Protein Database David Shiuan Department of Life Science

Protein Database David Shiuan Department of Life Science Institute of Biotechnology Interdisciplinary Program of Bioinformatics National Dong Hwa University

Proteome Bioinformatics/Databases n 1. Protein sequence databases - SWISS-PROT; Tr. EMBL 2. Nucleotide sequence databases - Hidden treasures: EST 3. Pattern and profile databases - PROSITE; BLOCKS; PRINTS; Pfam 4. 2 D-PAGE databases - SWISS-2 DPAGE; WORLD-2 DPAGE 5. 3 D structural databases - PDB; DSSP; HSSP

Proteome Bioinformatics/Databases n 6. Post-translational modification databases - O-GLYCBASE 7. Genomic databases - OMIM; GDB; MGD; Fly. Base 8. Metabolic databases - ENZYME; KEGG; EMP/WT 9. Interfacing and Integrating Databases - EXEx. PASy; SWISS-PROT; Cyber–Encyclopaedia of the Proteome

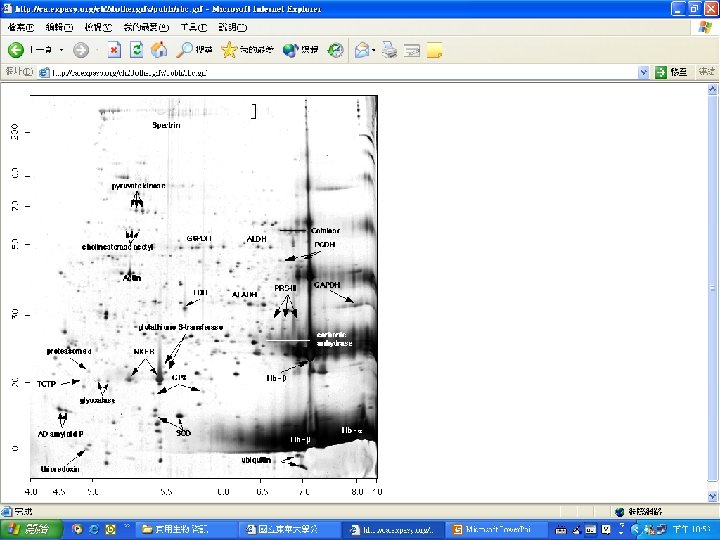
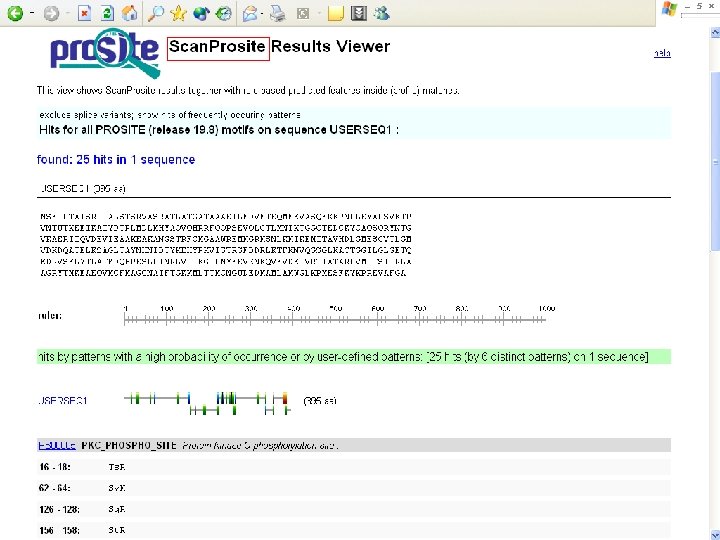
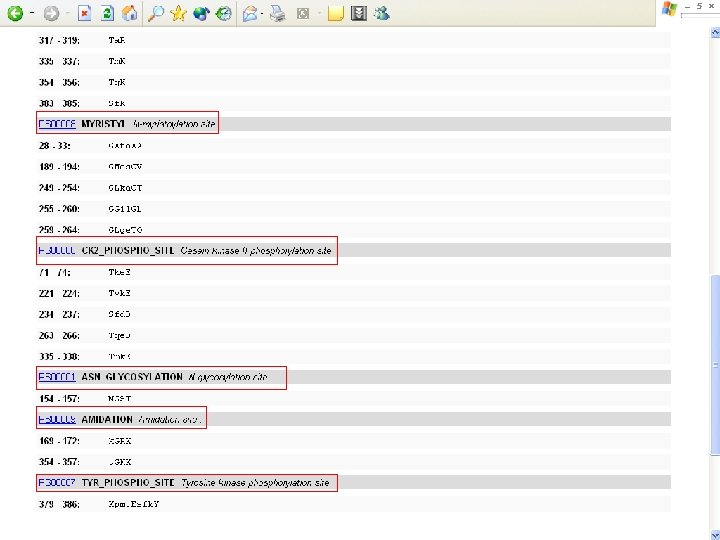
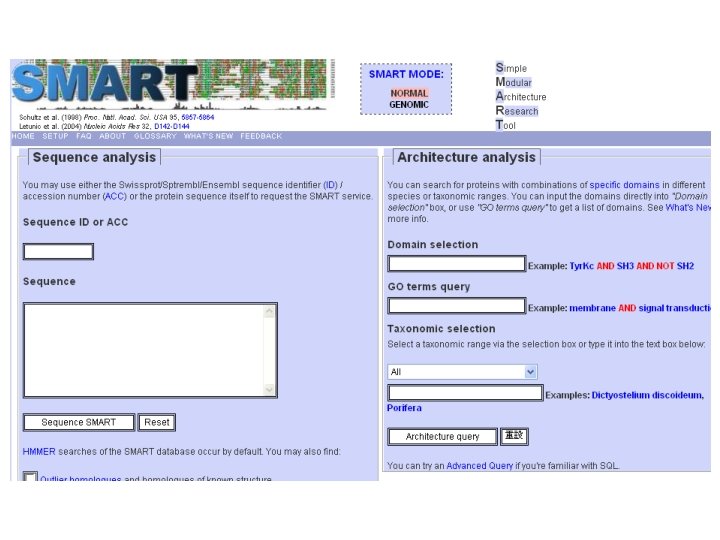
