The Pathway Tools Schema SRI International Bioinformatics Motivations

The Pathway Tools Schema

SRI International Bioinformatics Motivations for Understanding Schema l Pathway Tools visualizations and analyses depend upon the software being able to find precise information in precise places within a Pathway/Genome DB l When writing complex queries to PGDBs, those queries must name classes and slots within the schema l. A Pathway/Genome Database is a web of interconnected objects; each object represents a biological entity

Reference l Pathway SRI International Bioinformatics Tools User’s Guide, Volume I l Appendix A: Guide to the Pathway Tools Schema
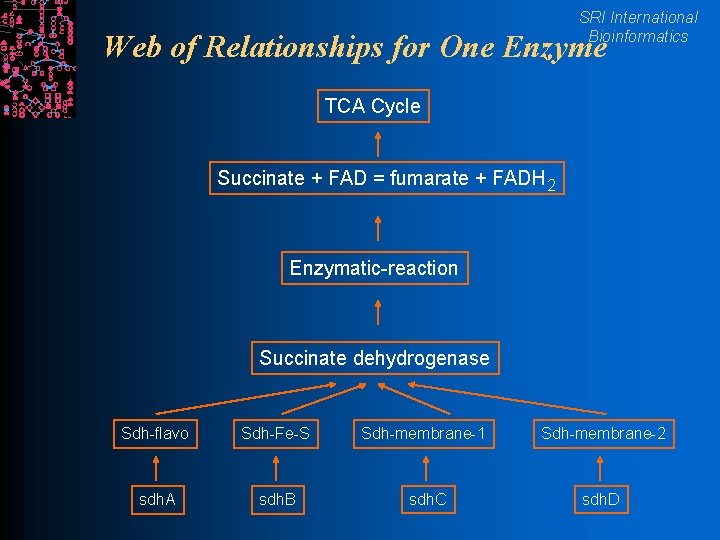
SRI International Bioinformatics Web of Relationships for One Enzyme TCA Cycle Succinate + FAD = fumarate + FADH 2 Enzymatic-reaction Succinate dehydrogenase Sdh-flavo Sdh-Fe-S Sdh-membrane-1 Sdh-membrane-2 sdh. A sdh. B sdh. C sdh. D

Frame Data Model l Frame PGDB SRI International Bioinformatics Data Model -- organizational structure for a l Knowledge base (KB, Database, DB) l Frames l Slots l Facets l Annotations

Knowledge Base l Collection SRI International Bioinformatics of frames and their associated slots, values, facets, and annotations l AKA: Database, PGDB l Can be stored within l An Oracle DB l A disk file l A Pathway Tools binary program

Frames SRI International Bioinformatics l Entities with which facts are associated l Kinds of frames: l Classes: Genes, Pathways, Biosynthetic Pathways l Instances (objects): trp. A, TCA cycle l Classes: l Superclass(es) l Subclass(es) l Instance(s) l A symbolic frame name (id, key) uniquely identifies each frame

Frame IDs SRI International Bioinformatics l Naming conventions for frame IDs l Uniqueness of frame IDs l Frame IDs must be unique within a PGDB l Goal: Same frame ID within different PGDBs should refer to the same biological entity l Because many frames are imported from Meta. Cyc, this helps ensure consistency of frame names l Frame IDs for newly created frames (not imported) are generated by Pathway Tools u u Those frame IDs contain a PGDB-specific identifier Example: CPLXzz-nnnn CPLXB 3 -0035

Slots SRI International Bioinformatics l Encode attributes/properties of a frame l Integer, real number, string, symbols l Represent relationships between frames l The value of a slot is the identifier of another frame l Every slot is described by a “slot frame” in a KB that defines meta information about that slot
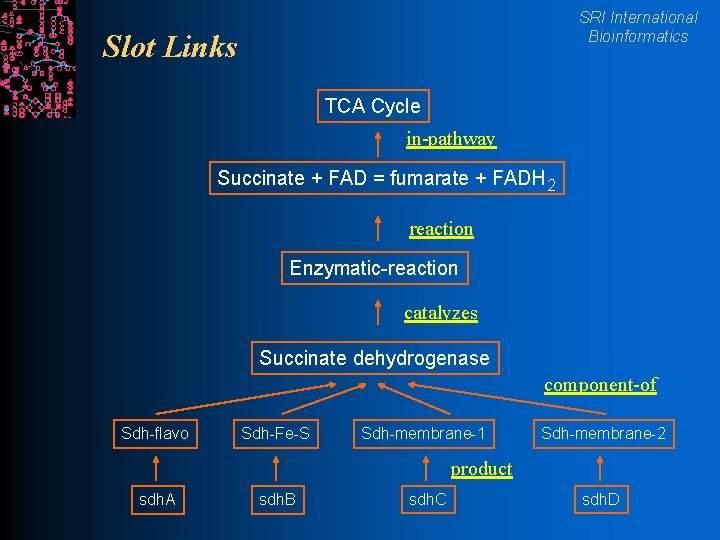
SRI International Bioinformatics Slot Links TCA Cycle in-pathway Succinate + FAD = fumarate + FADH 2 reaction Enzymatic-reaction catalyzes Succinate dehydrogenase component-of Sdh-flavo Sdh-Fe-S Sdh-membrane-1 Sdh-membrane-2 product sdh. A sdh. B sdh. C sdh. D

