Canadian Bioinformatics Workshops www bioinformatics ca Module Title

Canadian Bioinformatics Workshops www. bioinformatics. ca
Module #: Title of Module 2

Module 6 Metabolomic Data Analysis Using Metabo. Analyst David Wishart Informatics and Statistics for Metabolomics May 3 -4, 2012
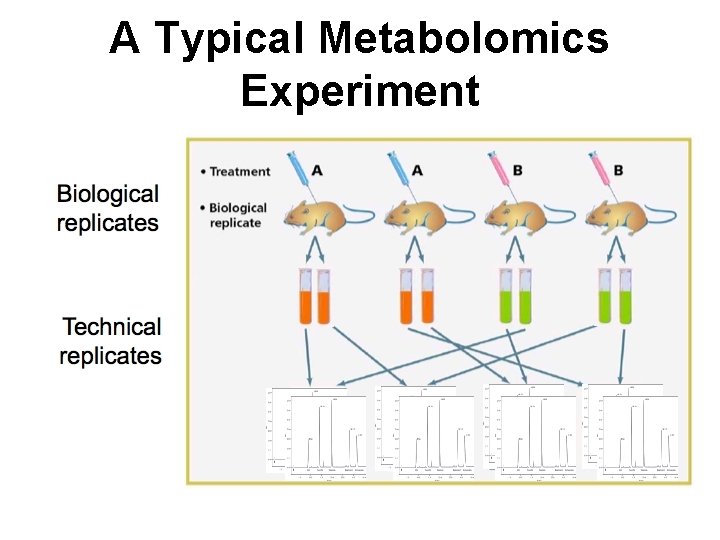
A Typical Metabolomics Experiment
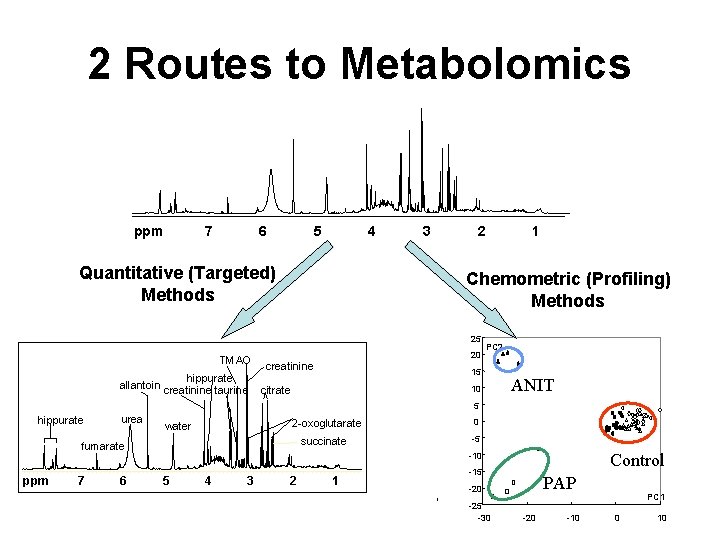
2 Routes to Metabolomics ppm 7 6 5 4 Quantitative (Targeted) Methods 3 2 Chemometric (Profiling) Methods 25 TMAO hippurate allantoin creatinine taurine 1 20 creatinine PC 2 15 10 citrate ANIT 5 hippurate urea 2 -oxoglutarate water succinate fumarate ppm 7 6 0 -5 Control -10 5 4 3 2 1 -15 PAP -20 -25 -30 -20 -10 PC 1 0 10