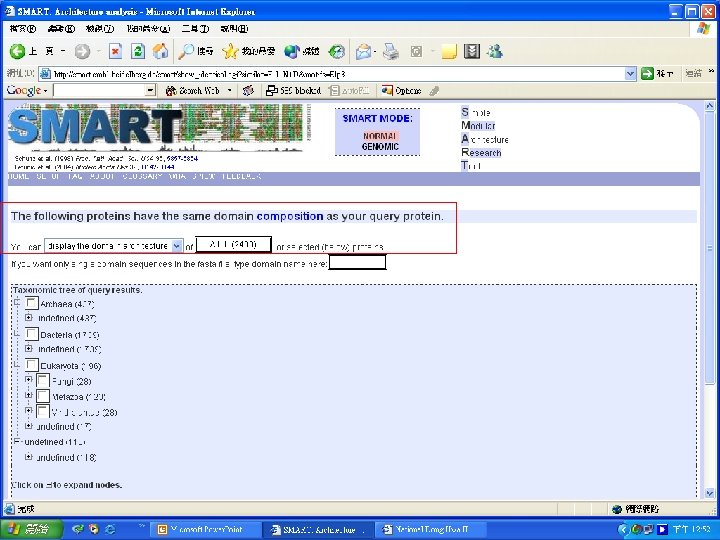
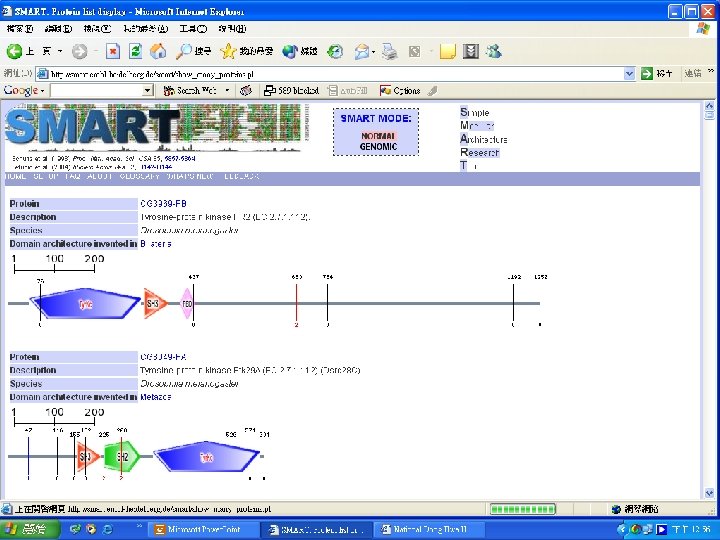
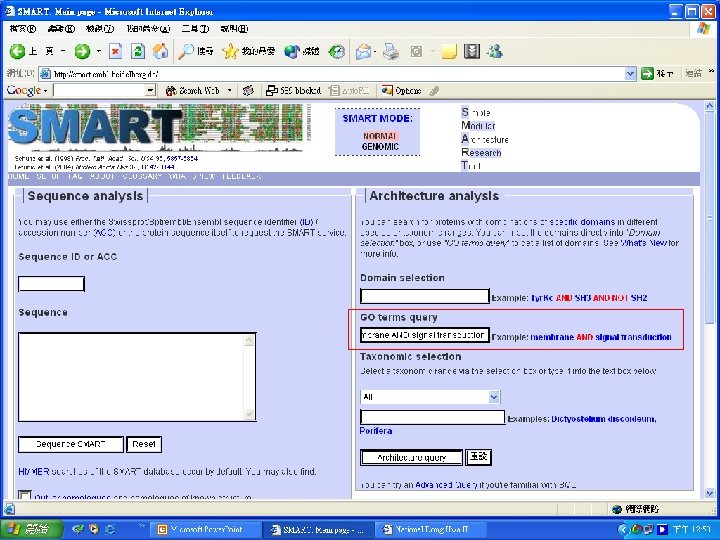
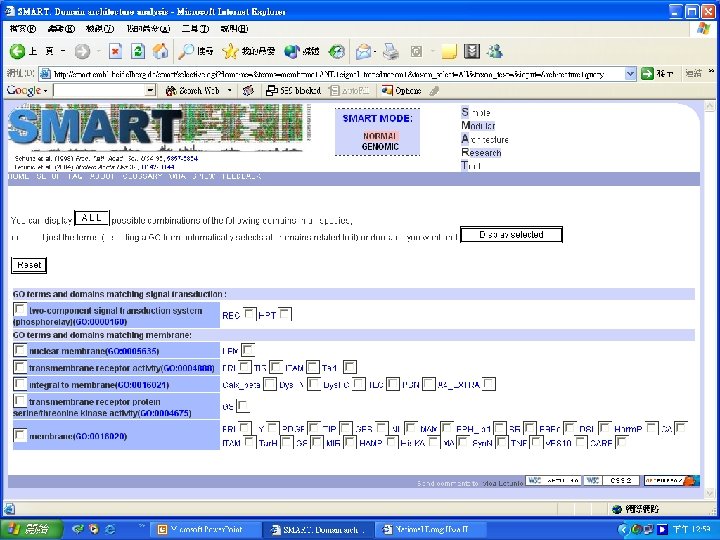
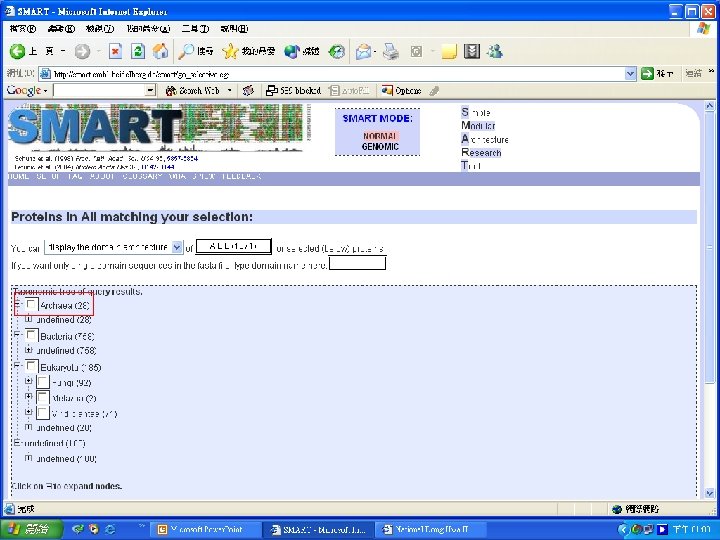
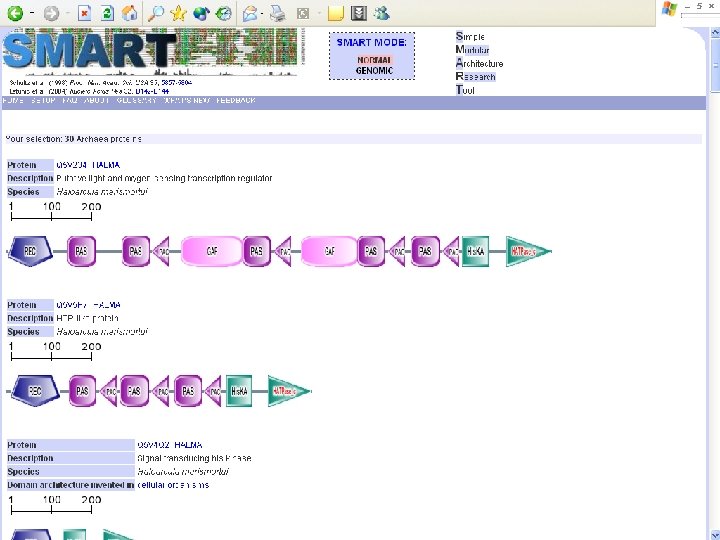
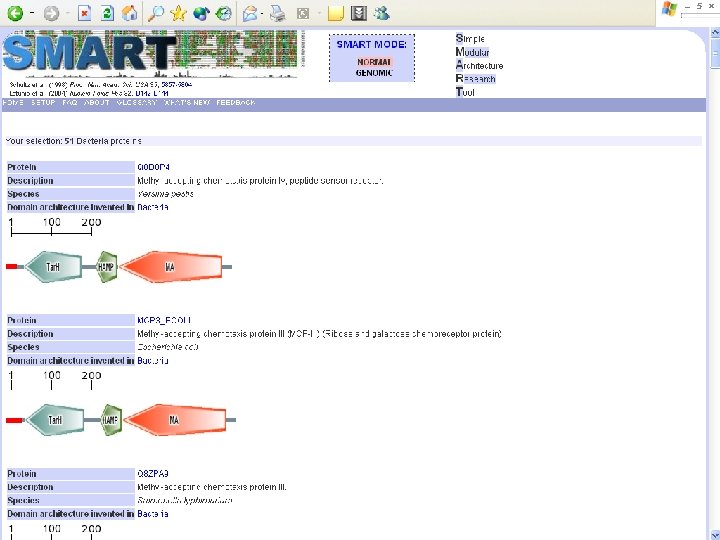
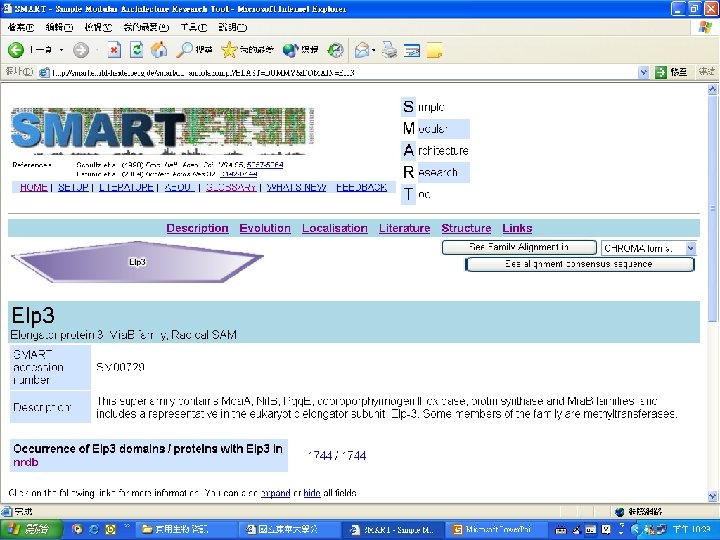
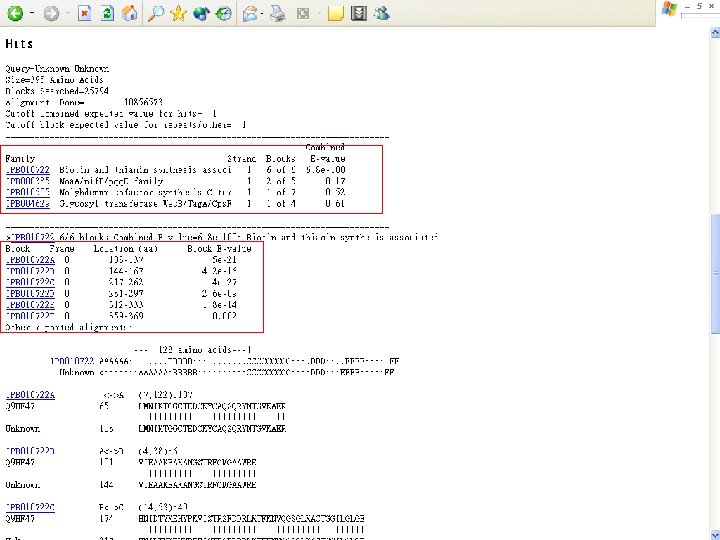
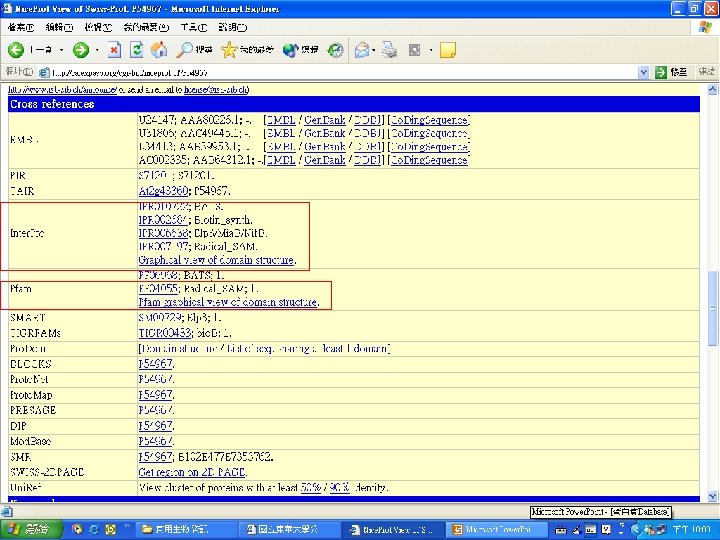
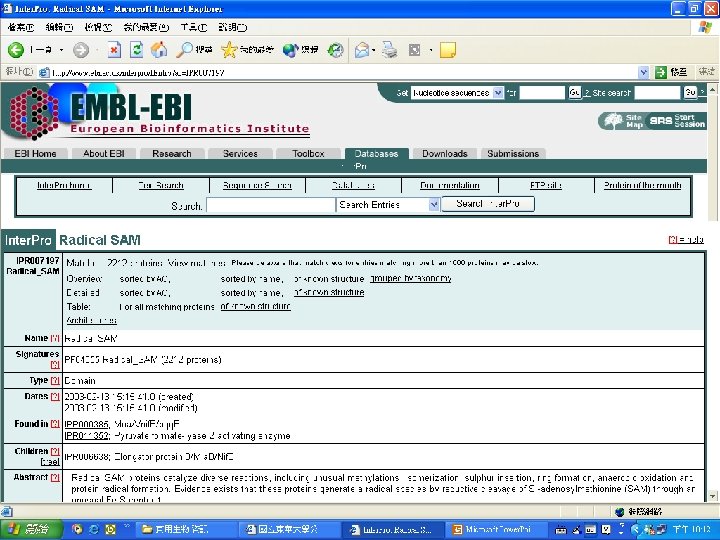
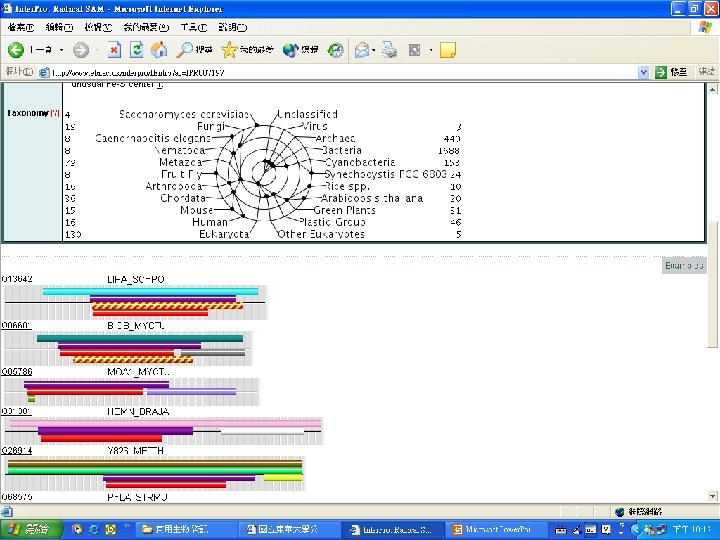
