operons

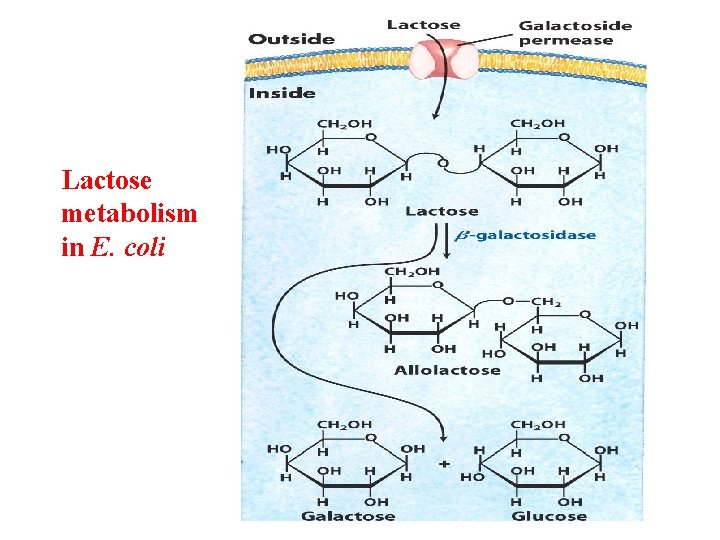
Lactose metabolism in E. coli

They published a paper - Coordinated regulation of lac operon, Proc. French Acad. Sci. (1960)

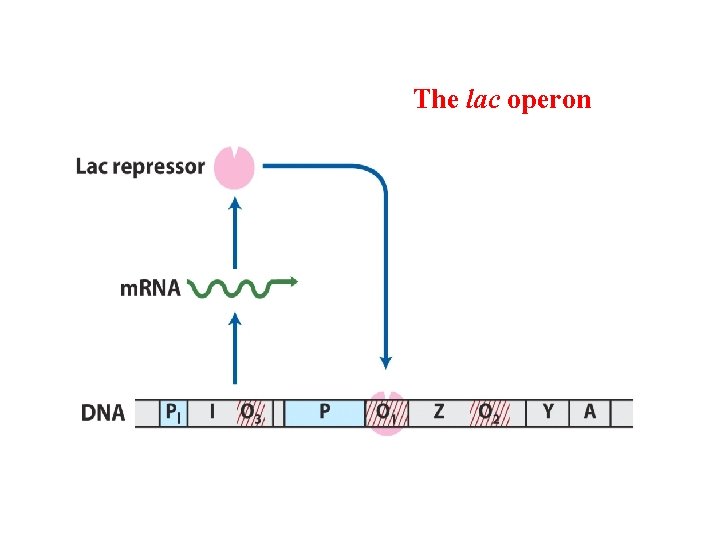
The lac operon

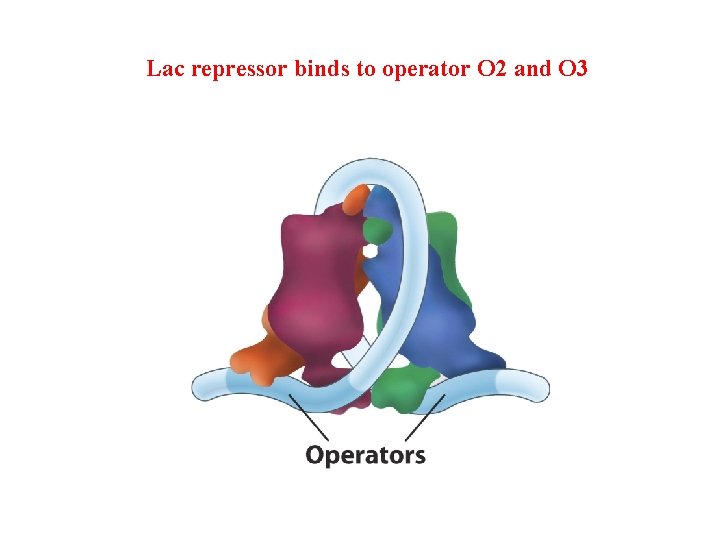
Lac repressor binds to operator O 2 and O 3

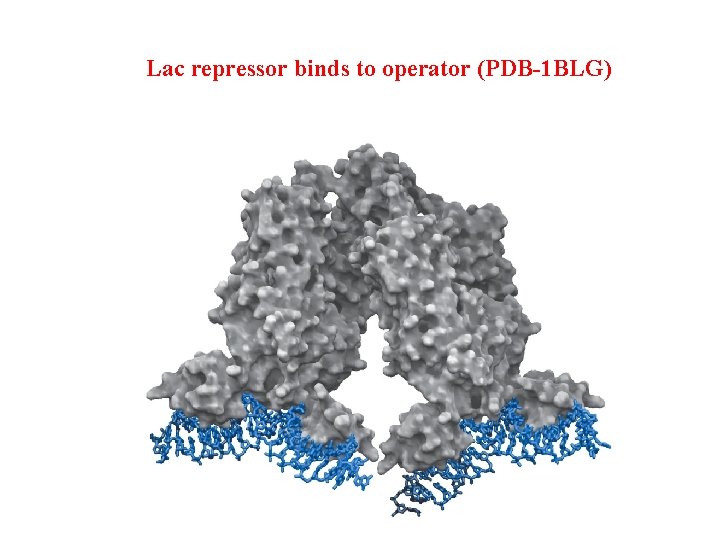
Lac repressor binds to operator (PDB-1 BLG)

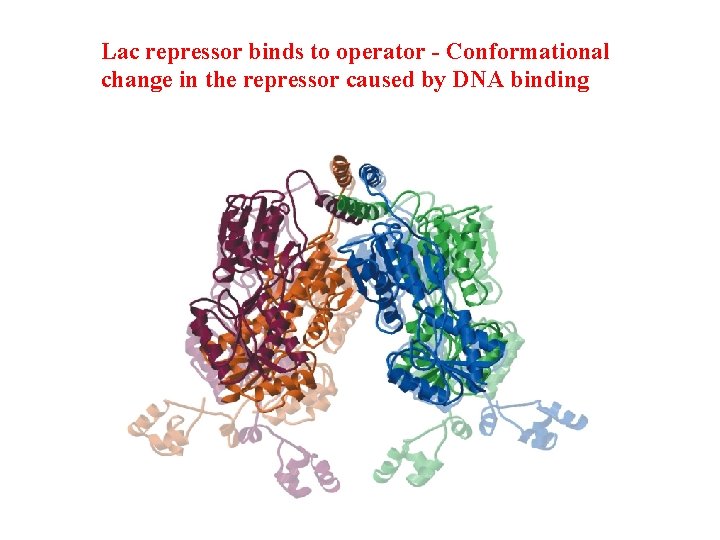
Lac repressor binds to operator - Conformational change in the repressor caused by DNA binding

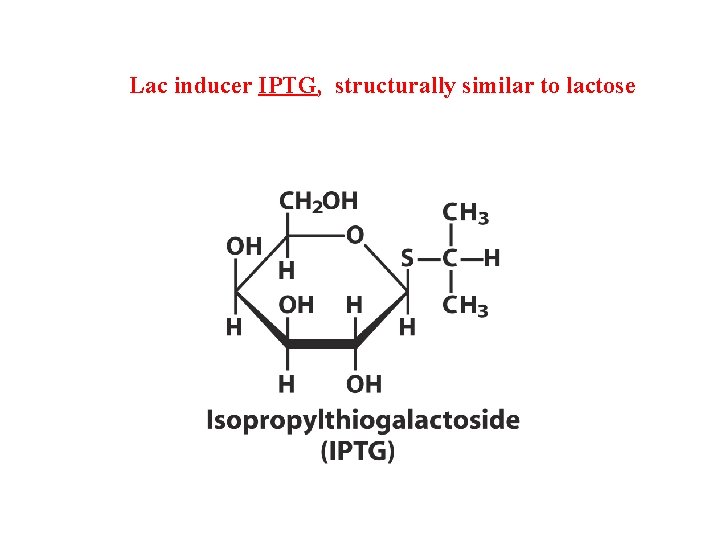
Lac inducer IPTG, structurally similar to lactose

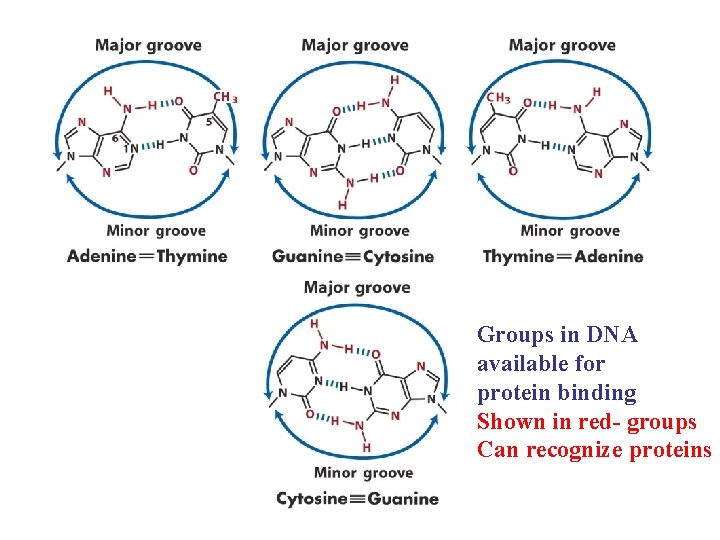
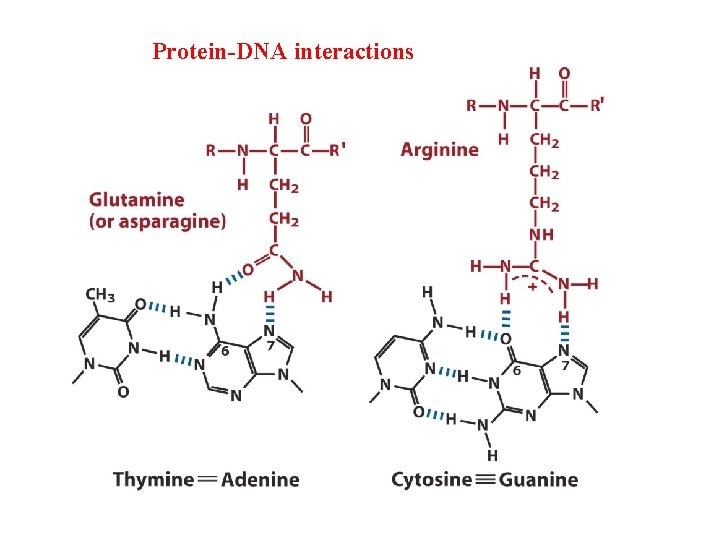
Groups in DNA available for protein binding Shown in red- groups Can recognize proteins

Protein-DNA interactions

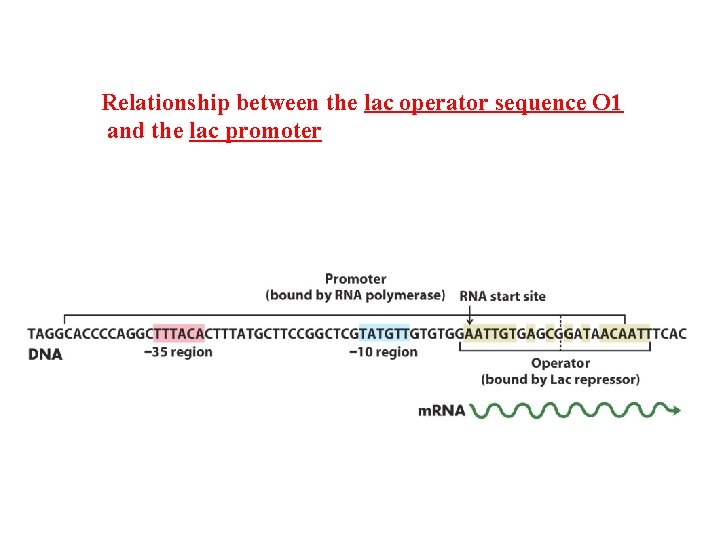
Relationship between the lac operator sequence O 1 and the lac promoter

