Chapter 9 Proteins and Their Synthesis Green Fluorescent





















- Slides: 58

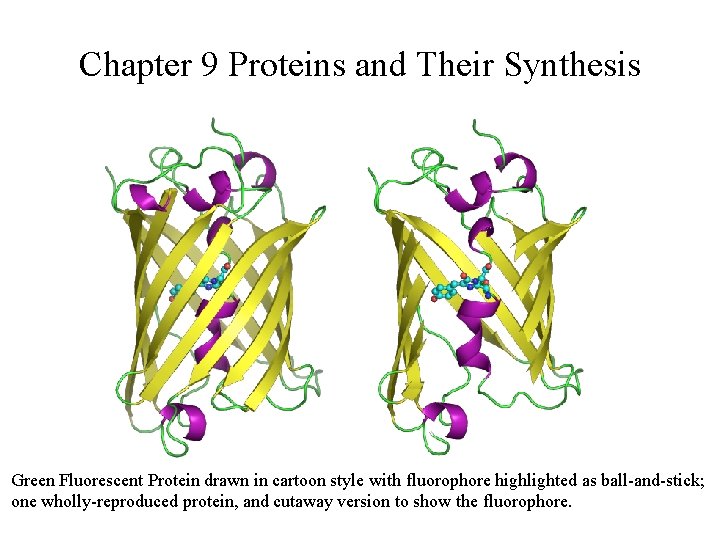
Chapter 9 Proteins and Their Synthesis Green Fluorescent Protein drawn in cartoon style with fluorophore highlighted as ball-and-stick; one wholly-reproduced protein, and cutaway version to show the fluorophore.

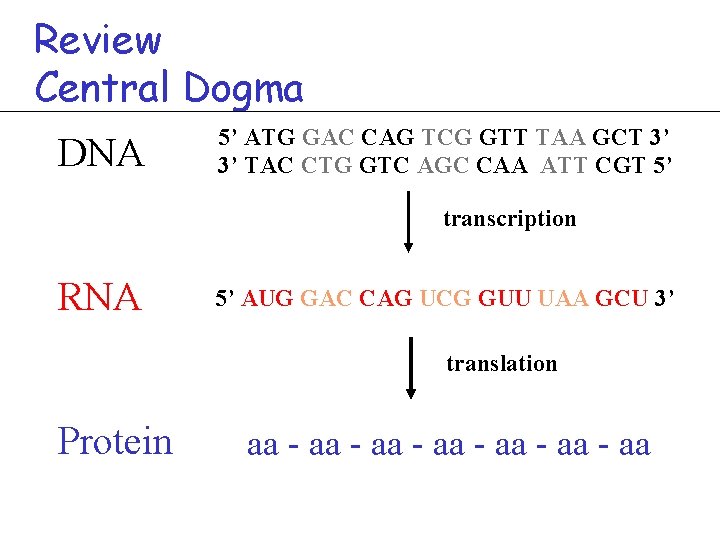
Review Central Dogma DNA 5’ ATG GAC CAG TCG GTT TAA GCT 3’ 3’ TAC CTG GTC AGC CAA ATT CGT 5’ transcription RNA 5’ AUG GAC CAG UCG GUU UAA GCU 3’ translation Protein aa - aa - aa

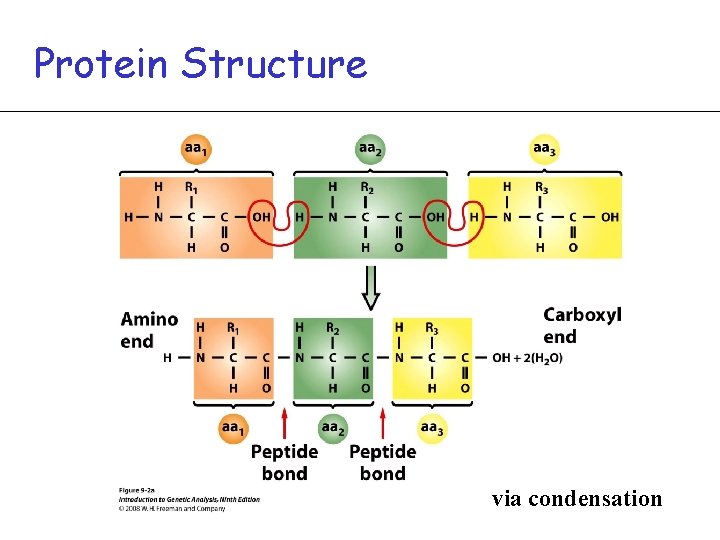
Protein Structure via condensation

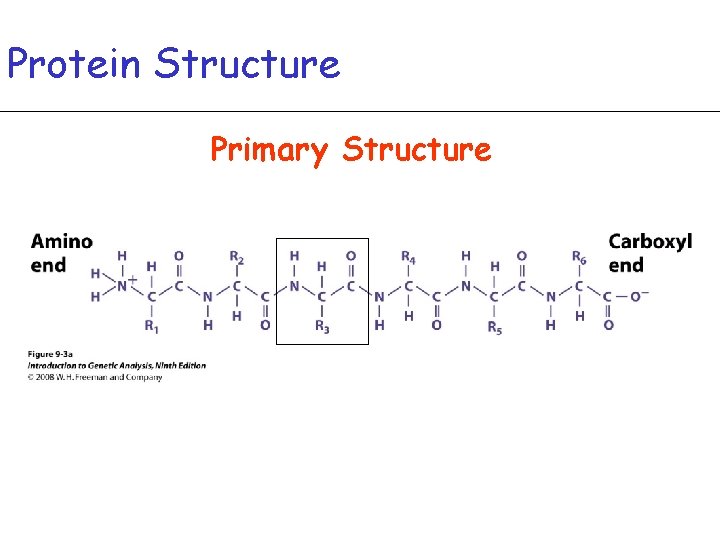
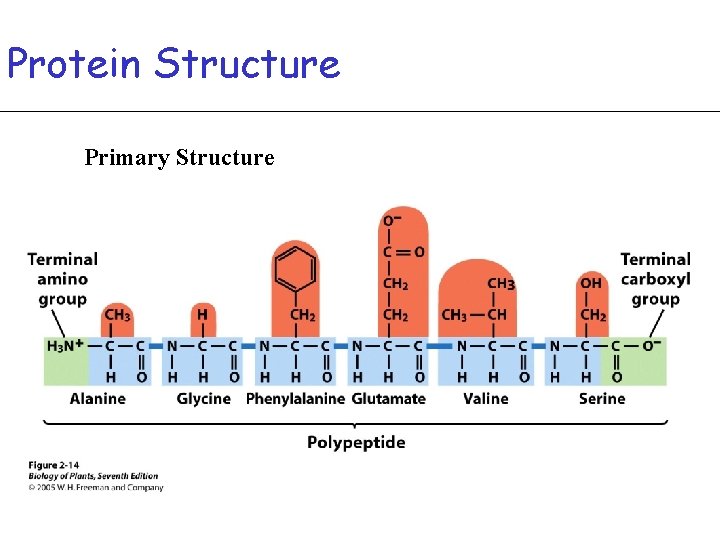
Protein Structure Primary Structure

Protein folding is dependent on the amino acid R groups R General Structure H 2 N C COOH H There are 20 amino acids. Their properties are determined by the R group.