Metabolomics Data Workflow Chemometric Methods • Data Integrity Check • Spectral alignment or binning • Data normalization • Data QC/outlier removal • Data reduction & analysis • Compound ID Targeted Methods • Data Integrity Check • Compound ID and quantification • Data normalization • Data QC/outlier removal • Data reduction & analysis
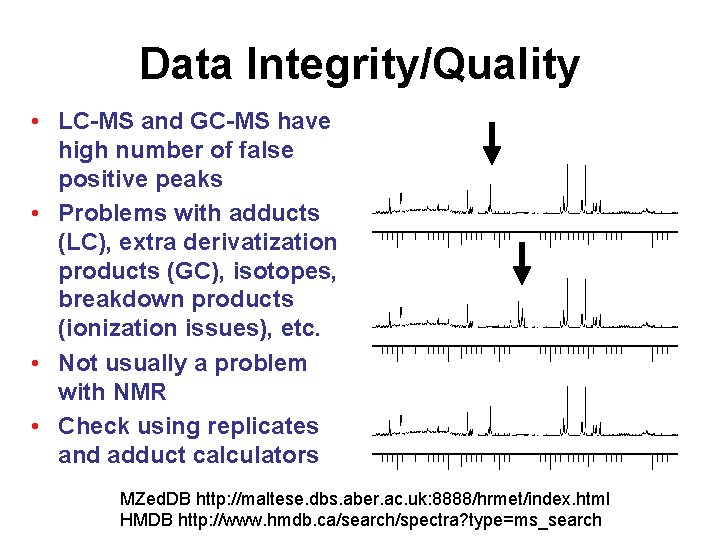
Data Integrity/Quality • LC-MS and GC-MS have high number of false positive peaks • Problems with adducts (LC), extra derivatization products (GC), isotopes, breakdown products (ionization issues), etc. • Not usually a problem with NMR • Check using replicates and adduct calculators MZed. DB http: //maltese. dbs. aber. ac. uk: 8888/hrmet/index. html HMDB http: //www. hmdb. ca/search/spectra? type=ms_search
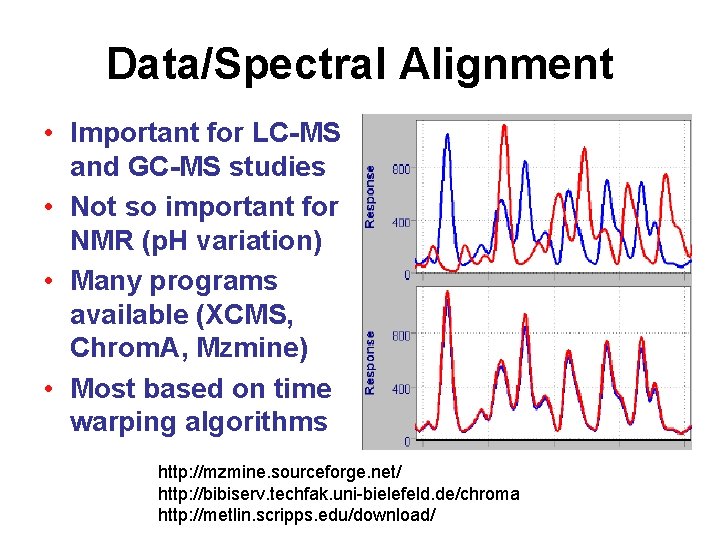
Data/Spectral Alignment • Important for LC-MS and GC-MS studies • Not so important for NMR (p. H variation) • Many programs available (XCMS, Chrom. A, Mzmine) • Most based on time warping algorithms http: //mzmine. sourceforge. net/ http: //bibiserv. techfak. uni-bielefeld. de/chroma http: //metlin. scripps. edu/download/