DNA Binding Domain of Lac Repressor - Helix-turn-helix

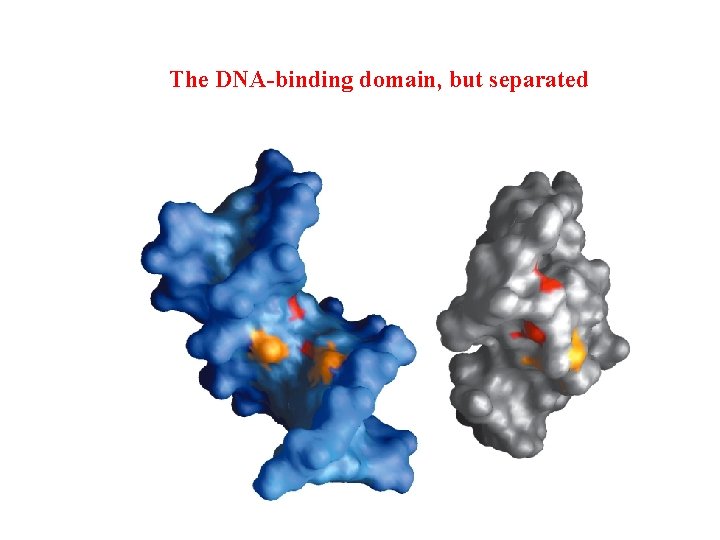
Surface rendering of the DNA-binding domain gray - lac repressor; blue - DNA

The DNA-binding domain, but separated

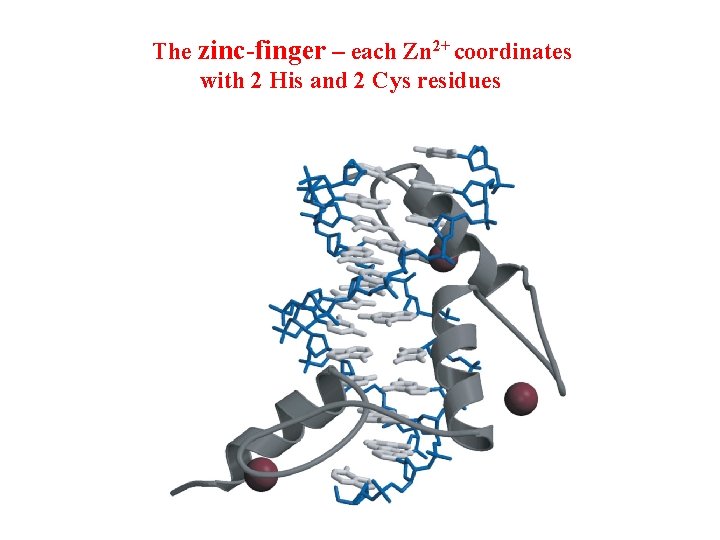
The zinc-finger – each Zn 2+ coordinates with 2 His and 2 Cys residues

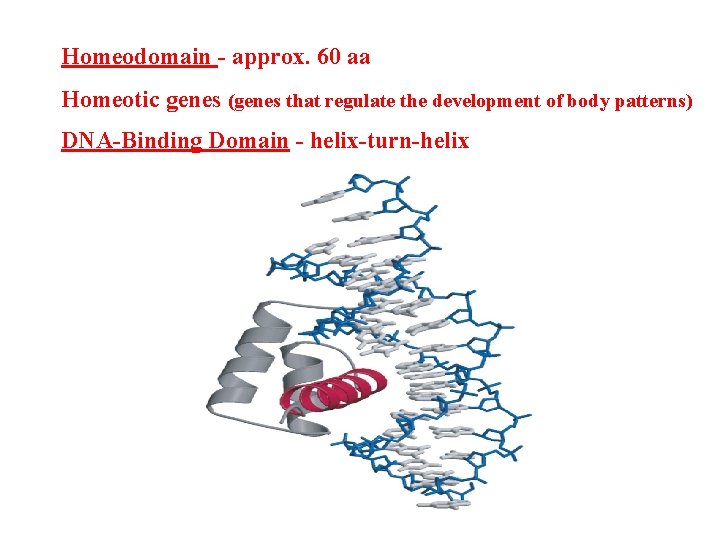
Homeodomain - approx. 60 aa Homeotic genes (genes that regulate the development of body patterns) DNA-Binding Domain - helix-turn-helix

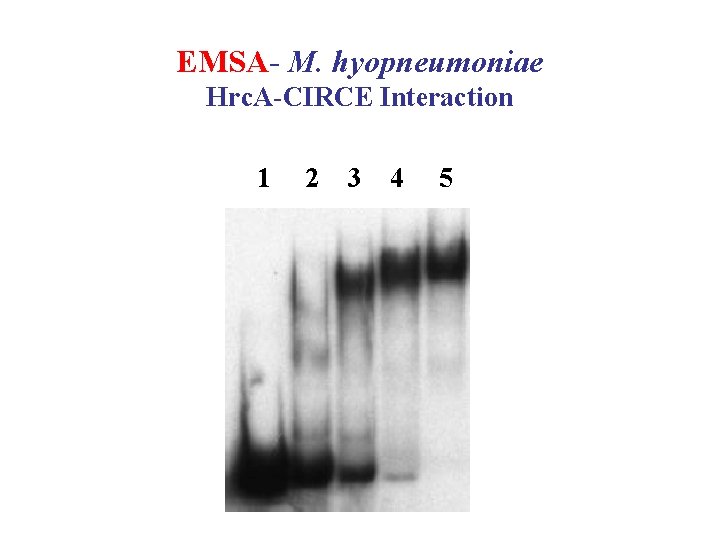
Studying DNA-Protein Interactions • EMSA (electrophoretic mobility shift assay); or gel retardation assay • DNase. I footprinting experiment • DNA affinity chromatography • SPR (Surface Plasmon Resonance)/BIACORE • CD/ORD; Spefctrofluorometry; NMR

EMSA- M. hyopneumoniae Hrc. A-CIRCE Interaction 1 2 3 4 5

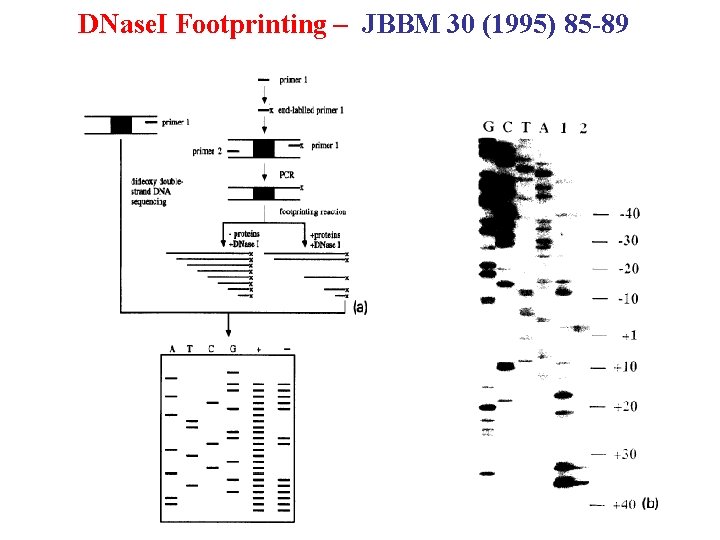
DNase. I Footprinting – JBBM 30 (1995) 85 -89

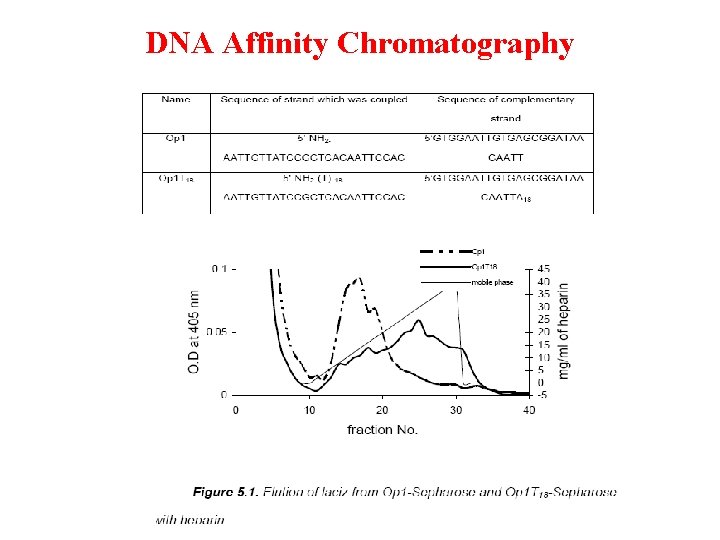
DNA Affinity Chromatography

Surface Plasmon Resonance (SPR) • SPR - Surface plasmon resonance is a phenomenon which occurs when light is reflected off thin metal films. A fraction of the light energy incident at a sharply defined angle can interact with the delocalized electrons in the metal film (plasmon) thus reducing the reflected light intensity


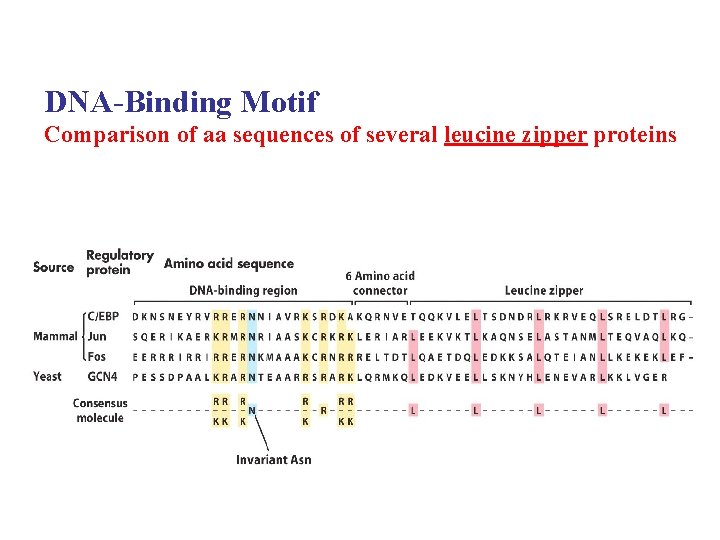
DNA-Binding Motif Comparison of aa sequences of several leucine zipper proteins

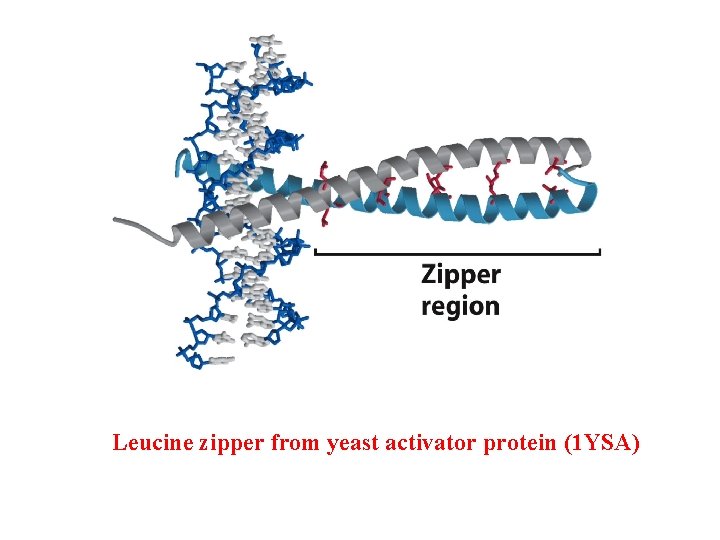
Leucine zipper from yeast activator protein (1 YSA)