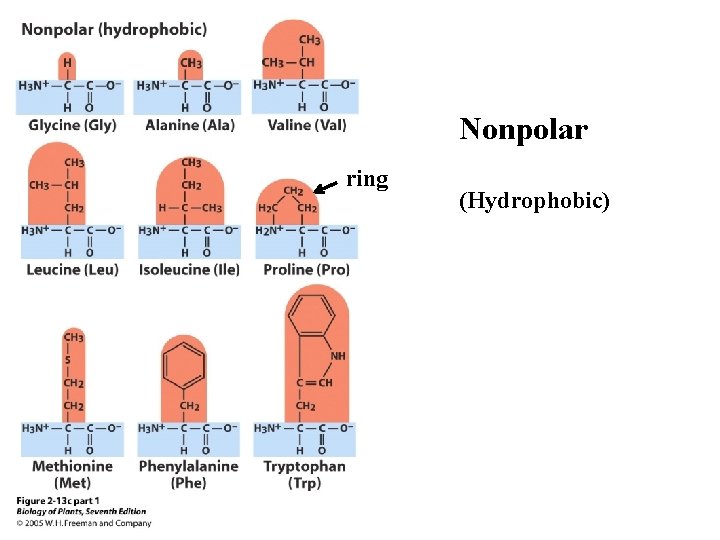
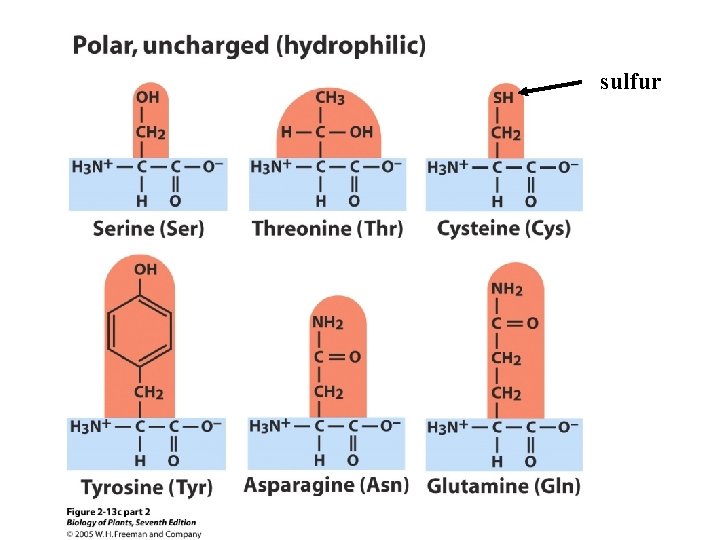
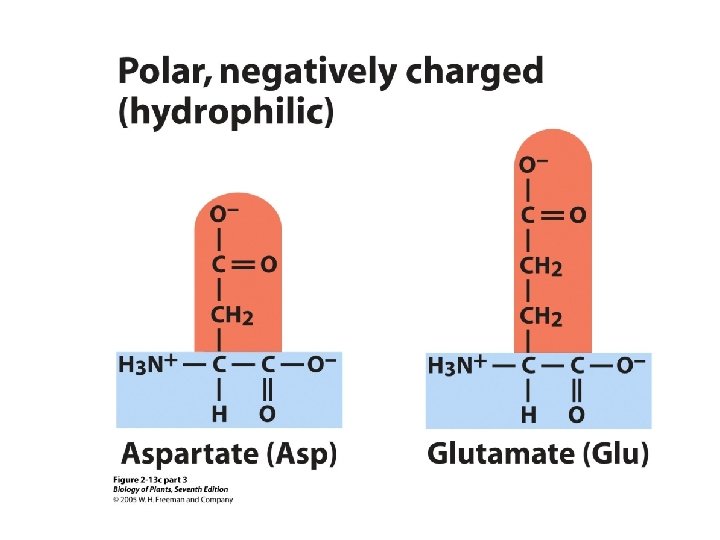
There are 20 amino acids. Ø Nonpolar or hydrophobic (9) Ø Polar (hydrophillic), but uncharged (6) Ø Polar (hydrophillic), but charged (5)

Nonpolar ring (Hydrophobic)

sulfur



Protein Structure Primary Structure

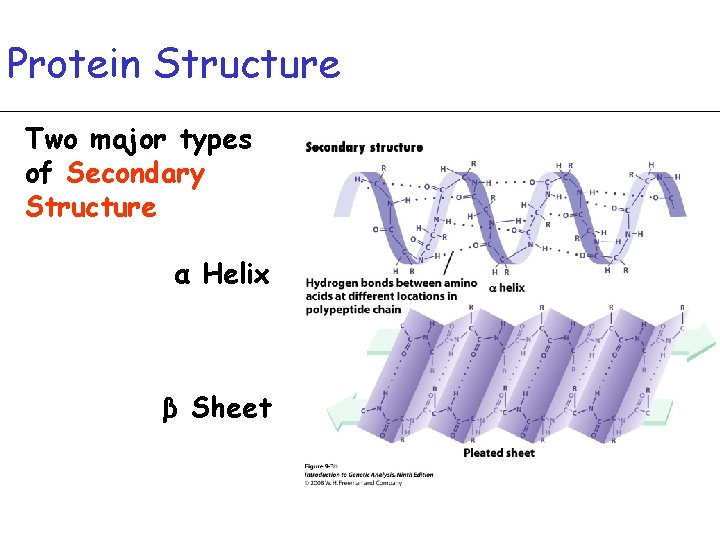
Protein Structure Two major types of Secondary Structure α Helix β Sheet