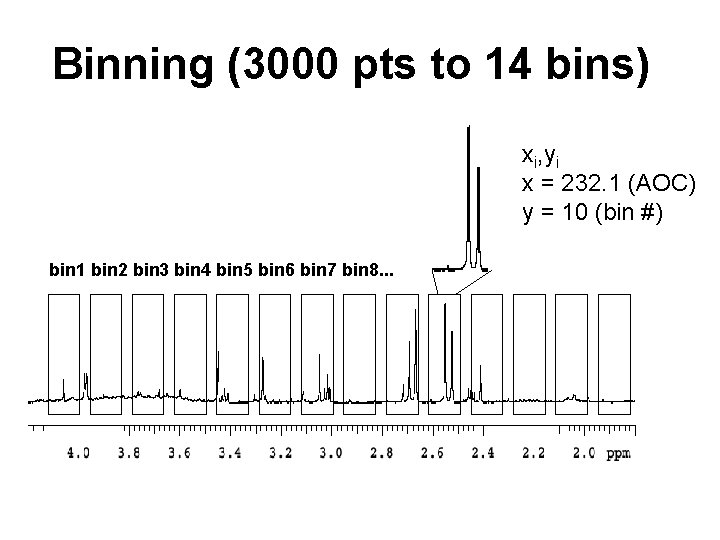
Binning (3000 pts to 14 bins) xi, yi x = 232. 1 (AOC) y = 10 (bin #) bin 1 bin 2 bin 3 bin 4 bin 5 bin 6 bin 7 bin 8. . .
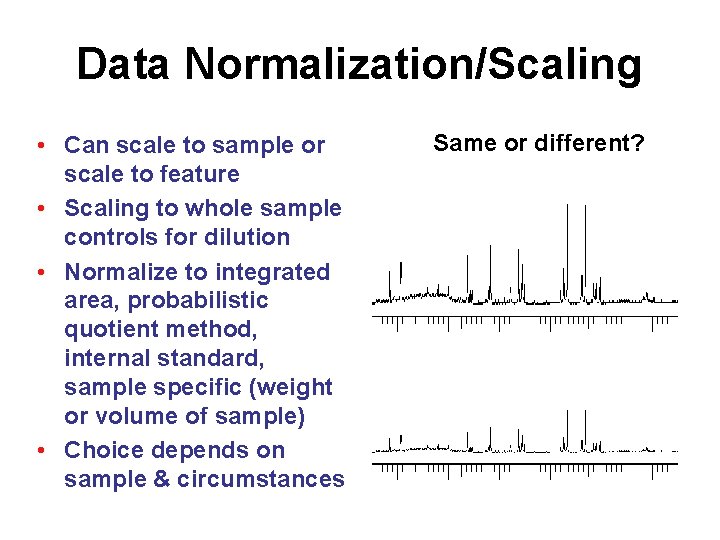
Data Normalization/Scaling • Can scale to sample or scale to feature • Scaling to whole sample controls for dilution • Normalize to integrated area, probabilistic quotient method, internal standard, sample specific (weight or volume of sample) • Choice depends on sample & circumstances Same or different?
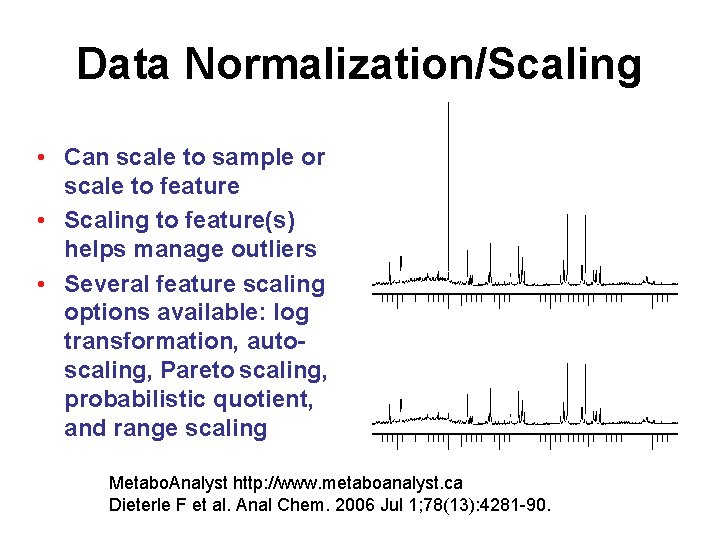
Data Normalization/Scaling • Can scale to sample or scale to feature • Scaling to feature(s) helps manage outliers • Several feature scaling options available: log transformation, autoscaling, Pareto scaling, probabilistic quotient, and range scaling Metabo. Analyst http: //www. metaboanalyst. ca Dieterle F et al. Anal Chem. 2006 Jul 1; 78(13): 4281 -90.

