II PROTEIN BIOCHEMISTRY 2 2 Protein Properties 2

II. PROTEIN BIOCHEMISTRY § 2. 2 Protein Properties § 2. 2 a Protein Diversity § 2. 2 b Protein Purification § 2. 2 c Protein Characterization § 2. 2 d Protein Sequencing

§ 2. 2 a Protein Diversity

Synopsis 2. 2 a - A polypeptide is a biopolymer comprised of amino acid residues covalently linked together via repetitive peptide bonds - In theory, the size and composition of a polypeptide chain are unlimited - In cells, such potential diversity is limited by the efficiency of protein synthesis and by the ability of the polypeptide to fold into a functional structure - A protein may be comprised of a: single polypeptide => monomeric protein multiple polypeptides => multi-chain/multi-subunit/multimeric protein
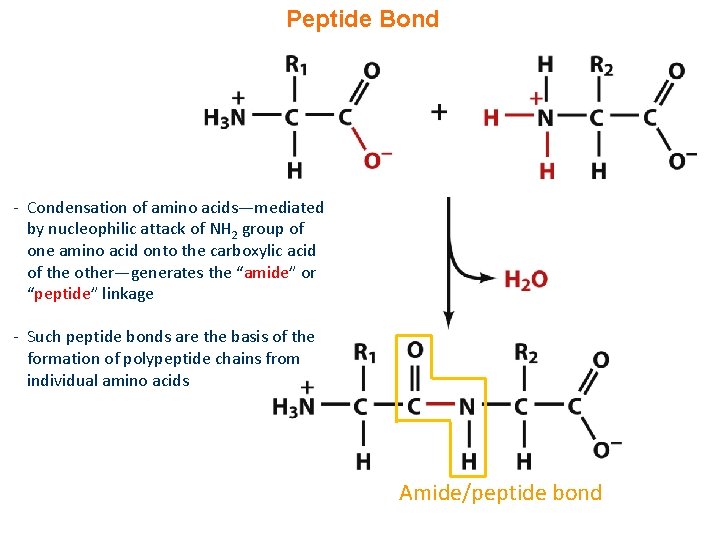
Peptide Bond - Condensation of amino acids—mediated by nucleophilic attack of NH 2 group of one amino acid onto the carboxylic acid of the other—generates the “amide” or “peptide” linkage - Such peptide bonds are the basis of the formation of polypeptide chains from individual amino acids Amide/peptide bond
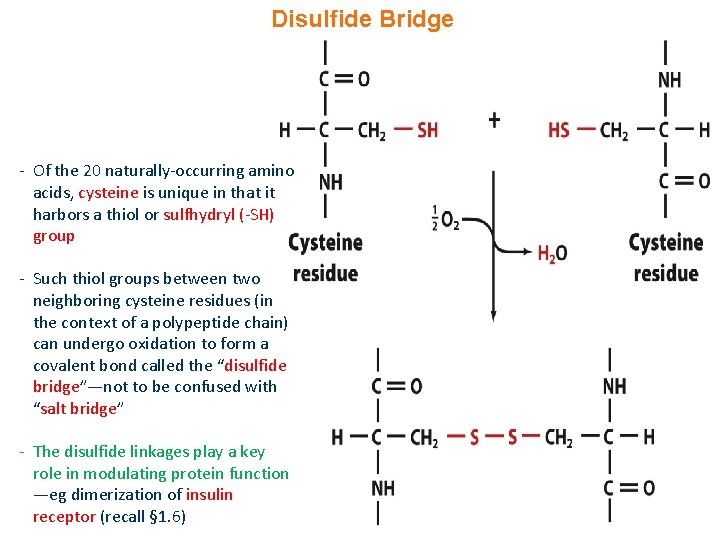
Disulfide Bridge - Of the 20 naturally-occurring amino acids, cysteine is unique in that it harbors a thiol or sulfhydryl (-SH) group - Such thiol groups between two neighboring cysteine residues (in the context of a polypeptide chain) can undergo oxidation to form a covalent bond called the “disulfide bridge”—not to be confused with “salt bridge” - The disulfide linkages play a key role in modulating protein function —eg dimerization of insulin receptor (recall § 1. 6)
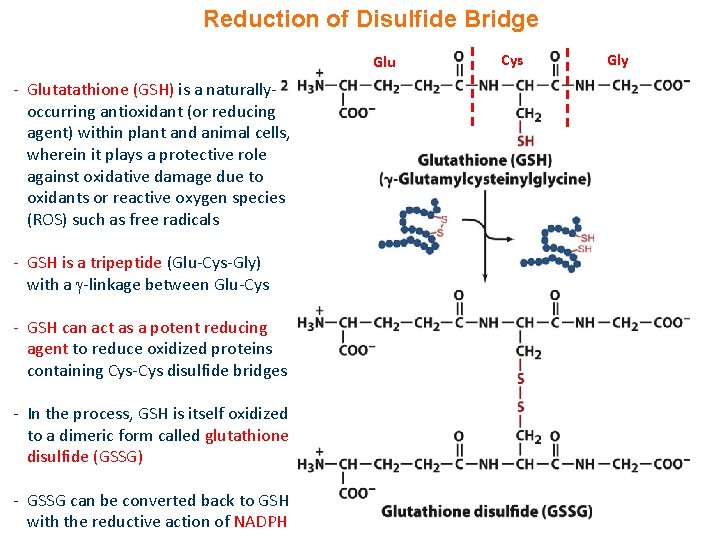
Reduction of Disulfide Bridge Glu - Glutatathione (GSH) is a naturallyoccurring antioxidant (or reducing agent) within plant and animal cells, wherein it plays a protective role against oxidative damage due to oxidants or reactive oxygen species (ROS) such as free radicals - GSH is a tripeptide (Glu-Cys-Gly) with a -linkage between Glu-Cys - GSH can act as a potent reducing agent to reduce oxidized proteins containing Cys-Cys disulfide bridges - In the process, GSH is itself oxidized to a dimeric form called glutathione disulfide (GSSG) - GSSG can be converted back to GSH with the reductive action of NADPH Cys Gly
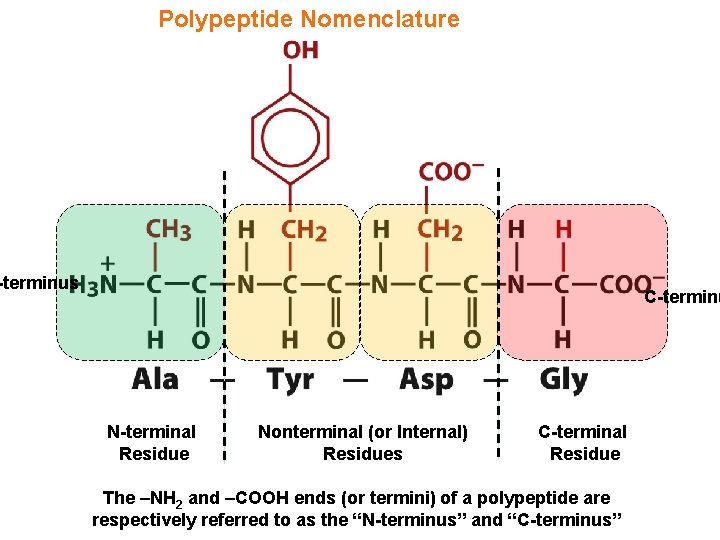
Polypeptide Nomenclature -terminus C-terminu N-terminal Residue Nonterminal (or Internal) Residues C-terminal Residue The –NH 2 and –COOH ends (or termini) of a polypeptide are respectively referred to as the “N-terminus” and “C-terminus”
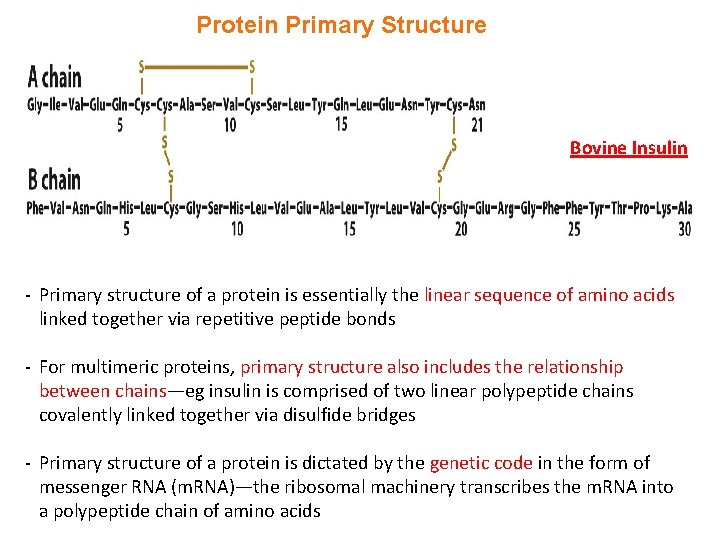
Protein Primary Structure Bovine Insulin - Primary structure of a protein is essentially the linear sequence of amino acids linked together via repetitive peptide bonds - For multimeric proteins, primary structure also includes the relationship between chains—eg insulin is comprised of two linear polypeptide chains covalently linked together via disulfide bridges - Primary structure of a protein is dictated by the genetic code in the form of messenger RNA (m. RNA)—the ribosomal machinery transcribes the m. RNA into a polypeptide chain of amino acids
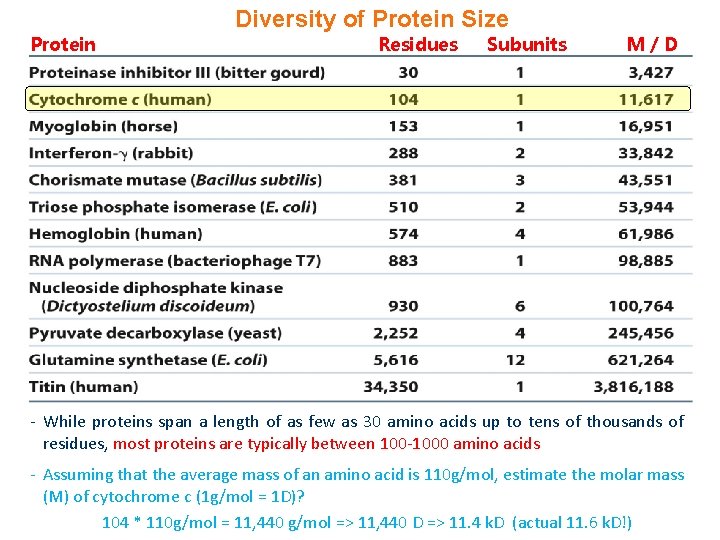
Diversity of Protein Size Protein Residues Subunits M/D - While proteins span a length of as few as 30 amino acids up to tens of thousands of residues, most proteins are typically between 100 -1000 amino acids - Assuming that the average mass of an amino acid is 110 g/mol, estimate the molar mass (M) of cytochrome c (1 g/mol = 1 D)? 104 * 110 g/mol = 11, 440 g/mol => 11, 440 D => 11. 4 k. D (actual 11. 6 k. D!)
![Isoeletric Point (p. I) of Proteins N p. I = (p. Ki/N) i=1 [1] Isoeletric Point (p. I) of Proteins N p. I = (p. Ki/N) i=1 [1]](http://slidetodoc.com/presentation_image_h/23e21b273fb291addd279132db4185a6/image-10.jpg)
Isoeletric Point (p. I) of Proteins N p. I = (p. Ki/N) i=1 [1] - Isoelectric point (p. I) is the p. H at which a molecule (or protein) carries no net charge (q)—eg myoglobin and hemoglobin are largely neutral @ p. H 7 - The net charge of a protein is highly sensitive to solution p. H due to the presence of ionizable residues such as Asp, Glu, Lys, Arg, and His - p. I can be estimated from amino acid sequence alone by summing up all constituent p. K (p. Ki) values (sidechain & terminal) and dividing by the total number (N) of ionizable residues according to Eq [1] - Knowledge of a protein’s p. I is critical to understanding its physical and chemical properties— eg acidic proteins (negatively charged) have a low p. I, while basic proteins (positively charged) harbor a rather high p. I

