The UCSC Genome Browser and other UCSC web

The UCSC Genome Browser …and other UCSC web tools Angie Hinrichs angie@soe. ucsc. edu UCSC Genome Bioinformatics Group May 9, 2005 UCSC Genome Browser 1

Outline n n UCSC web tools overview Genome Browser usage Quick usage examples How to learn more May 9, 2005 UCSC Genome Browser 2

http: //genome. ucsc. edu/ n n Visualizing annotations: Genome Browser Sequence search: BLAT, is. PCR Batch querying: Table Browser There’s more! May 9, 2005 UCSC Genome Browser 3
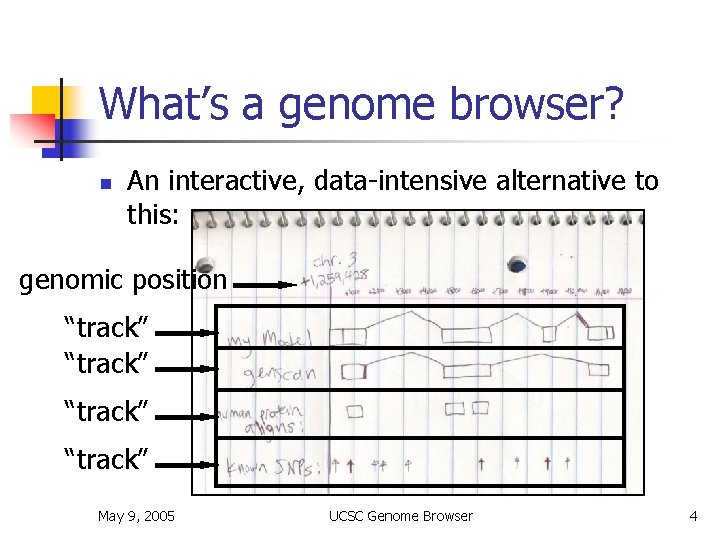
What’s a genome browser? n An interactive, data-intensive alternative to this: genomic position “track” May 9, 2005 UCSC Genome Browser 4
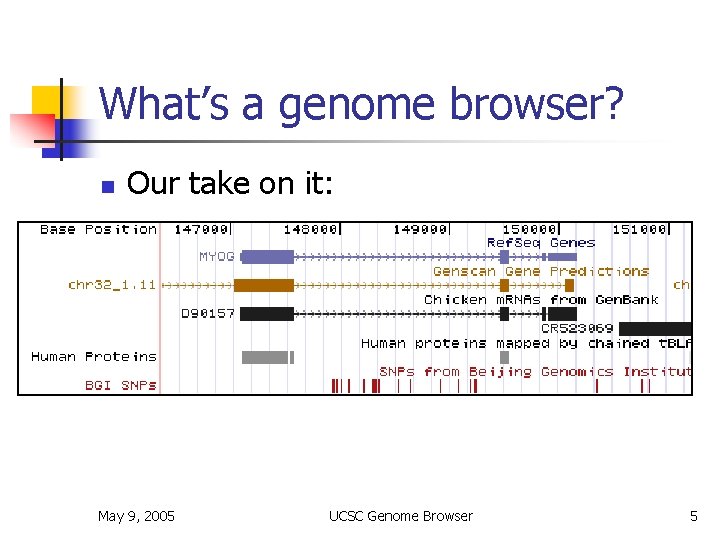
What’s a genome browser? n Our take on it: May 9, 2005 UCSC Genome Browser 5

What can I do with it? n n Align your sequence, look for homologs, conservation, c. DNA evidence, . . . Click through to another species’ homologous region Upload a custom track with locations of your features, compare with other annotations Download a piece (or all!) of the genome sequence/annotations May 9, 2005 UCSC Genome Browser 6

Genome Browser Features n n n Visually rich and speedy display of diverse types of data, organized by genomic position Provides access to more details, links out to other sites Broad gene/accession/keyword search May 9, 2005 UCSC Genome Browser 7