Slots SRI International Bioinformatics l Number of values l Single valued l Multivalued: sets, bags l Slot values l Any LISP object: Integer, real, string, symbol (frame name) l Slotunits define properties of slots: datatypes, classes, constraints l Two slots are inverses if they encode opposite relationships l Slot Product in class Genes
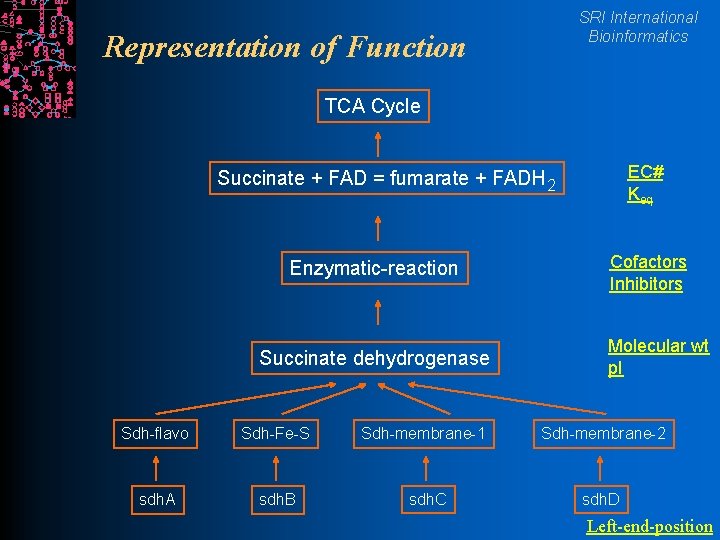
SRI International Bioinformatics Representation of Function TCA Cycle EC# Keq Succinate + FAD = fumarate + FADH 2 Enzymatic-reaction Succinate dehydrogenase Cofactors Inhibitors Molecular wt p. I Sdh-flavo Sdh-Fe-S Sdh-membrane-1 Sdh-membrane-2 sdh. A sdh. B sdh. C sdh. D Left-end-position
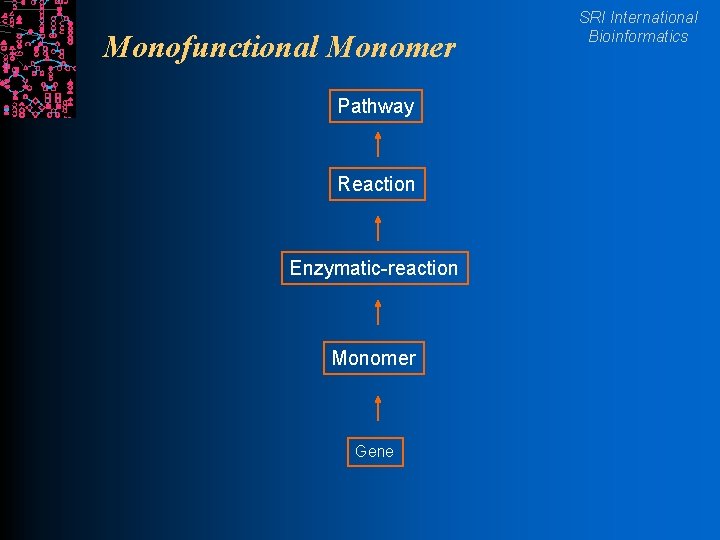
Monofunctional Monomer Pathway Reaction Enzymatic-reaction Monomer Gene SRI International Bioinformatics