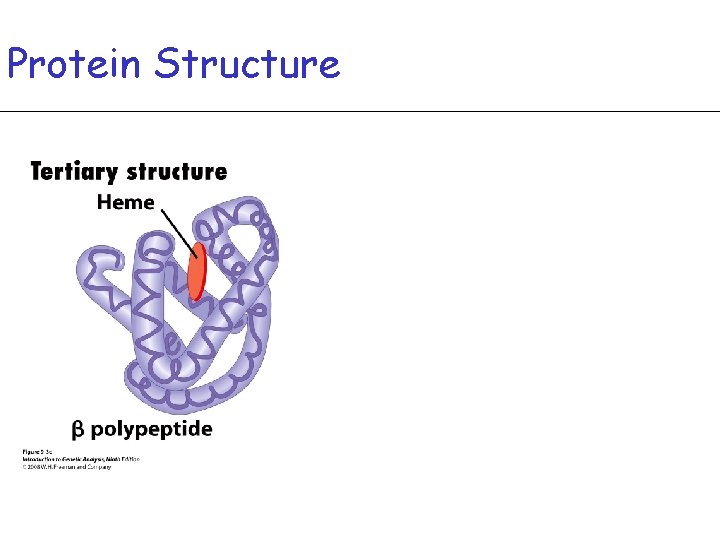
Protein Structure

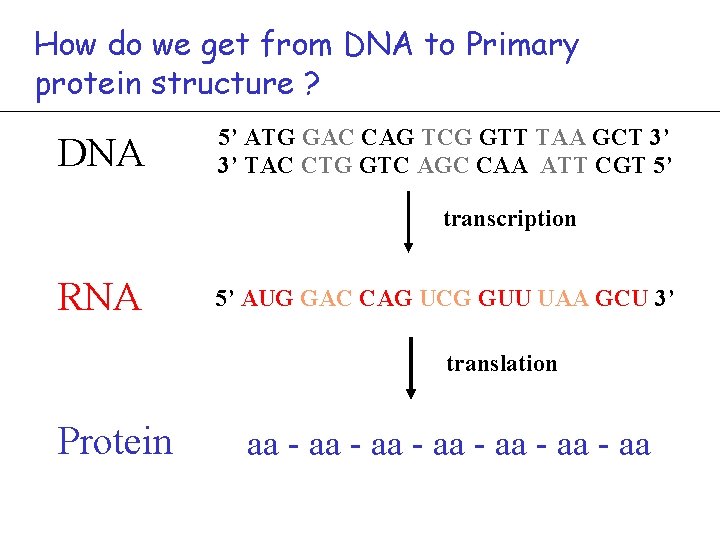
How do we get from DNA to Primary protein structure ? DNA 5’ ATG GAC CAG TCG GTT TAA GCT 3’ 3’ TAC CTG GTC AGC CAA ATT CGT 5’ transcription RNA 5’ AUG GAC CAG UCG GUU UAA GCU 3’ translation Protein aa - aa - aa

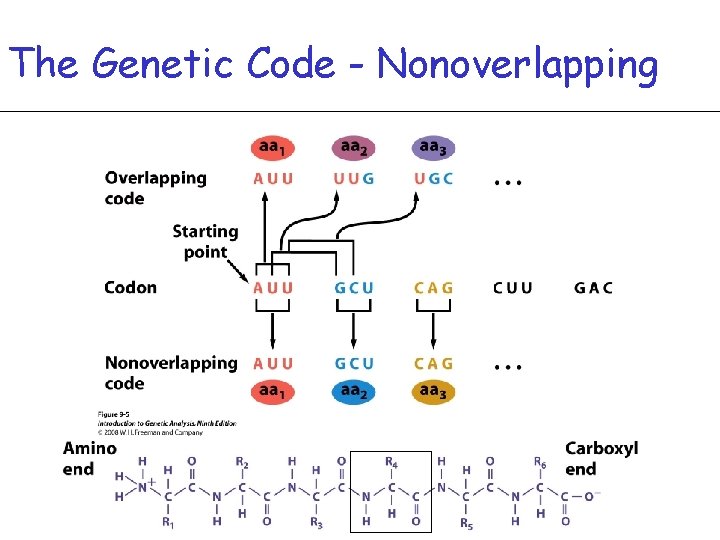
DNA (m. RNA) is read in Triplets -Codon – Group of 3 DNA bases codes for a specific amino acid Ex. ATG = methionine -This means the code is degenerate – more than one codon can specify one amino acid

The Genetic Code - Nonoverlapping

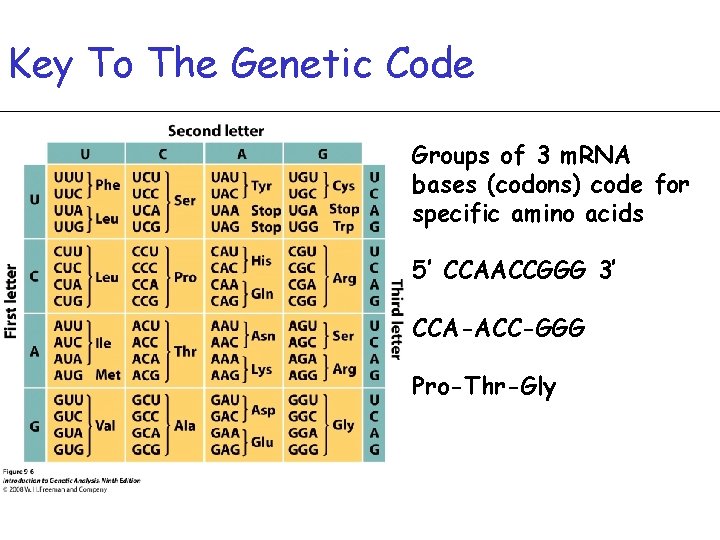
Key To The Genetic Code Groups of 3 m. RNA bases (codons) code for specific amino acids 5’ CCAACCGGG 3’ CCA-ACC-GGG Pro-Thr-Gly

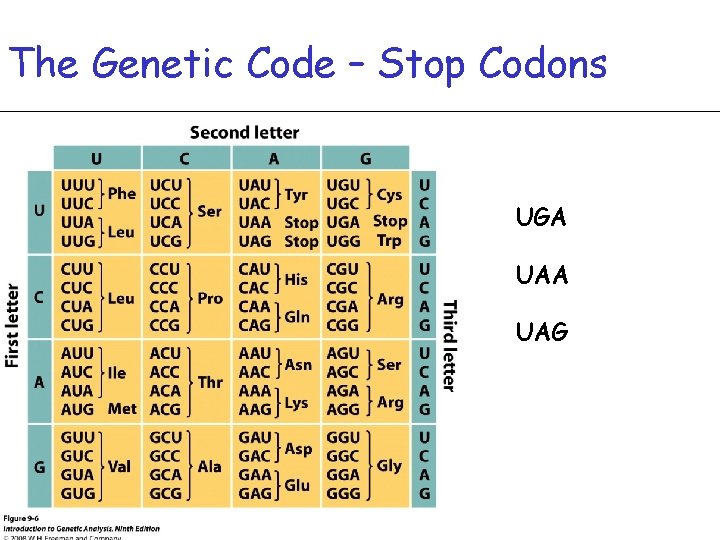
The Genetic Code – Stop Codons UGA UAG

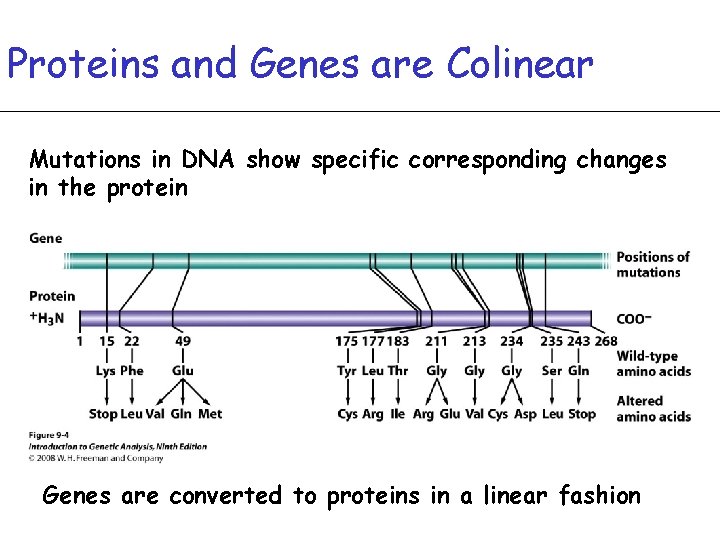
Proteins and Genes are Colinear Mutations in DNA show specific corresponding changes in the protein Genes are converted to proteins in a linear fashion

