The Pathway Tools Schema SRI International Bioinformatics Motivations

The Pathway Tools Schema

SRI International Bioinformatics Motivations for Understanding Schema l Pathway Tools visualizations and analyses depend upon the software being able to find precise information in precise places within a Pathway/Genome DB l When writing complex queries to PGDBs, those queries must name classes and slots within the schema l. A Pathway/Genome Database is a web of interconnected objects; each object represents a biological entity

Reference l Pathway SRI International Bioinformatics Tools User’s Guide, Volume I l Appendix A: Guide to the Pathway Tools Schema
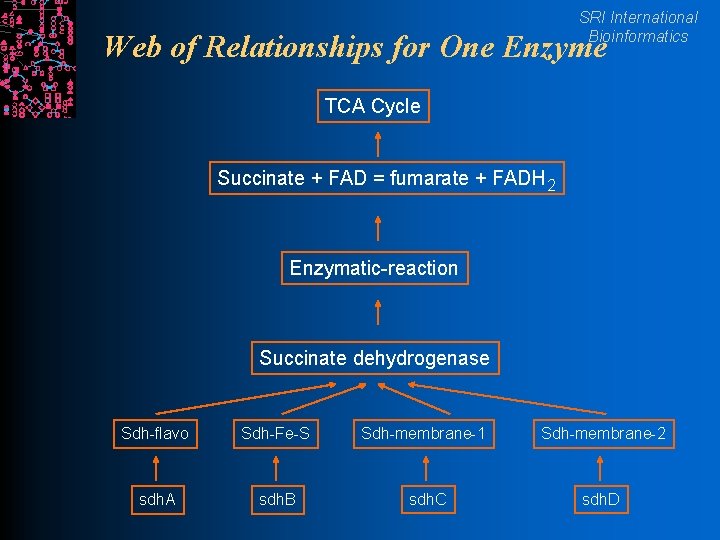
SRI International Bioinformatics Web of Relationships for One Enzyme TCA Cycle Succinate + FAD = fumarate + FADH 2 Enzymatic-reaction Succinate dehydrogenase Sdh-flavo Sdh-Fe-S Sdh-membrane-1 Sdh-membrane-2 sdh. A sdh. B sdh. C sdh. D

Frame Data Model l Frame PGDB SRI International Bioinformatics Data Model -- organizational structure for a l Knowledge base (KB, Database, DB) l Frames l Slots l Facets l Annotations

Knowledge Base l Collection SRI International Bioinformatics of frames and their associated slots, values, facets, and annotations l AKA: Database, PGDB l Can be stored within l An Oracle or My. SQL DB l A disk file l Pathway Tools binary program

Frames SRI International Bioinformatics l Entities with which facts are associated l Kinds of frames: l Classes: Genes, Pathways, Biosynthetic Pathways l Instances (objects): trp. A, TCA cycle l Classes: l Superclass(es) l Subclass(es) l Instance(s) l A symbolic frame name (id, key) uniquely identifies each frame

Slots SRI International Bioinformatics l Encode attributes/properties of a frame l Integer, real number, string l Represent relationships between frames l The value of a slot is the identifier of another frame l Every slot is described by a “slot frame” in a KB that defines meta information about that slot
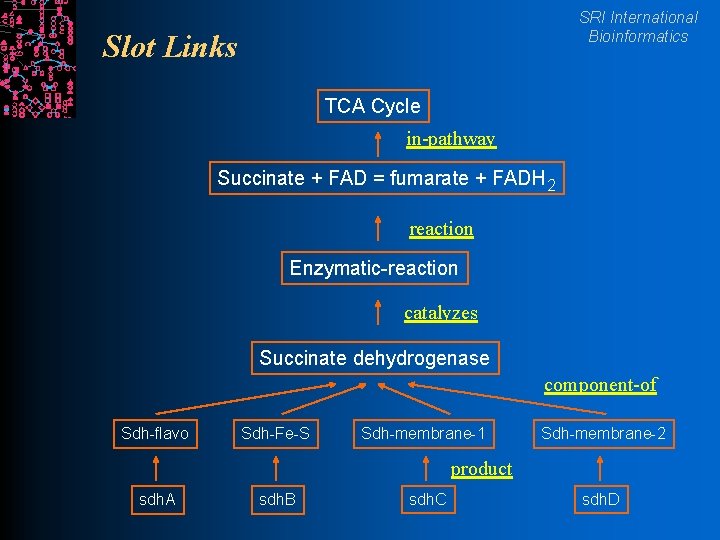
SRI International Bioinformatics Slot Links TCA Cycle in-pathway Succinate + FAD = fumarate + FADH 2 reaction Enzymatic-reaction catalyzes Succinate dehydrogenase component-of Sdh-flavo Sdh-Fe-S Sdh-membrane-1 Sdh-membrane-2 product sdh. A sdh. B sdh. C sdh. D

