SLRITools Project Providing a Platform for Bioinformatics Research

SLRITools Project: Providing a Platform for Bioinformatics Research Michel Dumontier Bioinformatics Technology Conference February 3 - 6, 2003

SLRITools Outline • • • Introduction Open-source Toolkit Foundation Toolkit Projects Future Prospects Michel Dumontier – SLRITools Project
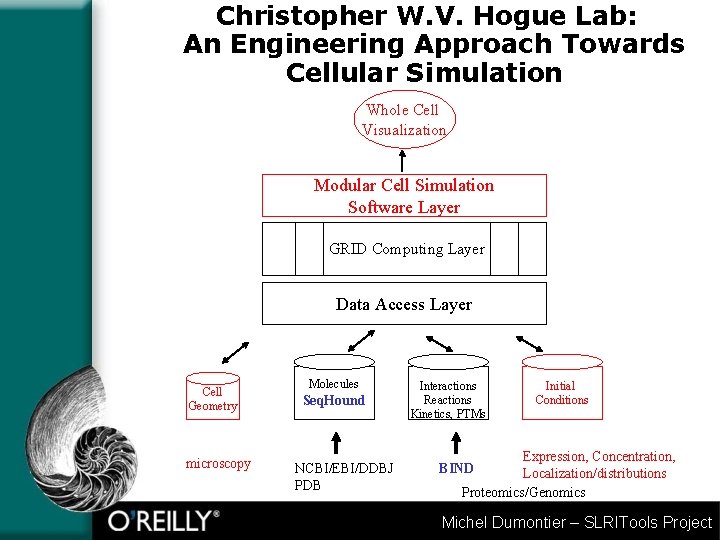
Christopher W. V. Hogue Lab: An Engineering Approach Towards Cellular Simulation Whole Cell Visualization Modular Cell Simulation Software Layer GRID Computing Layer Data Access Layer Cell Geometry microscopy Molecules Seq. Hound NCBI/EBI/DDBJ PDB Interactions Reactions Kinetics, PTMs Initial Conditions Expression, Concentration, Localization/distributions Proteomics/Genomics BIND Michel Dumontier – SLRITools Project

SLRITools Purpose • Make freely available our sequence and structure manipulation and analysis infrastructure and tool software to the greater benefit of the Bioinformatics community Michel Dumontier – SLRITools Project

SLRITools Description • Mainly C-based cross-platform toolkit for dealing with biological information, especially protein structure/function. • Extends the freely available NCBI C/C++ Toolkits and forms the basis for a number of powerful applications • GPL/LGPL/PAL licenses • Currently hosted at http: //sourceforge. net/projects/slritools • Training tutorials http: //bioinfo. mshri. on. ca/tkcourse/ • Canadian Bioinformatics Workshops http: //bioinformatics. ca Michel Dumontier – SLRITools Project

SLRITools Projects • SLRI lib - common library that extends NCBI Toolkit • Seq. Hound - Sequence and Structure Database Management System • • BIND - Biomolecular Interaction Network Database Text Indexer - ASN. 1 indexer NBLAST – Cluster variant of BLAST for Nx. N comparisons Kangaroo – Regular expression search of DNA/protein/CDR Michel Dumontier – SLRITools Project
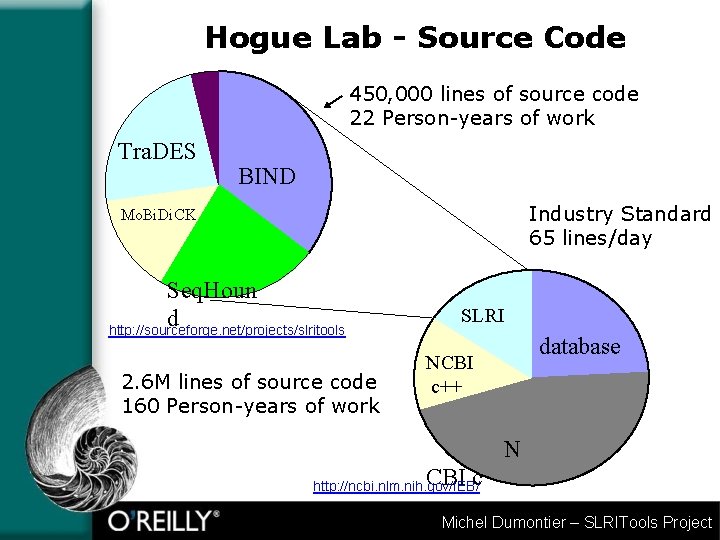
Hogue Lab - Source Code 450, 000 lines of source code 22 Person-years of work Tra. DES BIND Industry Standard 65 lines/day Mo. Bi. Di. CK Seq. Houn d http: //sourceforge. net/projects/slritools 2. 6 M lines of source code 160 Person-years of work SLRI database NCBI c++ N CBI c http: //ncbi. nlm. nih. gov/IEB/ Michel Dumontier – SLRITools Project

