Introduction to using the UCSC Genome browser Assembly

Introduction to using the UCSC Genome browser, Assembly Hubs and Table Browser intersections. IBM 029 keypunch BME 230 Winter 2014 11 February 2014 Hiram Clawson http: //genomewiki. ucsc. edu/index. php/User: Hiram Check for a most recent copy of this presentation at: http: //genomewiki. ucsc. edu/index. php/File: BME 230_Winter_2014. ppt

today’s topics: • • UCSC Genome Browser Track/Assembly Hubs Table Browser – example intersection not covered: C language programming with the ‘kent’ source tree

thank you Angie Hinrichs Matt Schwartz Terry Furey Chuck Sugnet Heather Trumbower Kayla Smith Brooke Rhead Ann Zweig Daryl Thomas Robert Baertsch Jakob Peterson Katherine Pollard Adam Siepel Rachel Harte Mark Diekhans Fan Hsu Bob Kuhn Andy Pohl Kate Rosenbloom Galt Barber Jorge Garcia Archana Thakkapallayil Ali Sultan-Qurraie Jennifer Jackson Krishna Roskin Andrew Kern Brian Raney Craig Lowe Yontao Lu Erich Weiler Ryan Weber Gary Moro Todd Lowe Josh Stewart Gill Bejerano Donna Karolchik Jim Kent David Haussler Jim Kent

1. genome browser • • • http: //genome. ucsc. edu/ (89 genomes) 14 primates 33 other mammals 18 other vertebrates 13 flies, mosquito, bee 6 nematodes 5 others, yeast, etc. see also: http: //genome-test. cse. ucsc. edu/ over 100 genomes at last count Just for fun: (search for “gateway” at genomewiki. ucsc. edu) http: //genomewiki. ucsc. edu/index. php/Genome_browser_photo_gateway

see also: • associated lectures: http: //genomewiki. ucsc. edu/index. php/Presentations • Open Helix training: genome. ucsc. edu/training. html • Galaxy analysis workflow management system: galaxy. psu. edu • “Powers of Ten” video by Charles & Ray Eames, 1977 http: //www. youtube. com/watch? v=0 f. KBhv. Djuy 0 • Genome browser documentation: genome. ucsc. edu/golden. Path/help/ • email help: genome at soe. ucsc. edu • genome-source. cse. ucsc. edu/gitweb/? p=kent. git; a=blob; f=src/user. Apps/README

UCSC Genome Browser • • A set of tools, more than just the browser: Genome Browser -> viewing annotations Blat Server -> find sequence Table Browser -> intersections, correlations Gene Sorter -> gene centric viewpoint In Silico PCR -> determine primer products Genome Graphs, Galaxy, Visi. Gene, etc … ~15, 000 users/day ~500, 000 pages/day ~1. 3 Tb/day browser statistics
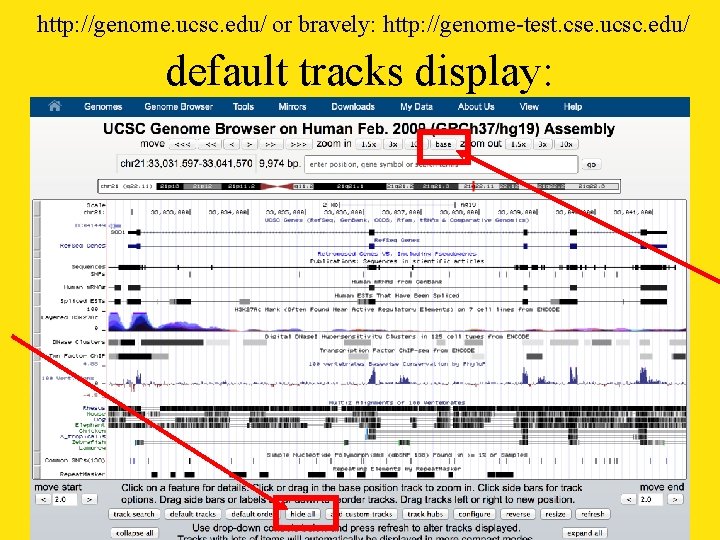
http: //genome. ucsc. edu/ or bravely: http: //genome-test. cse. ucsc. edu/ default tracks display:
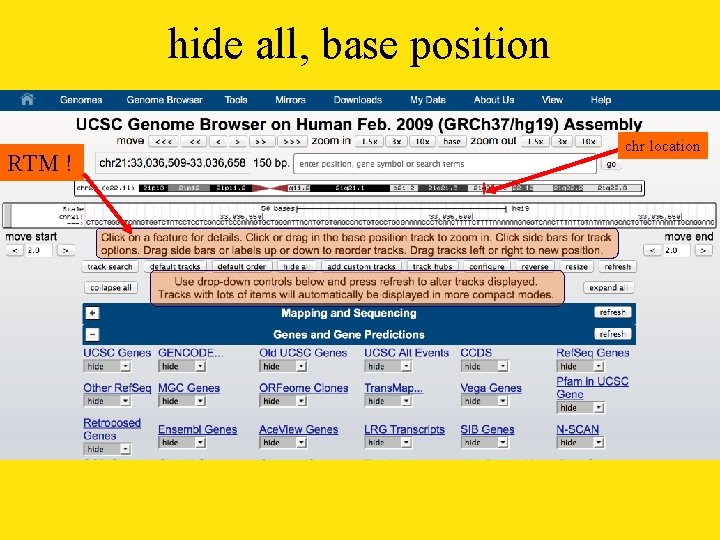
hide all, base position RTM ! chr location
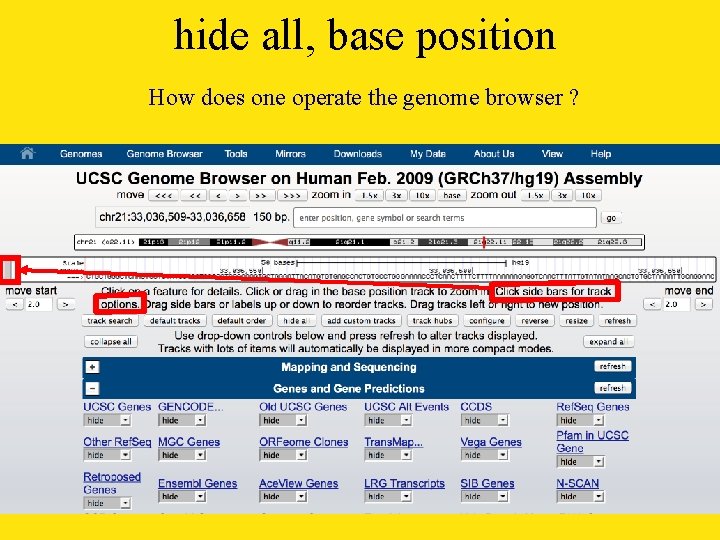
hide all, base position How does one operate the genome browser ?
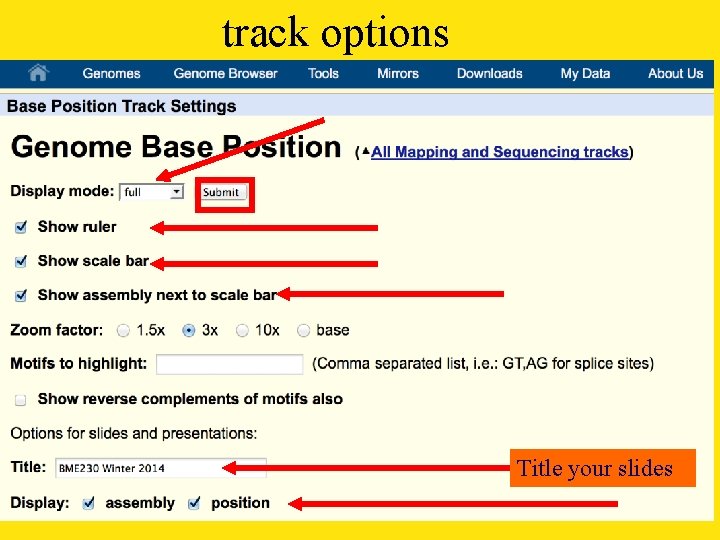
track options Title your slides
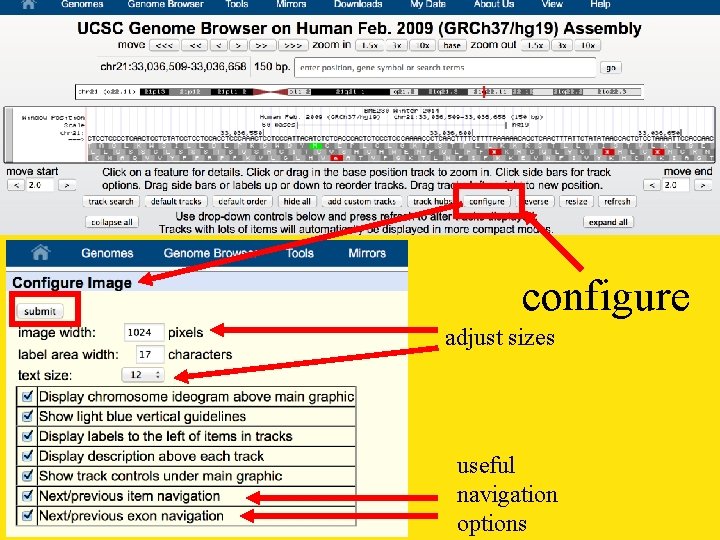
configure adjust sizes useful navigation options