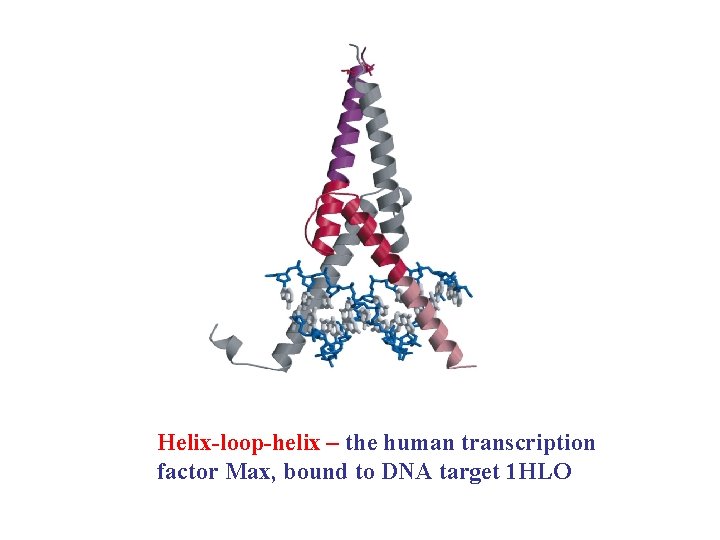
Helix-loop-helix – the human transcription factor Max, bound to DNA target 1 HLO

2. Regulation of Gene Expression in Prokaryotes

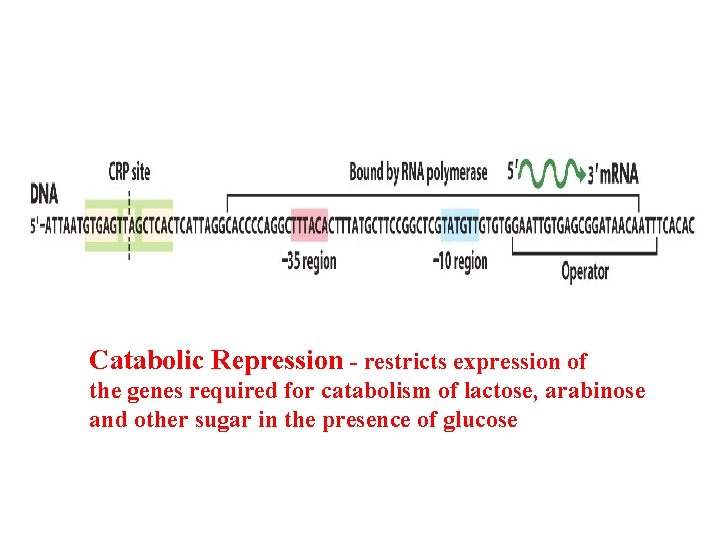
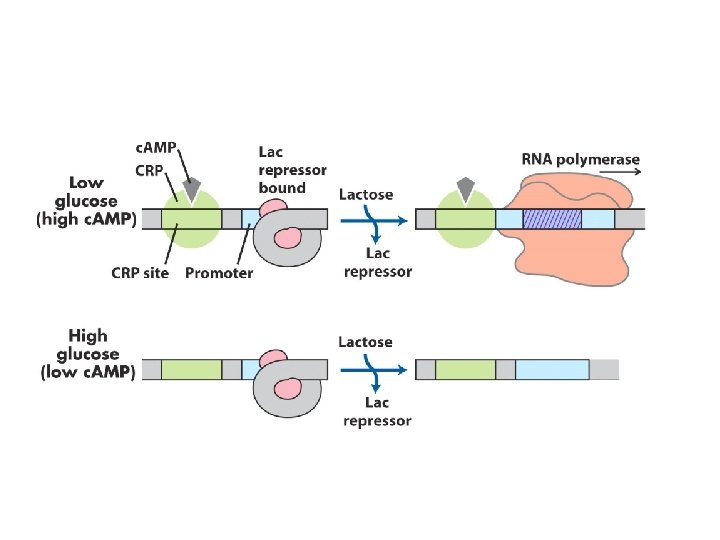
Catabolic Repression - restricts expression of the genes required for catabolism of lactose, arabinose and other sugar in the presence of glucose

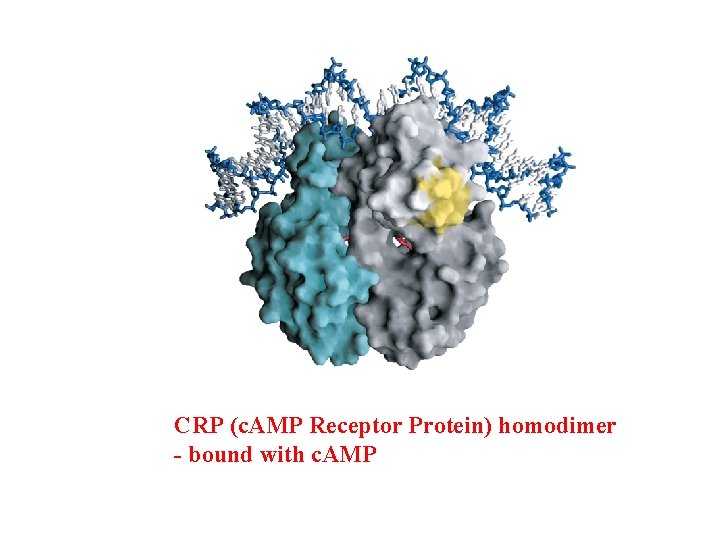
CRP (c. AMP Receptor Protein) homodimer - bound with c. AMP


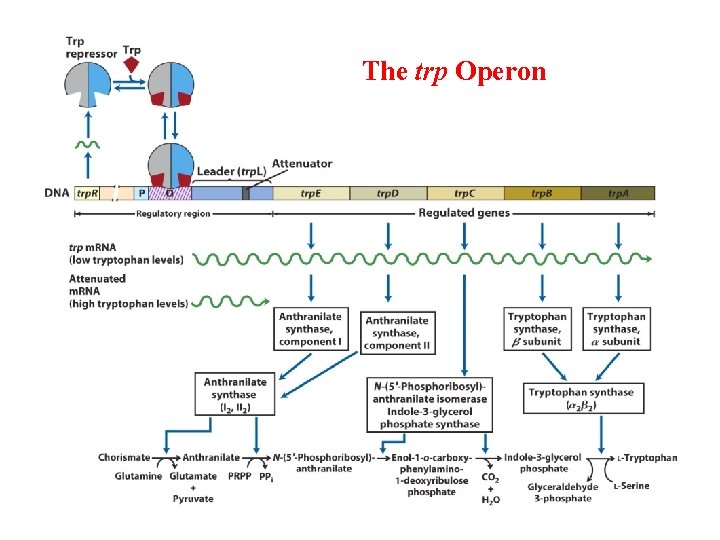
The trp Operon

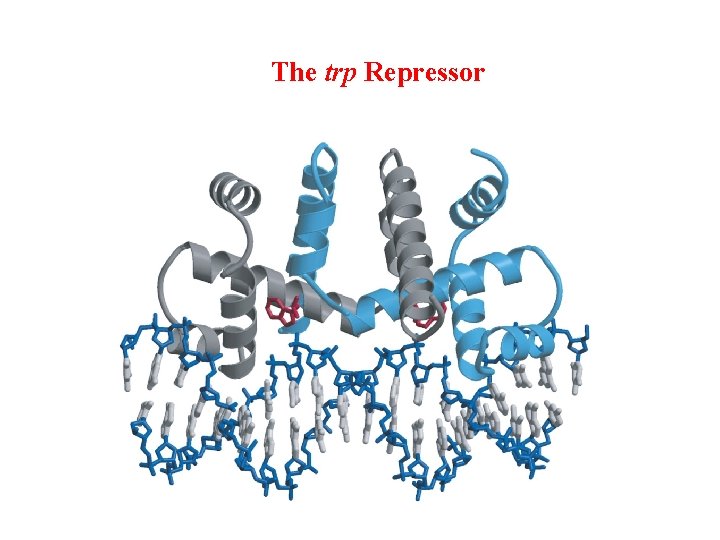
The trp Repressor

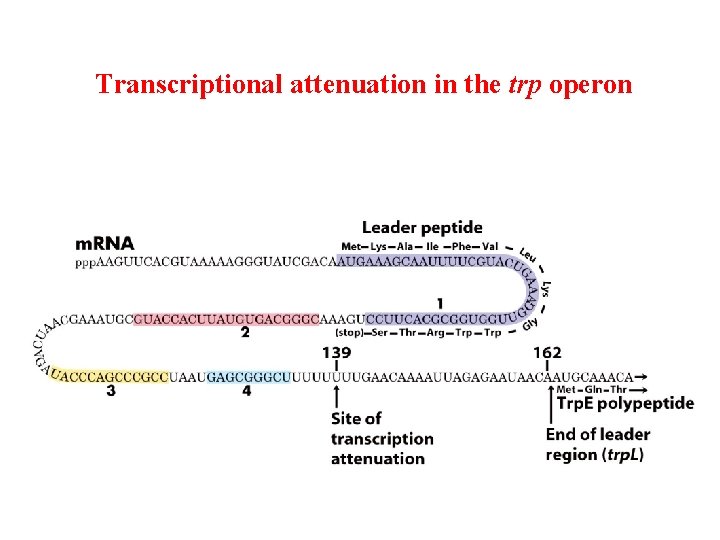
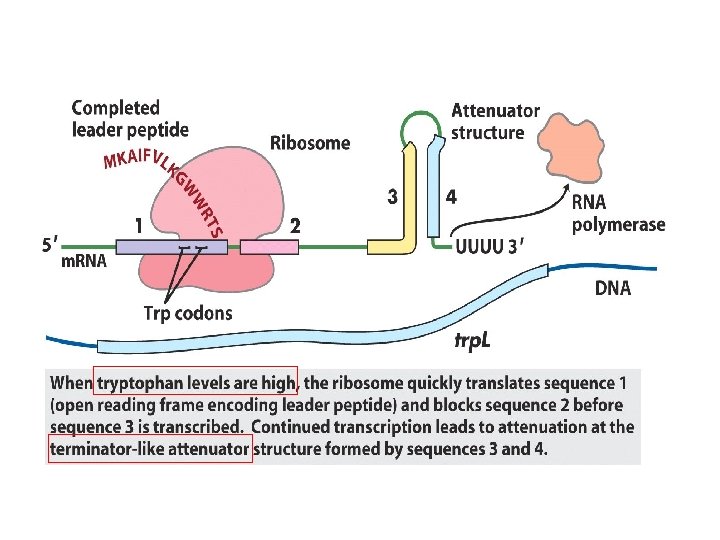
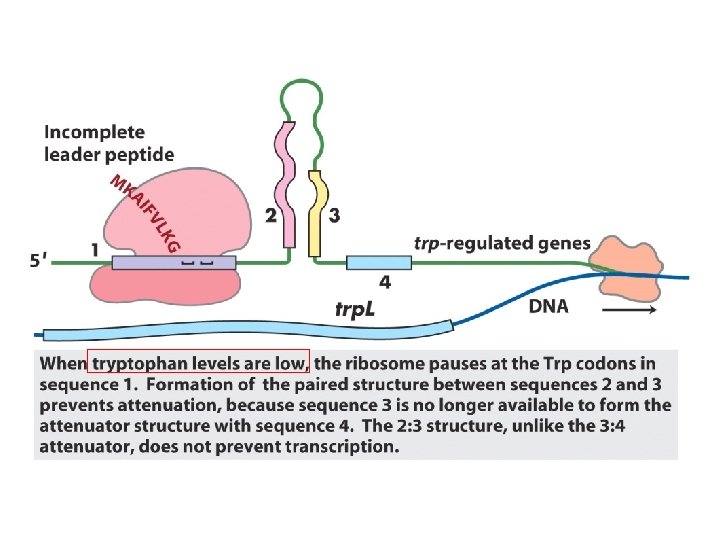
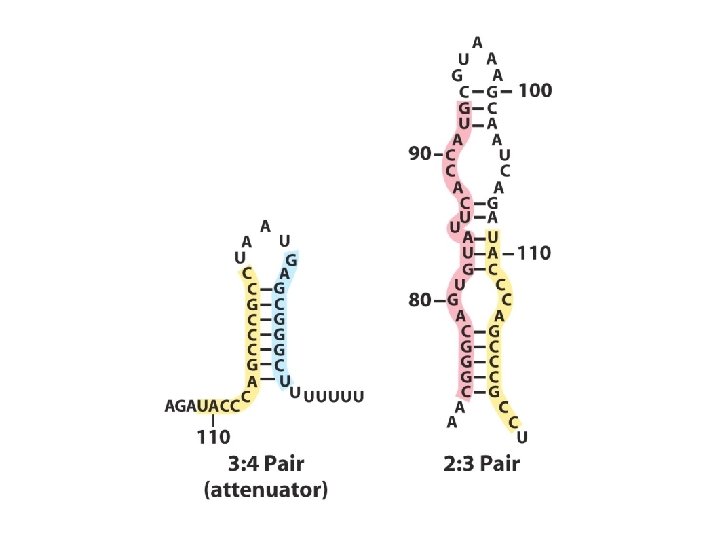
Transcriptional attenuation in the trp operon