Exercise 2. 2 a - Draw a Cys–Gly–Asn tripeptide. Identify the peptide bond and the N- and C-termini - Describe protein primary structure - What determines the protein primary structure? - What is a disulfide bridge? - What is a protein’s isoelectric point?

§ 2. 2 b Protein Purification

Synopsis 2. 2 b - The process by which a target protein (or protein-of-interest) is separated from a heterogeneous mixture of other proteins (and/or other impurities such as carbohydrates, lipids, or DNA/RNA) is called “protein purification” - Purified proteins are critical for understanding their mechanism of action—ie how do proteins do what they do—in terms of their thermodynamics, kinetics, and structure - Purification procedures take advantage of a protein’s unique structure and chemistry in order to separate it from other molecules—environmental factors such as p. H and temperature affect the stability of protein during purification - Protein purification in aqueous solution must be carried out at a p. H below or above its p. I by at least one unit—proteins are least soluble at their p. I! - Common purification methods include: 1. Affinity chromatography (AC)—primary method (workhorse of protein purification) 2. Size-exclusion chromatography (SEC)—secondary method 3. Ion exchange chromatography (IEC)—secondary method 4. Salting out (SO)—classical method rarely used today
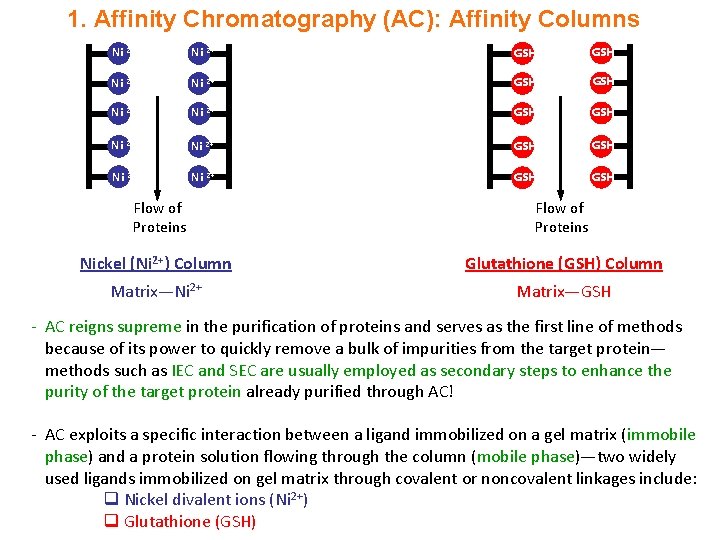
1. Affinity Chromatography (AC): Affinity Columns Ni 2+ Ni 2+ GSH GSH Ni 2+ GSH Flow of Proteins Nickel (Ni 2+) Column Glutathione (GSH) Column Matrix—Ni 2+ Matrix—GSH - AC reigns supreme in the purification of proteins and serves as the first line of methods because of its power to quickly remove a bulk of impurities from the target protein— methods such as IEC and SEC are usually employed as secondary steps to enhance the purity of the target protein already purified through AC! - AC exploits a specific interaction between a ligand immobilized on a gel matrix (immobile phase) and a protein solution flowing through the column (mobile phase)—two widely used ligands immobilized on gel matrix through covalent or noncovalent linkages include: q Nickel divalent ions (Ni 2+) q Glutathione (GSH)
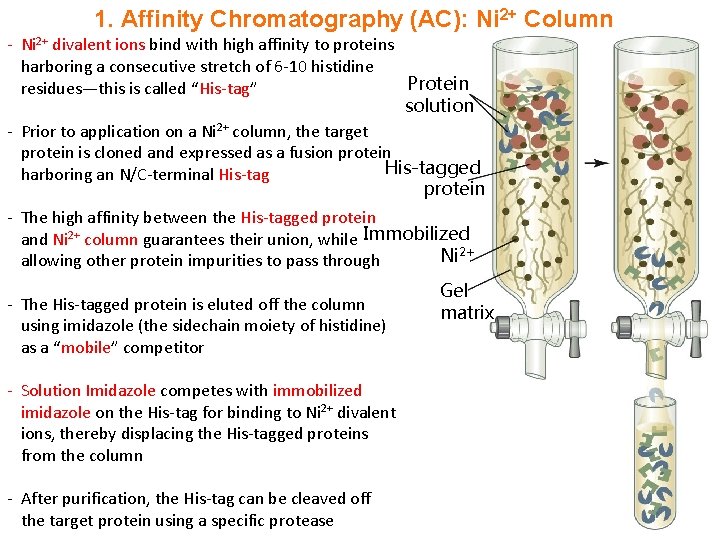
1. Affinity Chromatography (AC): Ni 2+ Column - Ni 2+ divalent ions bind with high affinity to proteins harboring a consecutive stretch of 6 -10 histidine Protein residues—this is called “His-tag” solution - Prior to application on a Ni 2+ column, the target protein is cloned and expressed as a fusion protein His-tagged harboring an N/C-terminal His-tag protein - The high affinity between the His-tagged protein and Ni 2+ column guarantees their union, while Immobilized Ni 2+ allowing other protein impurities to pass through - The His-tagged protein is eluted off the column using imidazole (the sidechain moiety of histidine) as a “mobile” competitor - Solution Imidazole competes with immobilized imidazole on the His-tag for binding to Ni 2+ divalent ions, thereby displacing the His-tagged proteins from the column - After purification, the His-tag can be cleaved off the target protein using a specific protease Gel matrix
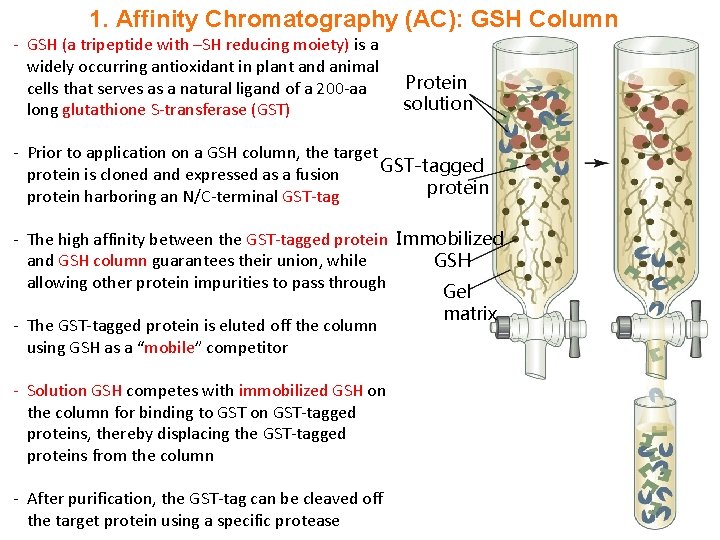
1. Affinity Chromatography (AC): GSH Column - GSH (a tripeptide with –SH reducing moiety) is a widely occurring antioxidant in plant and animal cells that serves as a natural ligand of a 200 -aa long glutathione S-transferase (GST) Protein solution - Prior to application on a GSH column, the target GST-tagged protein is cloned and expressed as a fusion protein harboring an N/C-terminal GST-tag - The high affinity between the GST-tagged protein Immobilized and GSH column guarantees their union, while GSH allowing other protein impurities to pass through Gel matrix - The GST-tagged protein is eluted off the column using GSH as a “mobile” competitor - Solution GSH competes with immobilized GSH on the column for binding to GST on GST-tagged proteins, thereby displacing the GST-tagged proteins from the column - After purification, the GST-tag can be cleaved off the target protein using a specific protease
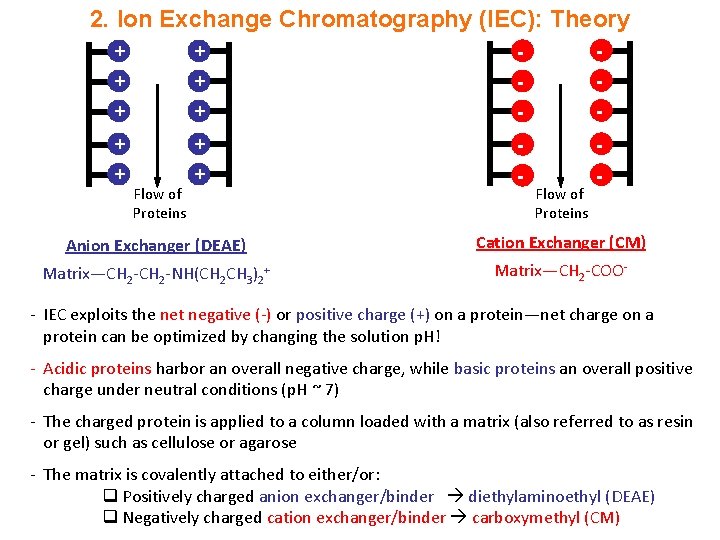
2. Ion Exchange Chromatography (IEC): Theory + + - - + + - - Flow of Proteins Anion Exchanger (DEAE) Cation Exchanger (CM) Matrix—CH 2 -NH(CH 2 CH 3)2+ Matrix—CH 2 -COO- - IEC exploits the net negative (-) or positive charge (+) on a protein—net charge on a protein can be optimized by changing the solution p. H! - Acidic proteins harbor an overall negative charge, while basic proteins an overall positive charge under neutral conditions (p. H ~ 7) - The charged protein is applied to a column loaded with a matrix (also referred to as resin or gel) such as cellulose or agarose - The matrix is covalently attached to either/or: q Positively charged anion exchanger/binder diethylaminoethyl (DEAE) q Negatively charged cation exchanger/binder carboxymethyl (CM)
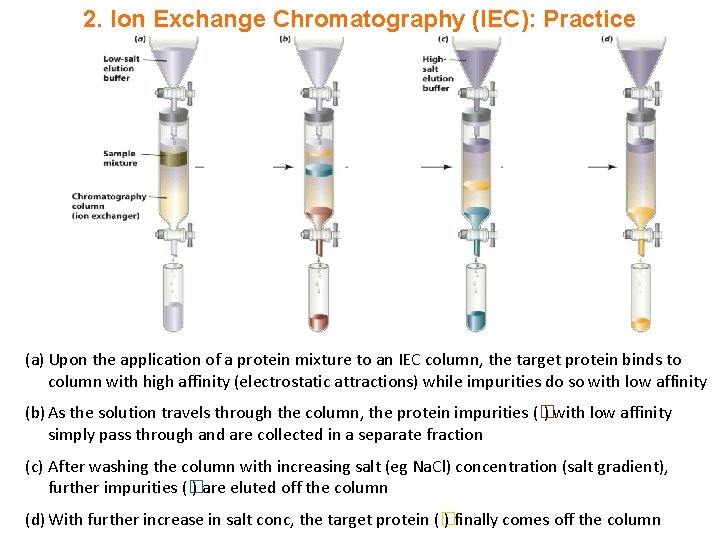
2. Ion Exchange Chromatography (IEC): Practice (a) Upon the application of a protein mixture to an IEC column, the target protein binds to column with high affinity (electrostatic attractions) while impurities do so with low affinity (b) As the solution travels through the column, the protein impurities (� ) with low affinity simply pass through and are collected in a separate fraction (c) After washing the column with increasing salt (eg Na. Cl) concentration (salt gradient), further impurities (� ) are eluted off the column (d) With further increase in salt conc, the target protein (� ) finally comes off the column