1024 width, 12 pt font readjust to base due to font size change space enough to show bases again
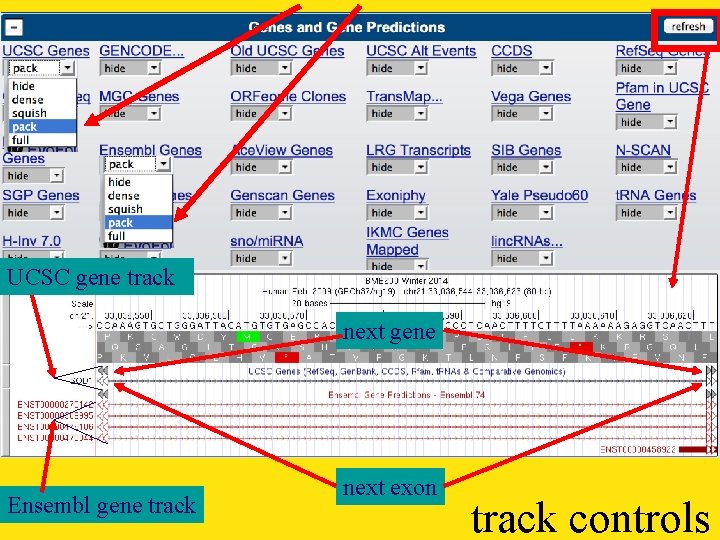
UCSC gene track next gene Ensembl gene track next exon track controls
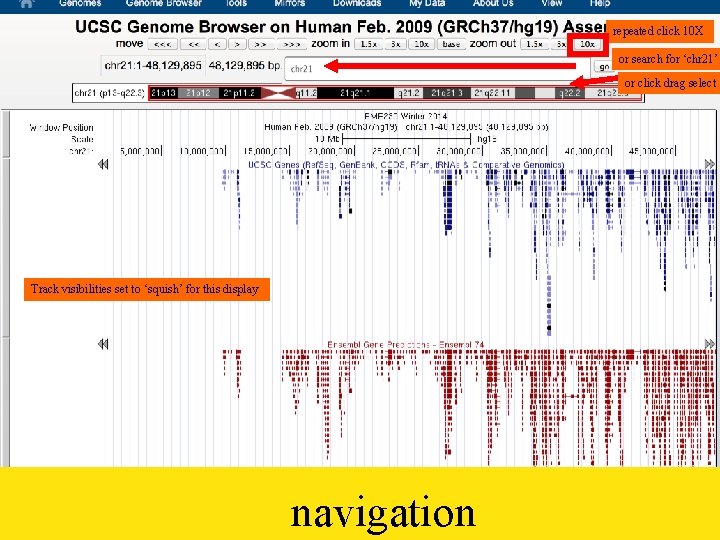
repeated click 10 X or search for ‘chr 21’ or click drag select Track visibilities set to ‘squish’ for this display navigation
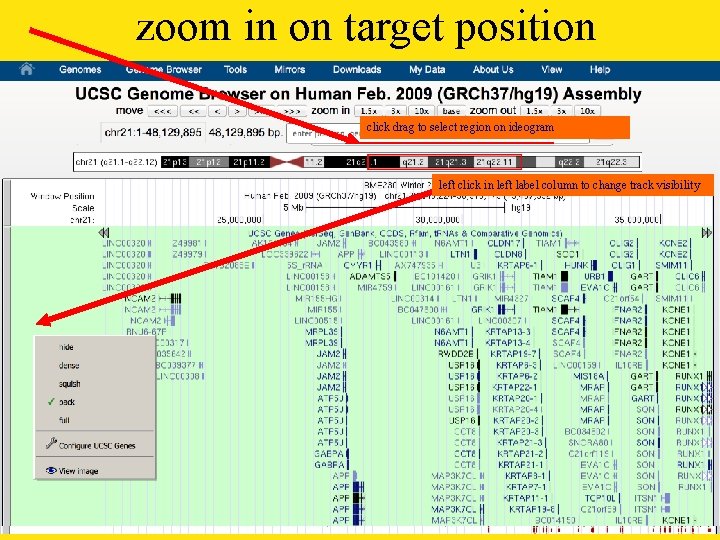
zoom in on target position click drag to select region on ideogram left click in left label column to change track visibility
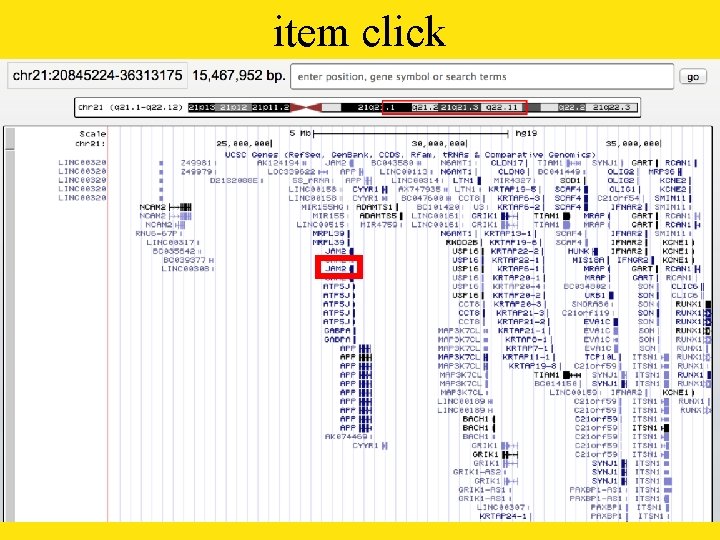
item click
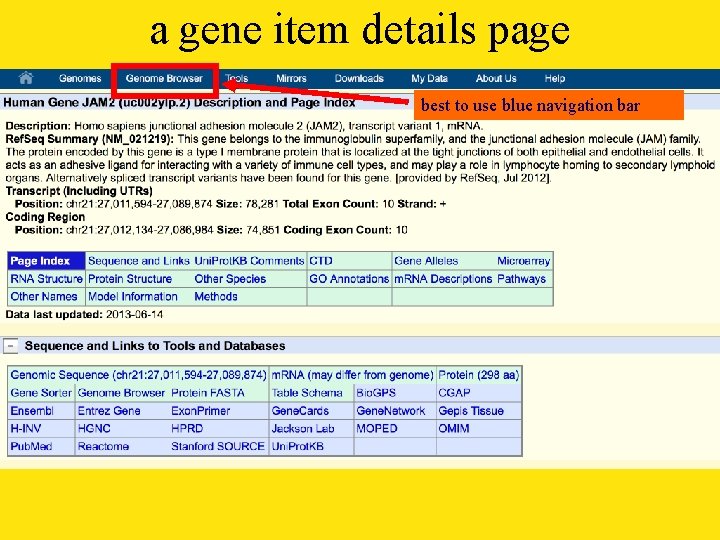
a gene item details page best to use blue navigation bar
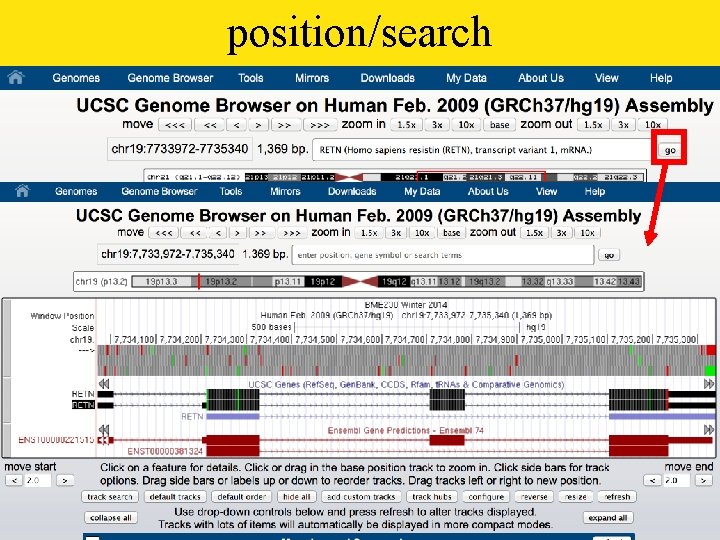
position/search
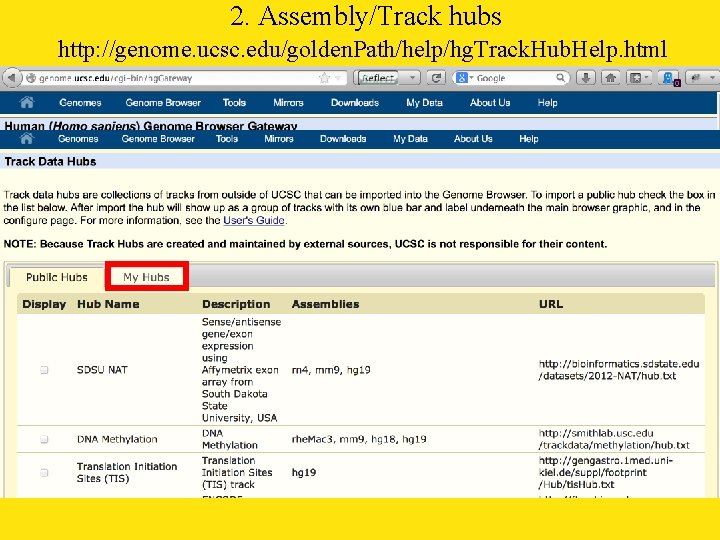
2. Assembly/Track hubs http: //genome. ucsc. edu/golden. Path/help/hg. Track. Hub. Help. html
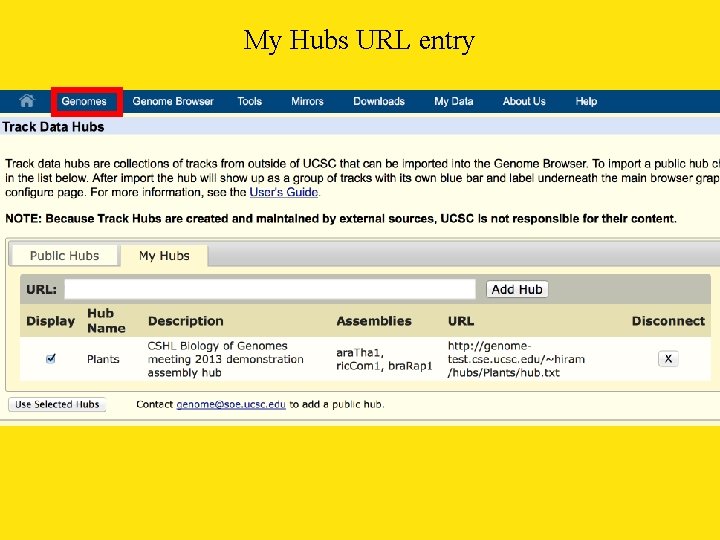
My Hubs URL entry
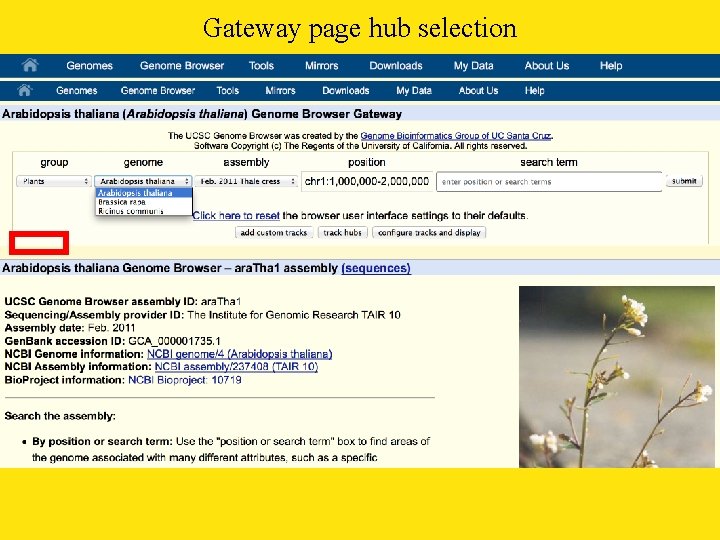
Gateway page hub selection