SLRITools Outline • • • Introduction Open-Source Toolkit Foundation Toolkit Projects Future Prospects Michel Dumontier – SLRITools Project

Going Open Source • Subject to the Intellectual Property Policy of Mt. Sinai Hospital • Does the software have the potential to improve patient care ? • Does the software have economic benefits that will fund new research and development? • Patents, Licenses & Publications Michel Dumontier – SLRITools Project

Software Licenses Stage 1) “Not Released” – “No license”– internal use only – Protects commercial interest of MSH SLRI Industrial Liasion Tech Transfer Office Patent IP • distributedfolding Stage 2) “Free to Academics” – Executables provided free, source upon request – Publication – Companies must license from MSH Board • MCODE, TRADES, SSSF Stage 3) “Public Use License” subcommittee on commercialization – GNU Public License • Seq. Hound, BIND Data Manager, BIND specification – Perl Artistic License/Lesser GNU Public License • Seq. Hound Remote Interfaces for Bio. PERL/ C, C++ API Michel Dumontier – SLRITools Project

Open Source Issues • Software Releases • Support Michel Dumontier – SLRITools Project

SLRITools Outline • • • Introduction Open-Source Toolkit Foundation Toolkit Projects Future Prospects Michel Dumontier – SLRITools Project

SLRITools Foundation • National Center for Biotechnology Information (NCBI) • NCBI Toolbox - Information Engineering Branch – http: //www. ncbi. nlm. nih. gov/IEB/ – Gen. Bank, Entrez, BLAST, Sequin, OMIM, Ref. Seq 1. Data Model – An explicit, complete data model of biological sequences, structures, bibliographic data, and associated annotations 2. Data Encoding - A formal specification and encoding rules. The telecommunications standard, ASN. 1, has been used for this. Recently it has been mapped to a similar language, XML. Provides automatic code generators. Michel Dumontier – SLRITools Project

SLRITools Foundation II 3. Programming Libraries – Originally written in a portable dialect of C. Recently a new generation is being written in C++. – Compiled and occasionally tested over 14 OS • • Linux, HPUX, Mac. OS 9/X, Irix, Solaris, Windows 3. 1/95/NT/2000/XP, Be. OS, QNX, alpha, BSD, AIX, parisc-Linux, Sony Play. Station 2 Linux 16/32/64 bit hardware – Open Source – Free License – ftp: //ftp. ncbi. nih. gov/toolbox/ Michel Dumontier – SLRITools Project

SLRITools Outline • • • Introduction Open-Source Toolkit Foundation Toolkit Projects Future Prospects Michel Dumontier – SLRITools Project