3. Size-Exclusion Chromatography (SEC): Theory Small molecules Large molecules - SEC (also called gel filtration) separates proteins solely on the basis of their mass (size) and conformation (shape) - The proteins are applied to a column comprised of a matrix (cross-linked gel beads) with tiny pores for the entry of small molecules, while allowing larger molecules to by-pass the gel matrix Gel bead - SEC essentially acts like a “molecular sieve”, whereby the movement of small molecules down the column (or gel matrix) is impeded due to their entry into the gel pores, while larger molecules are excluded from the gel matrix (they do not enter the gel pores) and thus traverse the column more rapidly - The overall effect is that the larger molecules elute off (or flow through) the column first, whilst smaller molecules emerge last Gel matrix (column)
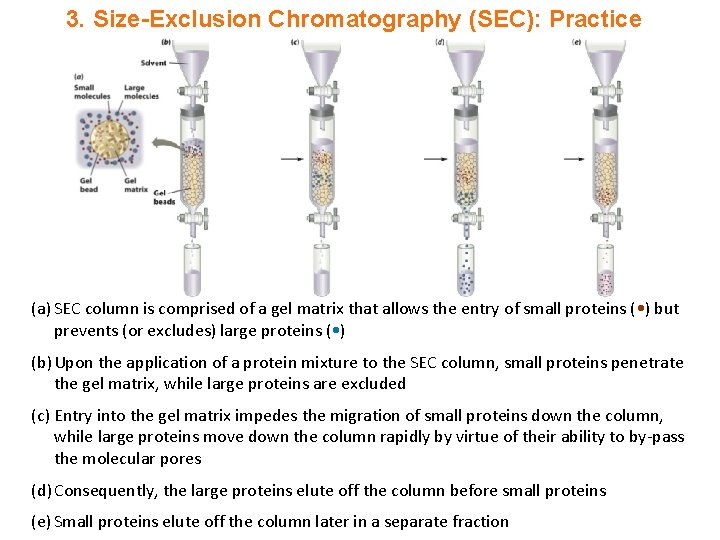
3. Size-Exclusion Chromatography (SEC): Practice (a) SEC column is comprised of a gel matrix that allows the entry of small proteins ( • ) but prevents (or excludes) large proteins ( • ) (b) Upon the application of a protein mixture to the SEC column, small proteins penetrate the gel matrix, while large proteins are excluded (c) Entry into the gel matrix impedes the migration of small proteins down the column, while large proteins move down the column rapidly by virtue of their ability to by-pass the molecular pores (d) Consequently, the large proteins elute off the column before small proteins (e) Small proteins elute off the column later in a separate fraction
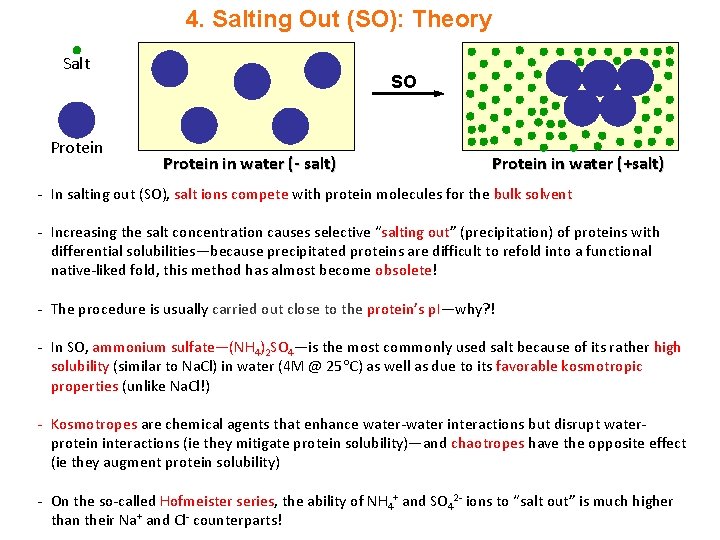
4. Salting Out (SO): Theory Salt Protein SO Protein in water (- salt) Protein in water (+salt) - In salting out (SO), salt ions compete with protein molecules for the bulk solvent - Increasing the salt concentration causes selective “salting out” (precipitation) of proteins with differential solubilities—because precipitated proteins are difficult to refold into a functional native-liked fold, this method has almost become obsolete! - The procedure is usually carried out close to the protein’s p. I—why? ! - In SO, ammonium sulfate—(NH 4)2 SO 4—is the most commonly used salt because of its rather high solubility (similar to Na. Cl) in water (4 M @ 25 C) as well as due to its favorable kosmotropic properties (unlike Na. Cl!) - Kosmotropes are chemical agents that enhance water-water interactions but disrupt waterprotein interactions (ie they mitigate protein solubility)—and chaotropes have the opposite effect (ie they augment protein solubility) - On the so-called Hofmeister series, the ability of NH 4+ and SO 42 - ions to “salt out” is much higher than their Na+ and Cl- counterparts!