Data QC, Outlier Removal & Data Reduction • Data filtering (remove solvent peaks, noise filtering, false positives, outlier removal -- needs justification) • Dimensional reduction or feature selection to reduce number of features or factors to consider (PCA or PLS-DA) • Clustering to find similarity

Metabo. Analyst http: //www. metaboanalyst. ca • Web server designed to handle large sets of LC-MS, GC-MS or NMR-based metabolomic data • Supports both univariate and multivariate data processing, including t-tests, ANOVA, PCA, PLS-DA • Identifies significantly altered metabolites, produces colorful plots, provides detailed explanations & summaries • Links sig. metabolites to pathways via SMPDB

Metabo. Analyst Workflow Data preprocessin g Data normalizatio n Data analysis Data annotation 14
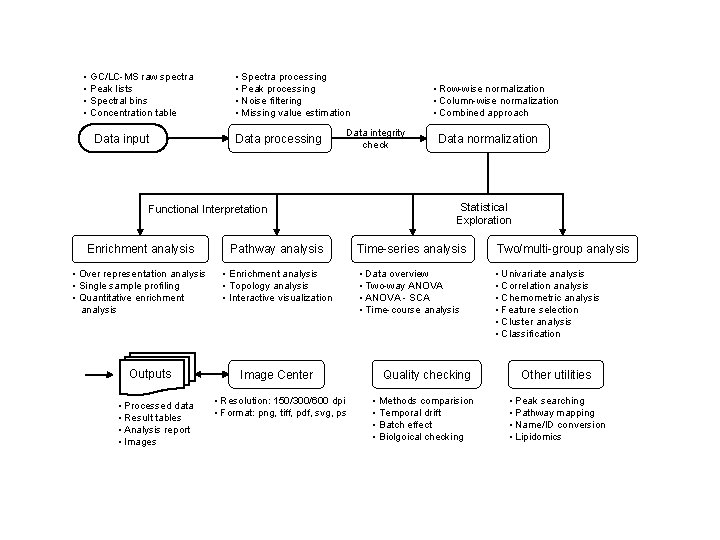
• GC/LC-MS raw spectra • Peak lists • Spectral bins • Concentration table • Spectra processing • Peak processing • Noise filtering • Missing value estimation Data processing Data input • Row-wise normalization • Column-wise normalization • Combined approach Data integrity check Functional Interpretation Data normalization Statistical Exploration Enrichment analysis Pathway analysis Time-series analysis Two/multi-group analysis • Over representation analysis • Single sample profiling • Quantitative enrichment analysis • Enrichment analysis • Topology analysis • Interactive visualization • Data overview • Two-way ANOVA • ANOVA - SCA • Time-course analysis • Univariate analysis • Correlation analysis • Chemometric analysis • Feature selection • Cluster analysis • Classification Outputs • Processed data • Result tables • Analysis report • Images Image Center • Resolution: 150/300/600 dpi • Format: png, tiff, pdf, svg, ps Quality checking • Methods comparision • Temporal drift • Batch effect • Biolgoical checking Other utilities • Peak searching • Pathway mapping • Name/ID conversion • Lipidomics

Metabo. Analyst Overview • Raw data processing – Using Metabo. Analyst • Data Reduction & Statistical analysis – Using Metabo. Analyst • Functional enrichment analysis – Using MSEA in Metabo. Analyst • Metabolic pathway analysis – Using Met. PA in Metabo. Analyst

