The Integrated Microbial Genome IMG systems Nikos Kyrpides

The Integrated Microbial Genome (IMG) systems Nikos Kyrpides
Data analysis Data Integration Comparative Analysis

What is the Matrix? Data management system for comparative analysis of biological data Genomes Functions Genes Metadata IMG SNPs M Clusters Proteomics G Transcriptomes Regulons I
![Integrated Microbial Genomes (IMG) [It’s easier to analyze 1000 genomes than a single one] Integrated Microbial Genomes (IMG) [It’s easier to analyze 1000 genomes than a single one]](http://slidetodoc.com/presentation_image_h/00735af738a64a596b5484b1a1fa7bc6/image-4.jpg)
Integrated Microbial Genomes (IMG) [It’s easier to analyze 1000 genomes than a single one] What is IMG: http: //img. jgi. doe. gov/ IMG is a data management system for comparative analysis and annotation of all publicly available genomes from three domains of life in a uniquely integrated context. • http: //img. jgi. doe. gov/ Mission: To become the Home of Microbial Genome and Metagenome Analysis Background: Launched on March 2005 3 Releases/Year >5, 000 unique visitors per month >300 citations Current Status: 8939 Genomes 24 Million Genes Bacteria: 3930 Archaea: 164 Eukarya: 177 Plasmids: 1205 Viruses: 2809 Gfragments: 654 USERS CAN ü Search data ü Browse data ü Compare data ü Export data
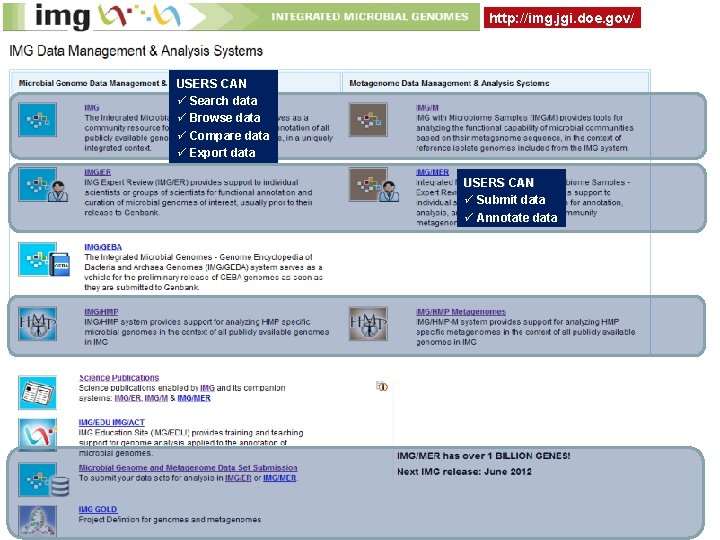
http: //img. jgi. doe. gov/ USERS CAN ü Search data ü Browse data ü Compare data ü Export data USERS CAN ü Submit data ü Annotate data
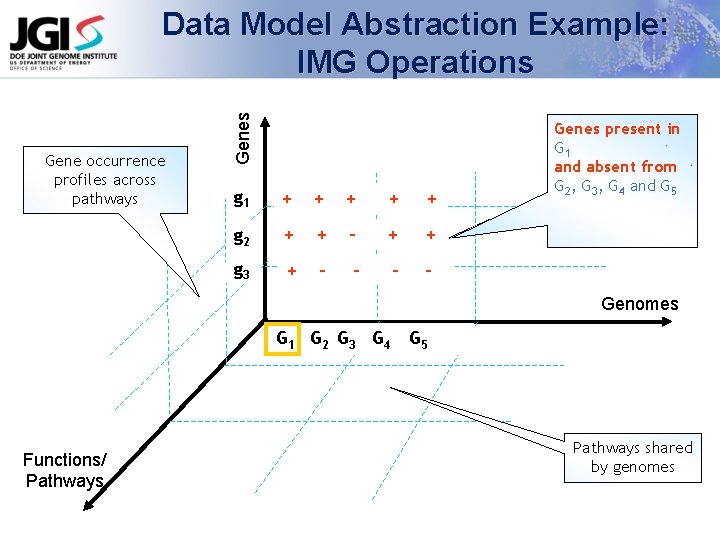
Gene occurrence profiles across pathways Genes Data Model Abstraction Example: IMG Operations g 1 + + + g 2 + + - + + g 3 + - - Genes Gene occurrence present in G 1 profile across and absent genomes from G 2, G 3, G 4 and G 5 Genomes G 1 G 2 G 3 G 4 Functions/ Pathways G 5 Pathways shared by genomes
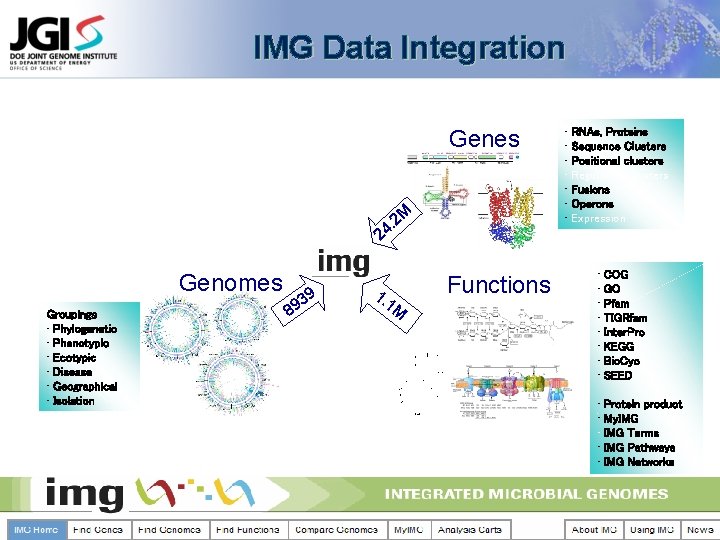
IMG Data Integration Genes 2 M . 24 Genomes Groupings • Phylogenetic • Phenotypic • Ecotypic • Disease • Geographical • Isolation 8 9 93 1. 1 M Functions • RNAs, Proteins • Sequence Clusters • Positional clusters • Regulatory clusters • Fusions • Operons • Expression • COG • GO • Pfam • TIGRfam • Inter. Pro • KEGG • Bio. Cyc • SEED • Protein product • My. IMG • IMG Terms • IMG Pathways • IMG Networks