External genome sequence display
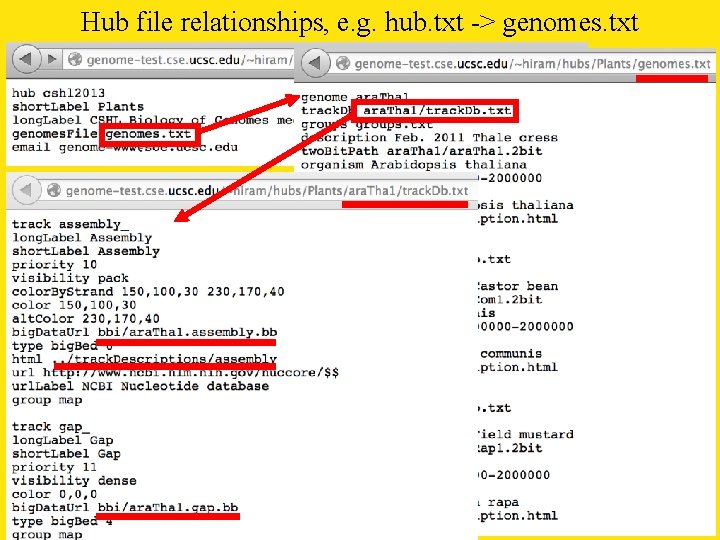
Hub file relationships, e. g. hub. txt -> genomes. txt