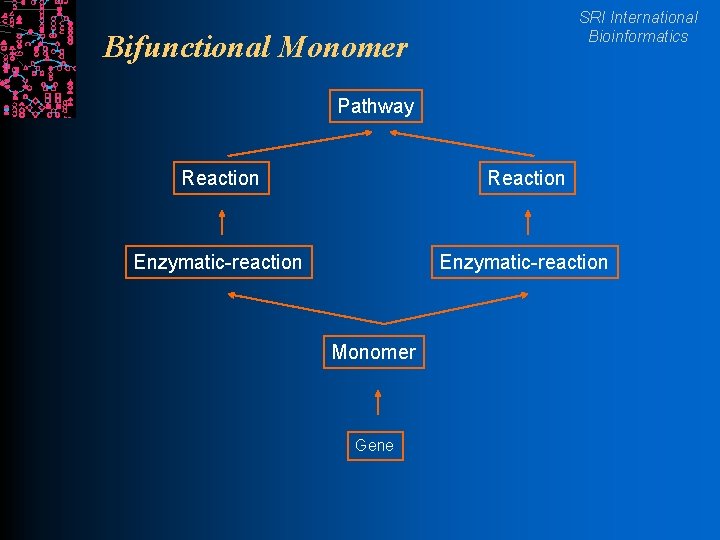
SRI International Bioinformatics Bifunctional Monomer Pathway Reaction Enzymatic-reaction Monomer Gene
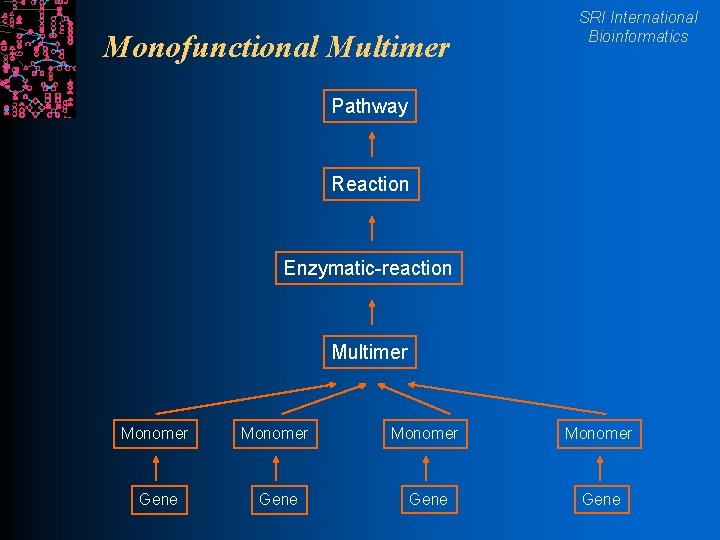
Monofunctional Multimer SRI International Bioinformatics Pathway Reaction Enzymatic-reaction Multimer Monomer Gene
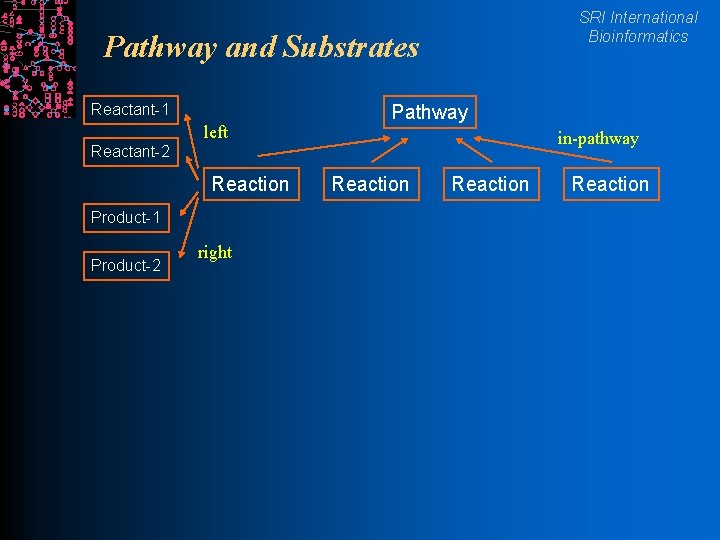
SRI International Bioinformatics Pathway and Substrates Reactant-1 left Pathway in-pathway Reactant-2 Reaction Product-1 Product-2 right Reaction
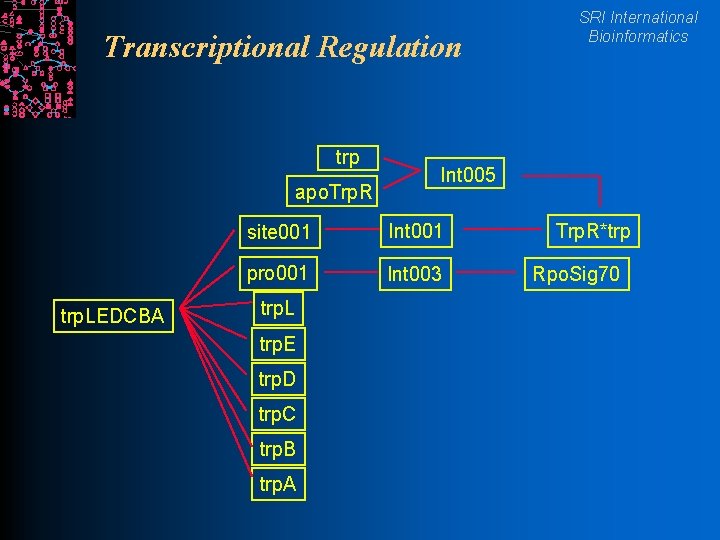
Transcriptional Regulation trp apo. Trp. R trp. LEDCBA Int 005 site 001 Int 001 pro 001 Int 003 trp. L trp. E trp. D trp. C trp. B trp. A SRI International Bioinformatics Trp. R*trp Rpo. Sig 70