Annotation Tracks from the Chicken Community n n n n BGI: SNPs, coverage, genes Ensembl: genes Gene predictions: Twinscan (WUSTL), SGP and Geneid (IMIM) Sanger Inst. : RNA Genes Genoscope: Tetraodon Ecores PSU: Isochores Got data? May 9, 2005 UCSC Genome Browser 8

http: //genome. ucsc. edu/ May 9, 2005 UCSC Genome Browser 9
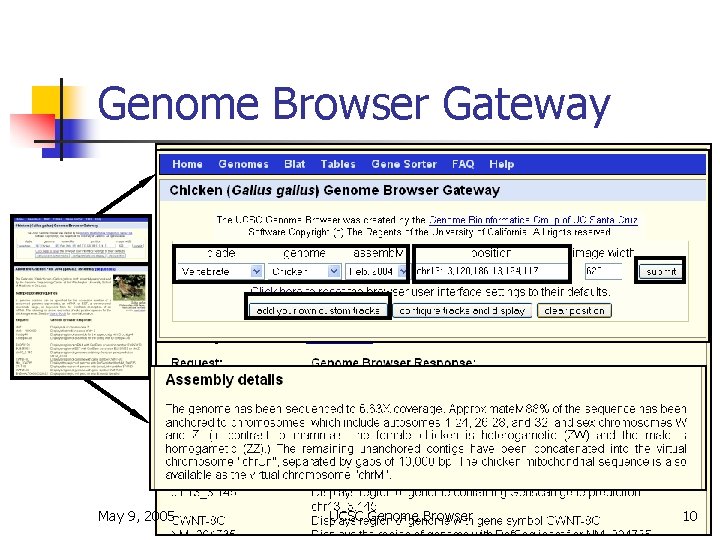
Genome Browser Gateway May 9, 2005 UCSC Genome Browser 10
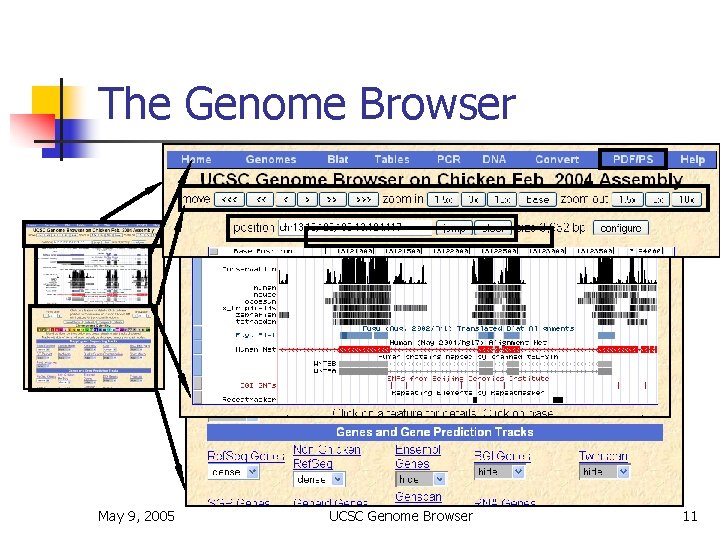
The Genome Browser May 9, 2005 UCSC Genome Browser 11
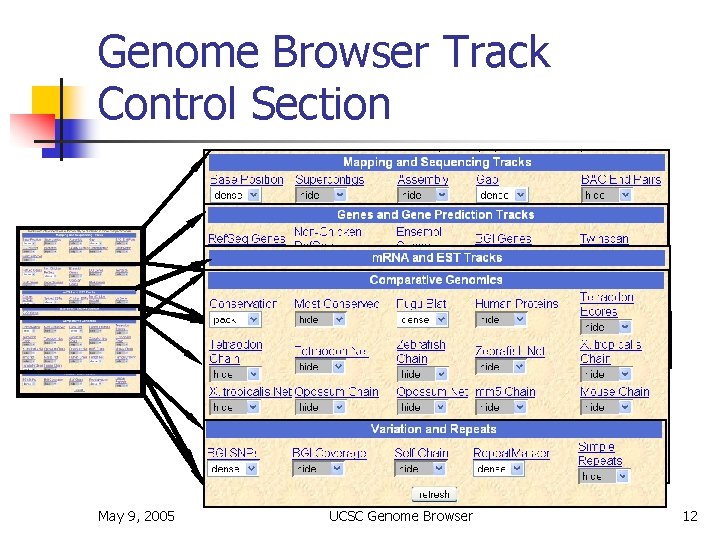
Genome Browser Track Control Section May 9, 2005 UCSC Genome Browser 12
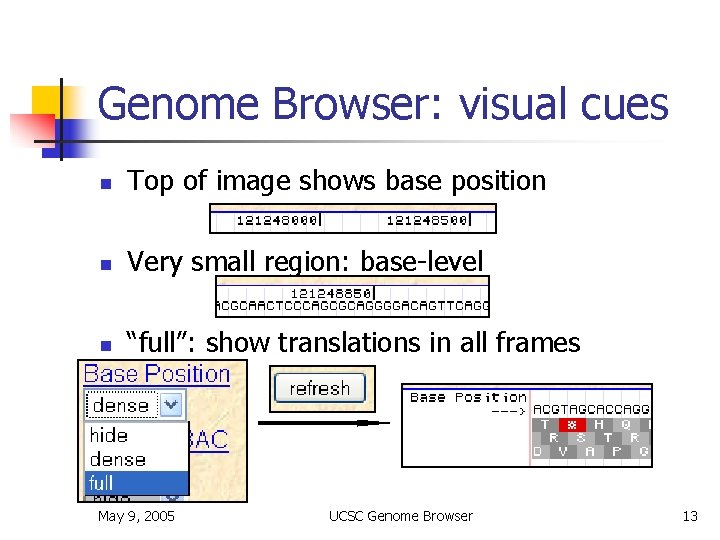
Genome Browser: visual cues n Top of image shows base position n Very small region: base-level n “full”: show translations in all frames May 9, 2005 UCSC Genome Browser 13