3. Table Browser DEC PDP 8/S desktop computer ASR 33 teletype, paper tape punch reader/writer Lear-Siegler ADM 3 A terminal
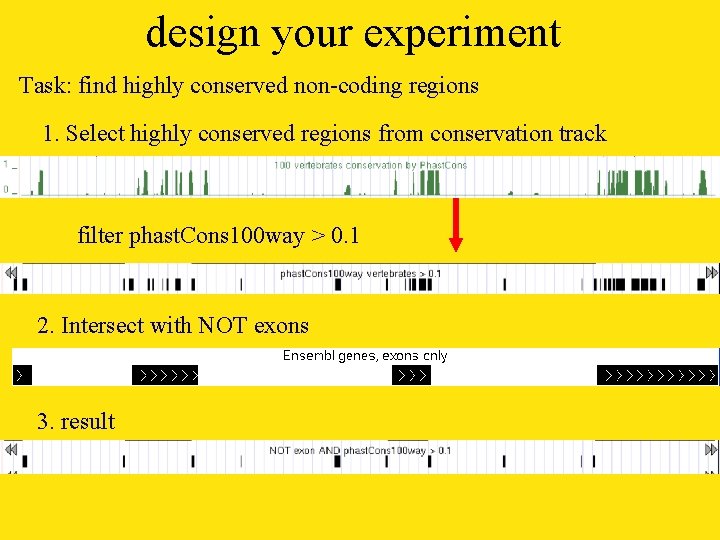
design your experiment Task: find highly conserved non-coding regions 1. Select highly conserved regions from conservation track filter phast. Cons 100 way > 0. 1 2. Intersect with NOT exons 3. result
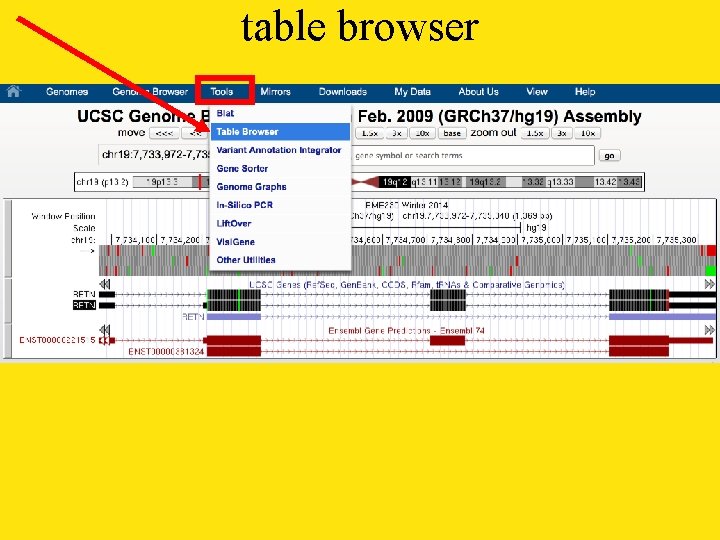
table browser
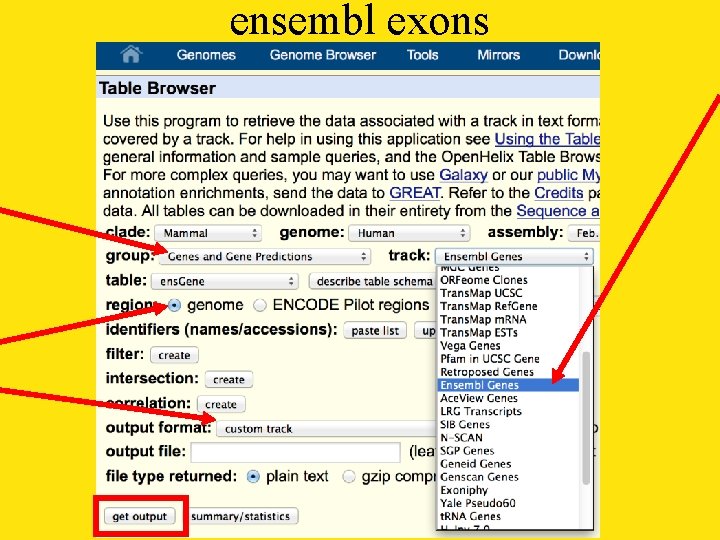
ensembl exons
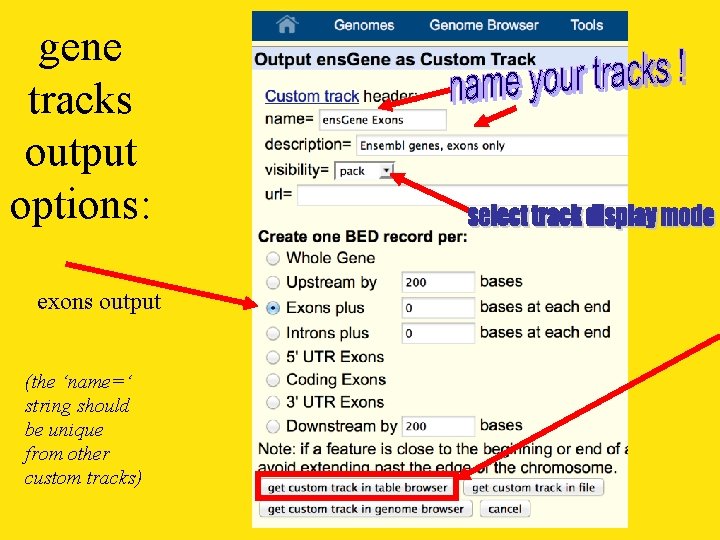
gene tracks output options: exons output (the ‘name=‘ string should be unique from other custom tracks)
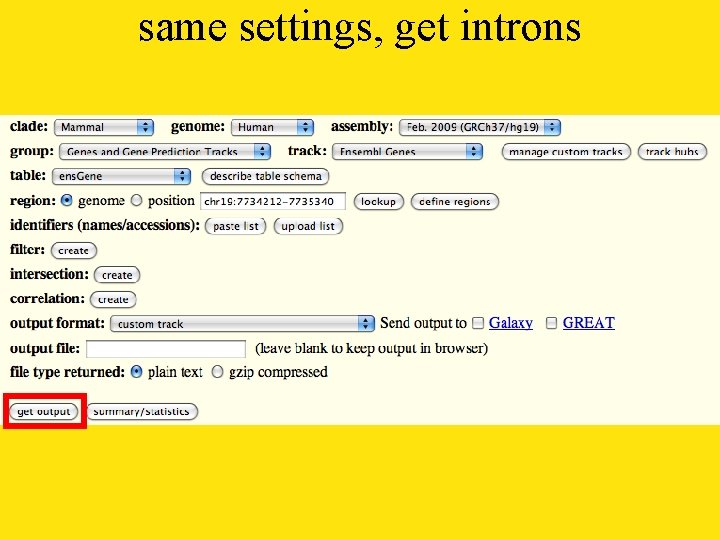
same settings, get introns
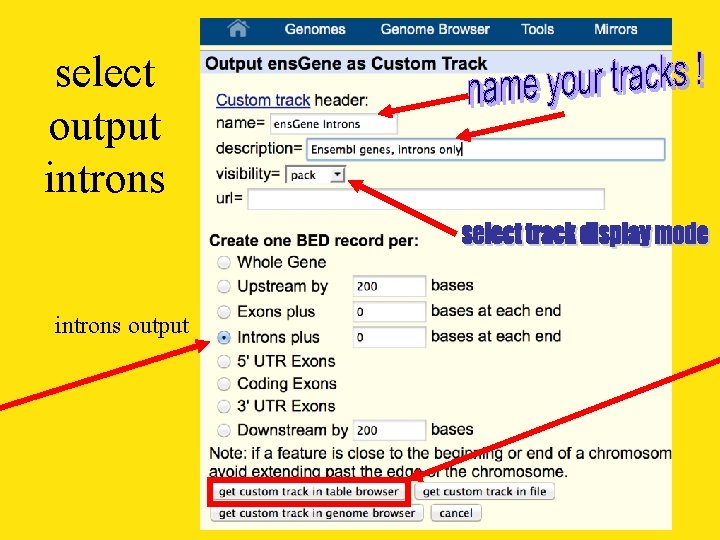