Principle Classes SRI International Bioinformatics l Class names are capitalized, plural, separated by dashes l Genetic-Elements, with subclasses: l Chromosomes l Plasmids Genes Transcription-Units RNAs l r. RNAs, sn. RNAs, t. RNAs, Charged-t. RNAs Proteins, with subclasses: l Polypeptides l Protein-Complexes l l

Principle Classes l Reactions, with subclasses: l Transport-Reactions l Enzymatic-Reactions l Pathways l Compounds-And-Elements SRI International Bioinformatics

Slots in Multiple Classes l Common-Name l Synonyms l Comment l Citations l DB-Links SRI International Bioinformatics

Genes Slots l Component-Of SRI International Bioinformatics (links to replicon, transcription unit) l Left-End-Position l Right-End-Position l Centisome-Position l Transcription-Direction l Product

Proteins Slots l Molecular-Weight-Seq l Molecular-Weight-Exp l p. I l Locations l Modified-Form l Unmodified-Form l Component-Of SRI International Bioinformatics

Polypeptides Slots l Gene SRI International Bioinformatics

Protein-Complexes Slots l Components SRI International Bioinformatics

Reactions Slots l EC-Number l Left, Right l Delta. G 0 l Keq l Spontaneous? SRI International Bioinformatics

Enzymatic-Reactions Slots l Enzyme l Reaction l Activators l Inhibitors l Physiologically-Relevant l Cofactors l Prosthetic-Groups l Alternative-Substrates l Alternative-Cofactors SRI International Bioinformatics

Pathways Slots l Reaction-List l Predecessors l Primaries SRI International Bioinformatics

GKB Editor l Browse l Tools l GKB SRI International Bioinformatics class hierarchy and slot definitions -> Ontology Browser Editor described at l http: //www. ai. sri. com/~gkb/user-man. html

Pathway Tools Data Access Mechanisms

SRI International Bioinformatics Introduction l MANY l APIs ways to access and update PGDBs in Java, Perl, and Lisp l Import/export l Registry l Import of files in many formats of Pathway/Genome Databases PGDB data into Bio. Warehouse l Updating a PGDB from an external genome DB

Pathway Tools APIs l Support SRI International Bioinformatics programmatic queries and updates to PGDBs l APIs in Java, Perl, and Lisp all provide access to a common set of procedures: l Generic Frame Protocol -- Ocelot object database API l Additional Pathway Tools functions l For more information see l http: //bioinformatics. ai. sri. com/ptoolsresources. html

Generic Frame Protocol (GFP) l. A SRI International Bioinformatics library of procedures for accessing Ocelot DBs l GFP specification: l http: //www. ai. sri. com/~gfp/spec/paper. html l. A small number of GFP functions are sufficient for most complex queries l Knowledge of Pathway Tools schema is critical for using the APIs: l Appendix I of Pathway Tools User’s Guide, Vol I

Generic Frame Protocol l get-class-all-instances (Class) l Returns the instances of Class l Key Pathway Tools classes: l Genetic-Elements l Genes l Proteins l Polypeptides (a subclass of Proteins) l Protein-Complexes (a subclass of Proteins) l Pathways l Reactions l Compounds-And-Elements l Enzymatic-Reactions l Transcription-Units l Promoters l DNA-Binding-Sites SRI International Bioinformatics