Genome Browser: visual cues n n n simple feature: box or tickmark default visibility: nameless features collapsed to one row click to expand: named features, multiple rows May 9, 2005 UCSC Genome Browser 14

Genome Browser: visual cues n Genes: n n Boxes are exons Lines are introns; arrows show direction of transcription Tall/thick boxes are CDS Thin/short boxes are UTR May 9, 2005 UCSC Genome Browser 15

Genome Browser: visual cues n n shade can indicate score or status comparative tracks: color key for other species’ chromosomes May 9, 2005 UCSC Genome Browser 16
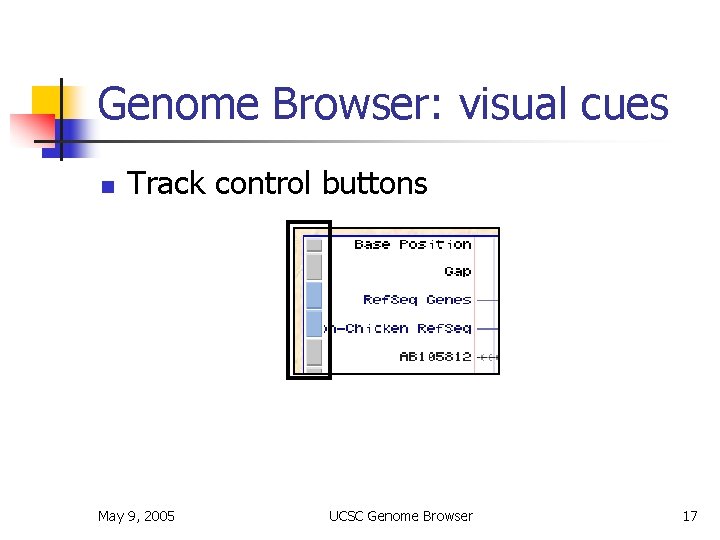
Genome Browser: visual cues n Track control buttons May 9, 2005 UCSC Genome Browser 17
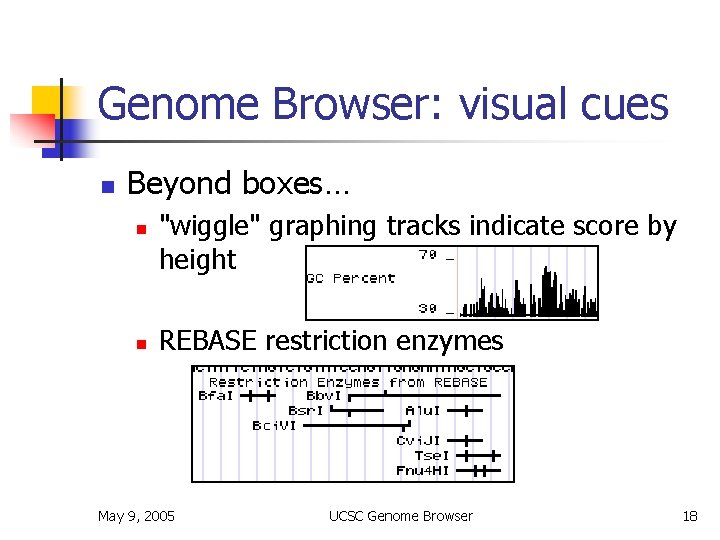
Genome Browser: visual cues n Beyond boxes… n n "wiggle" graphing tracks indicate score by height REBASE restriction enzymes May 9, 2005 UCSC Genome Browser 18
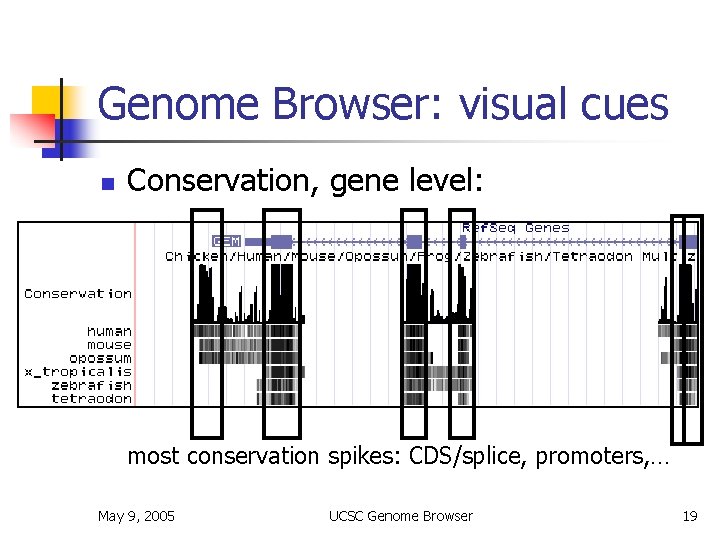
Genome Browser: visual cues n Conservation, gene level: n most conservation spikes: CDS/splice, promoters, … May 9, 2005 UCSC Genome Browser 19