select output introns output
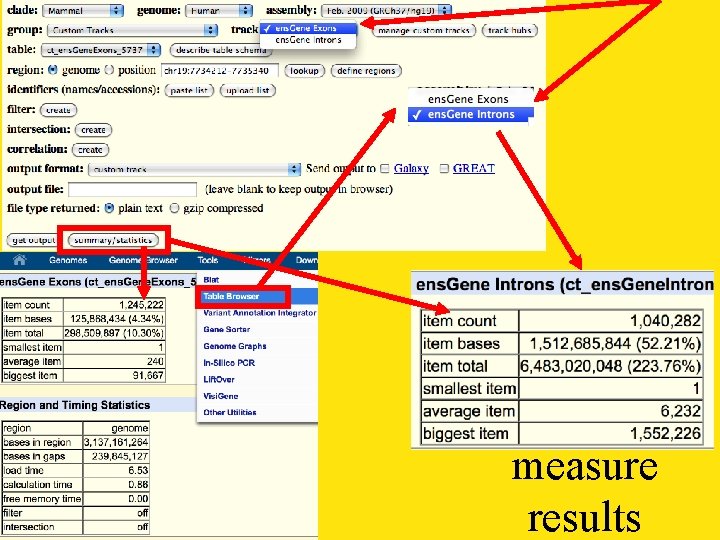
measure results
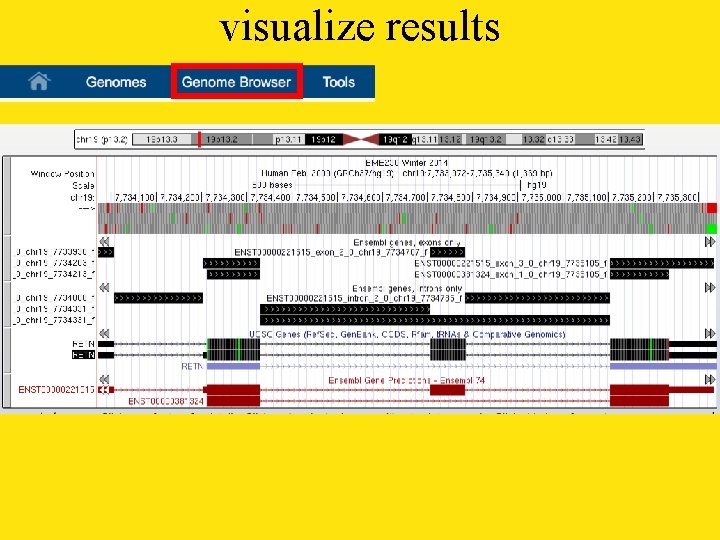
visualize results
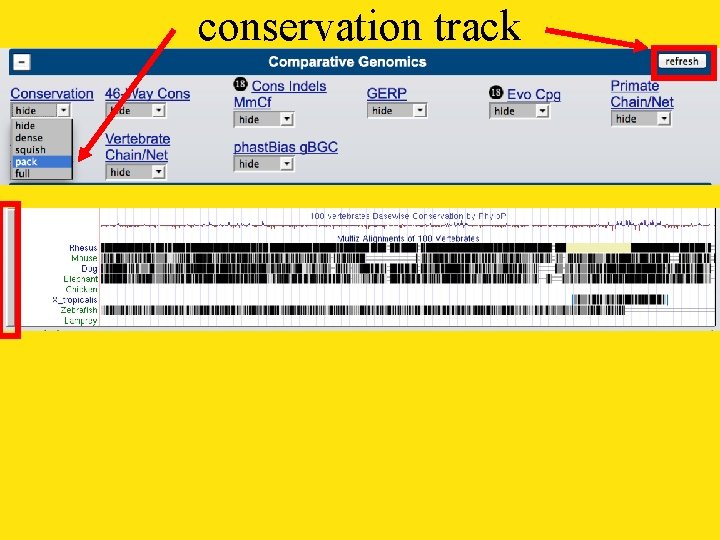
conservation track
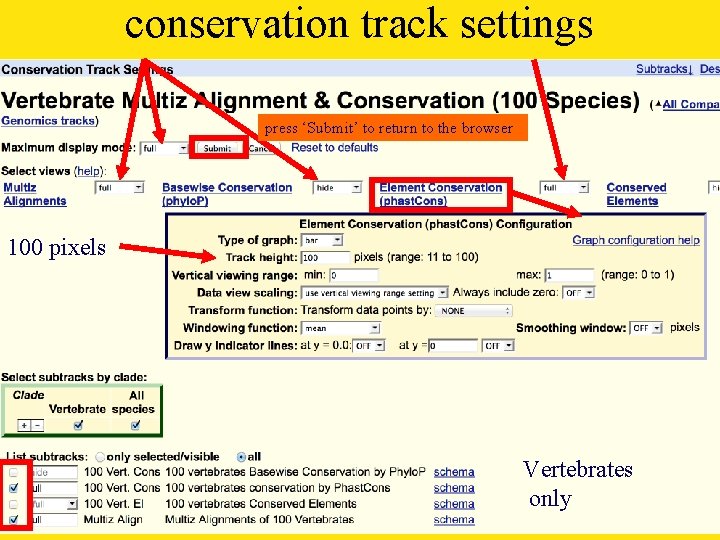
conservation track settings press ‘Submit’ to return to the browser 100 pixels Vertebrates only
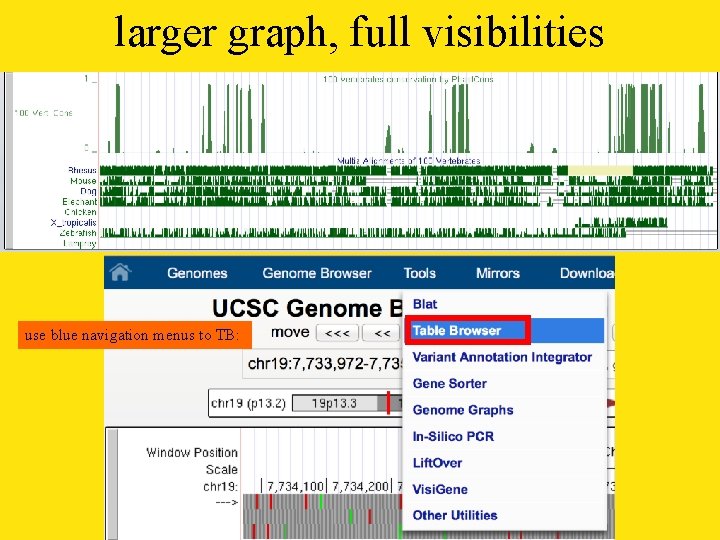
larger graph, full visibilities use blue navigation menus to TB:
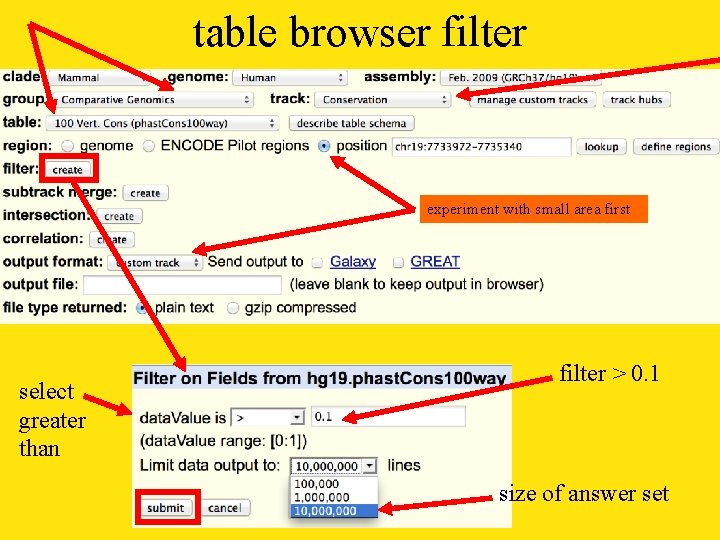
table browser filter experiment with small area first select greater than filter > 0. 1 size of answer set
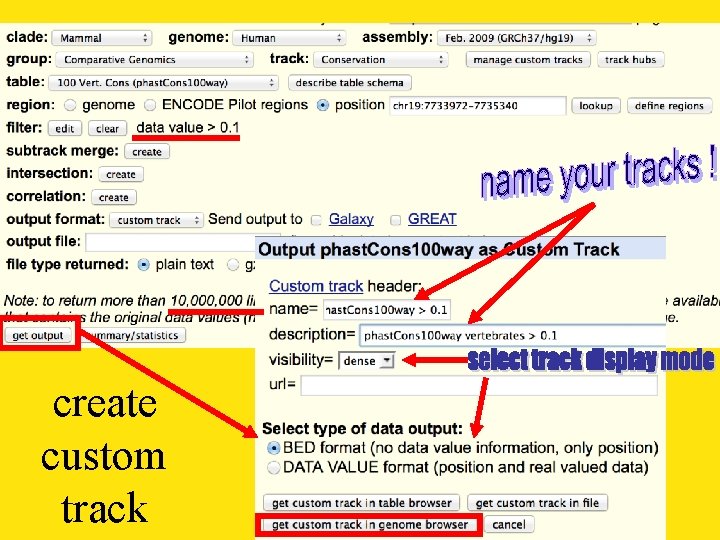
create custom track
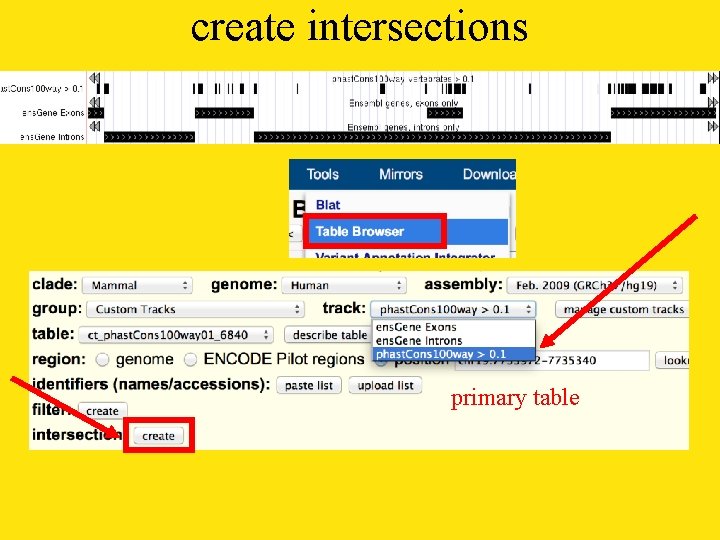
create intersections primary table