Seq. Hound • Seq. Hound is a sequence and structure database management system that inherits the NCBI data model and mirrors the NCBI core biological sequence and structure information • Why did we develop Seq. Hound? – Too many hits to NCBI server -> banned IP! – Data transmission & network connection issues – Generate more sophisticated API to access data currently only available within the NCBI – Faster, local or remote access with a variety of programming languages – Provide functionality necessary to retrieve specialized subsets of sequences, structures and structural domains. Michel Dumontier – SLRITools Project
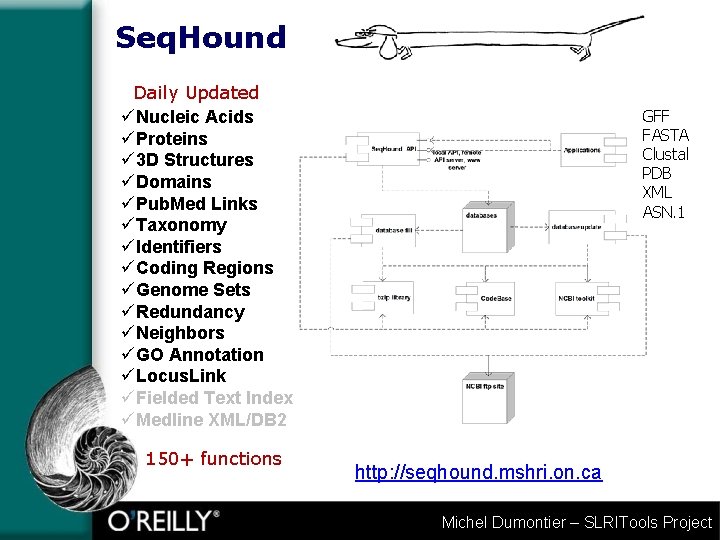
Seq. Hound Daily Updated üNucleic Acids üProteins ü 3 D Structures üDomains üPub. Med Links üTaxonomy üIdentifiers üCoding Regions üGenome Sets üRedundancy üNeighbors üGO Annotation üLocus. Link üFielded Text Index üMedline XML/DB 2 150+ functions GFF FASTA Clustal PDB XML ASN. 1 http: //seqhound. mshri. on. ca Michel Dumontier – SLRITools Project

Seq. Hound Resources • Seq. Hound is accessible via – http: //seqhound. mshri. on. ca – Simple web interface (under development) – C, C++, Java (new!), Perl remote API or an optimized local API. (->SOAP? ) • Timeline – Redundant fail-over server mid-summer – Concurrent with Bioperl release • Freely available article published in BMC Bioinformatics 2002, 3: 32 • http: //www. biomedcentral. com/14712105/3/32/ Michel Dumontier – SLRITools Project

BIND Biomolecular Interaction Network Database Motivation: • Massive influx of biomolecular interaction data requires repository, standards and access Goals: • Provide a standard, comprehensive and integrated interaction resource to the scientific community • Define protein function and mechanisms • Recover and integrate biomolecular interaction knowledge (backfilling) • Discover new knowledge through data mining Michel Dumontier – SLRITools Project

http: //bind. ca Result: • Database to archive and exchange molecular assembly information • Describes – Interactions – Complexes – Pathways • BIND has an extensive data model, GNU software tools and is based on the NCBI toolkit. • Recently funded for a 3 year effort at 25 M CDN – CIHR (1 M) OGI/Genome Canada (12. 5 M) Ontario R&D Challenge fund (5. 2 M) – IBM, MDS Proteomics and Foundry Networks – Sun Michel Dumontier – SLRITools Project