Slots SRI International Bioinformatics l Number of values l Single valued l Multivalued: sets, bags l Slot values l Any LISP object: Integer, real, string, symbol (frame name) l Slotunits define properties of slots: datatypes, classes, constraints l Two slots are inverses if they encode opposite relationships l Slot Product in class Genes
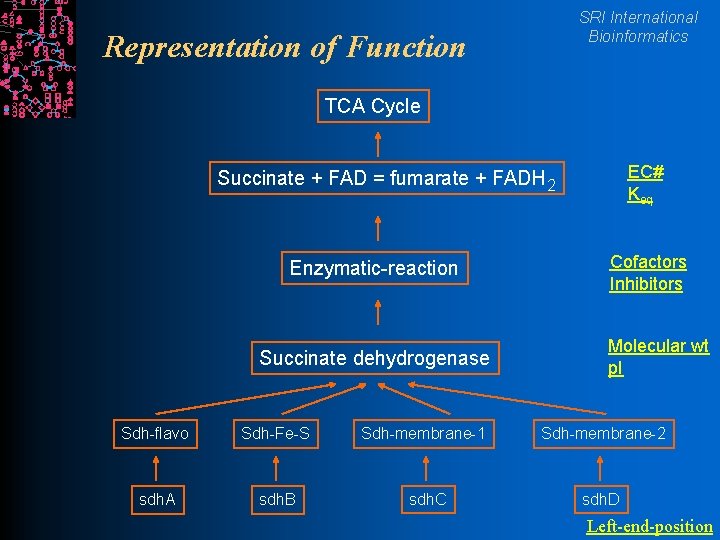
SRI International Bioinformatics Representation of Function TCA Cycle EC# Keq Succinate + FAD = fumarate + FADH 2 Enzymatic-reaction Succinate dehydrogenase Cofactors Inhibitors Molecular wt p. I Sdh-flavo Sdh-Fe-S Sdh-membrane-1 Sdh-membrane-2 sdh. A sdh. B sdh. C sdh. D Left-end-position
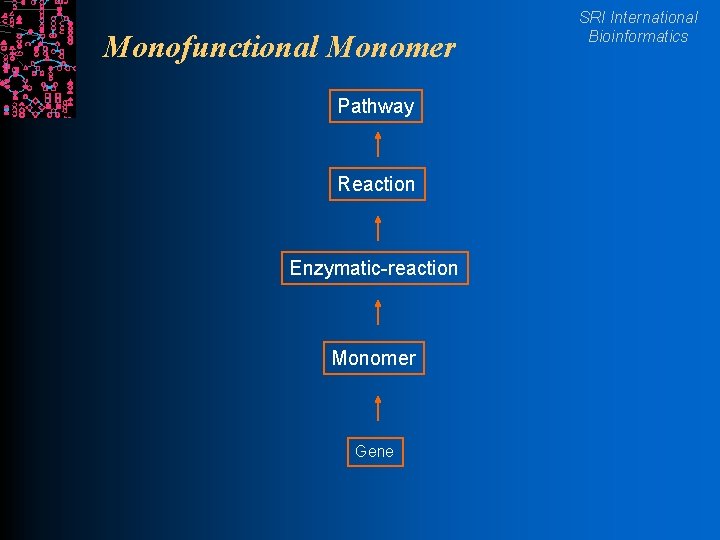
Monofunctional Monomer Pathway Reaction Enzymatic-reaction Monomer Gene SRI International Bioinformatics