Generic Frame Protocol SRI International Bioinformatics l Notation Frame. Slot means a specified slot of a specified frame l get-slot-value(Frame Slot) l Returns first value of Frame. Slot get-slot-values(Frame Slot) l Returns all values of Frame. Slot as a list l l slot-has-value-p(Frame Slot) l Returns T if Frame. Slot has at least one value member-slot-value-p(Frame Slot Value) l Returns T if Value is one of the values of Frame. Slot print-frame(Frame) l Prints the contents of Frame

Generic Frame Protocol l coercible-to-frame-p SRI International Bioinformatics (Thing) l Returns T if Thing is the name of a frame, or a frame object l save-kb l Saves the current KB

Generic Frame Protocol – Update Operations SRI International Bioinformatics l put-slot-value(Frame Slot Value) l Replace the current value(s) of Frame. Slot with Value l put-slot-values(Frame Slot Value-List) l Replace the current value(s) of Frame. Slot with Value-List, which must be a list of values l add-slot-value(Frame Slot Value) l Add Value to the current value(s) of Frame. Slot, if any l remove-slot-value(Frame Slot Value) l Remove Value from the current value(s) of Frame. slot l replace-slot-value(Frame Slot Old-Value New-Value) l In Frame. Slot, replace Old-Value with New-Value l remove-local-slot-values(Frame Slot) l Remove all of the values of Frame. Slot

SRI International Bioinformatics Additional Pathway Tools Functions – Semantic Inference Layer l Semantic inference layer defines built-in functions to compute commonly required relationships in a PGDB l http: //bioinformatics. ai. sri. com/ptoolsfns. html

Internal note l Note: SRI International Bioinformatics Refer to local copy of ptools-fns. html to go through the semantic inference layer fns

File Import/Export Capabilities SRI International Bioinformatics l PGDBs can be exported in whole or part to: l SBML – Systems Biology Markup Language – sbml. org l u Import supported by many simulation packages u File -> Export -> Selected Reactions to SBML File Pathway Tools Attribute-Value format and columndelimited format files u u u http: //brg. ai. sri. com/ptools/flatfile-format. shtml Dump entire PGDB to a suite of files: File -> Export -> Entire DB to Flat Files Dump selected frames to a single file: File -> Export -> Selected Frames to File

Import/Export l Import from attribute-value or column-delimited files l l File -> Import -> Frames From File Import/Export to/from internal Pathway Tools format that allows pathways, reactions, enzymes, and compounds to be easily moved between Pathway Tools installations u u u l SRI International Bioinformatics Edit -> Add Pathway to File Export List File -> Export -> Selected Pathways to File -> Import -> Pathways from File Import/Export to/from MDL molfile format l Edit -> Import compound structure from molfile l Edit -> Export compound structure to molfile

Miscellaneous Exports l l SRI International Bioinformatics Overview -> Highlight -> Save to File Overview -> Highlight -> Load from File Gene / Protein Sequence / Save to file Chromosome -> Show Sequence of a Segment of Replicon

Napster Comes to Bioinformatics l Public l SRI International Bioinformatics sharing of Pathway/Genome Databases PGDB registry maintained by SRI at URL http: //biocyc. org/registry. html l Registry operations l List contents of registry l Download PGDBs listed in the registry l Register PGDBs you have created

Registry Details SRI International Bioinformatics l Why register your PGDB? l Declare existence of your PGDB in a central location l Facilitate download by other scientists l Why download a PGDB? l Desktop Navigator provides more functionality than Web l Comparative operations l Programmatic querying and processing of PGDB l Registration process l Registered PGDBs have open availability by

Bio. Warehouse l Biospice. org SRI International Bioinformatics

New Import/Export Tools l Suggestions? l Volunteers? SRI International Bioinformatics

Updating a PGDB From an External Genome DB l Example: SRI International Bioinformatics Ara. Cyc forms a pathway module to the TAIR DB l TAIR is authoritative source for gene and geneproduct information l Update Ara. Cyc to reflect updates in TAIR

Proposed Approach l l l SRI International Bioinformatics Export TAIR to Patho. Logic files Build Ara. Cyc 2 from those Patho. Logic files – automated Patho. Logic only Compare Ara. Cyc 1 (A 1) to Ara. Cyc 2 (A 2) A. Import new genes/proteins from A 2 to A 1 B. Delete from A 1 genes/proteins not found in A 2 C. Rename genes/proteins whose names changed from A 2 to A 1 l Run name matcher on A 1’ l Check for pathways with no enzymes and report them so user can keep any that otherwise Patho. Logic will delete u What about enzymes that were assigned to a pathway by the hole filler? l Re-run pathway predictor Remember what pathways user deletes so they are not repredicted by Patho. Logic l Consider movement of genes from contig to chromosome l
- Slides: 47