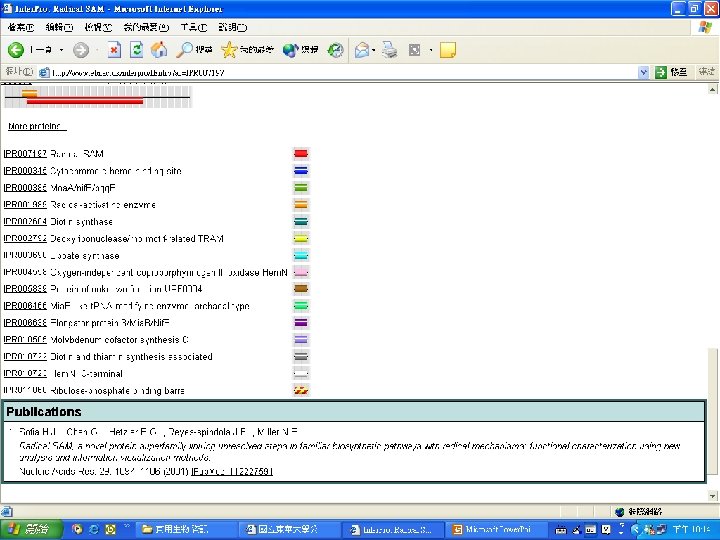
Protein Database n n n Uni. Pro - protein knowledge database Swiss 2 DPAGE - 2 D PAGE Pfam - protein family and domain Prosite - protein family and domain SMART - protein module BLOCK - protein conserved regions






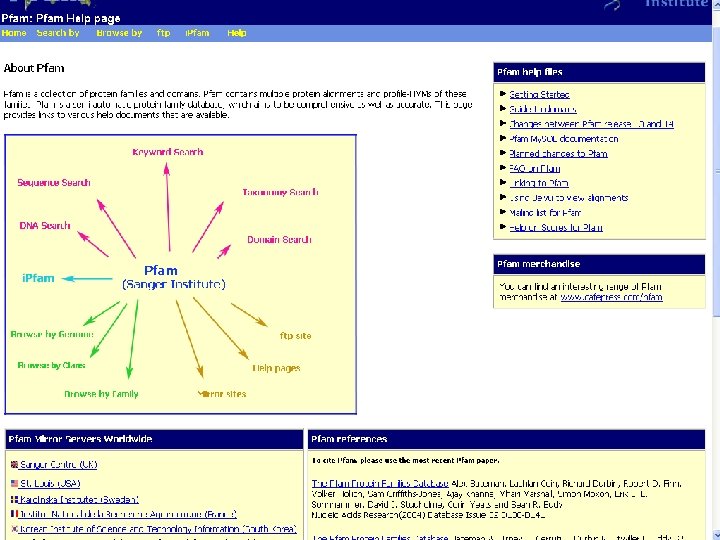
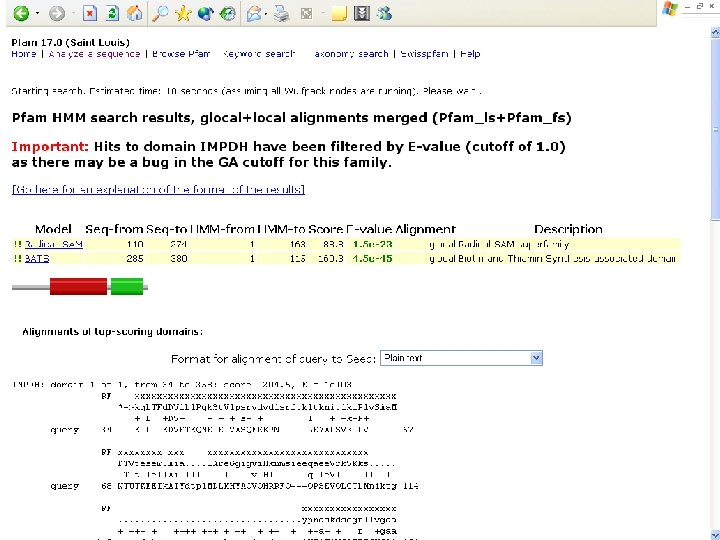
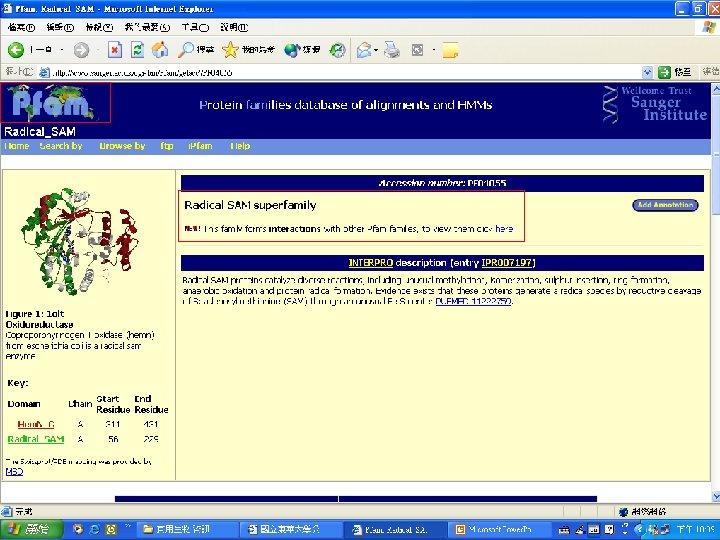
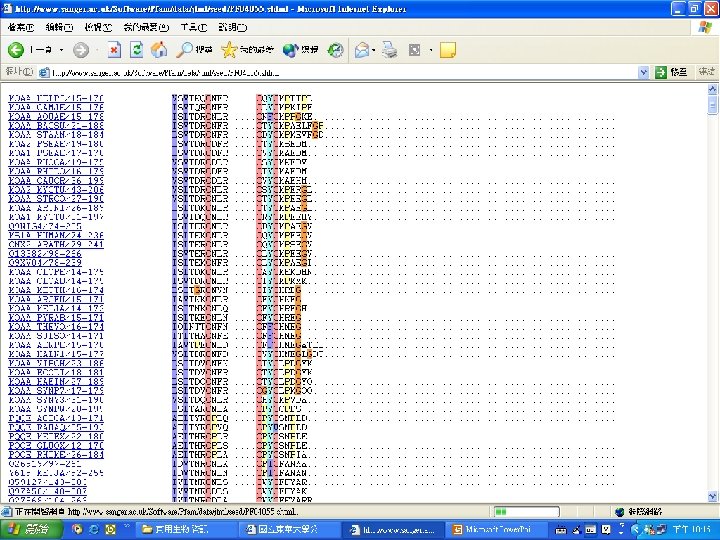
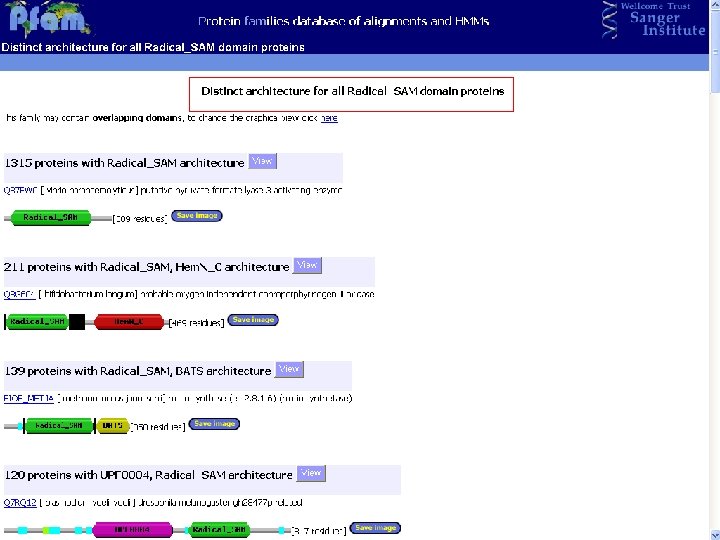
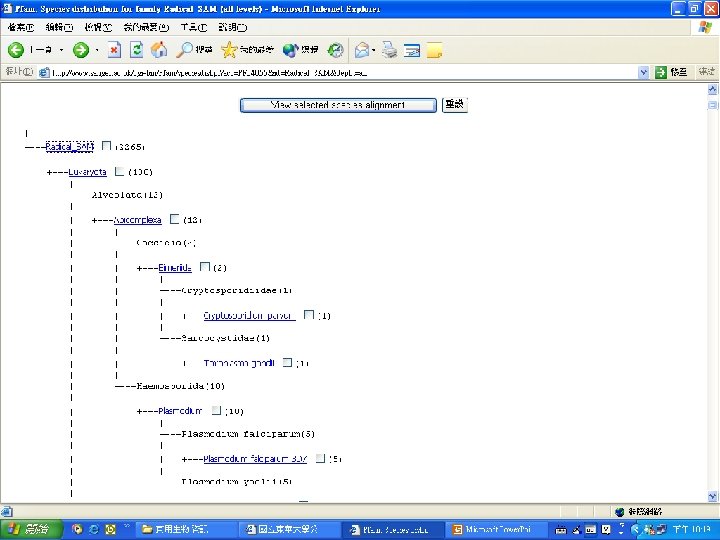