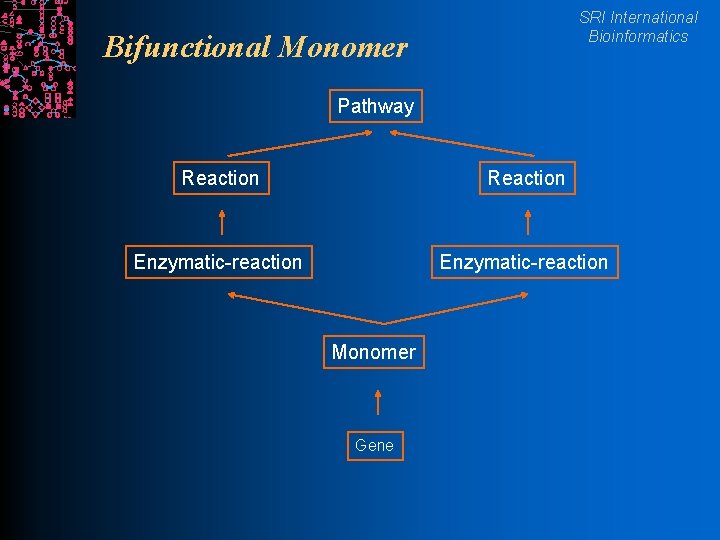
SRI International Bioinformatics Bifunctional Monomer Pathway Reaction Enzymatic-reaction Monomer Gene
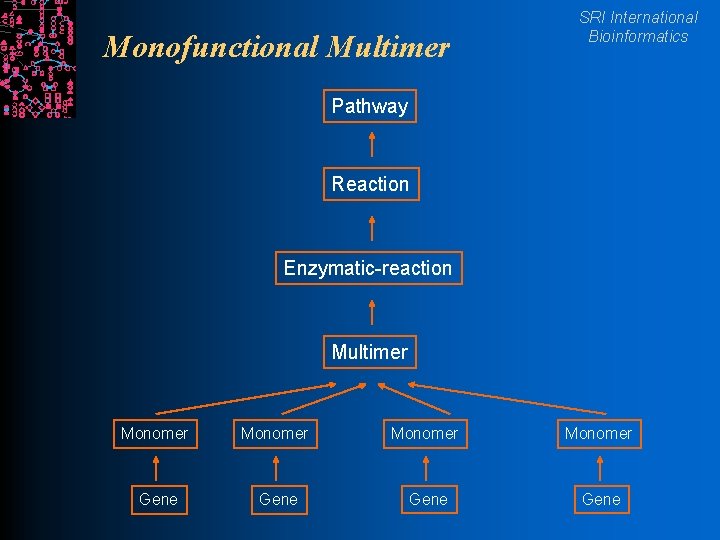
Monofunctional Multimer SRI International Bioinformatics Pathway Reaction Enzymatic-reaction Multimer Monomer Gene
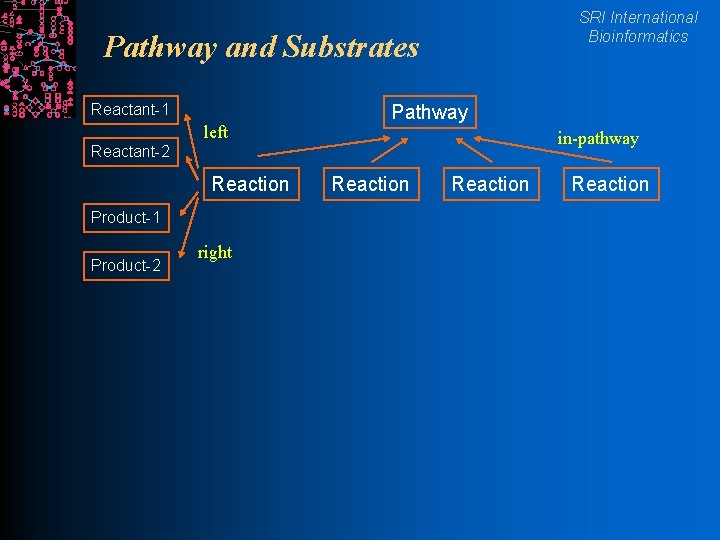
SRI International Bioinformatics Pathway and Substrates Reactant-1 left Pathway in-pathway Reactant-2 Reaction Product-1 Product-2 right Reaction
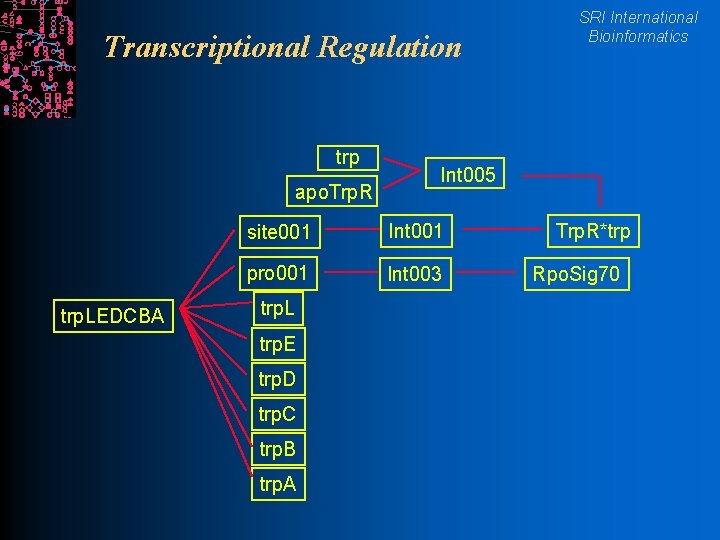
Transcriptional Regulation trp apo. Trp. R trp. LEDCBA Int 005 site 001 Int 001 pro 001 Int 003 trp. L trp. E trp. D trp. C trp. B trp. A SRI International Bioinformatics Trp. R*trp Rpo. Sig 70