BIND Data Policies Gen. Bank Policy – BIND data is freely available for any purpose Direct Submission – Submitters cannot limit the intended use of submitted BIND data – Submitters have the right to edit/alter their records over time – Suggestions made by a third party will be forwarded by us to the submitters to seek approval for any changes or corrections Availability – ftp: //ftp. bind. ca – ASN. 1/XML data+specification Michel Dumontier – SLRITools Project
Browsing BINDhttp: //bind. ca Michel Dumontier – SLRITools Project
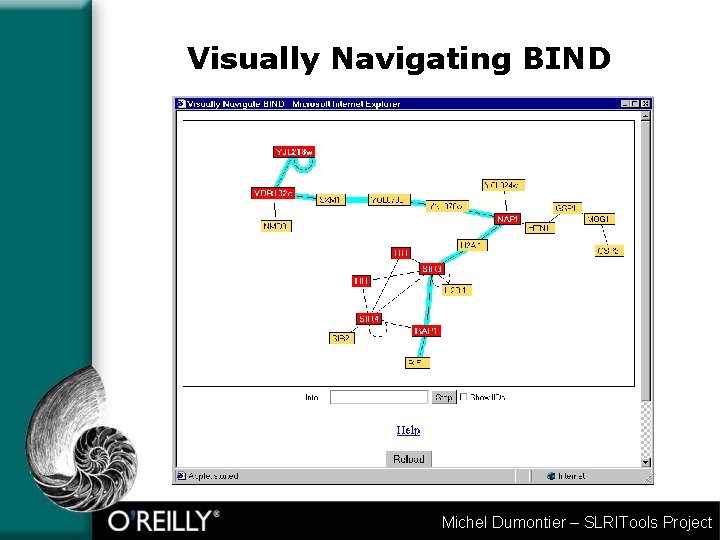
Visually Navigating BIND Michel Dumontier – SLRITools Project
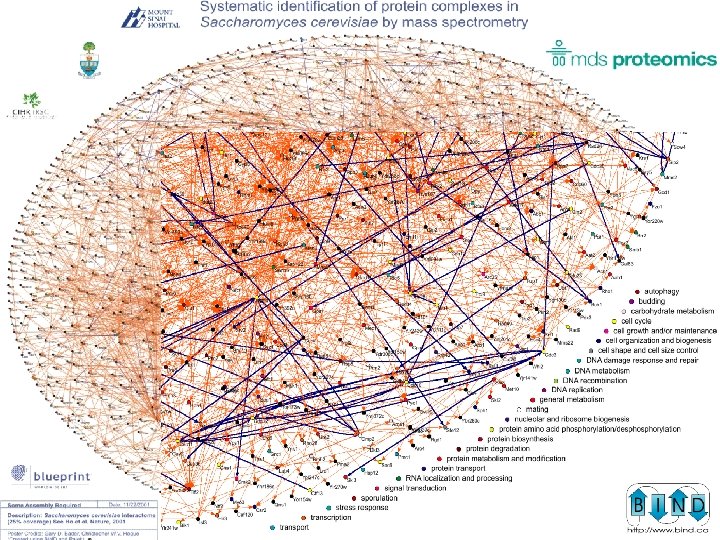
Michel Dumontier – SLRITools Project

Molecular Complex Detection (MCODE) • Assume densely connected regions of a heterogeneous interaction network represent molecular complexes • MCODE finds densely connected regions of a graph • Weight nodes by local density (scoring function) • From highest weighted node, recursively add neighbours above threshold score to complex • Evaluation (Yeast): • 88/221 Cell. Zome hand annotated complexes • 64/208 MIPS complexes (166 predicted) • 200 complexes predicted in 15, 143 protein interactions from yeast Published: BMC Bioinformatics 2003. 4: 2. • http: //www. biomedcentral. com/1471 -2105/4/2 Michel Dumontier – SLRITools Project
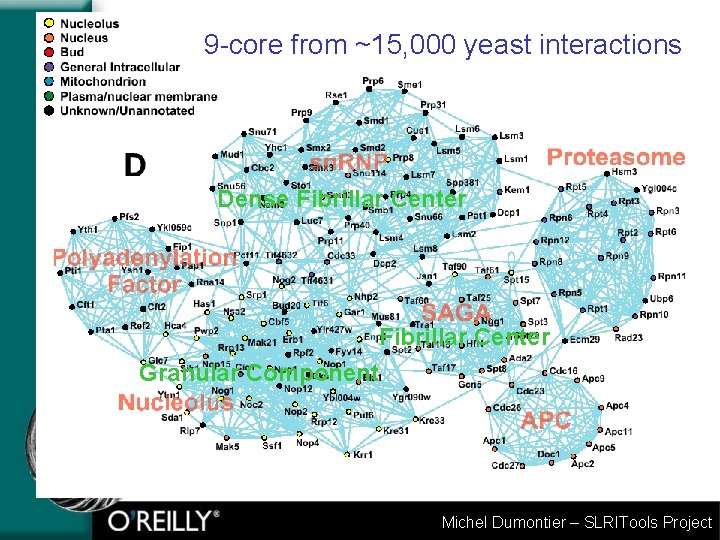
9 -core from ~15, 000 yeast interactions Dense Fibrillar Center Granular Component Michel Dumontier – SLRITools Project