secondary table base pair AND function NOT exons
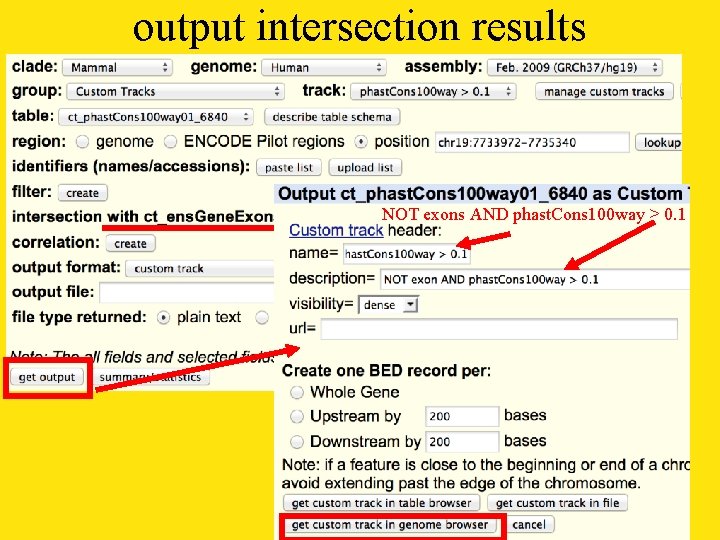
output intersection results NOT exons AND phast. Cons 100 way > 0. 1