Genome Browser: visual cues n Conservation, base level: May 9, 2005 UCSC Genome Browser 20
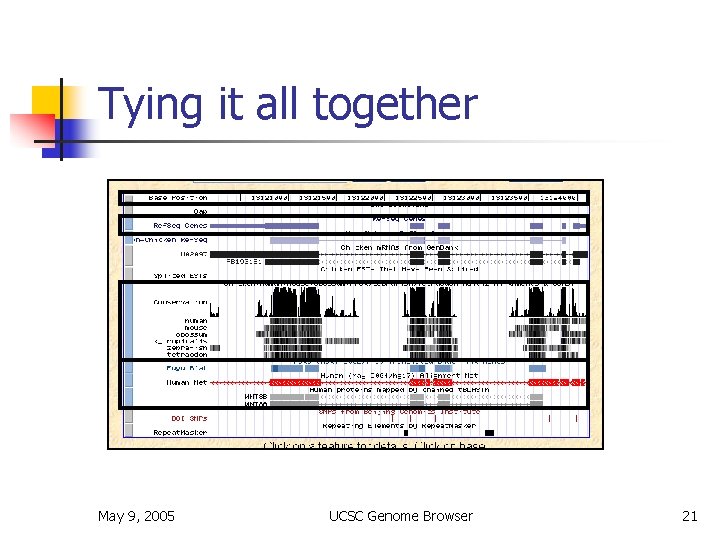
Tying it all together May 9, 2005 UCSC Genome Browser 21
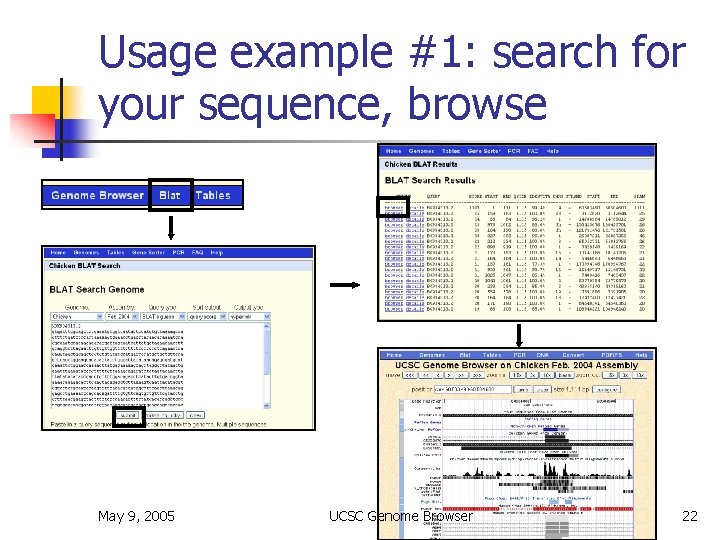
Usage example #1: search for your sequence, browse May 9, 2005 UCSC Genome Browser 22

Usage example #2: click through to other species (part 1 of 3) May 9, 2005 UCSC Genome Browser 23
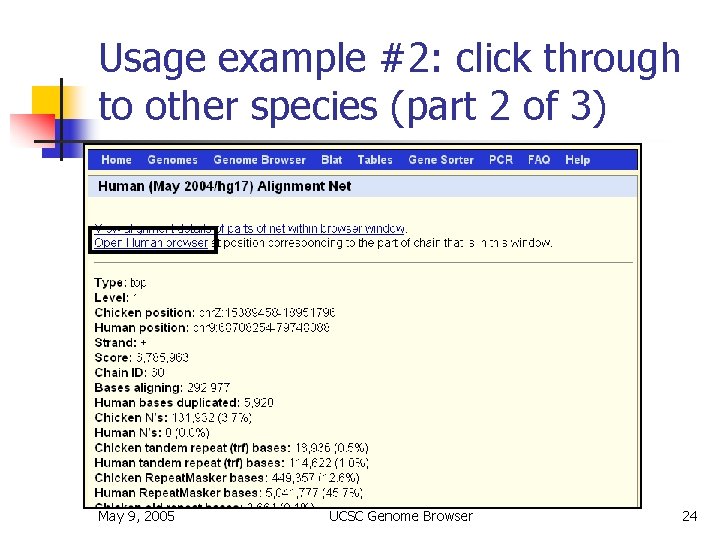
Usage example #2: click through to other species (part 2 of 3) May 9, 2005 UCSC Genome Browser 24

Usage example #2: click through to other species (part 3 of 3) May 9, 2005 UCSC Genome Browser 25