Example Datasets

Example Datasets

Metabolomic Data Processing

Common Tasks • Purpose: to convert various raw data forms into data matrices suitable for statistical analysis • Supported data formats – Concentration tables (Targeted Analysis) – Peak lists (Untargeted) – Spectral bins (Untargeted) – Raw spectra (Untargeted)
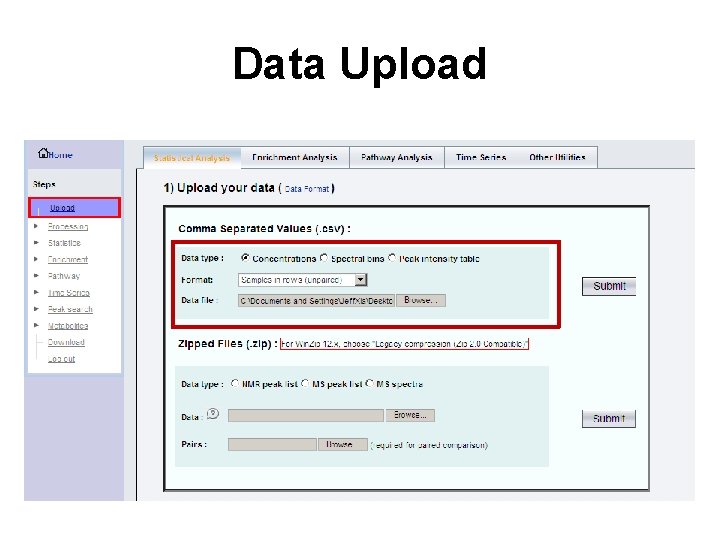
Data Upload
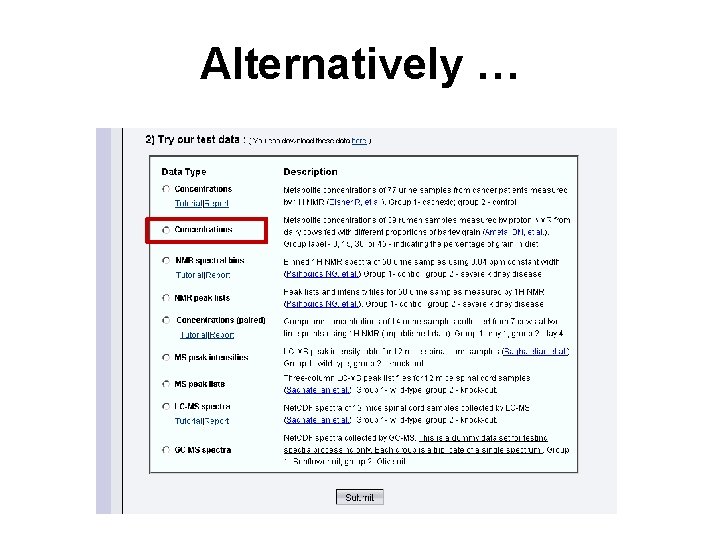
Alternatively …

Data Set Selected • Here we will be selecting a data set from dairy cattle fed different proportions of cereal grains (0%, 15%, 30%, 45%) • The rumen was analyzed using NMR spectroscopy using quantitative metabolomic techniques • High grain diets are thought to be stressful on cows
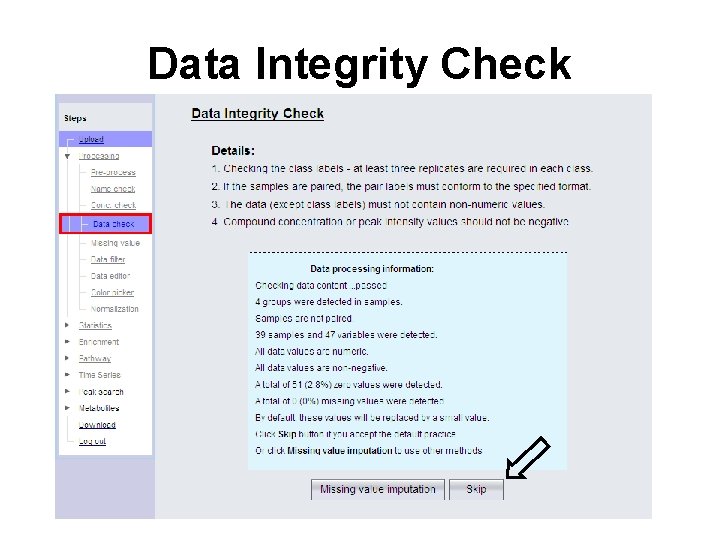
Data Integrity Check
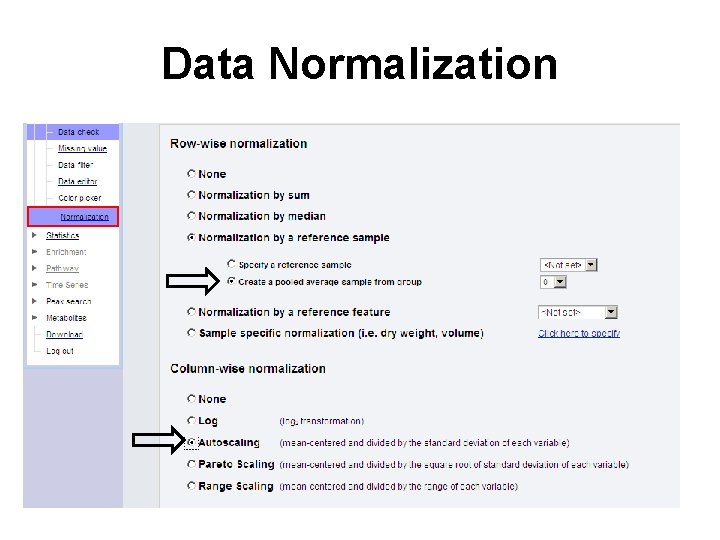
Data Normalization