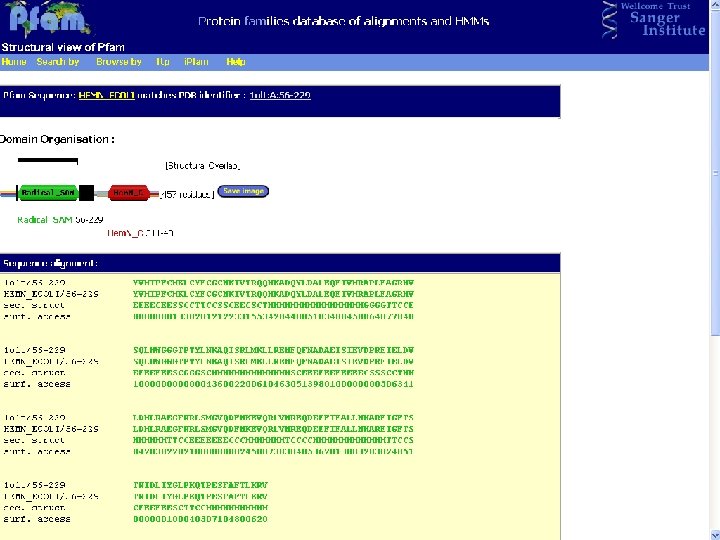
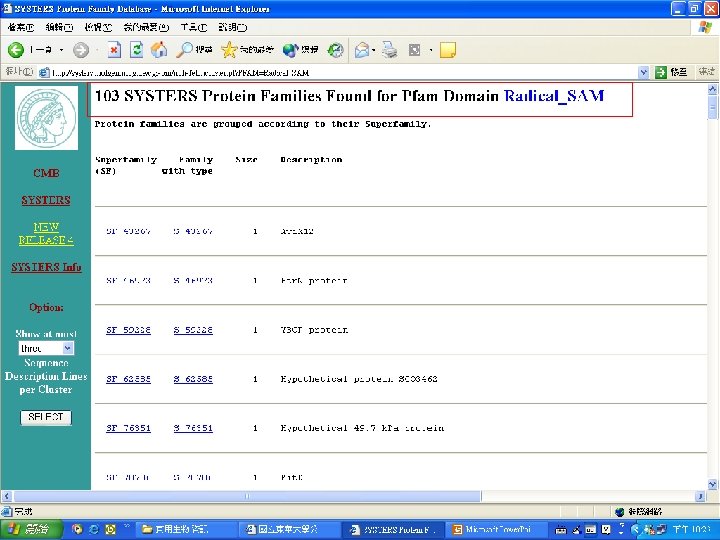
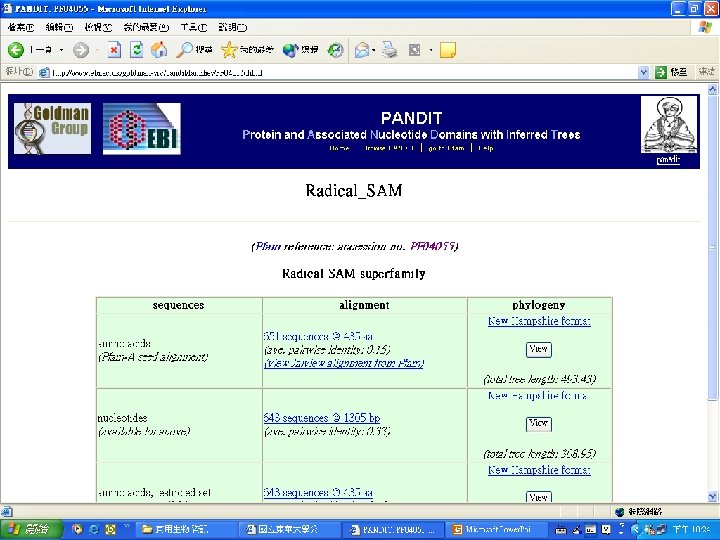
Pfam : : Home The Pfam database of protein families and HMMs n n Pfam is a large collection of multiple sequence alignments and hidden Markov models covering many common protein families. Pfam version 17. 0 (March 2005) contains alignments and models for 7868 protein families, based on the Swissprot 46. 0 and SP-Tr. EMBL 29. 0 protein sequence databases.