4. Salting Out (SO): Practice Solubility: • < • Target Protein (a) A mixture comprised of target protein ( • ) and impurities ( • / • ) in low salt concentrations is subjected to centrifugation to spin down or sediment the least soluble proteins (b) After centrifugation, the precipitate containing the least soluble impurities ( • ) is discarded and the supernatant is recovered (c) After the addition of further salt to the supernatant, the mixture is subjected to centrifugation again to precipitate out the target protein ( • )

Exercise 2. 2 b - Describe environmental conditions that must be controlled while purifying a protein? - Describe the basis for separating proteins by salting out, ion exchange chromatography, size-exclusion chromatography, and affinity chromatography - Describe the basis of SDS-PAGE as used in protein visualization - Describe how absorbance spectroscopy can be used to determine protein concentration

§ 2. 2 c Protein Characterization

Synopsis 2. 2 c - The process of quality control by which a purified protein is analyzed to assess its “purity”, “yield”, and “concentration” has come to be known as “protein characterization” - The purity of a protein is defined as the ratio of the mass of target protein over the total mass of all proteins in a purified sample (impurities due to carbohydrates and nucleic acids are usually ignored unless there is a strong reason to believe that such macromolecules interact with the target protein): - The yield of a protein is defined as the mass of target protein obtained from a given volume of bacterial-plant-or-animal cell culture harvested: - The concentration of a protein is defined as the moles of target protein in a known amount of solution (usually an aqueous buffer): - SDS-PAGE and UVS reign supreme among the first line of methods available in a standard biochemical laboratory for the characterization of newly purified proteins: SDS Sodium dodecyl sulfate PAGE Polyacrylamide gel electrophoresis UVS Ultraviolet-Visible spectrophotometry

SDS-PAGE: Electrophoresis Illustration of an electric field generated by opposite charges - In electrophoresis, charged molecules migrate in a fluid medium under the influence of an electric field (E)—ie application of a potential difference (voltage) across a medium to move charged molecules—the greater the charge, the greater the distance traveled! - In PAGE, the goal is to separate proteins on the basis of their size (mass) as they migrate through a porous polyacrylamide gel matrix across when a potential difference is applied - However, neither charge (determined by amino acid composition) nor shape (determined by 3 D structure) is dependent upon the molar mass of a protein - Thus, two proteins of same size (or molar mass) may experience differential migration in PAGE and vice versa—this methodology is called “Native-PAGE”—and is rarely used because of its poor resolution—enter SDS-PAGE!

SDS-PAGE: SDS Chemistry Sodium dodecyl sulfate (SDS) Apolar Tail (Hydrophobic) Charged Head (Hydrophilic) - SDS-PAGE is one of the most widely used laboratory methods to visualize purified proteins so as to determine their purity in a qualitative manner - SDS is an anionic detergent—detergents (eg hand soaps) are amphiphilic molecules that are used as “cleaning agents” to remove substances such as grease and oil from surfaces - Owing to its amphiphilic character, SDS reigns supreme in its ability to quickly denature proteins in aqueous solutions—a pre-requisite for visualization of proteins on SDS-PAGE - Accordingly, SDS-PAGE is an “invasive” technique in that it destroys the native structure of proteins—in an irreversible manner for all intents and purposes
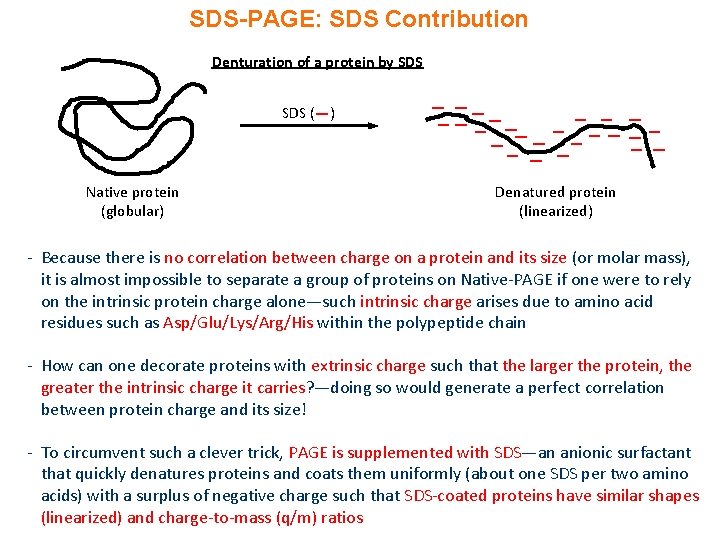
SDS-PAGE: SDS Contribution Denturation of a protein by SDS ( ) Native protein (globular) Denatured protein (linearized) - Because there is no correlation between charge on a protein and its size (or molar mass), it is almost impossible to separate a group of proteins on Native-PAGE if one were to rely on the intrinsic protein charge alone—such intrinsic charge arises due to amino acid residues such as Asp/Glu/Lys/Arg/His within the polypeptide chain - How can one decorate proteins with extrinsic charge such that the larger the protein, the greater the intrinsic charge it carries? —doing so would generate a perfect correlation between protein charge and its size! - To circumvent such a clever trick, PAGE is supplemented with SDS—an anionic surfactant that quickly denatures proteins and coats them uniformly (about one SDS per two amino acids) with a surplus of negative charge such that SDS-coated proteins have similar shapes (linearized) and charge-to-mass (q/m) ratios