IMG Toolkit Chromosome Map Function Profile Gene Synteny IMG Pathway Profile Metadata Search Phylogenetic Profile KEGG Maps Phylogenetic Distribution Chromosomal Map Abundance Profiles Functional Categories Projects Map Genome Clustering Compare Annotations VISTA Artemis Recruitme nt Plot Fragment Recruitme nt
Challenges and Opportunities Annotations Quality Metadata Data Analysis Integration Scaling Genes Functions New data types and tools # genes and genomes
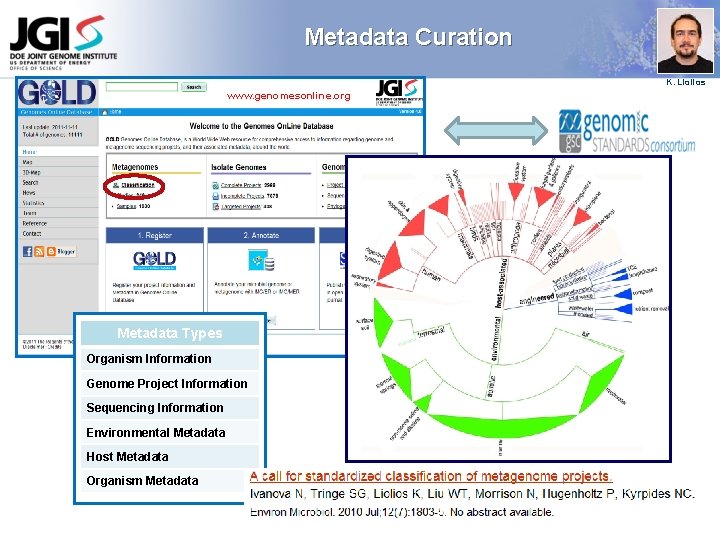
Metadata Curation K. Liolios www. genomesonline. org Metadata Types Organism Information Genome Project Information Sequencing Information Environmental Metadata Host Metadata Organism Metadata

Metagenome Classification Genomes vs Metagenomes
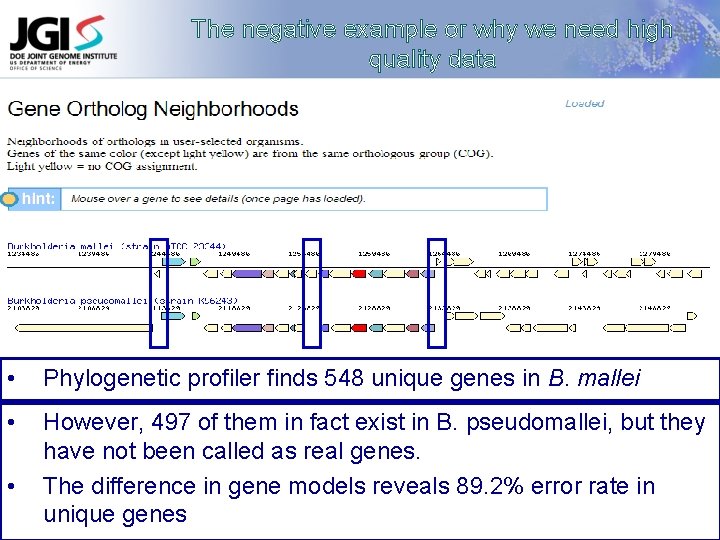
The negative example or why we need high quality data • Phylogenetic profiler finds 548 unique genes in B. mallei • However, 497 of them in fact exist in B. pseudomallei, but they have not been called as real genes. The difference in gene models reveals 89. 2% error rate in unique genes •