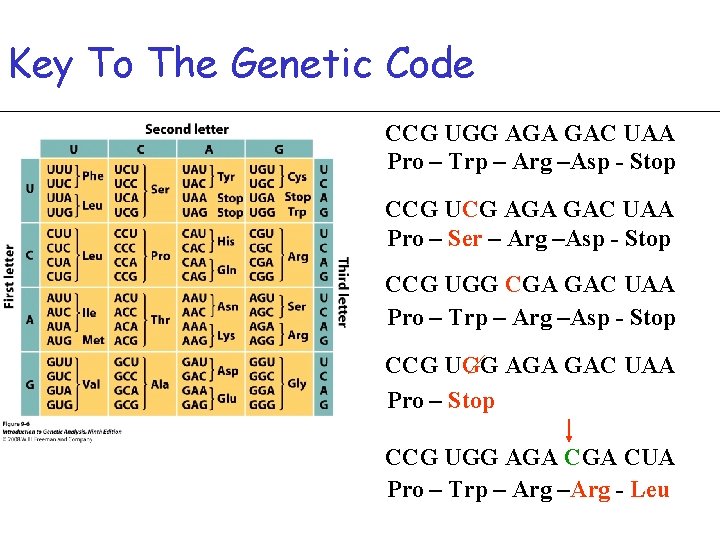
Key To The Genetic Code CCG UGG AGA GAC UAA Pro – Trp – Arg –Asp - Stop CCG UCG AGA GAC UAA Pro – Ser – Arg –Asp - Stop CCG UGG CGA GAC UAA Pro – Trp – Arg –Asp - Stop CCG UGG AGA GAC UAA Pro – Stop CCG UGG AGA CUA Pro – Trp – Arg –Arg - Leu

The Genetic Code - Mutations 4 Types of Mutations 1. Silent mutations 2. Missense mutations 3. Nonsense mutations 4. Frameshift mutations

The Genetic Code m. RNA has 3 potential “reading frames” 5’ CUUACAGUUUAUUGAUACGGAGAAGG 3’ 3’ GAAUGUCAAAUAACUAUGCCUCUUCC 5’ 5’ CUU ACA GUU UAU UGA UAC GGA GAA GG 3’ 3’ GAA UGU CAA AUA ACU AUG CCU CUU CC 5’ 5’ C UUA CAG UUU AUU GAU ACG GAG AAG G 3’ 3’ G AAU GUC AAA UAA CUA UGC CUC UUC C 5’ 5’ CU UAC AGU UUA UUG AUA CGG AGA AGG 3’ 3’ GA AUG UCA AAU AAC UAU GCC UCU UCC 5’ Stop UAA UGA UAG

The Genetic Code m. RNA has 3 potential reading frames 5’ CUUACAGUUUAUUGAUACGGAGAAGG 3’ 3’ GAAUGUCAAAUAACUAUGCCUCUUCC 5’ 5’ CUU ACA GUU UAU UGA UAC GGA GAA GG 3’ 3’ GAA UGU CAA AUA ACU AUG CCU CUU CC 5’ 5’ C UUA CAG UUU AUU GAU ACG GAG AAG G 3’ 3’ G AAU GUC AAA UAA CUA UGC CUC UUC C 5’ 5’ CU UAC AGU UUA UUG AUA CGG AGA AGG 3’ 3’ GA AUG UCA AAU AAC UAU GCC UCU UCC 5’ Stop UAA UGA UAG

Review - RNA m. RNA- messenger RNA t. RNA- transfer RNA r. RNA- Ribosomal RNA

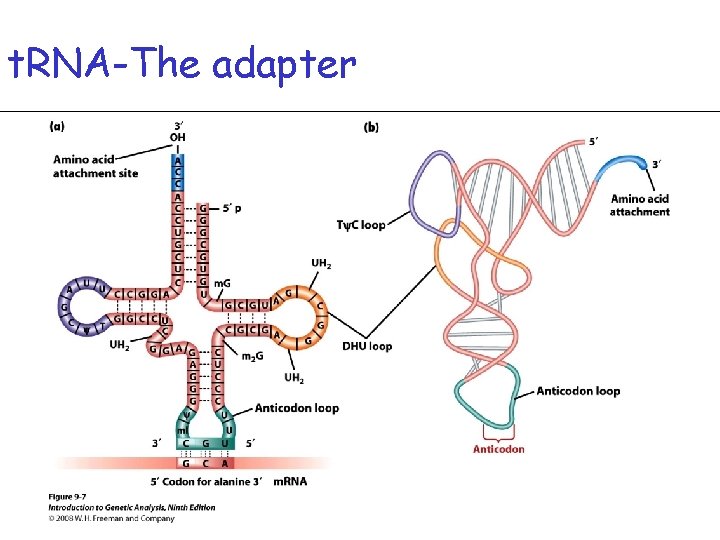
t. RNA-The adapter

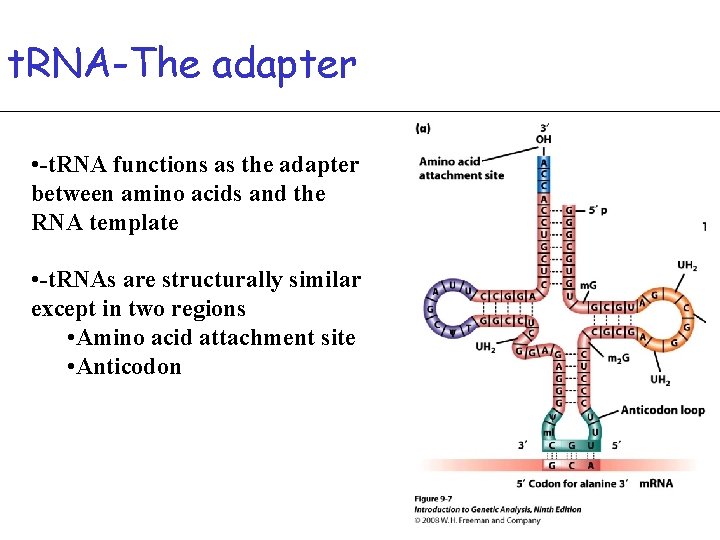
t. RNA-The adapter • -t. RNA functions as the adapter between amino acids and the RNA template • -t. RNAs are structurally similar except in two regions • Amino acid attachment site • Anticodon

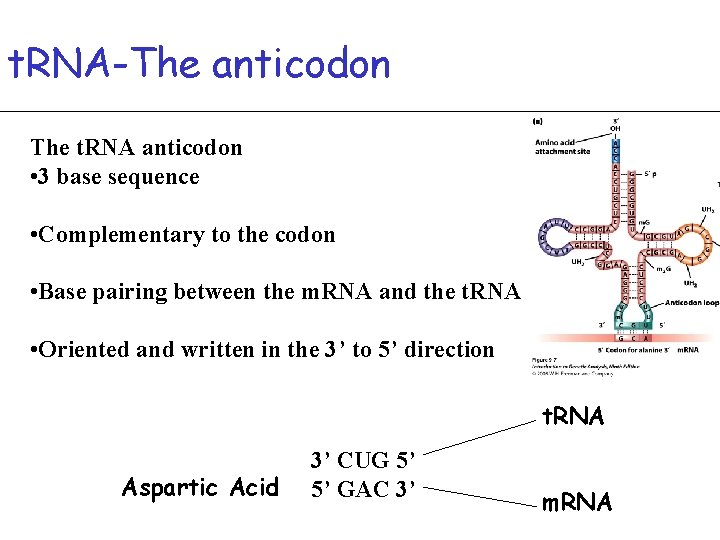
t. RNA-The anticodon The t. RNA anticodon • 3 base sequence • Complementary to the codon • Base pairing between the m. RNA and the t. RNA • Oriented and written in the 3’ to 5’ direction t. RNA Aspartic Acid 3’ CUG 5’ 5’ GAC 3’ m. RNA