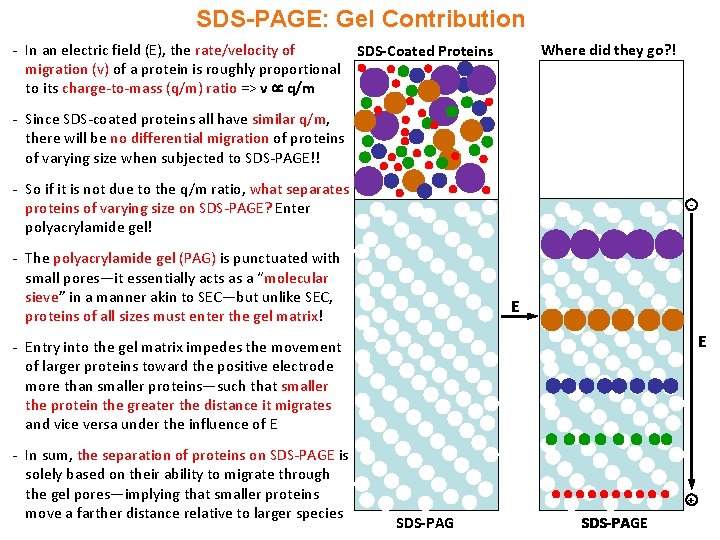
SDS-PAGE: Gel Contribution - In an electric field (E), the rate/velocity of migration (v) of a protein is roughly proportional to its charge-to-mass (q/m) ratio => v q/m Where did they go? ! SDS-Coated Proteins - Since SDS-coated proteins all have similar q/m, there will be no differential migration of proteins of varying size when subjected to SDS-PAGE!! - So if it is not due to the q/m ratio, what separates proteins of varying size on SDS-PAGE? Enter polyacrylamide gel! - - The polyacrylamide gel (PAG) is punctuated with small pores—it essentially acts as a “molecular sieve” in a manner akin to SEC—but unlike SEC, proteins of all sizes must enter the gel matrix! E E - Entry into the gel matrix impedes the movement of larger proteins toward the positive electrode more than smaller proteins—such that smaller the protein the greater the distance it migrates and vice versa under the influence of E - In sum, the separation of proteins on SDS-PAGE is solely based on their ability to migrate through the gel pores—implying that smaller proteins move a farther distance relative to larger species + SDS-PAGE SDS-PAG
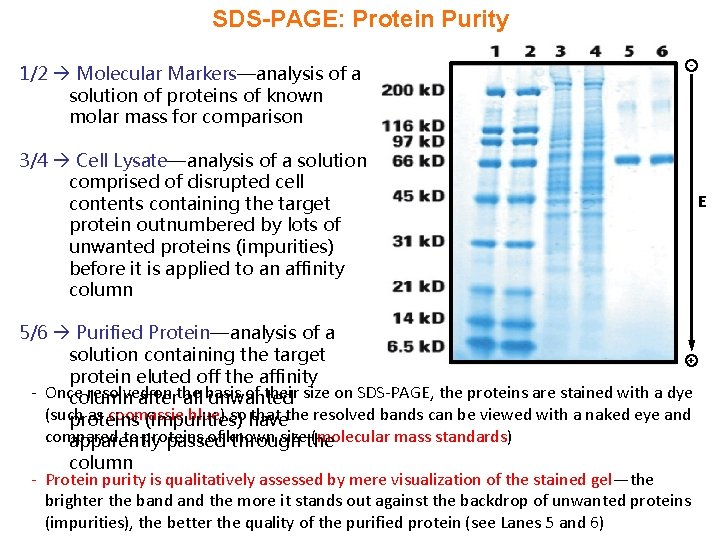
SDS-PAGE: Protein Purity 1/2 Molecular Markers—analysis of a solution of proteins of known molar mass for comparison - 3/4 Cell Lysate—analysis of a solution comprised of disrupted cell contents containing the target protein outnumbered by lots of unwanted proteins (impurities) before it is applied to an affinity column 5/6 Purified Protein—analysis of a solution containing the target + protein eluted off the affinity - Once resolved on the of their size on SDS-PAGE, the proteins are stained with a dye column after all basis unwanted (such as coomassie blue) so have that the resolved bands can be viewed with a naked eye and proteins (impurities) compared to proteins of known sizethe (molecular mass standards) apparently passed through column - Protein purity is qualitatively assessed by mere visualization of the stained gel—the brighter the band the more it stands out against the backdrop of unwanted proteins (impurities), the better the quality of the purified protein (see Lanes 5 and 6) E
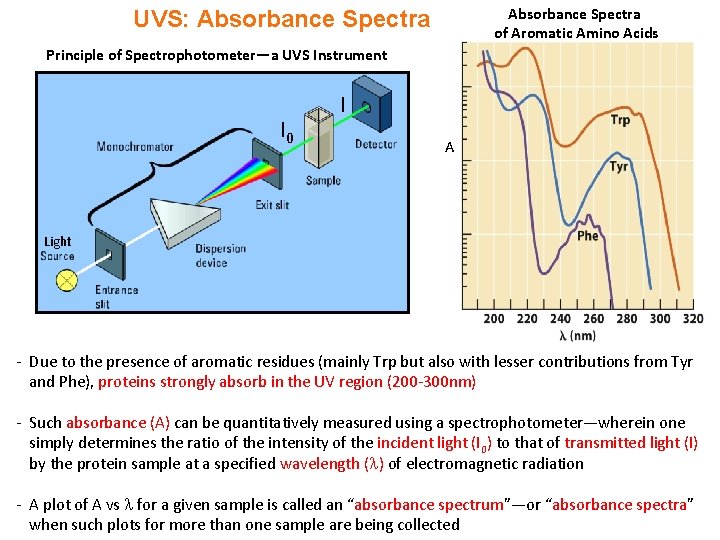
Absorbance Spectra of Aromatic Amino Acids UVS: Absorbance Spectra Principle of Spectrophotometer—a UVS Instrument I 0 I A Light - Due to the presence of aromatic residues (mainly Trp but also with lesser contributions from Tyr and Phe), proteins strongly absorb in the UV region (200 -300 nm) - Such absorbance (A) can be quantitatively measured using a spectrophotometer—wherein one simply determines the ratio of the intensity of the incident light (I 0) to that of transmitted light (I) by the protein sample at a specified wavelength ( ) of electromagnetic radiation - A plot of A vs for a given sample is called an “absorbance spectrum”—or “absorbance spectra” when such plots for more than one sample are being collected
![UVS: Protein Concentration N = i i=1 August Beer (1825 -1863) [2] Johann Lambert UVS: Protein Concentration N = i i=1 August Beer (1825 -1863) [2] Johann Lambert](http://slidetodoc.com/presentation_image_h/23e21b273fb291addd279132db4185a6/image-32.jpg)
UVS: Protein Concentration N = i i=1 August Beer (1825 -1863) [2] Johann Lambert (1728 -1777) - The spectroscopic properties of proteins—ie their ability to absorb in the UV region—can be exploited to quantify their concentrations according to the Beer-Lambert law: A = log(Io/I) = c. L [1] A = Protein absorbance (unitless) @ 280 nm = Protein extinction coefficient (M-1 cm-1) @ 280 nm c = Protein concentration (M) => M = mol. L-1 L = Cuvette (light) pathlength (cm) I 0 = Intensity of incident light (cd) I = Intensity of transmitted light (cd) - For most proteins, can be estimated from amino acid sequence alone using Eq[2]—via summation of all the constituent ( i) values of individual residues (eg Trp and Tyr) @ 280 nm
![UVS: Units of - Prove that the units of are: [ ] = M-1. UVS: Units of - Prove that the units of are: [ ] = M-1.](http://slidetodoc.com/presentation_image_h/23e21b273fb291addd279132db4185a6/image-33.jpg)
UVS: Units of - Prove that the units of are: [ ] = M-1. cm-1 = L. mol-1. cm-1 - From Beer-Lambert law: A = cl [1] - Rearranging Eq [1] for gives: = A/cl [2] [ ] = [A]/[c][l] [3] where the square brackets in this context indicate the units of the enclosed parameter - But, the units of A, c and l are: [A] = 1 (unitless) [c] = mol. L-1 [l] = cm - Now, substituting into Eq [3] gives: [ ] = 1 / [(mol. L-1). (cm)] => [ ] = L. mol-1. cm-1 => [ ] = M-1. cm-1