Principle Classes SRI International Bioinformatics l Class names are capitalized, plural, separated by dashes l Genetic-Elements, with subclasses: l Chromosomes l Plasmids Genes Transcription-Units RNAs l r. RNAs, sn. RNAs, t. RNAs, Charged-t. RNAs Proteins, with subclasses: l Polypeptides l Protein-Complexes l l

Principle Classes l Reactions, with subclasses: l Transport-Reactions l Enzymatic-Reactions l Pathways l Compounds-And-Elements SRI International Bioinformatics

Frame IDs of Instances l Instance frame ID conventions have evolved over time l Examples: l Pathways u l l TRPSYN-PWY, P 23 -PWY Genes u AG 10045 Monomers u SRI International Bioinformatics TRPA-MONOMER, AG 10045 -MONOMER

Slots in Multiple Classes SRI International Bioinformatics l Common-Name l Synonyms l Names (computed as union of Common-Name, Synonyms) l Comment l Citations l DB-Links

Genes Slots l Component-Of SRI International Bioinformatics (links to replicon, transcription unit) l Left-End-Position l Right-End-Position l Centisome-Position l Transcription-Direction l Product

Proteins Slots l Molecular-Weight-Seq l Molecular-Weight-Exp l p. I l Locations l Modified-Form l Unmodified-Form l Component-Of SRI International Bioinformatics

Polypeptides Slots l Gene SRI International Bioinformatics

Protein-Complexes Slots l Components SRI International Bioinformatics

Reactions Slots SRI International Bioinformatics l EC-Number l Left, Right l Substrates (computed as union of Left, Right) l Delta. G 0 l Keq l Spontaneous?

Enzymatic-Reactions Slots l Enzyme l Reaction l Activators l Inhibitors l Physiologically-Relevant l Cofactors l Prosthetic-Groups l Alternative-Substrates l Alternative-Cofactors SRI International Bioinformatics

Pathways Slots l Reaction-List l Predecessors l Primaries SRI International Bioinformatics
- Slides: 27