Usage example #3: get upstream sequence for gene May 9, 2005 UCSC Genome Browser 26
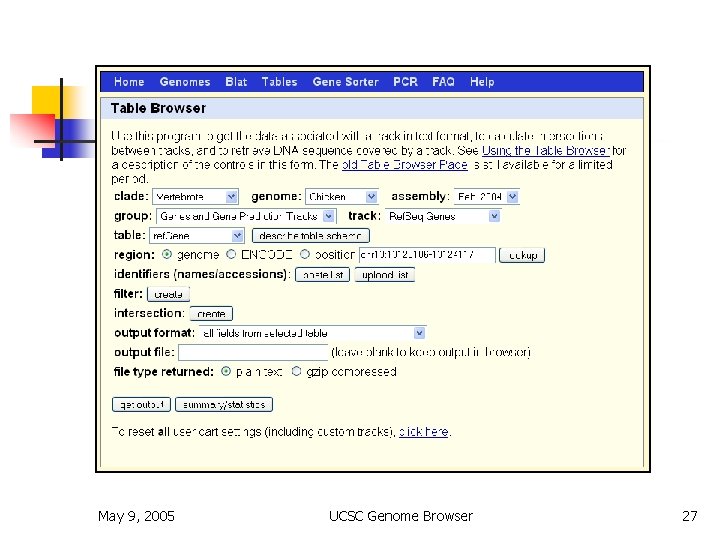
Table Browser n n n bulk download or batch query filtering by values intersection by position joining by related columns various output formats May 9, 2005 UCSC Genome Browser 27
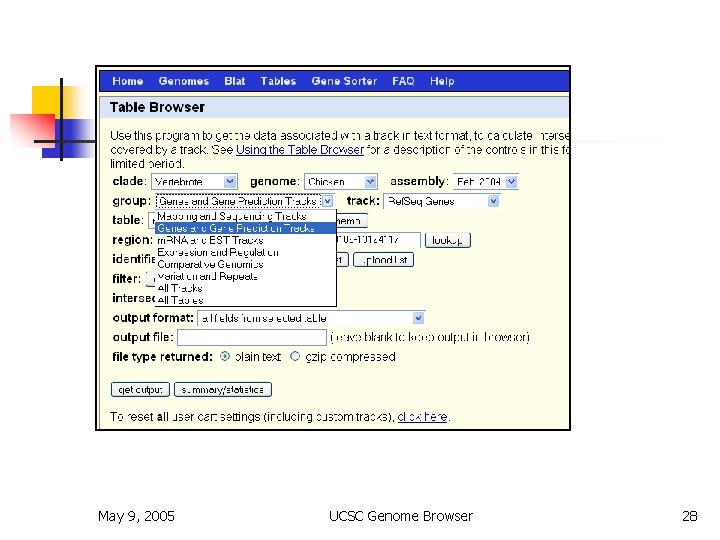
Table Browser May 9, 2005 UCSC Genome Browser 28