UVS: Molar vs Mass Concentration - Protein concentration is usually expressed in terms of moles per unit volume: - Protein concentration can also be expressed in terms of mass per unit volume: - Given the molar mass of a protein, the molar and mass concentrations can also be easily interchanged:

UVS: Protein Yield - Protein yield—ie how much purified protein is obtained from a given amount of cell culture harvested—can be determined from the knowledge of protein concentration - From the Beer-Lambert law, protein concentration is calculated in the units of mol/L—ie the concentration (c) is simply the number of moles (n) of a protein in a known volume (V) of solution (the purified protein sample): c[mol/L] = n[mol] / V[L] [1] - In terms of the absolute mass (m) and molar mass (M) of a protein in the purified sample, n can be expressed as: n[mol] = m[g] / M[g/mol] [2] - Combining Eqs [1] and [2] gives: c [mol/L] = m[g] / V[L]. M[g/mol] => m[g] = c[mol/L]. V[L]. M[g/mol] [3] [4] - Using eq[4], we can thus calculate milligrams (mg) of protein in a purified protein solution—usually a few milliliters (ml) - Protein yield is thus simply obtained by dividing the mass (mg) of purified protein by the volume (L) of cell culture harvested

Exercise 2. 2 c - Describe environmental conditions that must be controlled while purifying a protein? - Describe the basis for separating proteins by salting out, ion exchange chromatography, size-exclusion chromatography, and affinity chromatography - Describe the basis of SDS-PAGE as used in protein visualization - Describe how UVS can be used to determine protein concentration