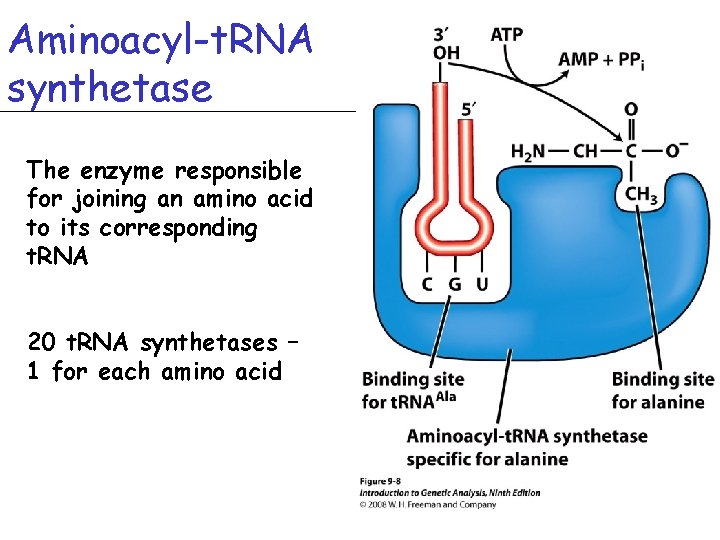
Aminoacyl-t. RNA synthetase The enzyme responsible for joining an amino acid to its corresponding t. RNA 20 t. RNA synthetases – 1 for each amino acid

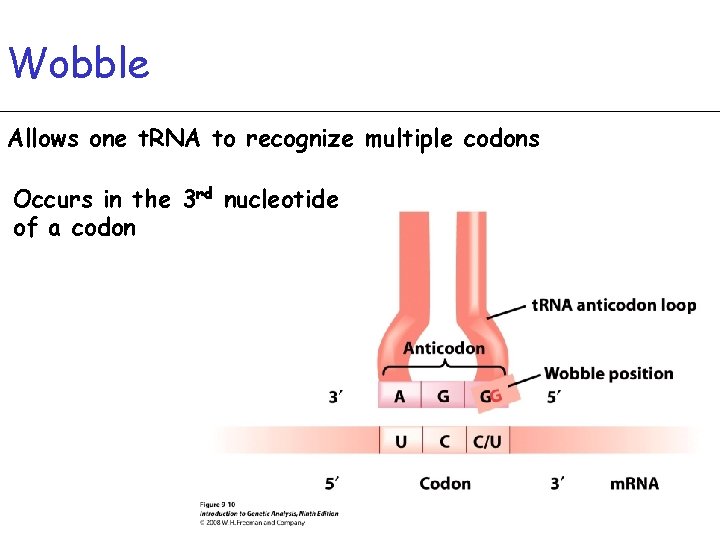
Wobble Allows one t. RNA to recognize multiple codons Occurs in the 3 rd nucleotide of a codon

Wobble – A new set pairing of rules I = Inosine: A rare base found in t. RNA

Wobble – A new set pairing of rules Isoaccepting t. RNAs: t. RNAs that accept the same amino acid but are transcribed from different genes

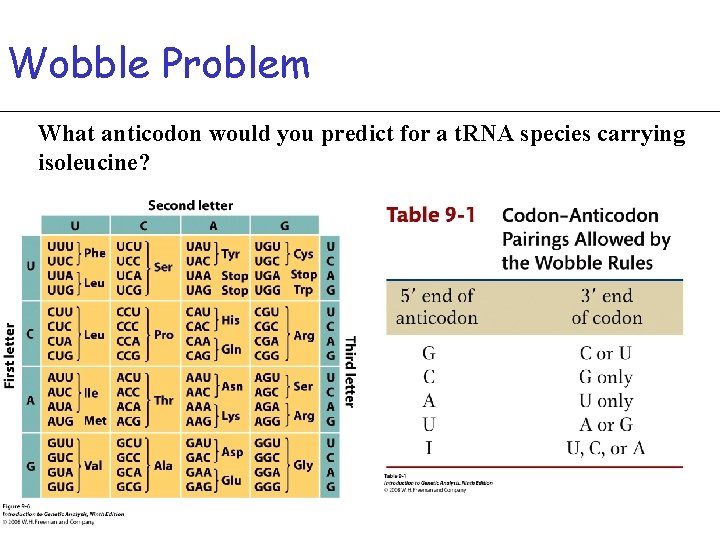
Wobble Problem What anticodon would you predict for a t. RNA species carrying isoleucine?

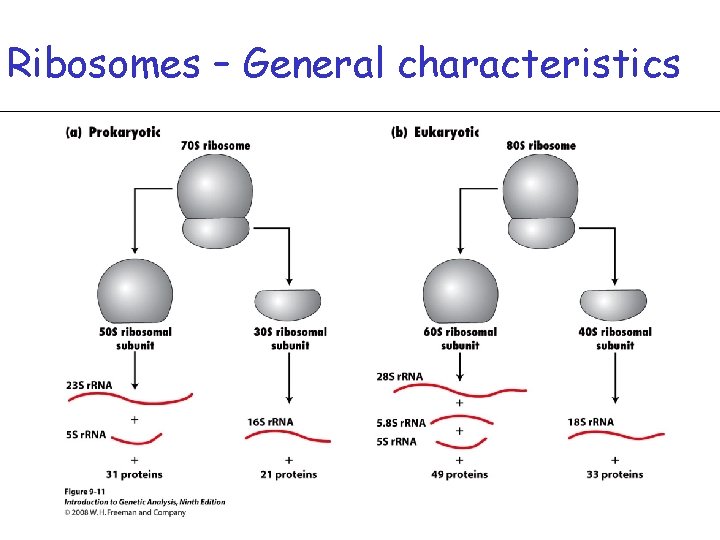
Ribosomes – General characteristics • Come together with t. RNA and m. RNA to create protein • Ribosome consist of one small and one large subunit • In prokaryotes, 30 S and 50 S subunits form a 70 S particle • In Eukaryotes, 40 S and 60 S subunits form an 80 S particle • Each subunit is composed of 1 to 3 types of r. RNA and up to 49 proteins