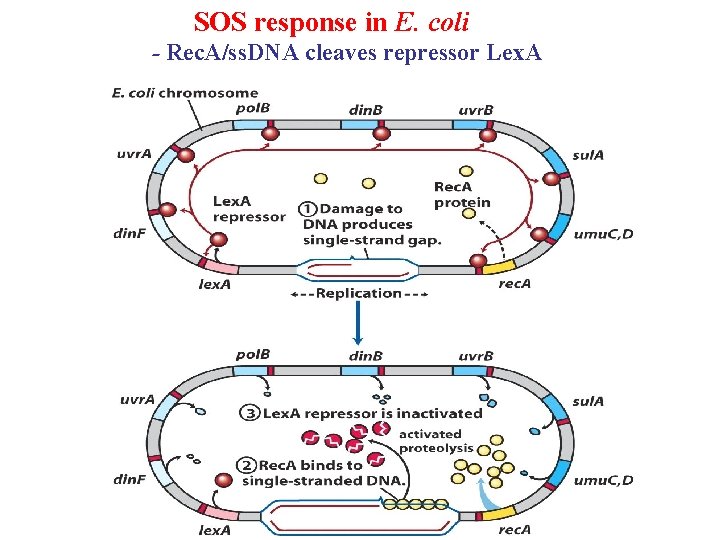
SOS response in E. coli - Rec. A/ss. DNA cleaves repressor Lex. A

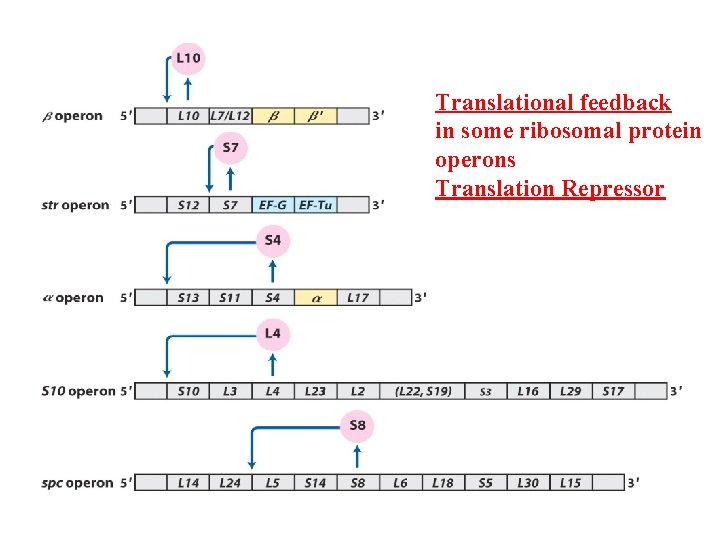
Translational feedback in some ribosomal protein operons Translation Repressor

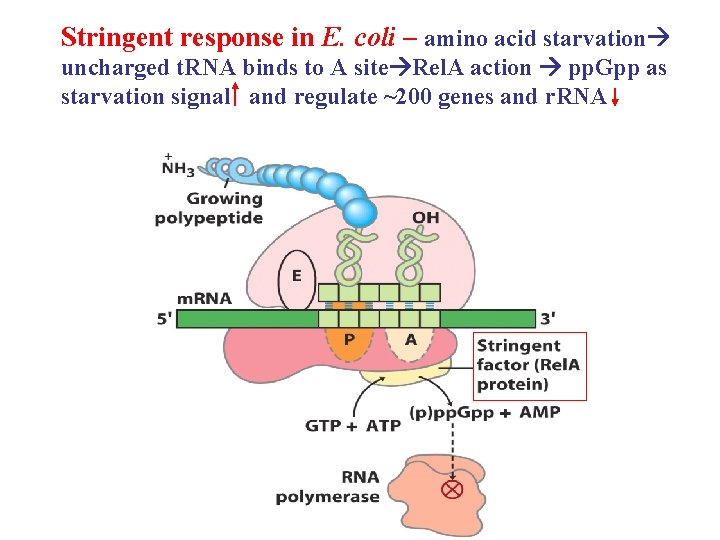
Stringent response in E. coli – amino acid starvation uncharged t. RNA binds to A site Rel. A action pp. Gpp as starvation signal and regulate ~200 genes and r. RNA

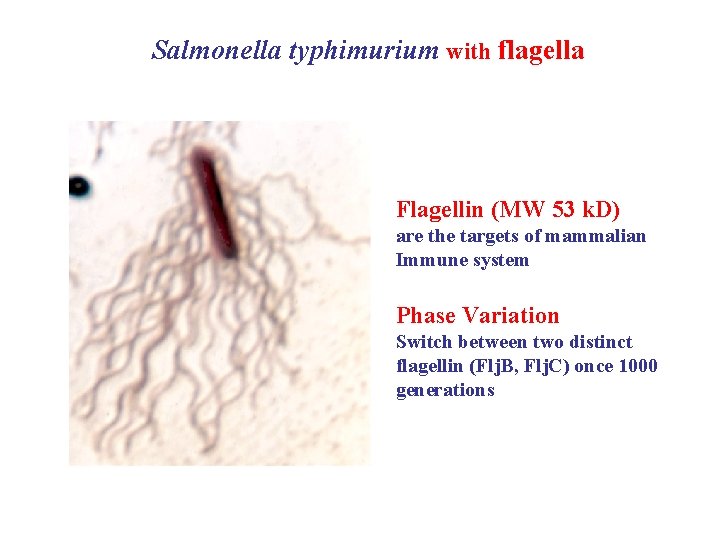
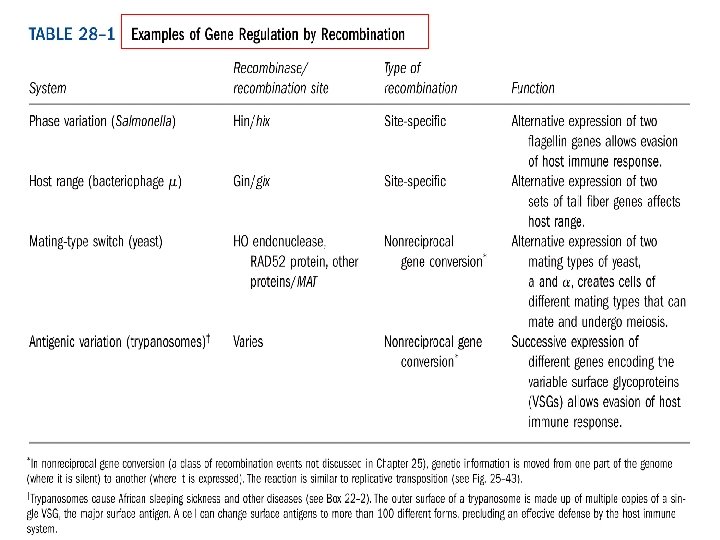
Salmonella typhimurium with flagella Flagellin (MW 53 k. D) are the targets of mammalian Immune system Phase Variation Switch between two distinct flagellin (Flj. B, Flj. C) once 1000 generations

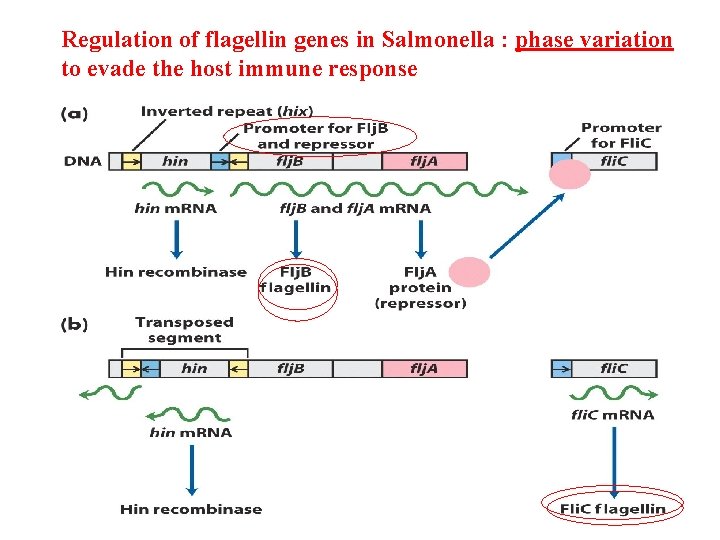
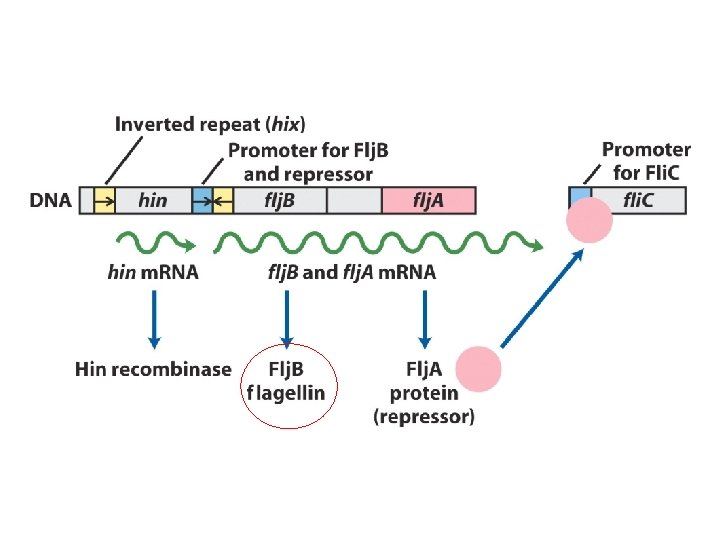
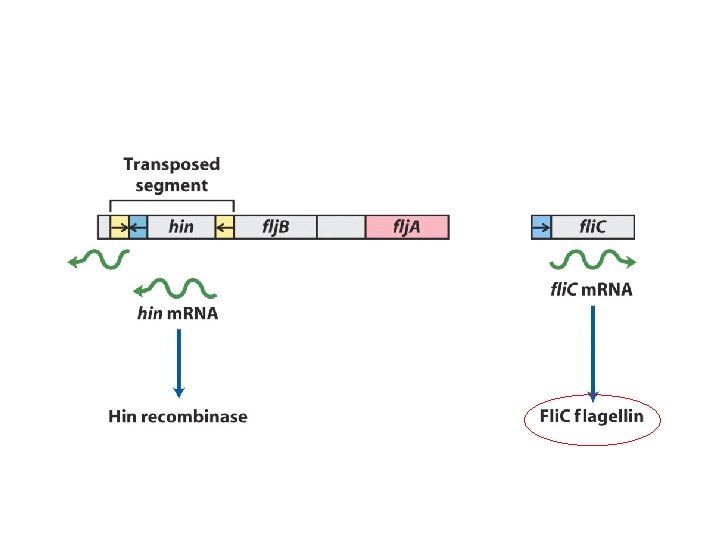
Regulation of flagellin genes in Salmonella : phase variation to evade the host immune response