§ 2. 2 d Protein Sequencing

Synopsis 2. 2 d - To be sequenced, a protein must be separated into individual polypeptides that can be cleaved into sets of overlapping fragments - Protein sequence data are deposited in online databases such as Uni. Prot @ http: //uniprot. org - In practice, protein sequencing is an ancient art that is rarely exercised in the 21 st century—why sequence a protein when one can simply obtain the primary structure from its corresponding coding sequence (gene) on the basis of genetic codon (see § 4. 5) - Nonetheless, as students of biochemistry, we must at least understand the rationale underlying the science of protein sequencing
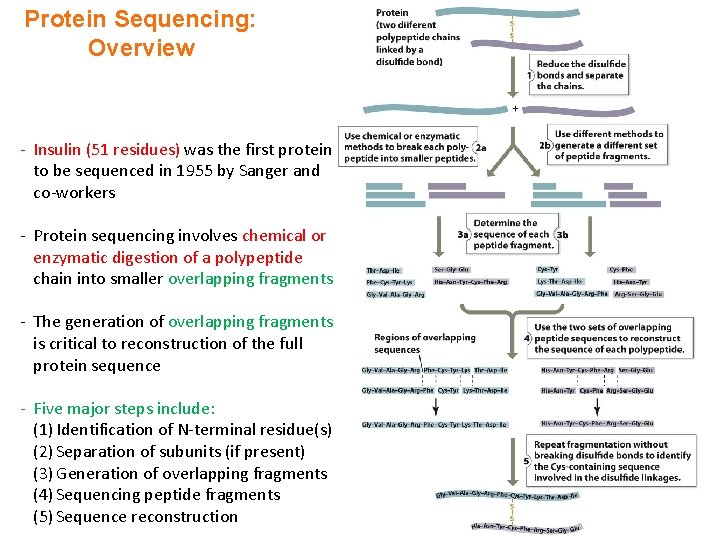
Protein Sequencing: Overview - Insulin (51 residues) was the first protein to be sequenced in 1955 by Sanger and co-workers - Protein sequencing involves chemical or enzymatic digestion of a polypeptide chain into smaller overlapping fragments - The generation of overlapping fragments is critical to reconstruction of the full protein sequence - Five major steps include: (1) Identification of N-terminal residue(s) (2) Separation of subunits (if present) (3) Generation of overlapping fragments (4) Sequencing peptide fragments (5) Sequence reconstruction

(1) Identification of N-terminal Residue(s) - N-terminal analysis allows identification of a distinct number of N-terminal residues and hence the total number of distinct polypeptide chains in a multi-subunit protein - Reaction of a polypeptide chain with dansyl chloride under alkaline conditions generates a dansyl polypeptide - Subsequent acid treatment of dansyl polypeptide yields the dansylamino acid— the dansylated derivative of N-terminal residue - Dansylamino acid can be identified by a combination of chromatographic and spectroscopic methods - Above analysis for insulin yields equal amounts of Gly and Phe—thereby implying that insulin is comprised of two distinct polypeptide chains
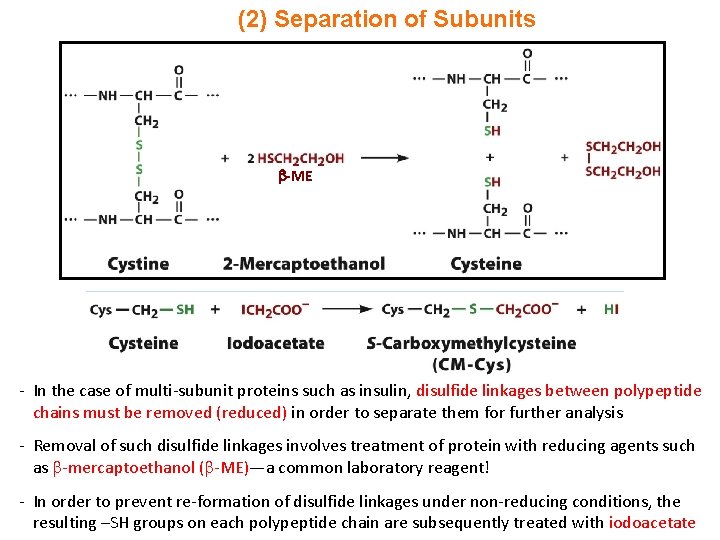
(2) Separation of Subunits -ME - In the case of multi-subunit proteins such as insulin, disulfide linkages between polypeptide chains must be removed (reduced) in order to separate them for further analysis - Removal of such disulfide linkages involves treatment of protein with reducing agents such as -mercaptoethanol ( -ME)—a common laboratory reagent! - In order to prevent re-formation of disulfide linkages under non-reducing conditions, the resulting –SH groups on each polypeptide chain are subsequently treated with iodoacetate
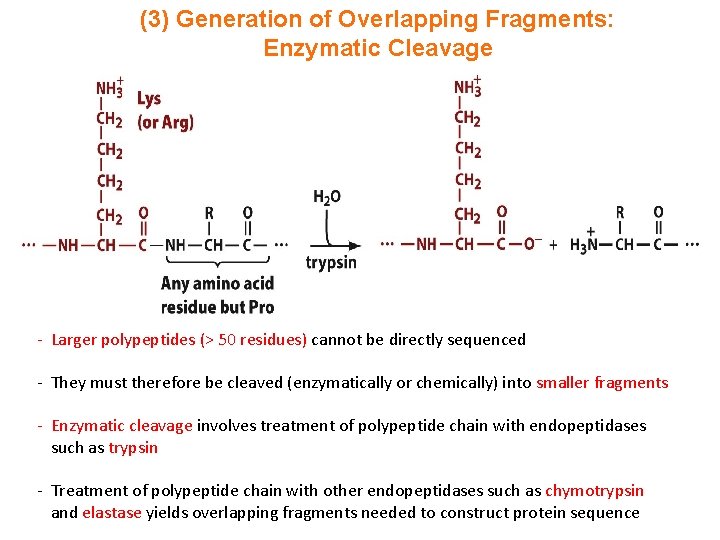
(3) Generation of Overlapping Fragments: Enzymatic Cleavage - Larger polypeptides (> 50 residues) cannot be directly sequenced - They must therefore be cleaved (enzymatically or chemically) into smaller fragments - Enzymatic cleavage involves treatment of polypeptide chain with endopeptidases such as trypsin - Treatment of polypeptide chain with other endopeptidases such as chymotrypsin and elastase yields overlapping fragments needed to construct protein sequence