Ribosomes – General characteristics

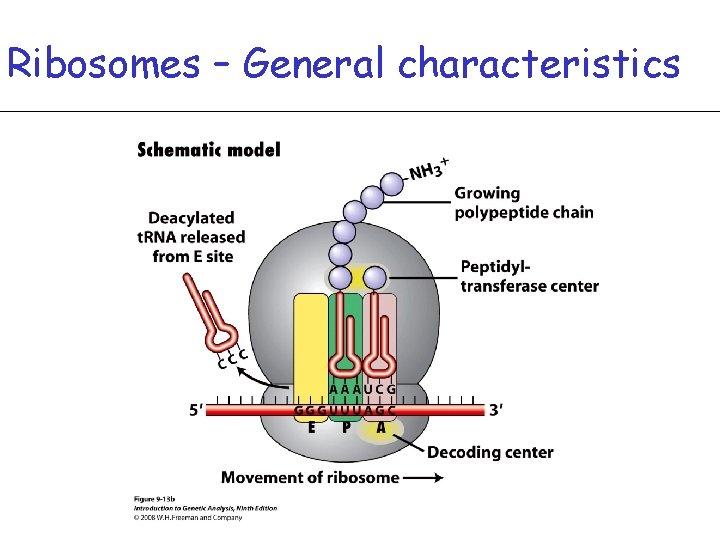
Ribosomes – General characteristics • r. RNA folds up by intramolecular base pairing

Ribosomes – General characteristics

Translation Synthesizing Protein

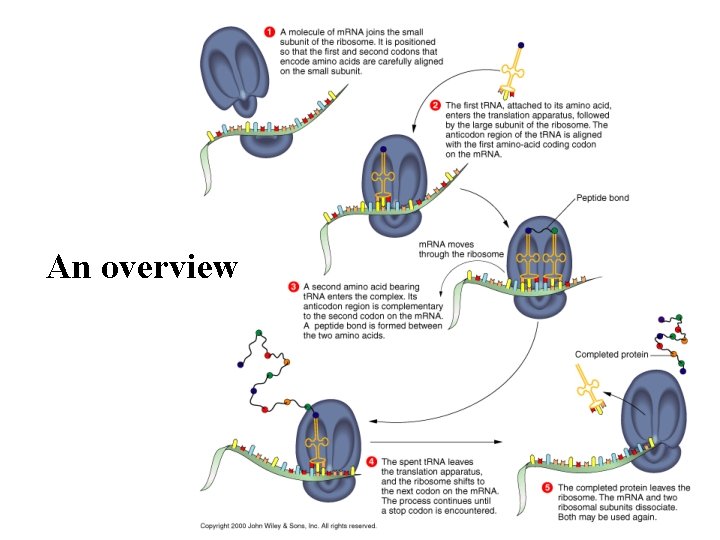
An overview

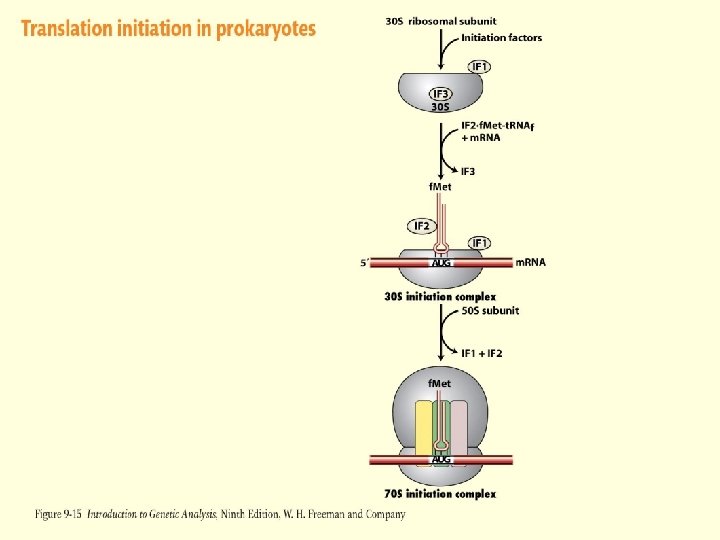
Translation Initiation - Prokaryotes Translation begins at an AUG codon – Methionine Requires a special “initiator” t. RNA charged with Met – t. RNAMeti This involves the addition of a formyl group to methionine while it is attached to the initiator

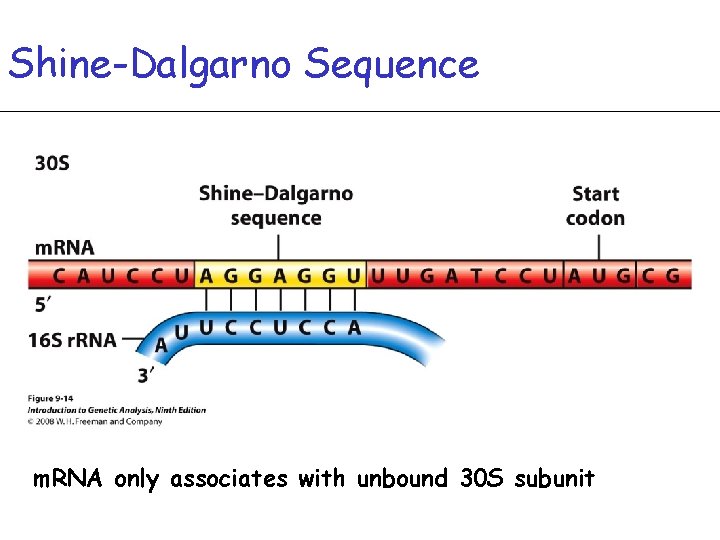
Shine-Dalgarno Sequence m. RNA only associates with unbound 30 S subunit

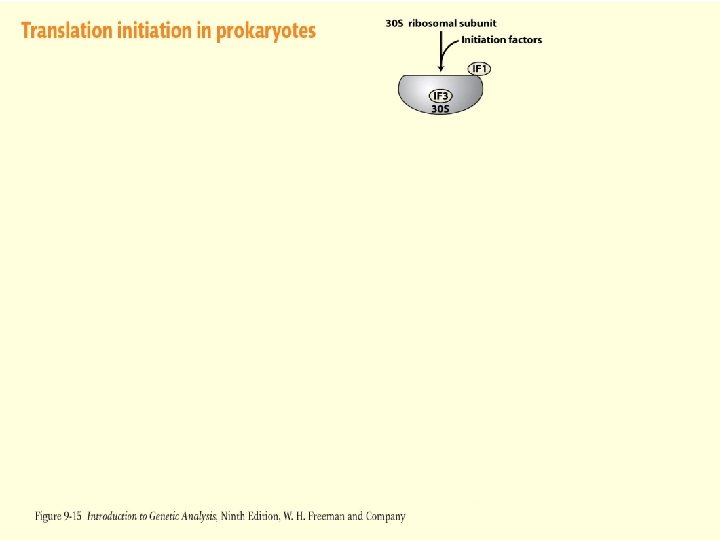
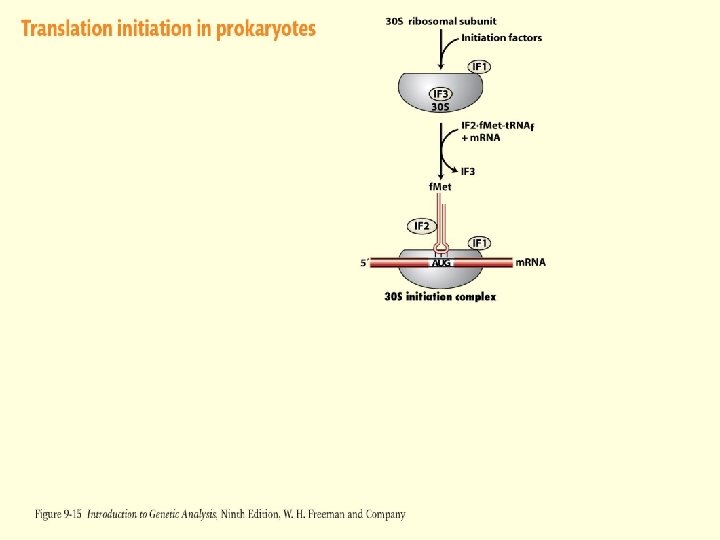
Translation Initiation – Prokaryotes Initiation Factors 3 initiation factor proteins are required for the start of translation in prokaryotes IF 1 – Binds to 30 S subunit as part of the complete initiation complex. Could be involved in stability IF 2 – Binds to charged initiator t. RNA and insures that other t. RNAS do not enter initiation complex IF 3 – Keeps the 30 S subunit disassociated from the 50 S subunit and allows binding of m. RNA