and the answer is:

4. C language programming http: //genome-source. cse. ucsc. edu/gitweb/? p=kent. git; a=blob; f=src/user. Apps/README IBM System 360 Model 40, late 1960’s, 256 Kb, 10, 000 FLOPS SWARM cluster, UCSC 2008, 1024 CPU cores, Intel Xeon 2. 3 Ghz

Fetch and build kent source • • • GIT: http: //genome. ucsc. edu/admin/git. html - hourly updated ZIP file: http: //hgdownload. cse. ucsc. edu/admin/jksrc. zip - ~biweekly updated build instructions: http: //genome. ucsc. edu/admin/jk-install. html see also, in the source tree kent/src/product/README. * BME 230 Winter 2011 presentation (similar to this one) http: //genomewiki. ucsc. edu/index. php/File: BME 230_Winter_2011. ppt • Angie’s presentation of this material: http: //genomewiki. ucsc. edu/index. php/Image: Bejerano_Lab_2008_03_31. ppt • • Jim Kent source tree design discussions, several at: http: //genomewiki. ucsc. edu/index. php/Presentations BME 230 Winter 2008 presentation: http: //genomewiki. ucsc. edu/index. php/Image: Baertsch-code-talk. ppt The presentations mentioned at the genomewiki are complementary to this presentation. Please take a look at them.

source tree organization kent/src/ - top level - see README here kent/src/inc/*. h - API manual for common functions kent/src/utils/ - file manipulation utilities kent/src/lib/$MACHTYPE/ - built libraries kent/src/lib/ - builds lib jkweb. a kent/src/hg/ - database manipulation commands (mostly) kent/src/hg/lib/ - builds lib jkhgap. a kent/src/jk. Own. Lib/ - blat support – COPYRIGHT © J. Kent see also www. kentinformatics. com

using the database utilities 1. Create public My. SQL user/password access file: $ cd $HOME $ cat << ‘_EOF_’ >. hg. conf db. host=genome-mysql. cse. ucsc. edu db. user=genomep db. password=password ‘_EOF_’ $ chmod 600. hg. conf # will not work without this security 2. That file is used by kent src commands to access My. SQL: $ feature. Bits hg 19 gap 239845127 bases of 2897316137 (8. 278%) in intersection (try this same command with the argument -count. Gaps)

writing a new command Use ‘new. Prog’ to establish the skeleton of a new command: $ new. Prog - make a new C source skeleton. usage: new. Prog prog. Name description words This will make a directory 'prog. Name' and a file in it 'prog. Name. c' with a standard skeleton Options: -jkhgap - include jkhgap. a and mysql libraries as well as jkweb. a archives -cgi - create shell of a CGI script for web -cvs - obsolete option, needs to update to git commands (note: this works only within the source tree hierarchy)

command line arguments #include "common. h" // see also: src/inc/common. h "options. h" // see also: src/inc/options. h static long offset = 0; static char * chrom = NULL; static double min. Val = 0; static double max. Val = BIGNUM; static boolean do. Nothing = FALSE; static struct option. Specs[] = { {"offset", OPTION_LONG}, {"chrom", OPTION_STRING}, {"min. Val", OPTION_DOUBLE}, {"max. Val", OPTION_DOUBLE}, {"do. Nothing", OPTION_BOOLEAN}, {NULL, 0} };

command line argument processing #include "common. h" // see also: src/inc/common. h "options. h" // see also: src/inc/options. h int main( int argc, char *argv[] ) /* program demonstrates fetching options from the command line */ { … offset = option. Long("offset", 0); chrom = option. Val("chrom", NULL); min. Val = option. Double("min. Val", -1 * INFINITY); max. Val = option. Double("max. Val", INFINITY); do. Nothing = option. Exists(”do. Nothing"); … exit(0); }

line oriented file I/O #include "common. h" // see also: src/inc/common. h #include "linefile. h" // see also: src/inc/linefile. h void some. Function(char *file. Name) /* function demonstrates reading a file line by line */ { struct line. File *lf = line. File. Open(file. Name, TRUE); char * line = NULL; int size = 0; while (line. File. Next(lf, &line, &size)) { char *words[128]; int word. Count = chop. By. White(line, words, Array. Size(words)); } line. File. Close(&lf); }