HMM: A Hidden Markov Model n HMM: A Hidden Markov Model, or HMM, is a statistical model for any system that can be represented as a succession of transitions between discrete states.
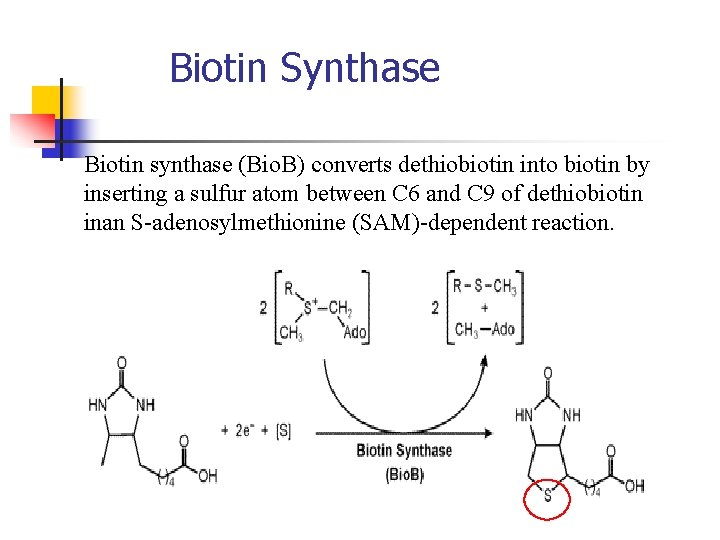
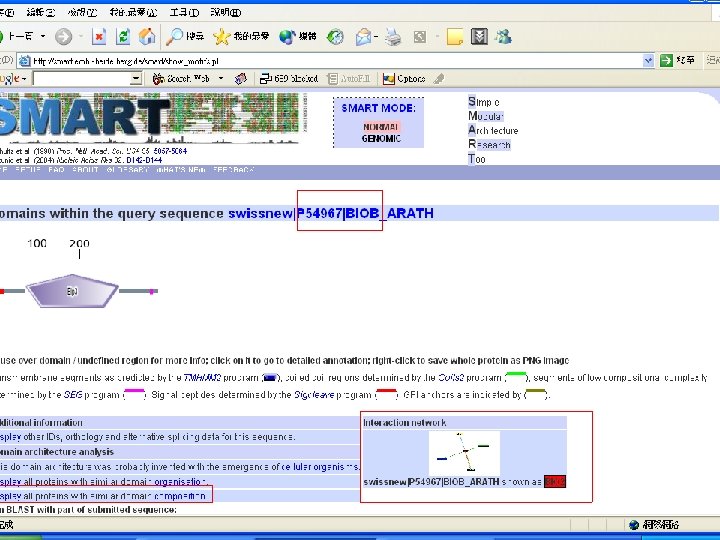
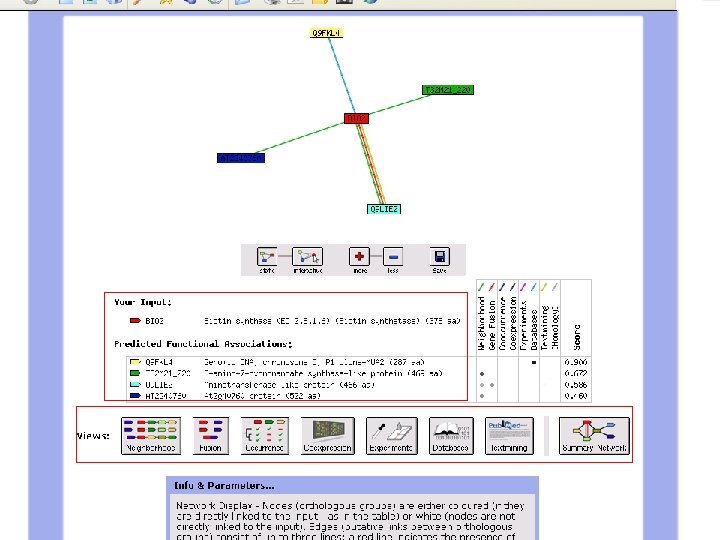
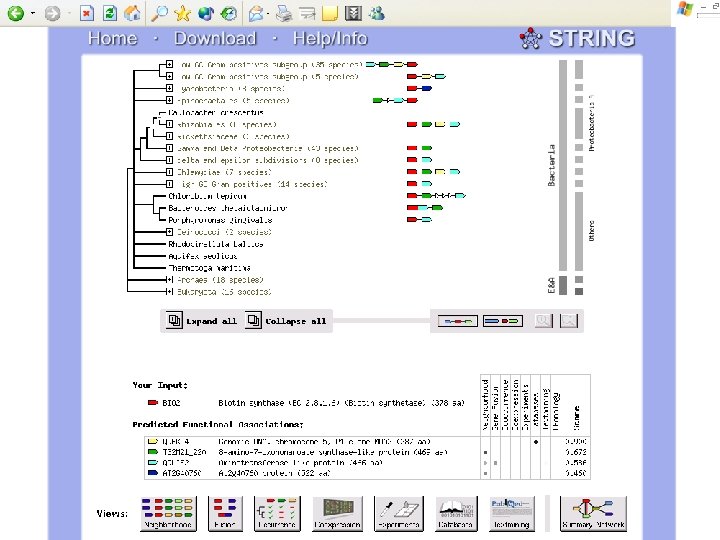
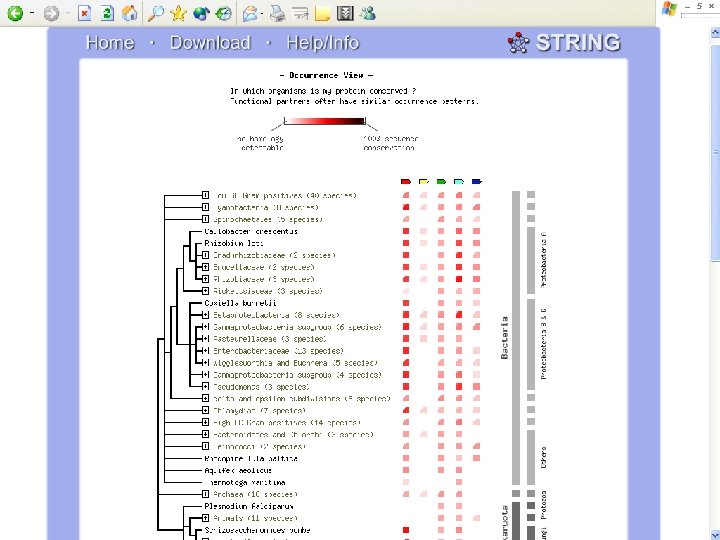
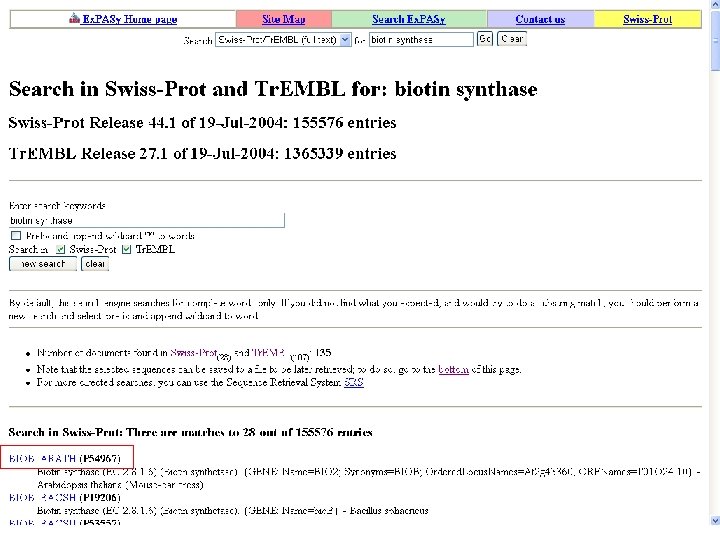