3. Regulation of Gene Expression in Eukaryotes

Eukaryote Gene Regulation - Four Different Features 1. Eukaryotic promoter is restricted by the structure of chromatid 2. Positive regulation 3. More multimeric regulatory proteins 4. Transcription is separated from translation in both space and time

Transcriptionally Active Chromatin is Structurally Distinct from Inactive Chromatin • Heterochromatin - ~10% in eukaryotic cells, more condensed, transcriptionally inactive, generally associated with chromosome structure such as centormeres • Euchromatin - the remaining, less condensed chromatin • Hypersensitive Sites - in actively transcribed regions; many bind to regulatory proteins • Histones – different modifications in different regions

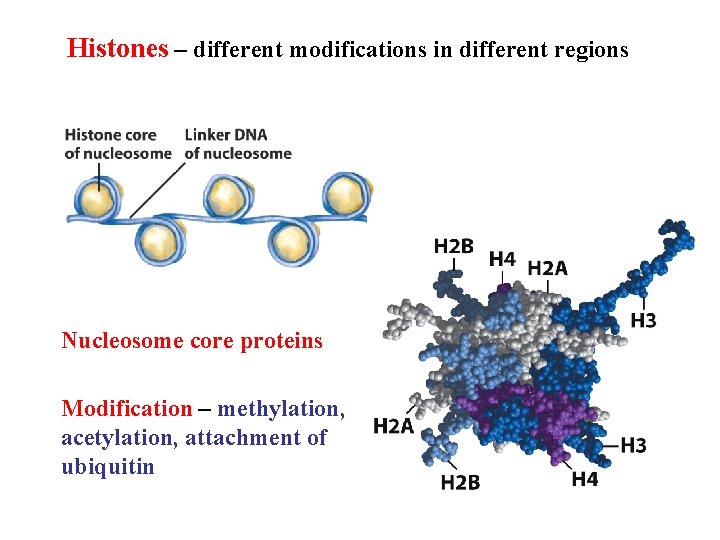
Histones – different modifications in different regions Nucleosome core proteins Modification – methylation, acetylation, attachment of ubiquitin

Histones act as a general repressor of transcription, because they interfere with protein binding to DNA 1. Histones form nucleosomes on TATA boxes, blocking transcription. Promoter-binding proteins cannot disrupt the nucleosomes. Enhancer-binding proteins bind to enhancers, displacing any histones, and then cause the histones at the TATA box to free the DNA. 2. Histone Acetylation with increased transcription. Histone are acetylated on lysines in regions on the outside of the nucleosome. Acetylation destabilizes higher-order chromatin structure. DNA becomes more accessible to transcription factors, and overcoming histone repression of transcription.

DNA Methylation 1. DNA methylation and transcription are correlated, with lower levels of methylated DNA in transcriptionally active genes. 2. Other recent observations also indicate a role for methylation in gene expression: (a) A methylase is essential for development in mice. (b) Methylation is involved in fragile X syndrome, where expansion of a triplet repeat and abnormal methylation in the FMR-1 gene silence its expression.

Chromatin Remodeling – detailed mechanisms for transcription-associated structural changes in chromatin

Acetylation in histone H 3 globular domain regulate gene expression in Yeast Cell 121 (2005) 375 • Lys 56 in histone H 3 : in the globular domain and extends toward the DNA major groove/nucleosome • K 56 acetylation : enriched at certain active genes, such as histones

Acetylation in histone H 3 globular domain regulates gene expression in yeast Cell 121(2005) 375 • SPT 10, a putative acetyltransferase: required for cell cycle-specific K 56 acetylation at histone genes • Histone H 3 K 56 acetylation at the entry-exit gate enables recruitment of the SWI/SNF nucleosome remodeling complex and so regulates gene activity

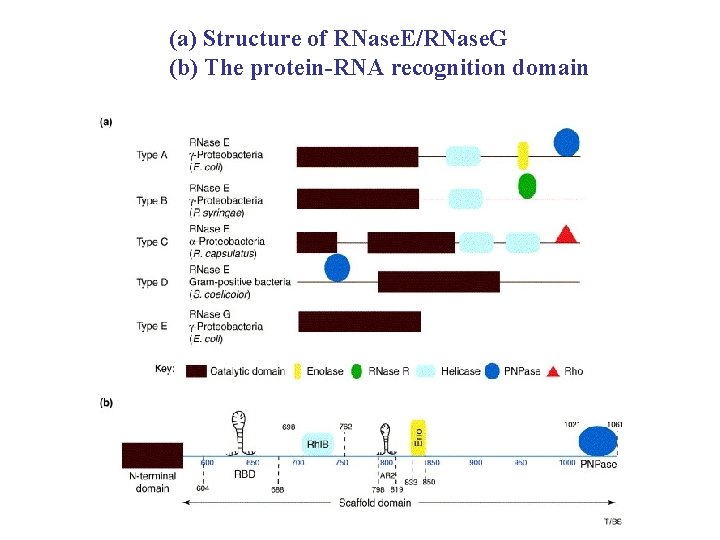
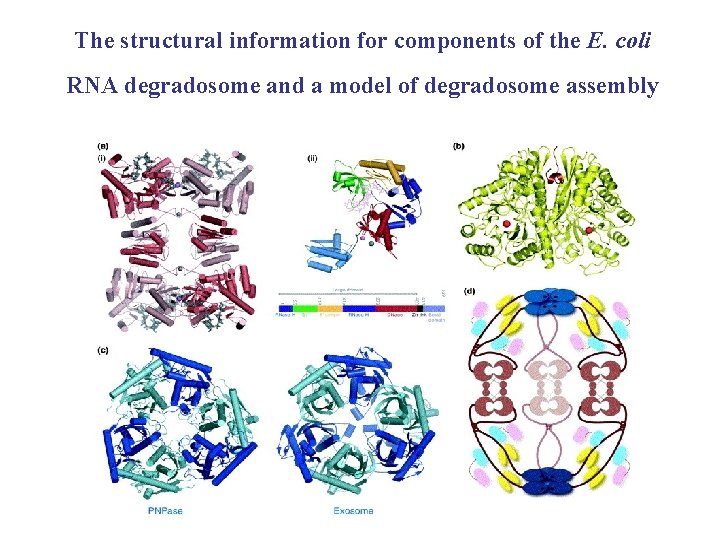
The RNA degradosome TIBS 31 (2006) 359 -365 • Most m. RNA molecules are destroyed shortly after synthesized • In E. coli, a multi-enzyme complex RNA degradosome can drive the energy-dependent turnover of m. RNA and trim RNA species into their active forms • Degradosome comprises : 1. endoribonuclease RNase E : initiates the m. RNA turnover 2. ATP-dependent RNA helicase Rhl. B : unwinds and translocates RNA 3. glycolytic enzyme : Enolase 4. phosphorolytic exoribonuclease : PNPase

(a) Structure of RNase. E/RNase. G (b) The protein-RNA recognition domain

The structural information for components of the E. coli RNA degradosome and a model of degradosome assembly

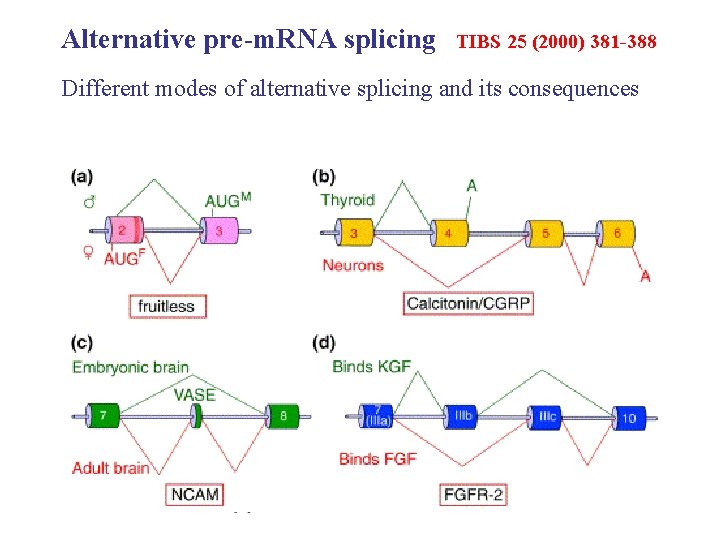
Alternative pre-m. RNA splicing TIBS 25 (2000) 381 -388 Different modes of alternative splicing and its consequences

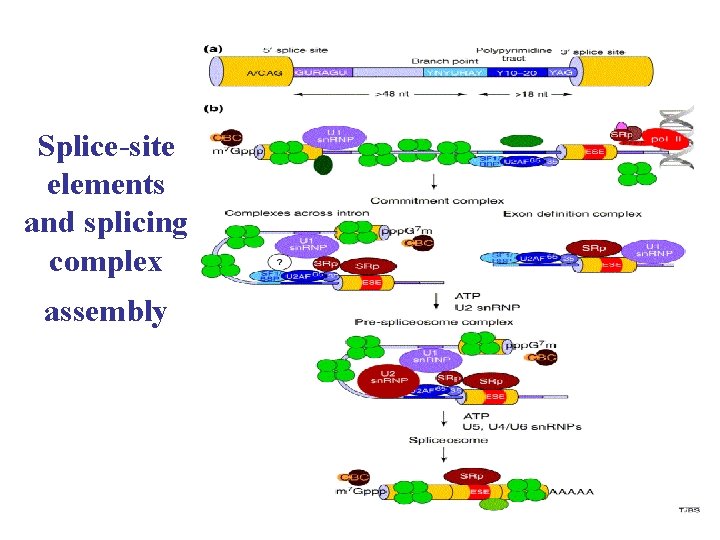
Splice-site elements and splicing complex assembly

Packing and Remodelling RNA • Primary transcripts associate with a family of polypeptides known as hn. RNP proteins • They contain RNA-binding motifs, and Gly-rich domains for protein–protein interactions and RNA transport • hn. RNP packaging can also bring together distant regions of the pre-m. RNA and therefore assist splicesite pairing

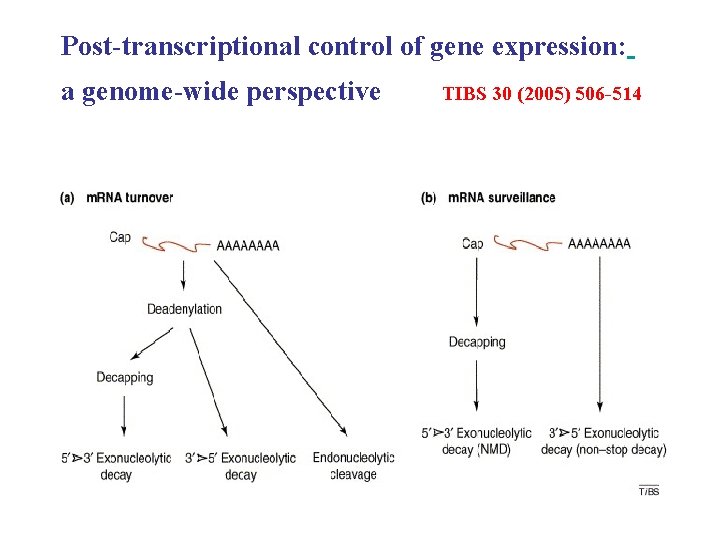
Post-transcriptional control of gene expression: a genome-wide perspective TIBS 30 (2005) 506 -514