Data Normalization • At this point, the data has been transformed to a matrix with the samples in rows and the variables (compounds/peaks/bins) in columns • Metabo. Analyst offers three types of normalization, row-wise normalization, column-wise normalization and combined normalization • Row-wise normalization aims to make each sample (row) comparable to each other (i. e. urine samples with different dilution effects)

Data Normalization • Column-wise normalization aims to make each variable (column) comparable to each other • This procedure is useful when variables are of very different orders of magnitude • Four methods have been implemented for this purpose – log transformation, autoscaling, Pareto scaling and range scaling
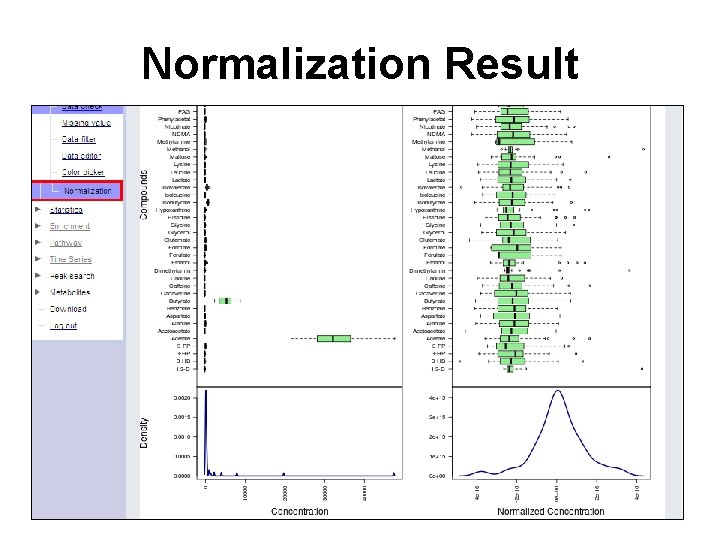
Normalization Result

Quality Control • Dealing with outliers – Detected mainly by visual inspection – May be corrected by normalization – May be excluded • Noise reduction – More of a concern for spectral bins/ peak lists – Usually improves downstream results
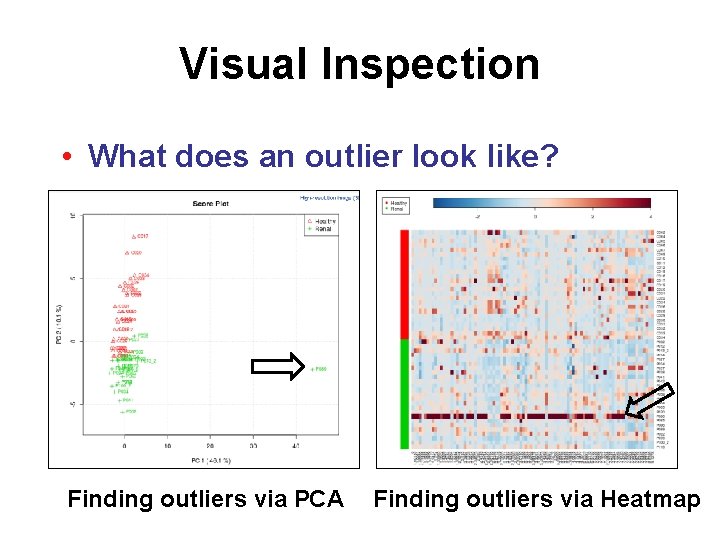
Visual Inspection • What does an outlier look like? Finding outliers via PCA Finding outliers via Heatmap
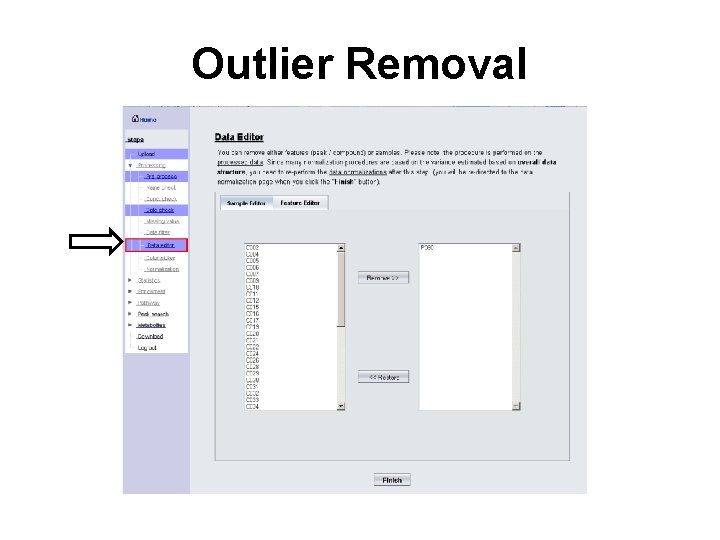
Outlier Removal