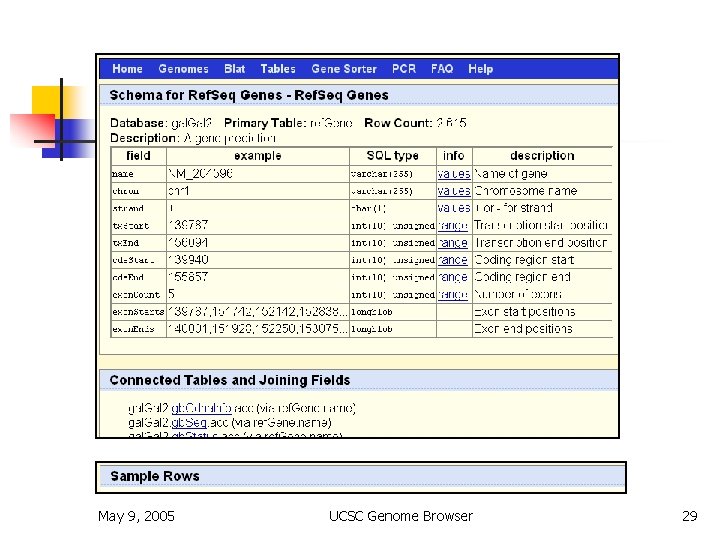
Table Browser May 9, 2005 UCSC Genome Browser 29
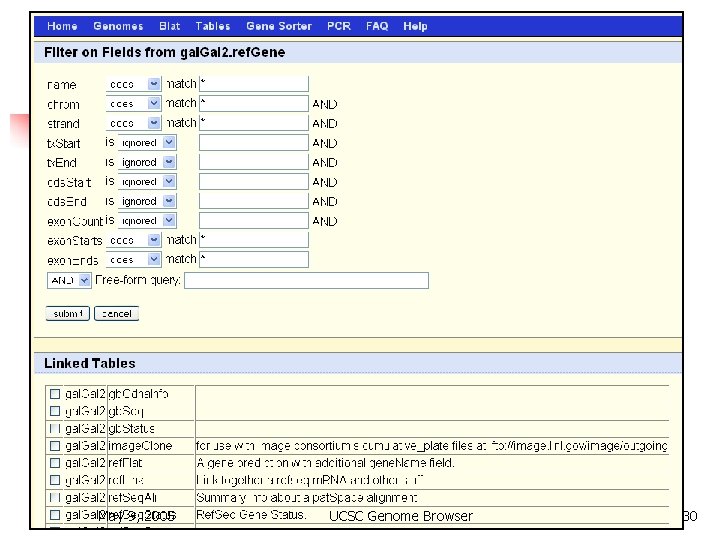
Table Browser May 9, 2005 UCSC Genome Browser 30
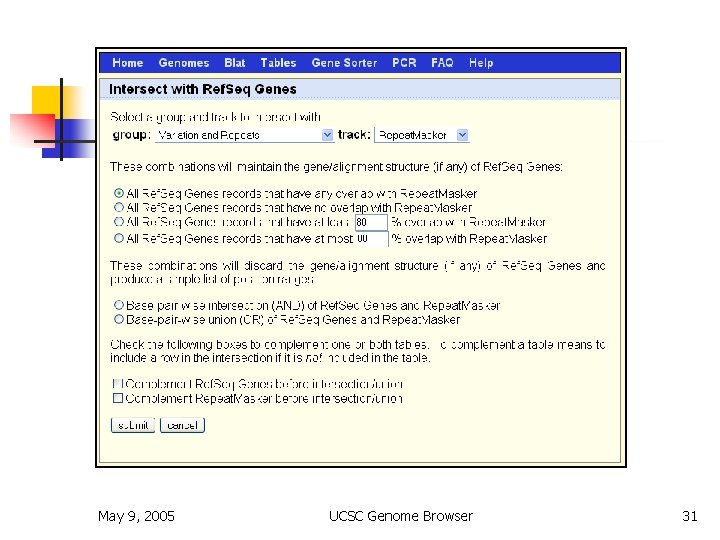
Table Browser May 9, 2005 UCSC Genome Browser 31
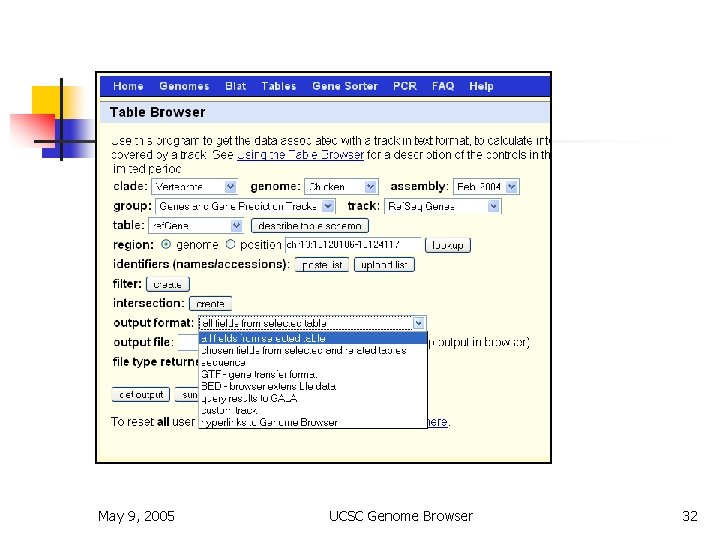
Table Browser May 9, 2005 UCSC Genome Browser 32
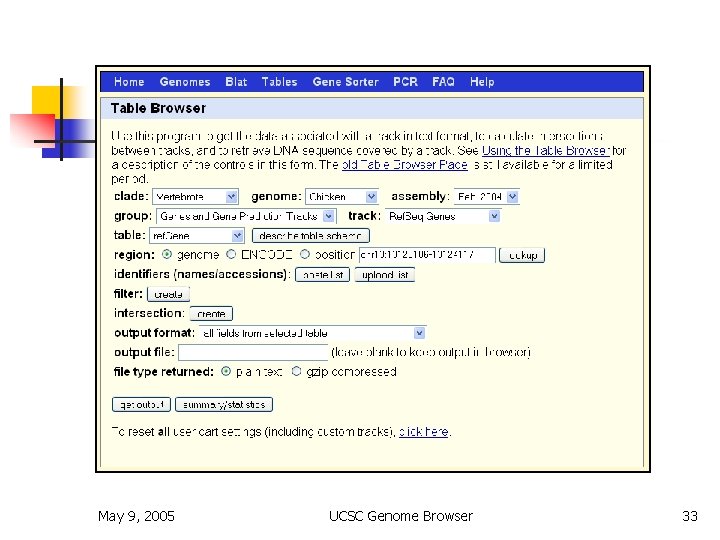