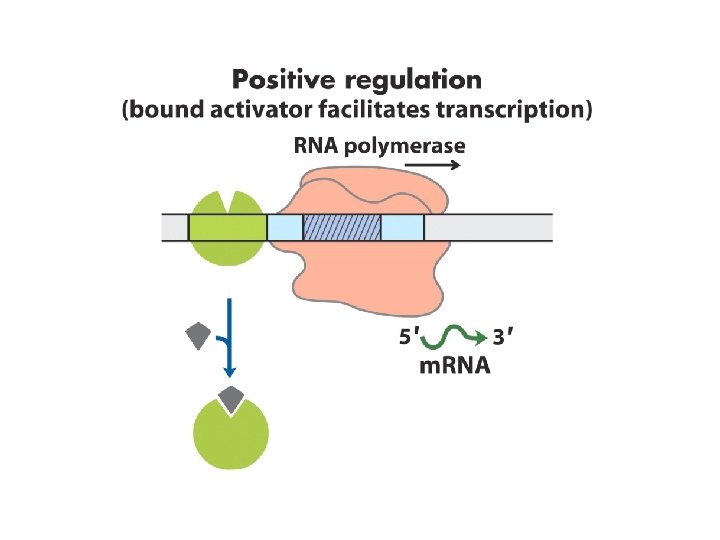
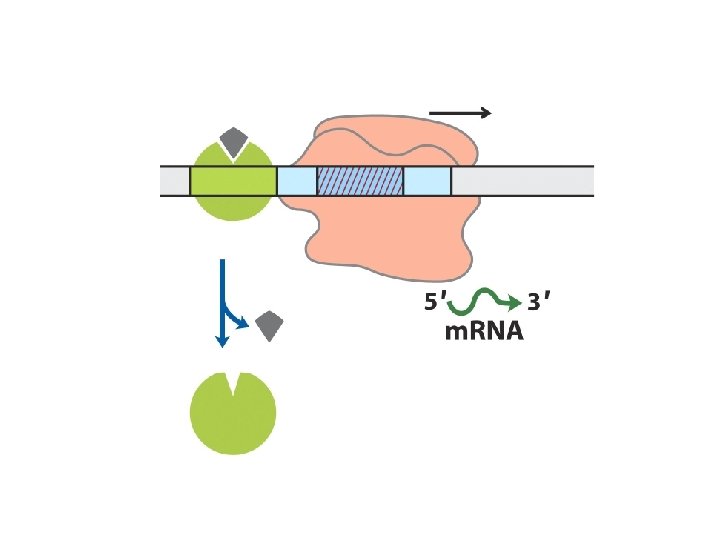
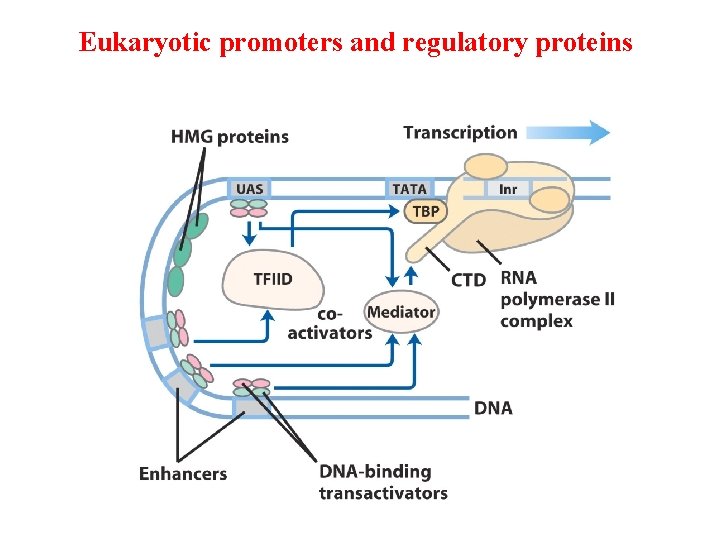
Many eukaryotic promoters are positively regulated • RNA polymerases have little or no intrinsic affinity for promoters • Transcription initiation depends on activator proteins • Enhancer; Upstream Activator Sequence (UAS, Yeast) • Basal transcription factors – RNA pol II DNA-binding transactivator – enhancer Coactivator – interconnections Repressors -

Eukaryotic promoters and regulatory proteins

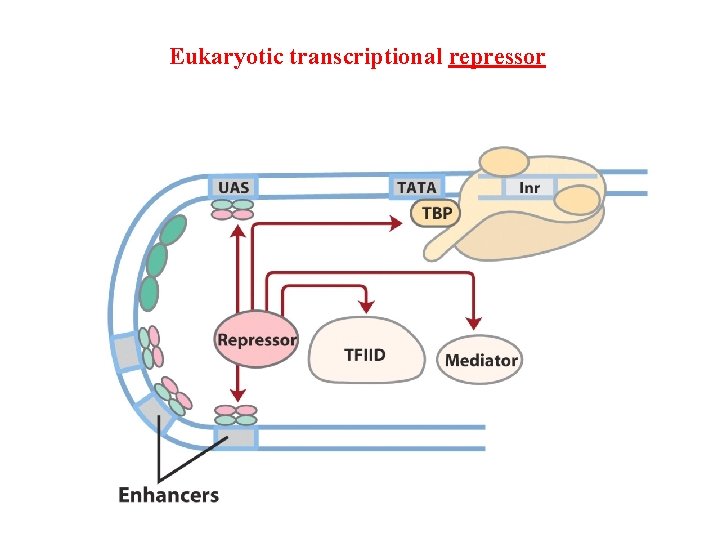
Eukaryotic transcriptional repressor

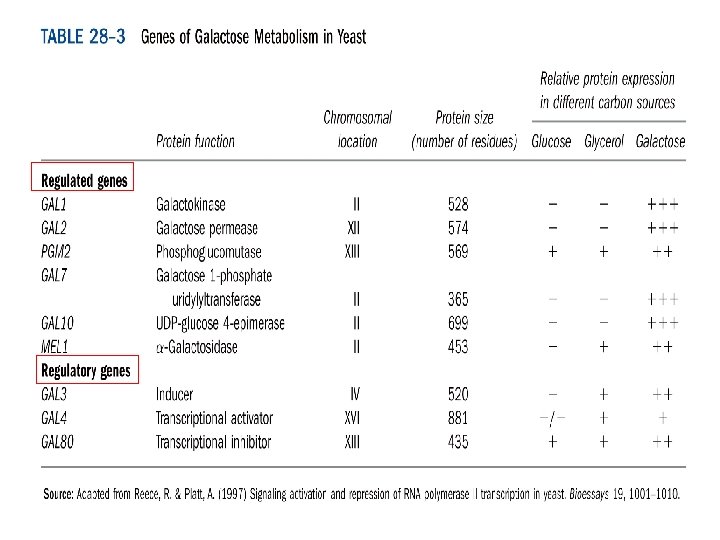
Galactose-utilization genes of yeast GAL 1 galactokinase. GAL 7 galactose transferase. GAL 10 galactose epimerase


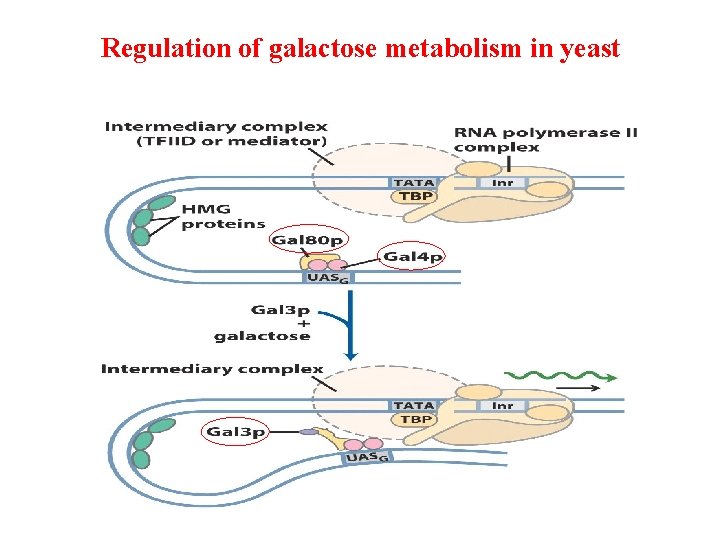
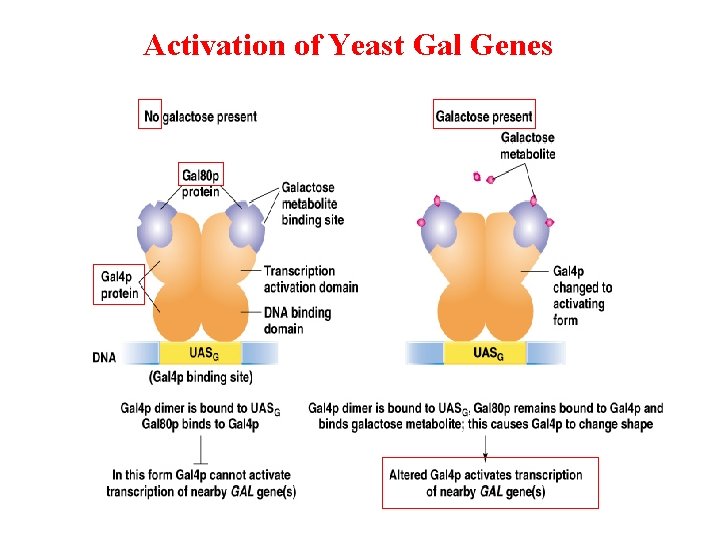
Regulation of galactose metabolism in yeast

Activation of Yeast Gal Genes

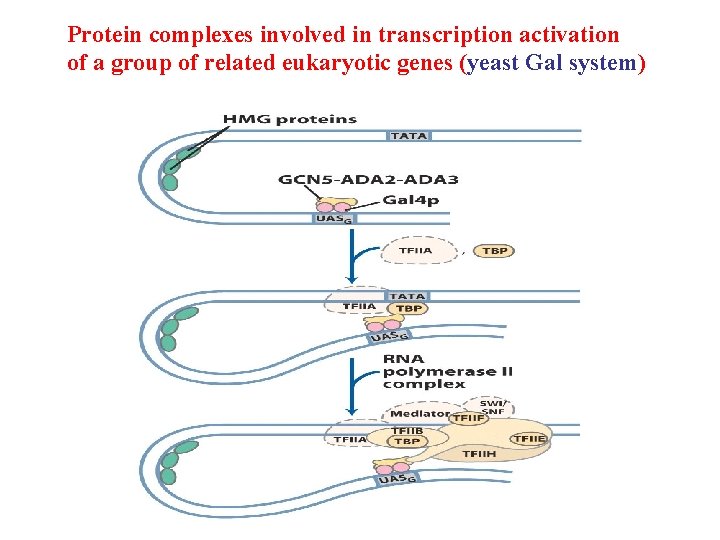
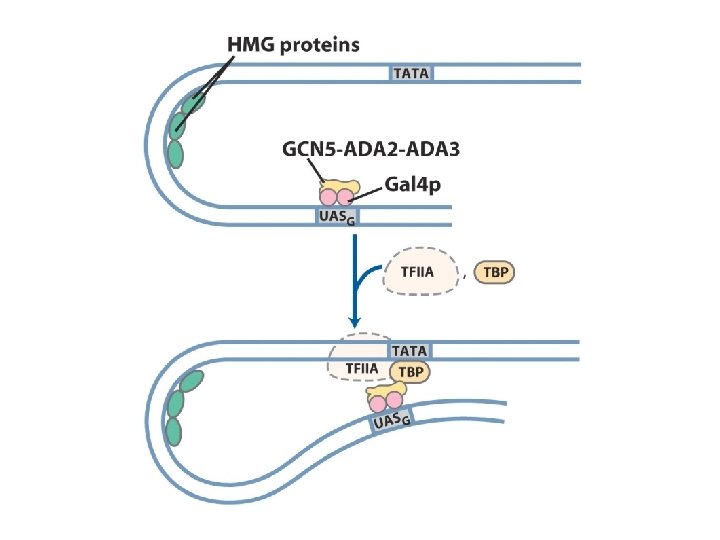
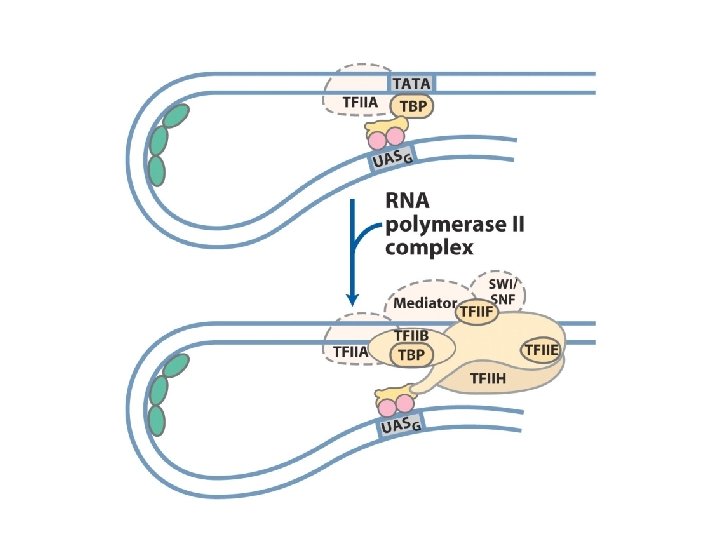
Protein complexes involved in transcription activation of a group of related eukaryotic genes (yeast Gal system)