structured file I/O #include "common. h" #include "maf. h" // see also: src/inc/common. h // see also: src/inc/maf. h void scan. Maf(char *maf. File) /* function demonstrates reading a MAF file, record by record */ { struct maf. File *mf = maf. Open(maf. File); struct maf. Ali *ali = NULL; while ((ali = maf. Next(mf)) != NULL) { int c. Count = sl. Count(ali->components); … processing maf alignment record … maf. Ali. Free(&ali); // need to free the maf record structure } // maf. Next() just happens to close the file on the last record } // see also: src/hg/mouse. Stuff/maf. Coverage/

reading DNA sequence files Example is from src/utils/find. Motif. c Include files would be, from src/inc/ common. h dnaseq. h dna. Load. h These functions read fasta files, nib files, or 2 bit files. DNA sequence is any of: ACGTacgt. Nn lower case is ‘masked’ static void find. Motif(char *input) /* find. Motif - find specified motif in sequence file. */ { struct dna. Load *dl = dna. Load. Open(input); struct dna. Seq *seq; while ((seq = dna. Load. Next(dl)) != NULL) { verbose(2, "#tprocessing: %sn", seq->name); DNA *dna = seq->dna; for (i=0; i < seq->size; ++i) { val = nt. Val[(int)dna[i]]; switch (val) { case T_BASE_VAL: case C_BASE_VAL: case A_BASE_VAL: case G_BASE_VAL: … etc … } } }

memory management #include "common. h" // see also: src/inc/common. h #include "memalloc. h" // see also: src/inc/memalloc. h #include "bed. h" // see also: src/hg/inc/bed. h void some. Function(int count) /* demonstrate use of need. Mem(), free. Mem(), Alloc. Var() */ { int *pile. Of. Ints = need. Mem(sizeof(int) * count); … use ‘em or lose ‘em … free. Mem(pile. Of. Ints); // or: freez(&pile. Of. Ints); struct bed *bed. Item; Alloc. Var(bed. Item); // macro defined in src/inc/common. h bed. Item->chrom = clone. String("chr 23"); … etc … }

linked lists #include "common. h" struct sl. List { struct sl. List *next; } // see also: src/inc/common. h int sl. Count(void *list) /* count elements in list. */ { struct sl. List *pt = (struct sl. List *)list; int len = 0; while (pt != NULL) // or more concisely: for ( ; pt = pt->next) { // ++len; len += 1; pt = pt->next; } return len; } HP 2100 A mini computer

sorting a linked list #include "common. h" // see also: src/inc/common. h #include ”bed. h" // see also: src/inc/bed. h int bed. Cmp(const void *va, const void *vb) /* Compare to sort based on chrom, chrom. Start. */ { const struct bed *a = *((struct bed **)va); const struct bed *b = *((struct bed **)vb); int dif; dif = strcmp(a->chrom, b->chrom); if (dif == 0) dif = a->chrom. Start - b->chrom. Start; return dif; } /* somewhere in code later */ struct bed *bed. List; // a linked list of bed items sl. Sort(&bed. List, bed. Cmp) // voilà, sorted IBM 082 card sorter

inheritance via C structures struct axt. Score. Scheme /* A scoring scheme or DNA alignment. */ { struct score. Matrix *next; int matrix[256]; /* Look up with letters. */ int gap. Open; /* Gap open cost. */ int gap. Extend; /* Gap extension. */ char *extra; /* extra parameters */ }; This structure “inherits” the methods of the sl. List structure Because the first element(s) in the structure are the same as in struct sl. List --> ‘*next’. Therefore, sl. Count() will work on this list too.

hashed storage structures #include "common. h" #include "hash. h" // see also: src/inc/common. h // see also: src/inc/hash. h struct hash. El *hash. Add(struct hash *hash, char *name, void *val); /* Add new element to hash table. If an item with name, already exists, a new * item is added in a LIFO manner. The last item added for a given name is * the one returned by the hash. Lookup functions. hash. Lookup. Next must be used * to find the preceding entries for a name. */ The given ‘name’ becomes the key to this item. The item to save in the hash is in the pointer ‘val’, which can be any type of structured item.

creating a hash #include "common. h" // see also: src/inc/common. h #include "hash. h" // see also: src/inc/hash. h #include "hg. Tracks. h" // see also: src/hg/hg. Tracks. h DEC PDP 11/40 struct maf. Item *some. Function(struct track *tg) /* create a hash of maf. Items from tg->items simple list */ { struct maf. Item *mi. List = tg->items, *mi; struct hash *mi. Hash = new. Hash(9); /* Make hash of items keyed by database. */ int i = 0; for (mi = mi. List; mi != NULL; mi = mi->next) { mi->ix = i++; if (mi->db != NULL) hash. Add(mi. Hash, mi->db, mi); } return mi. Hash; }

scanning through a hash #include "common. h" #include "hash. h" // see also: src/inc/common. h // see also: src/inc/hash. h void hash. Verify(struct hash *bed. Name. Hash) /* verify that bed element name is identical to hash name */ { struct hash. Cookie cookie; struct hash. El *el; cookie = hash. First(bed. Name. Hash); while ((el = hash. Next(&cookie)) != NULL) { struct bed *bed. El = (struct bed *)el->val; if (different. String(bed. El->name, el->name)) err. Abort("hash broken '%s' != ‘'%s'", bed. El->name, el->name); } }

dynamic strings #include "common. h" #include "hash. h" #include "dystring. h" // see also: src/inc/common. h // see also: src/inc/hash. h // see also: src/inc/dystring. h struct dy. String *concat. Names(struct hash *bed. Name. Hash) /* return comma delimited string of all bed element names */ { struct hash. Cookie cookie; struct hash. El *el; cookie = hash. First(bed. Name. Hash); struct dy. String *big. String = dy. String. New(256); while ((el = hash. Next(&cookie)) != NULL) { struct bed *bed. El = (struct bed *)el->val; dy. String. Printf(big. String, "%s, ", bed. El->name, ); } return big. String; }

thank you Angie Hinrichs Matt Schwartz Terry Furey Chuck Sugnet Heather Trumbower Kayla Smith Brooke Rhead Ann Zweig Daryl Thomas Robert Baertsch Jakob Peterson Katherine Pollard Adam Siepel Rachel Harte Mark Diekhans Fan Hsu Bob Kuhn Andy Pohl Kate Rosenbloom Galt Barber Jorge Garcia Archana Thakkapallayil Ali Sultan-Qurraie Jennifer Jackson Krishna Roskin Andrew Kern Brian Raney Craig Lowe Yontao Lu Erich Weiler Ryan Weber Gary Moro Todd Lowe Josh Stewart Gill Bejerano Donna Karolchik Jim Kent David Haussler Jim Kent
- Slides: 60