Figure 2 -12 -1 Figure 9 -15 -1

Figure 2 -12 -1 Figure 9 -15 -2

Figure 2 -12 -1 Figure 9 -15 -3

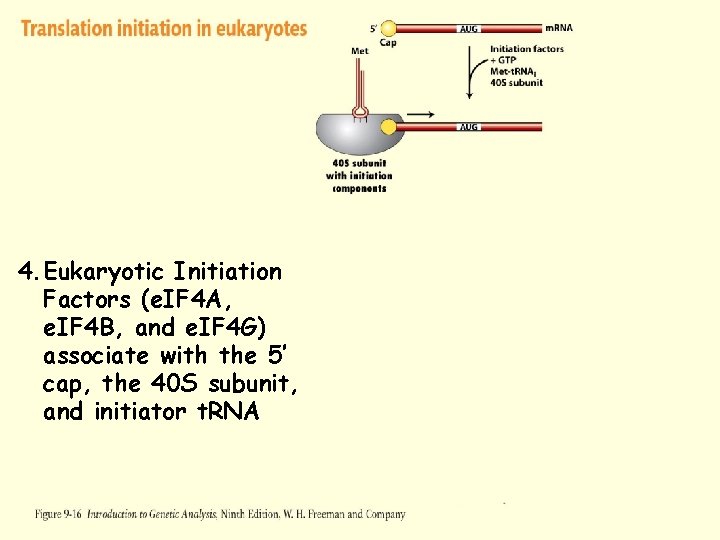
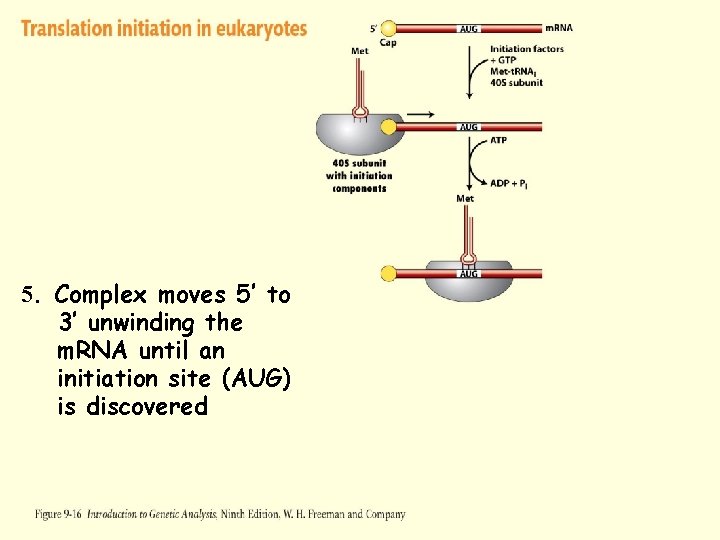
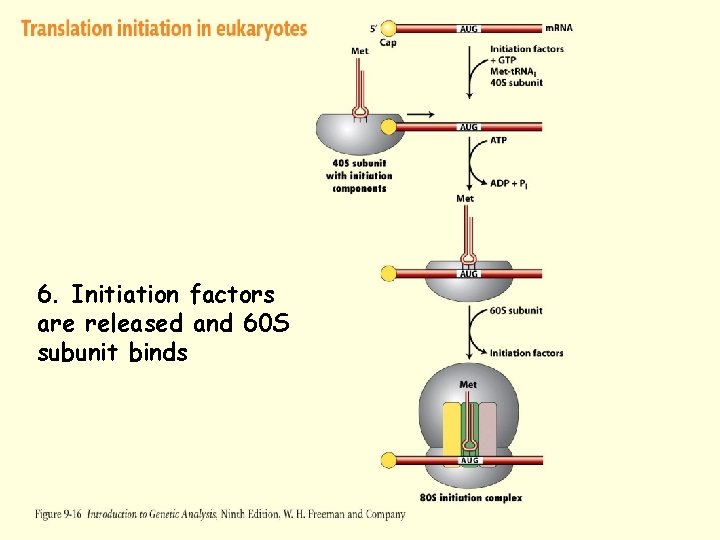
Translation Initiation – Eukaryotes 1. m. RNA is produced in the nucleus and transported to the cytoplasm 2. 5’ end of the m. RNA is “capped” to prevent degradation 3. Eukaryotic Initiation Factors (e. IF 4 A, e. IF 4 B, and e. IF 4 G) associate with the 5’ cap, the 40 S subunit, and initiator t. RNA 4. Complex moves 5’ to 3’ unwinding the m. RNA until an initiation site (AUG) is discovered 5. Initiation factors are released and 60 S subunit binds

Figure 2 -12 -1 Figure 9 -16 -1 1. m. RNA is produced in the nucleus and transported to the cytoplasm 2. m. RNA is covered with proteins and often folds on itself 3. 5’ end of the m. RNA is “capped” to prevent degradation

Figure 2 -12 -1 Figure 9 -16 -2 4. Eukaryotic Initiation Factors (e. IF 4 A, e. IF 4 B, and e. IF 4 G) associate with the 5’ cap, the 40 S subunit, and initiator t. RNA

Figure 9 -16 -3 5. Complex moves 5’ to 3’ unwinding the m. RNA until an initiation site (AUG) is discovered

Figure 9 -16 -4 6. Initiation factors are released and 60 S subunit binds

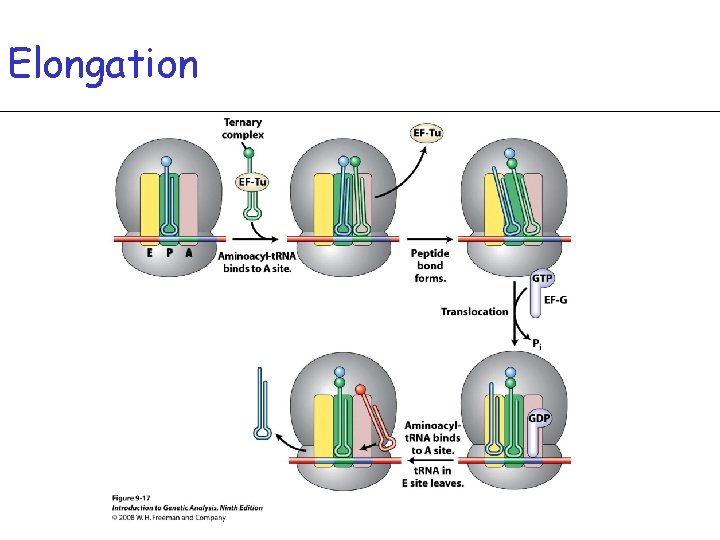
Elongation • Requires two protein Elongation Factors: EF-Tu and EF-G • Amino acids are added to the growing peptide chain at the rate of 2 -15 amino acids per second