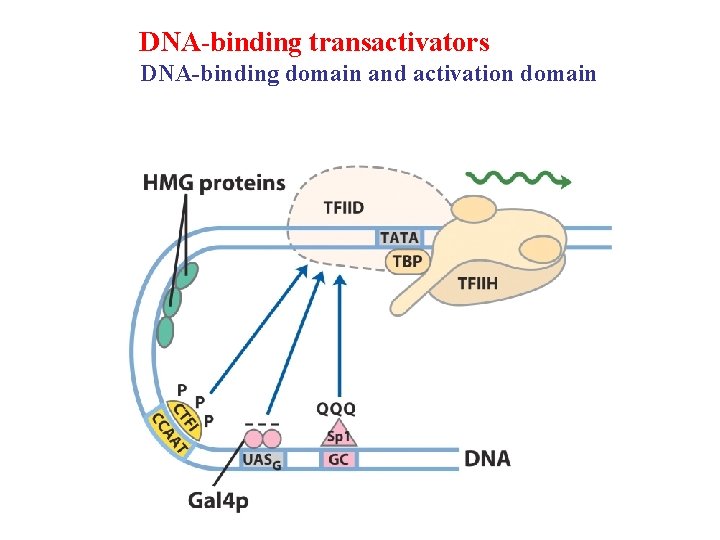
DNA-binding transactivators DNA-binding domain and activation domain

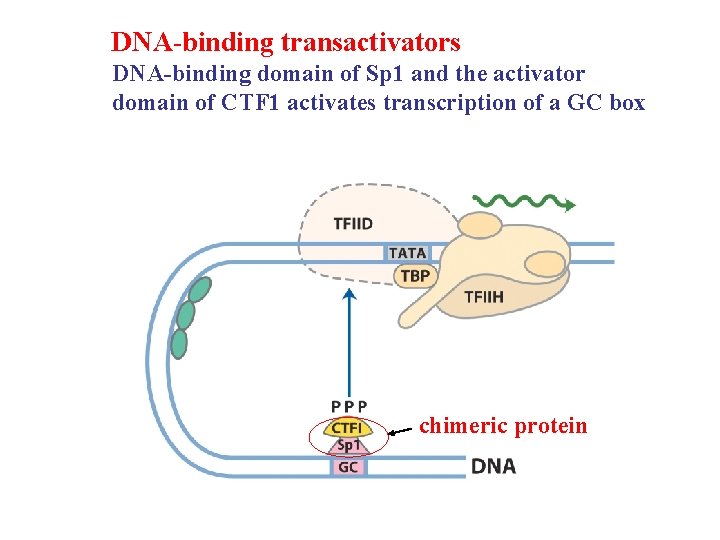
DNA-binding transactivators DNA-binding domain of Sp 1 and the activator domain of CTF 1 activates transcription of a GC box chimeric protein

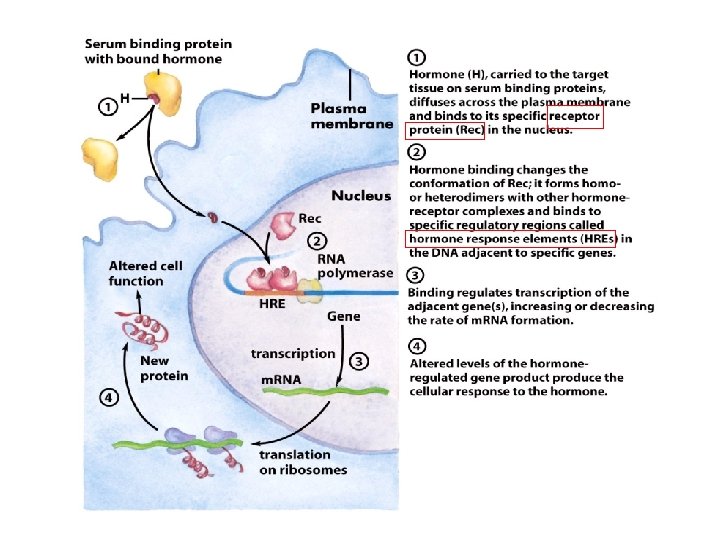
Eukaryotic gene expression can be regulated by intracellular and extracellular signals • Steroid hormone – extracellular signal bind to intracellular receptor hormone-receptor complex binds to HRE (hormone-response elements) • Regulation through phosphorylation of transcription factors – intracellular signal


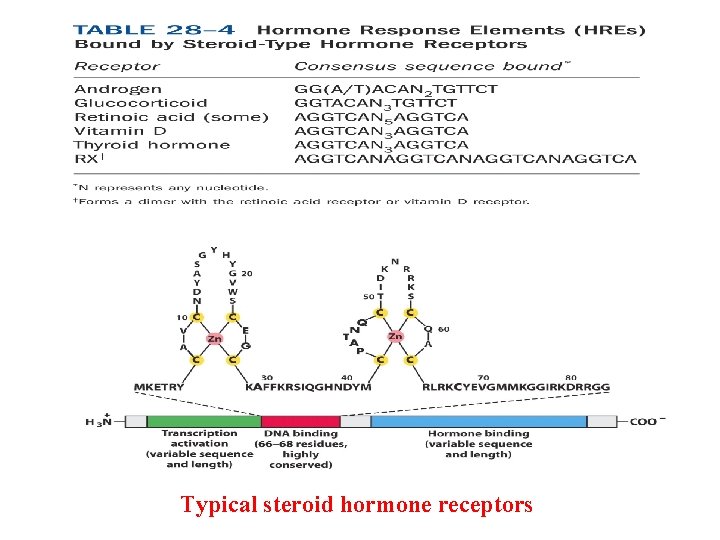
Typical steroid hormone receptors

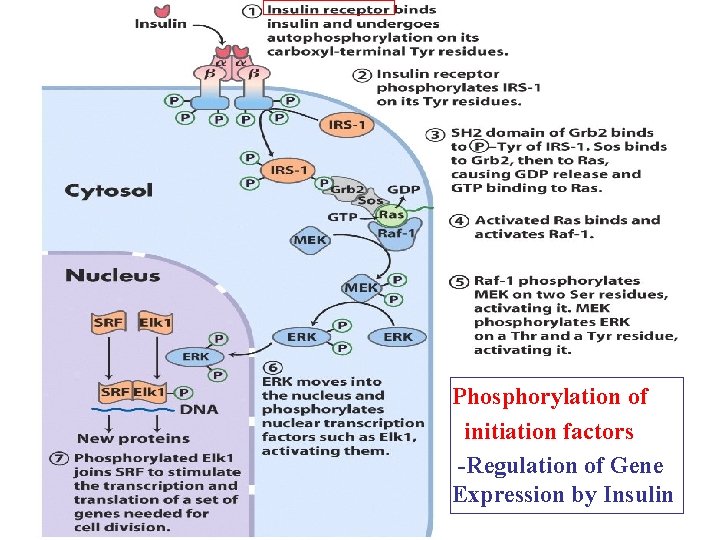
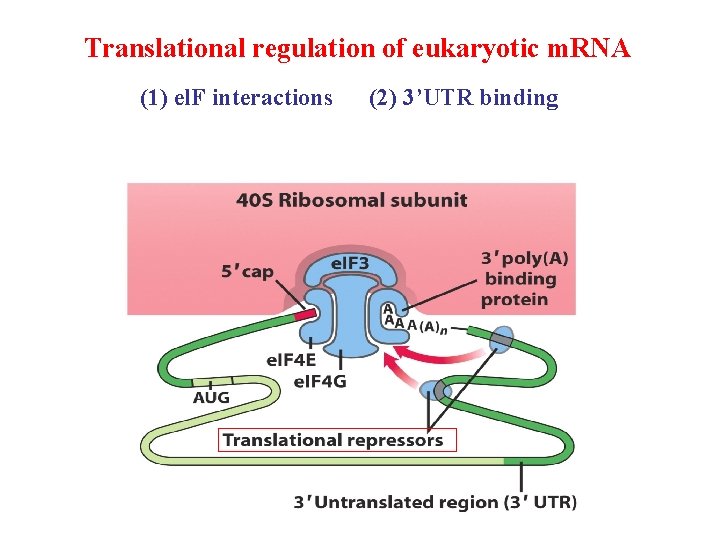
Translational Regulation 1. Phosphorylation of initiation factor less active 2. Protein repressor bind to 3’UTR of m. RNA to prevent translation initiation 3. Binding proteins disrupt the interaction of el. F 4 E and el. F 4 G to prevent the formation of eukaryotic initiation complex

Phosphorylation of initiation factors -Regulation of Gene Expression by Insulin

Translational regulation of eukaryotic m. RNA (1) el. F interactions (2) 3’UTR binding

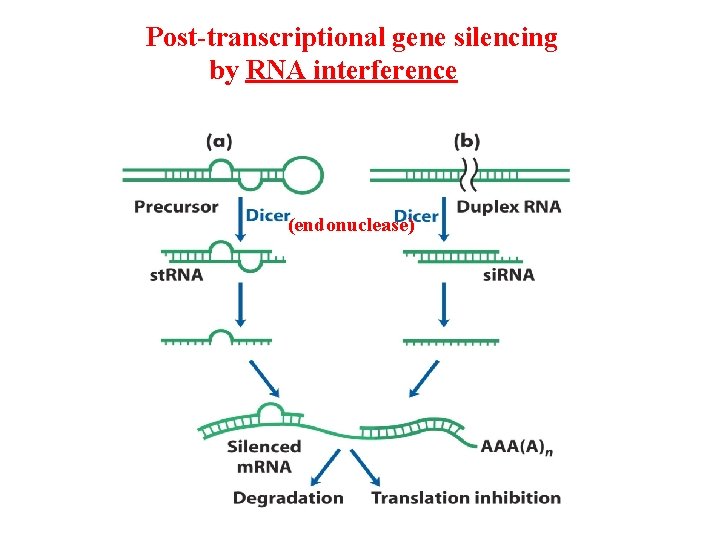
Post-transcriptional gene silencing by RNA interference (endonuclease)

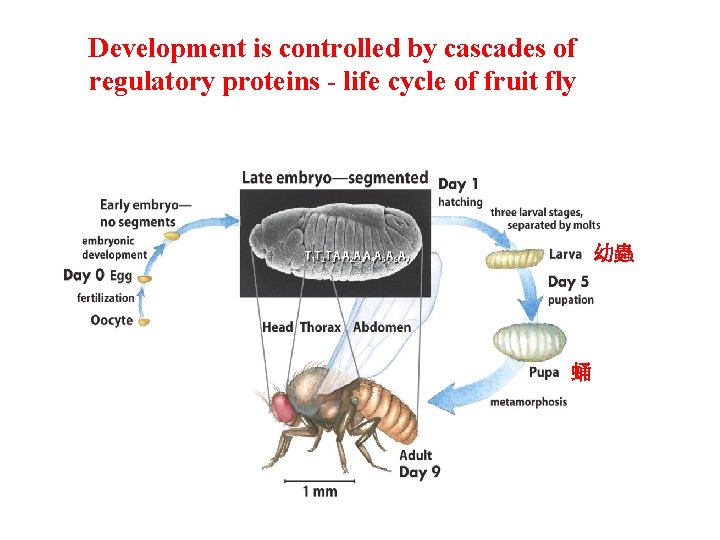
Development is controlled by cascades of regulatory proteins - life cycle of fruit fly 幼蟲 蛹