Table Browser n n n bulk download or batch query filtering by values intersection by position joining by related columns various output formats May 9, 2005 UCSC Genome Browser 33
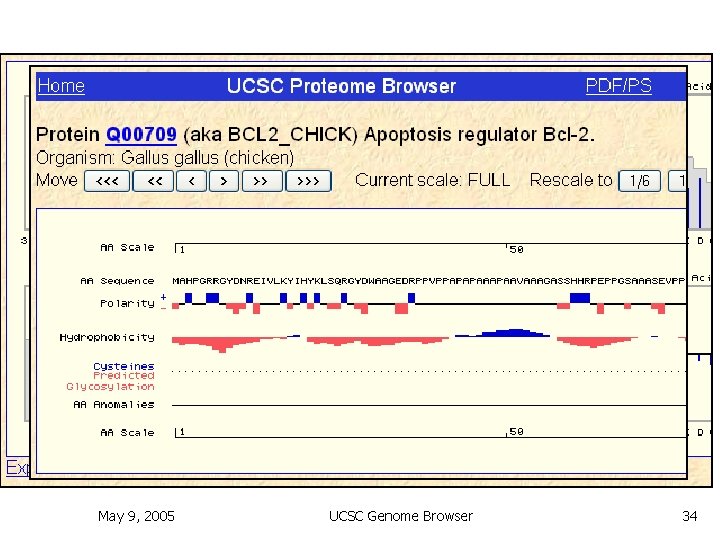
Proteome Browser n n Enter a Uni. Prot accession Horizontal axis: amino acid sequence Tracks: several protein properties Histograms for protein properties May 9, 2005 UCSC Genome Browser 34