Elongation

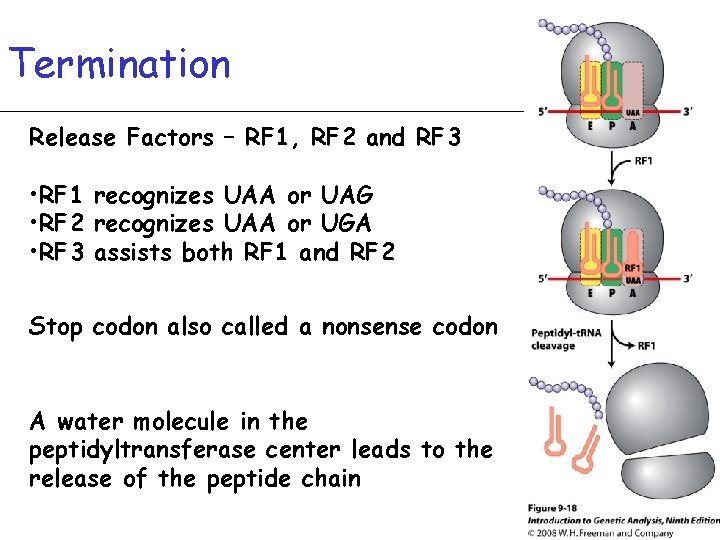
Termination Release Factors – RF 1, RF 2 and RF 3 • RF 1 recognizes UAA or UAG • RF 2 recognizes UAA or UGA • RF 3 assists both RF 1 and RF 2 Stop codon also called a nonsense codon A water molecule in the peptidyltransferase center leads to the release of the peptide chain

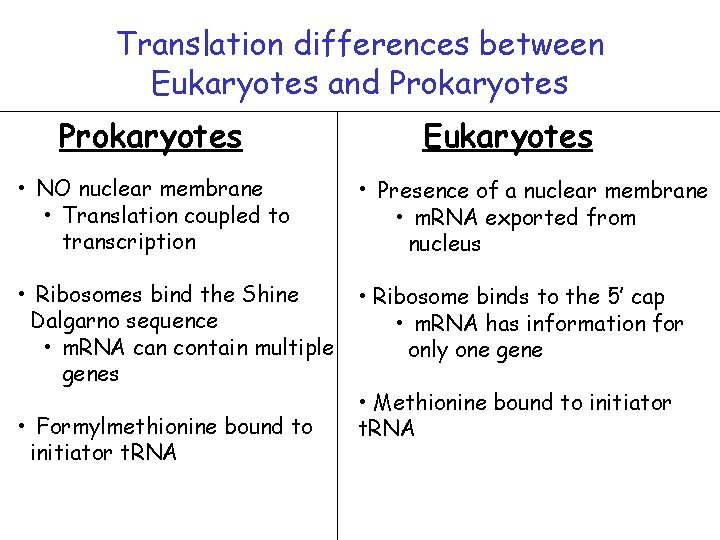
Translation differences between Eukaryotes and Prokaryotes Eukaryotes • NO nuclear membrane • Translation coupled to transcription • Presence of a nuclear membrane • Ribosomes bind the Shine Dalgarno sequence • m. RNA can contain multiple genes • Ribosome binds to the 5’ cap • m. RNA has information for only one gene • Formylmethionine bound to initiator t. RNA • m. RNA exported from nucleus • Methionine bound to initiator t. RNA

Posttranslational Folding Proteins must fold correctly to be functional Correct folding is not always energetically favorable in the cytoplasm Chaperones (including Gro. E chaperonins) bind to nascent peptides and facilitate correct folding

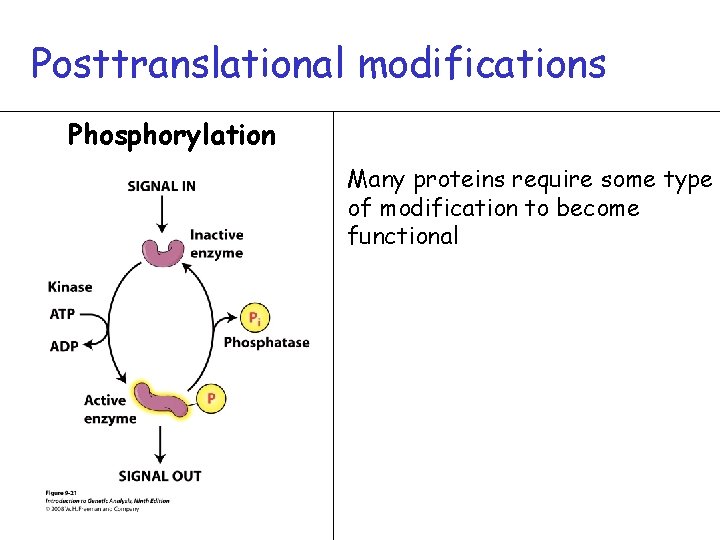
Posttranslational modifications Phosphorylation Many proteins require some type of modification to become functional

Posttranslational modifications Glycosylation – adding sugars Signaling molecules Cell wall proteins Glycoproteins

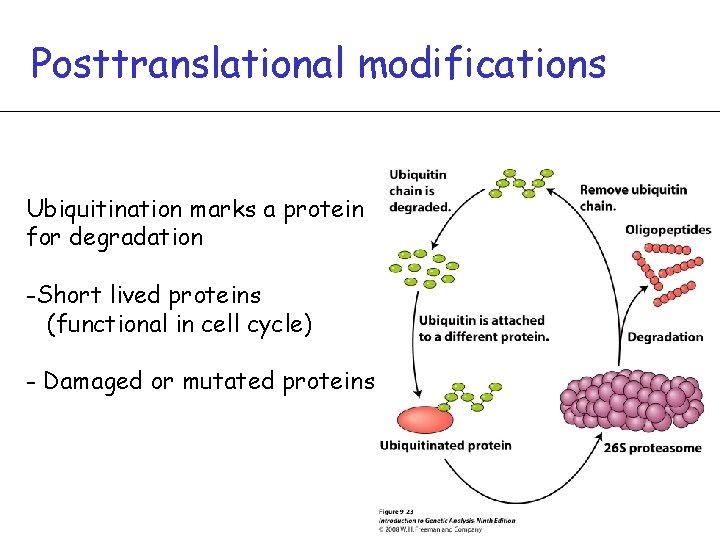
Posttranslational modifications Ubiquitination marks a protein for degradation -Short lived proteins (functional in cell cycle) - Damaged or mutated proteins

Summary • Translation – Prokaryote – Eukaryote • Post translational modifications – Phosphorylation – Glycosylation – Ubiquitination