Gene Prediction - Standards Usage of Reference Genomes • Annotation of isolate genomes, single cells and metagenomes • Computation of Gene cassettes • Computation of Pangenomes Problems with current Public Reference Genomes • lack of provenance for the predicted features • presence of artificial (non-biological) variation between the genomes including variation in gene content and variation within protein families Re-Annotation workshops - January 19 -20, 2012 @ JGI - February 29, 2012, @NCBI Ivanova
Constant Benchmarking Evaluation of Annotation Quality with constantly changing: Sequencing technologies Similarity methods Read lengths Clustering methods K. Mavrommatis Gene calling methods Functional annotation methods
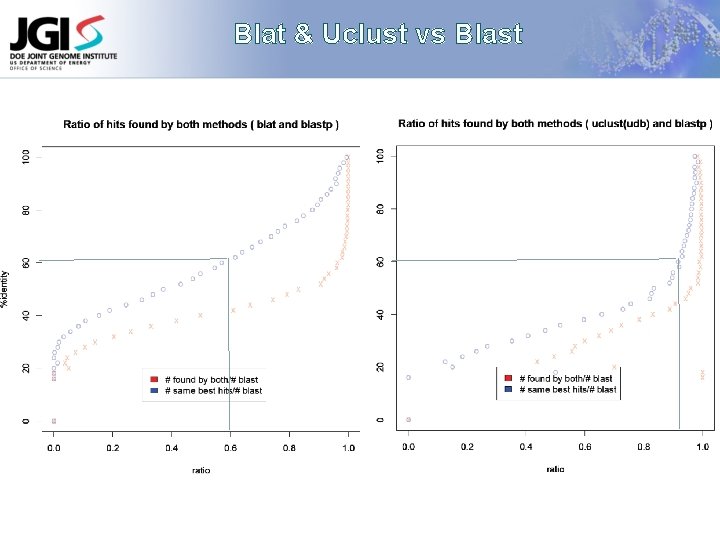
Blat & Uclust vs Blast
Program Informatics Challenges and Opportunities Quality Data Analysis Integration Scaling Annotations New data types and tools ##genesand andgenomes

Why annotate unassembled reads? Kansas soil Total size 102, 722, 384 (2 x 150) reads Assembled contigs 1, 375, 950 contigs Assembled (reported by the CLC workbench assembler) 38, 094, 033 reads Assembled reads Mapped (by bwa) 11, 778, 925 reads Genes called on unassembled reads 64, 737, 444 genes 5060 different pfams 7481 different pfams 8, 373, 641 (12%) genes Similar to genes on contigs 1 Genes with similarity to isolate genomes 40, 778, 854 genes Assembled only More accurate statistics based on unassembled + assembled Unassembled + real metagenome Additional information about functions and phylogeny
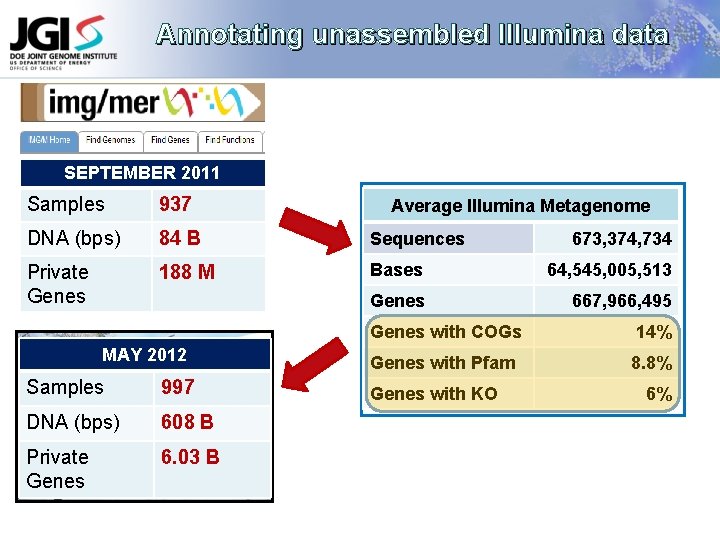
Annotating unassembled Illumina data SEPTEMBER 2011 Samples 937 DNA (bps) 84 B Sequences Private Genes 188 M Bases 64, 545, 005, 513 Genes 667, 966, 495 MAY 2012 Samples 997 DNA (bps) 608 B Private Genes 6. 03 B Average Illumina Metagenome 673, 374, 734 Genes with COGs 14% Genes with Pfam 8. 8% Genes with KO 6%

Where do we go from here?

DELUGE AVALANCHE FLOOD TSUNAMI OPPORTUNITY
Program Informatics Challenges and Opportunities Quality Data Analysis Annotations / Publications Integration Scaling New data types and tools # genes and genomes
Challenges and Opportunities Gene Clustering Metagenome Classification Data Analysis

Unique properties of IMG Largest metadata integration from GOLD Largest integration of Genes (> 4 Billion) Clustering of all metagenomic genes Metagenome classification scheme IMG QC of all gene predictions from isolate genomes Large array of function & pathway analysis tools
- Slides: 23