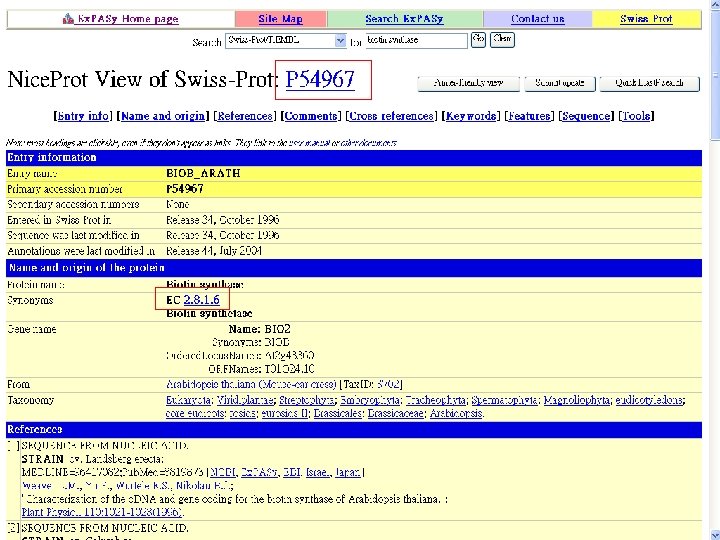
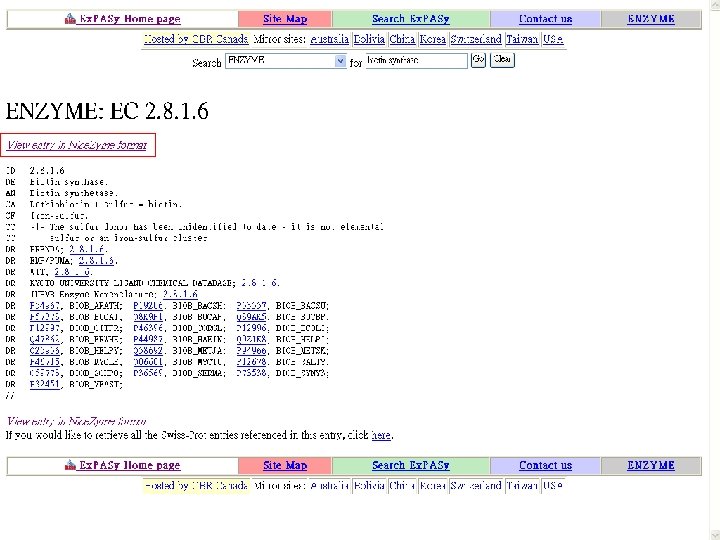
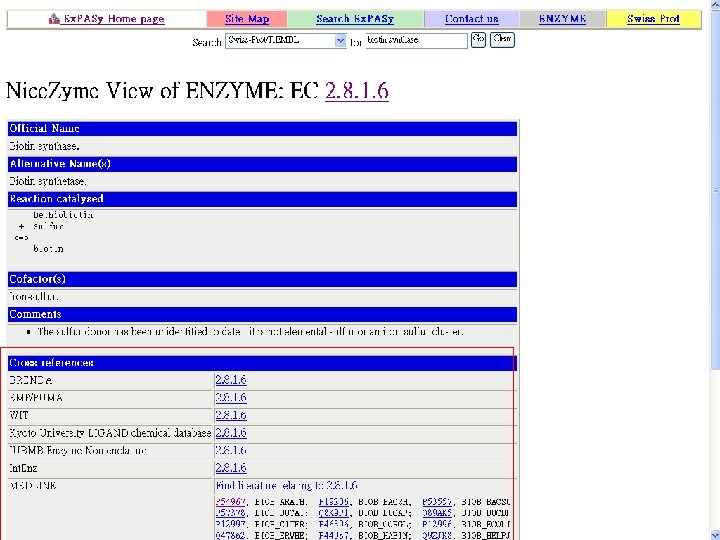
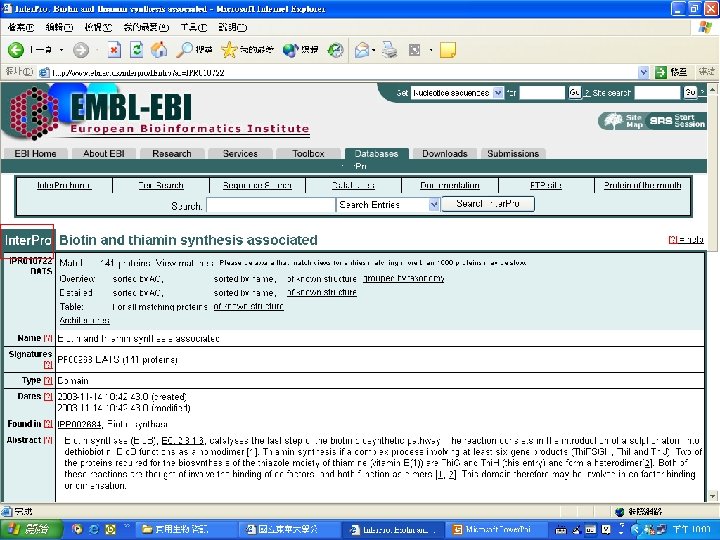
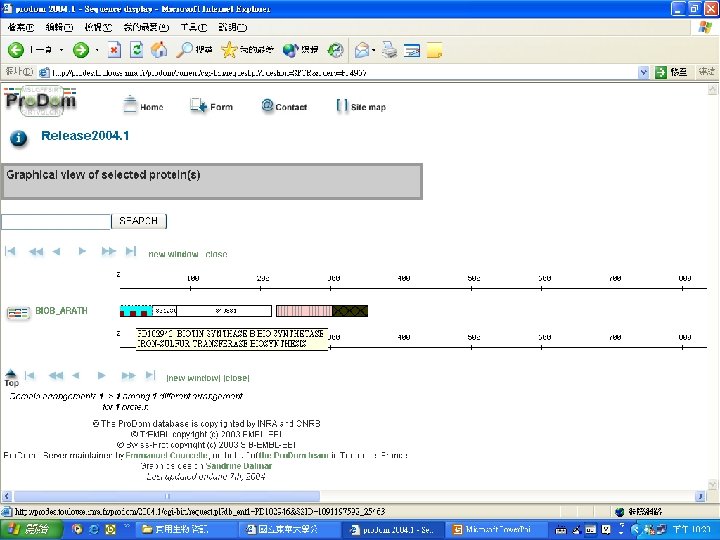
Biotin Synthase Biotin synthase (Bio. B) converts dethiobiotin into biotin by inserting a sulfur atom between C 6 and C 9 of dethiobiotin inan S-adenosylmethionine (SAM)-dependent reaction.