FAST = “parallel” RPS BLAST Used to spot domain similarities in a protein interaction cluster Server-generated scalable FLASH graphics – zoomable, printable. Followed-up by zoom in on FASTA formatted sequences to see domain superposition and links to SMART/PFAM Michel Dumontier – SLRITools Project
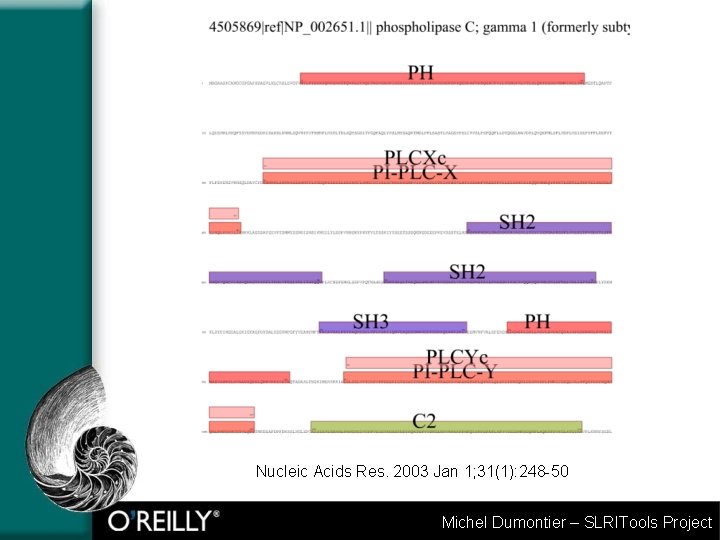
Nucleic Acids Res. 2003 Jan 1; 31(1): 248 -50 Michel Dumontier – SLRITools Project

NBLAST Description: • NBLAST is a cluster computer variant of BLAST • It performs the minimum number of sequence comparisons and stores sequence alignments and the list of similar sequences (neighbours) as binary ASN. 1 (XML) • NBLAST is written in C using the NCBI C Toolkit. • Separate function and database layers Accessibility: via Seq. Hound • http: //seqhound. mshri. on. ca Neighbours DB (codebase) • ftp: //ftp. mshri. on. ca/pub/nblast Published: BMC Bioinformatics 2002, 3: 13 • http: //www. biomedcentral. com/14712105/3/13/ Michel Dumontier – SLRITools Project

Ookpik CFI/ORDCF Funded. 216 P-III 450 64 GB 1. 2 TB disk NBLAST RPS-BLAST TRADES Mo. Bi. Di. CK http: //bioinfo. mshri. on. ca/yac/ http: //sourceforge. net/projects/slritools/

Kangaroo Description: • Kangaroo is implemented to facilitate a wide range of queries with no restriction on the length or complexity of the query expression • Uses regular expression • Search DNA, protein, or coding region • Web-based form and results • Links to Seq. Hound Accessibility: • http: //bioinfo. mshri. on. ca/kangaroo currently supports searches on 10 organisms (including human, mouse) Published: BMC Bioinformatics 2002, 3: 20 http: //www. biomedcentral. com/14712105/3/20 Michel Dumontier – SLRITools Project

Summary • Robust tools and services based on the NCBI data model • Flexible licensing Future Prospects • BIND/Seq. Hound Web Services (SOAP) • Seq. Hound – Web Interface – Inter. Pro|COG • Larger & more sophisticated BIND (JAVA) • Grid Engine & Cell Simulation Michel Dumontier – SLRITools Project

Christopher W. V. Hogue Lab Projects/Graduate Students • BIND – Gary Bader – Doron Betel • Seq. Hound – Katerina Michalickova • Protein Folding/CASP Predictions – Howard Feldman • Species Specific Protein Scoring Functions – Michel Dumontier • Cell Simulation/Systems Biology – Adrian Heilbut – Ken Lau • FPGA Hardware Database Search Engines – Ruth Isserlin Michel Dumontier – SLRITools Project

BIND = “Blueprint Initiative” • Database Curation – – Vicki Lay Susan Moore Brigitte Tuekam Cheryl Wolting • Software Engineering – – – – Neil Bahroos Ian Donaldson Marc Dumontier Vladimir Grytsan Hao Lieu Greg Pintile John Salama • Administration – – Eric Andrade Marianne Rukavina Sue Sroka Greg Van Volkenburg • IT – Greg Clark – Edward Lee Michel Dumontier – SLRITools Project

- Slides: 35