BLAT: BLAST-Like Alignment Tool n n n FAST! Scan whole genome in seconds Nucleotide (95%id) or protein (80%id) Output: text or link to Genome Browser UCSC BLAT server: free for all, but some usage limits (see BLAT page) Install locally: free for academic, nonprofit use; license for commercial May 9, 2005 UCSC Genome Browser 35

How to learn more n n n http: //genome. ucsc. edu/ - play! questions? genome@soe. ucsc. edu Open. Helix: http: //www. openhelix. com/ n n n Classes, seminars Free online tutorial Quick reference cards (ask me for one!) May 9, 2005 UCSC Genome Browser 36

Thanks! n n n Funding: NHGRI, HHMI Data: WUSTL, NCBI, numerous contributors UCSC team: n n n David Haussler – PI Jim Kent – Browser Concept, BLAT, Team Leader Mark Diekhans, Fan Hsu, Angie Hinrichs, Kate Rosenbloom, Hiram Clawson, Rachel Harte, Andy Pohl - Engineering Donna Karolchik, Heather Trumbower, Robert Kuhn, Galt Barber, Jen Jackson, Ali Sultan-Qurraie – QA/Doc/Support Jorge Garcia, Patrick Gavin, Paul Tatarsky – Kilo. Kluster, Sysadmin May 9, 2005 UCSC Genome Browser 37
- Slides: 37