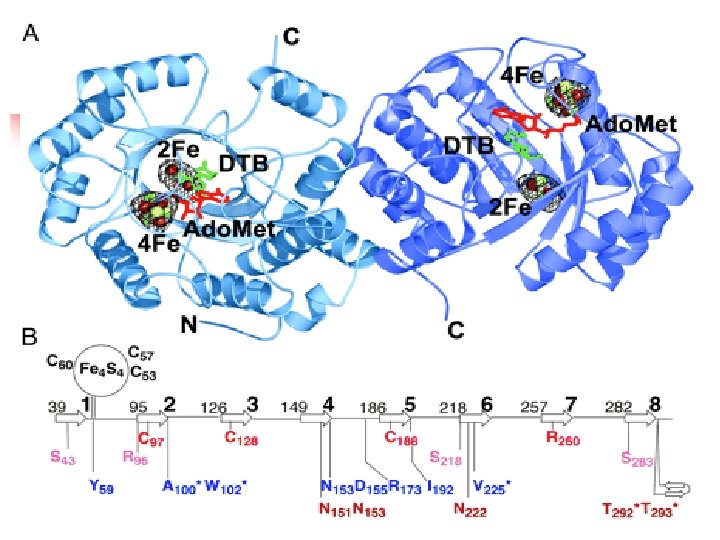
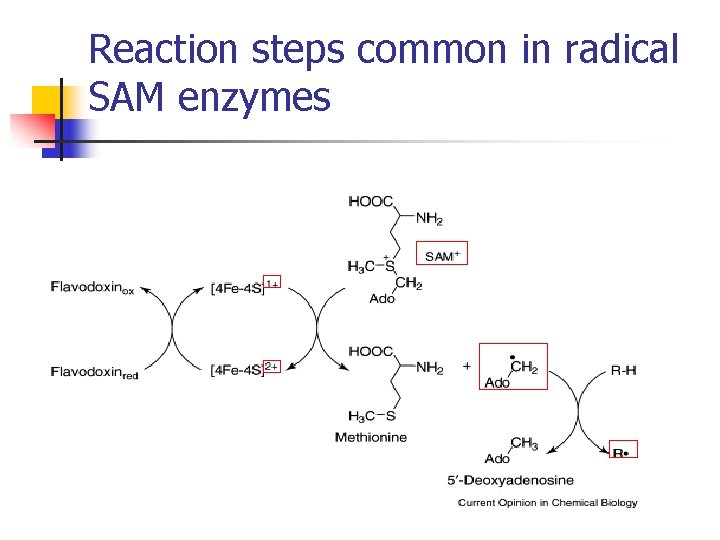
Reaction steps common in radical SAM enzymes


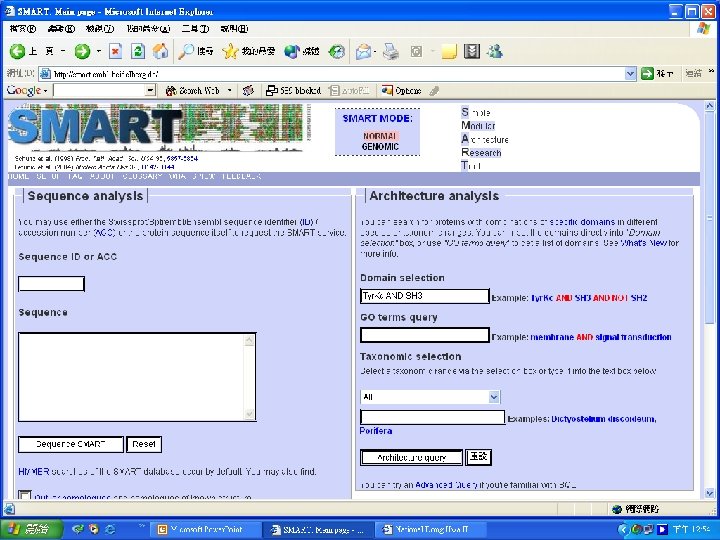
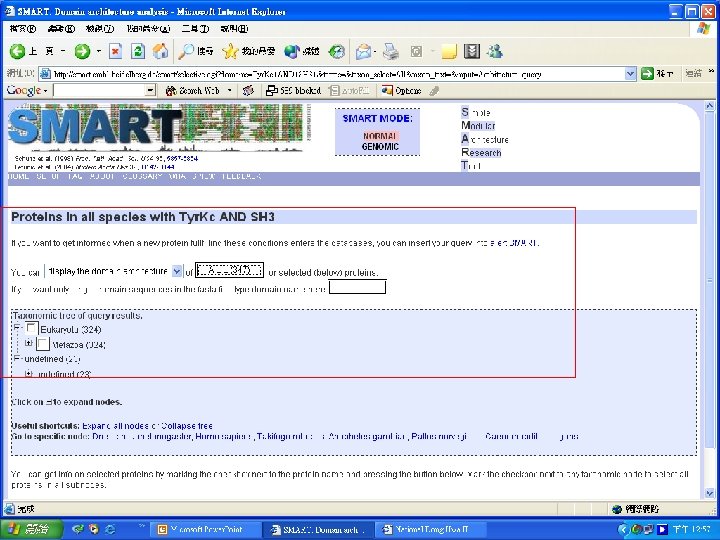
Simple n Modular n Architecture n Research n Tool n


- Slides: 76