(3) Generation of Overlapping Fragments: Endopeptidase Specificity - Endopeptidases catalyze the hydrolysis of internal peptide bonds in a highly specific manner —of all endopeptidases, trypsin has the greatest specificity! - Specificity of endopeptidases is derived from their requirement of residues flanking the scissile peptide bond—ie the peptide bond undergoing scission/cleavage
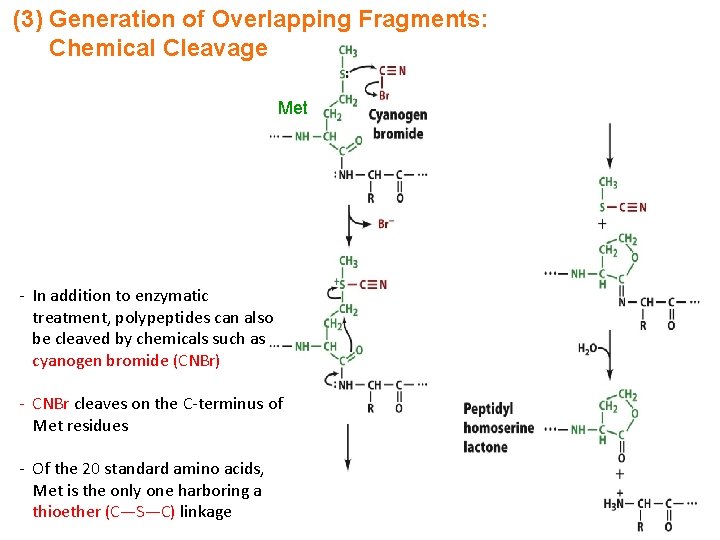
(3) Generation of Overlapping Fragments: Chemical Cleavage Met - In addition to enzymatic treatment, polypeptides can also be cleaved by chemicals such as cyanogen bromide (CNBr) - CNBr cleaves on the C-terminus of Met residues - Of the 20 standard amino acids, Met is the only one harboring a thioether (C—S—C) linkage
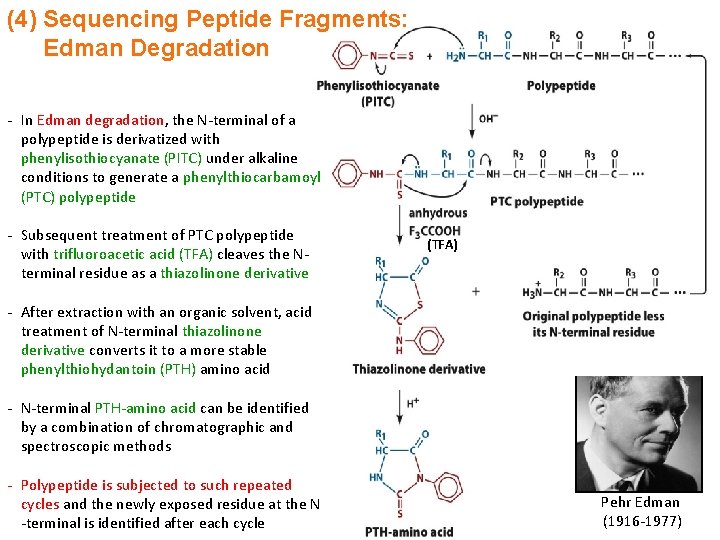
(4) Sequencing Peptide Fragments: Edman Degradation - In Edman degradation, the N-terminal of a polypeptide is derivatized with phenylisothiocyanate (PITC) under alkaline conditions to generate a phenylthiocarbamoyl (PTC) polypeptide - Subsequent treatment of PTC polypeptide with trifluoroacetic acid (TFA) cleaves the Nterminal residue as a thiazolinone derivative (TFA) - After extraction with an organic solvent, acid treatment of N-terminal thiazolinone derivative converts it to a more stable phenylthiohydantoin (PTH) amino acid - N-terminal PTH-amino acid can be identified by a combination of chromatographic and spectroscopic methods - Polypeptide is subjected to such repeated cycles and the newly exposed residue at the N -terminal is identified after each cycle Pehr Edman (1916 -1977)
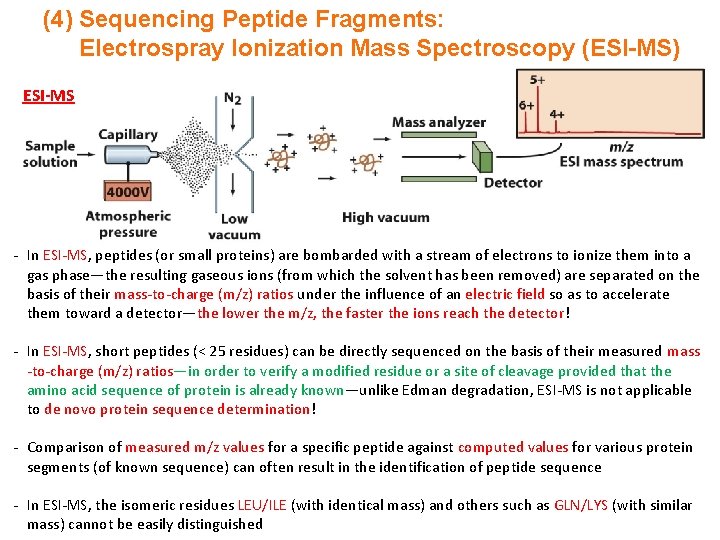
(4) Sequencing Peptide Fragments: Electrospray Ionization Mass Spectroscopy (ESI-MS) ESI-MS - In ESI-MS, peptides (or small proteins) are bombarded with a stream of electrons to ionize them into a gas phase—the resulting gaseous ions (from which the solvent has been removed) are separated on the basis of their mass-to-charge (m/z) ratios under the influence of an electric field so as to accelerate them toward a detector—the lower the m/z, the faster the ions reach the detector! - In ESI-MS, short peptides (< 25 residues) can be directly sequenced on the basis of their measured mass -to-charge (m/z) ratios—in order to verify a modified residue or a site of cleavage provided that the amino acid sequence of protein is already known—unlike Edman degradation, ESI-MS is not applicable to de novo protein sequence determination! - Comparison of measured m/z values for a specific peptide against computed values for various protein segments (of known sequence) can often result in the identification of peptide sequence - In ESI-MS, the isomeric residues LEU/ILE (with identical mass) and others such as GLN/LYS (with similar mass) cannot be easily distinguished
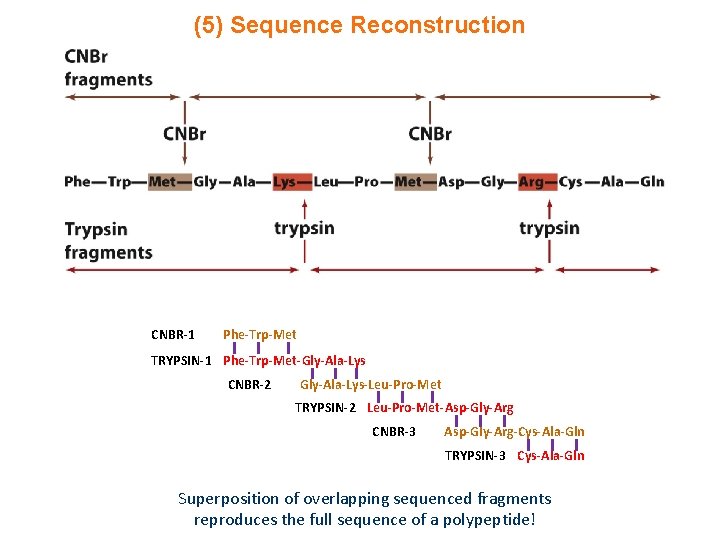
(5) Sequence Reconstruction CNBR-1 Phe-Trp-Met TRYPSIN-1 Phe-Trp-Met-Gly-Ala-Lys CNBR-2 Gly-Ala-Lys-Leu-Pro-Met TRYPSIN-2 Leu-Pro-Met-Asp-Gly-Arg CNBR-3 Asp-Gly-Arg-Cys-Ala-Gln TRYPSIN-3 Cys-Ala-Gln Superposition of overlapping sequenced fragments reproduces the full sequence of a polypeptide!

Exercise 2. 2 d - Summarize the steps involved in sequencing a protein - Why is it important to identify the N-terminal residue(s) of a protein? - Explain why long polypeptides must be broken into at least two different sets of peptide fragments for sequencing
- Slides: 48