Development is controlled by Cascades of Regulatory Proteins • Polarity (anterior/posterior; dorsal/ventral) • Metamerism (serially repeating segments) • Pattern regulating genes – morphogens 1. maternal genes (expressed in unfertilized eggs) 2. segmentation genes (gap; pair-rule; segment polarity) 3. homeotic genes (expressed later to organs)

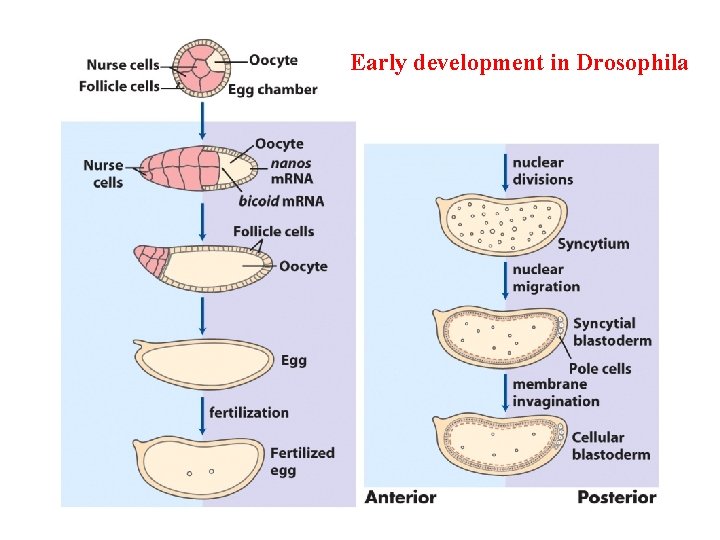
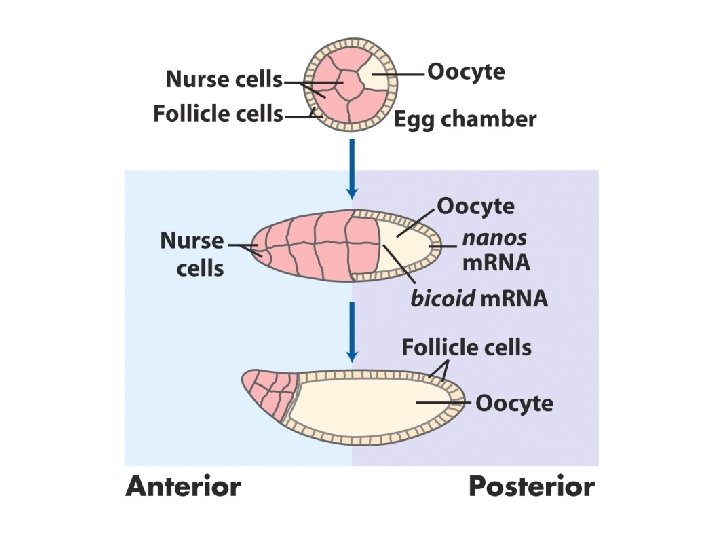
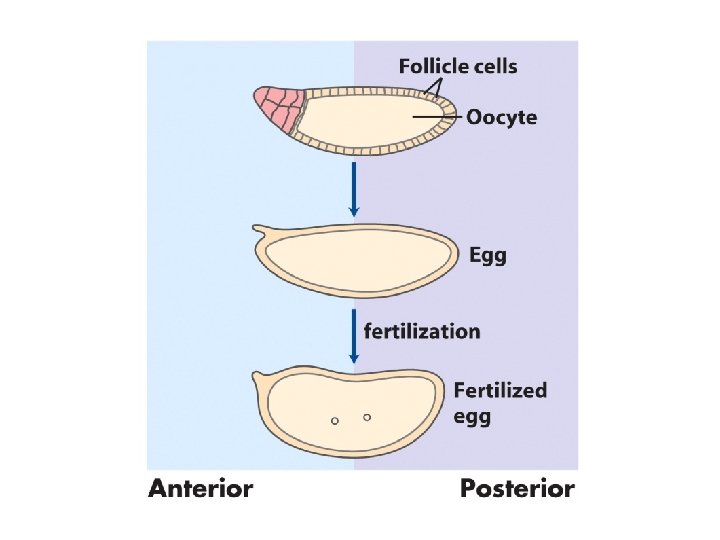
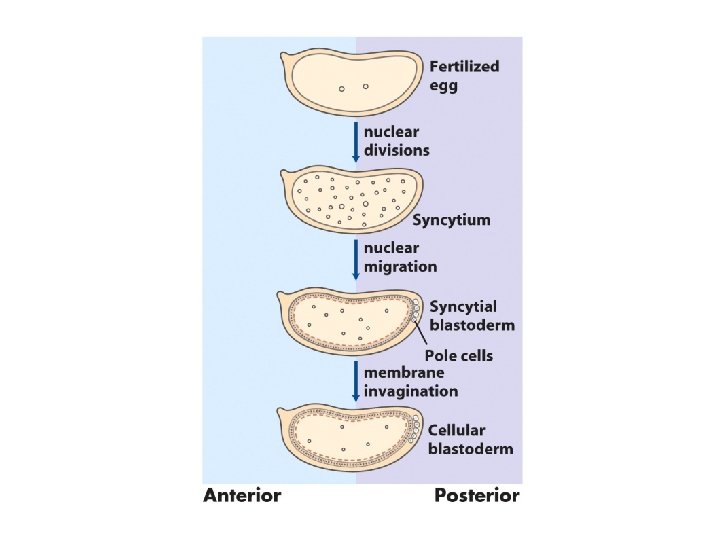
Early development in Drosophila




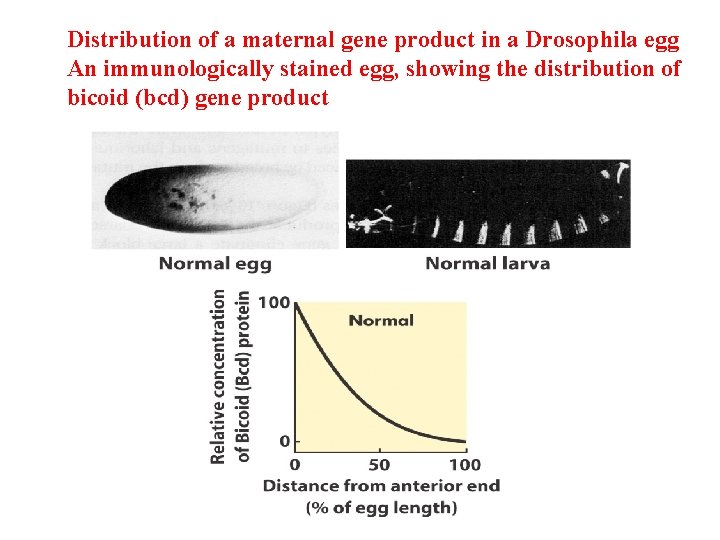
Distribution of a maternal gene product in a Drosophila egg An immunologically stained egg, showing the distribution of bicoid (bcd) gene product

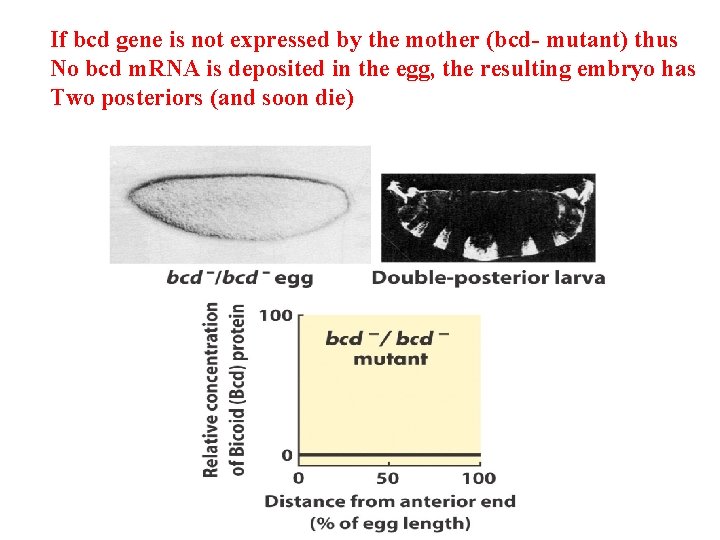
If bcd gene is not expressed by the mother (bcd- mutant) thus No bcd m. RNA is deposited in the egg, the resulting embryo has Two posteriors (and soon die)

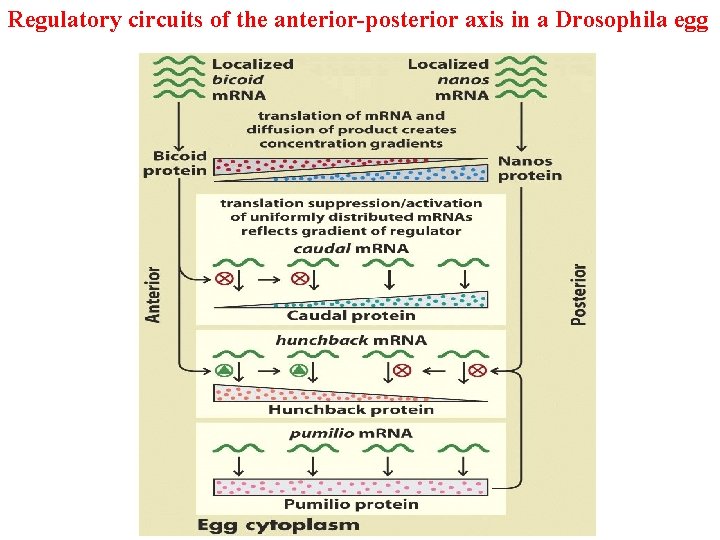
Regulatory circuits of the anterior-posterior axis in a Drosophila egg

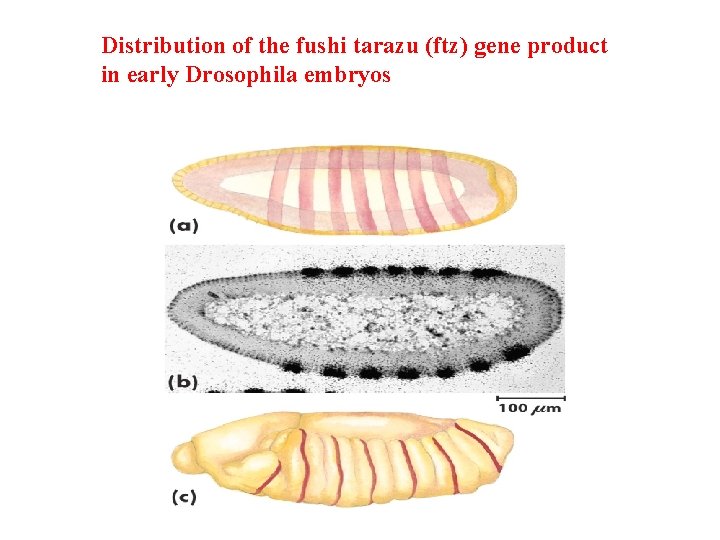
Distribution of the fushi tarazu (ftz) gene product in early Drosophila embryos

Homeotic Genes • Homeotic genes - genes that regulate the development of body patterns • Homeodomain - approx. 60 aa; helix-turn-helix; • Homeobox - DNA part • Ultrabithrax (ubx) gene: 76 kb (73 kb intron) Ubx protein is transcriptional activator

Effects of mutation in homeotic genes in Drosophila