Noise Reduction

Noise Reduction (cont. ) • Characteristics of noise & uninformative features – Low intensities – Low variances (default)

Data Reduction and Statistical Analysis

Common tasks • To identify important features; • To detect interesting patterns; • To assess difference between the phenotypes • To facilitate classification / prediction
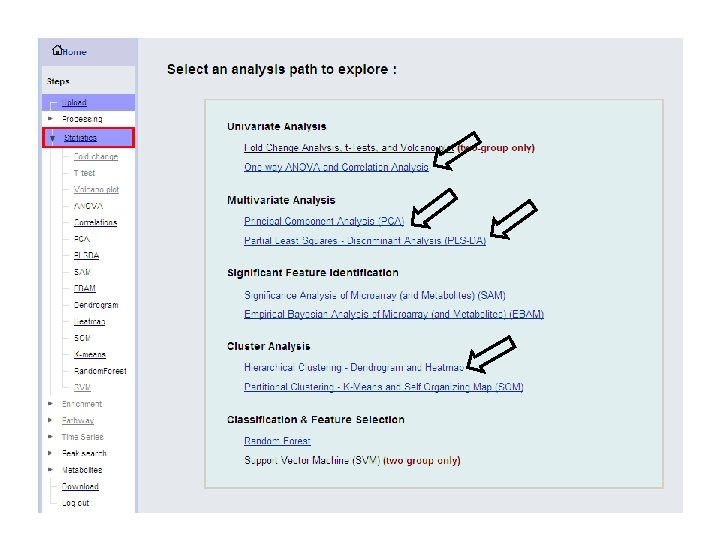
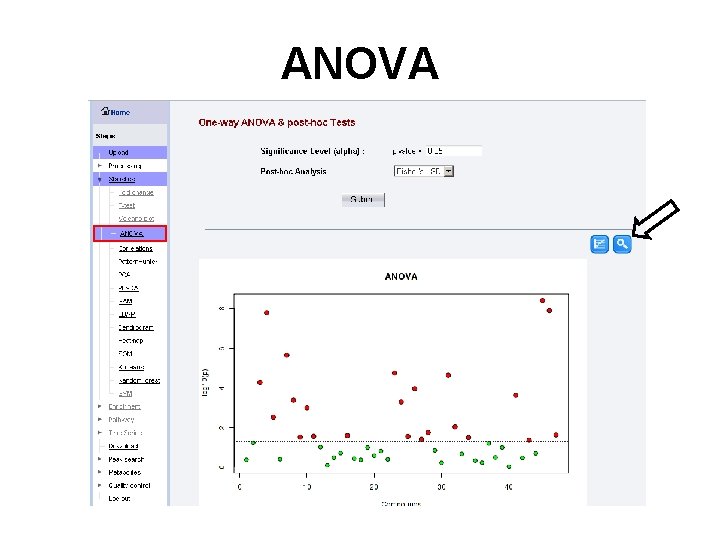
ANOVA
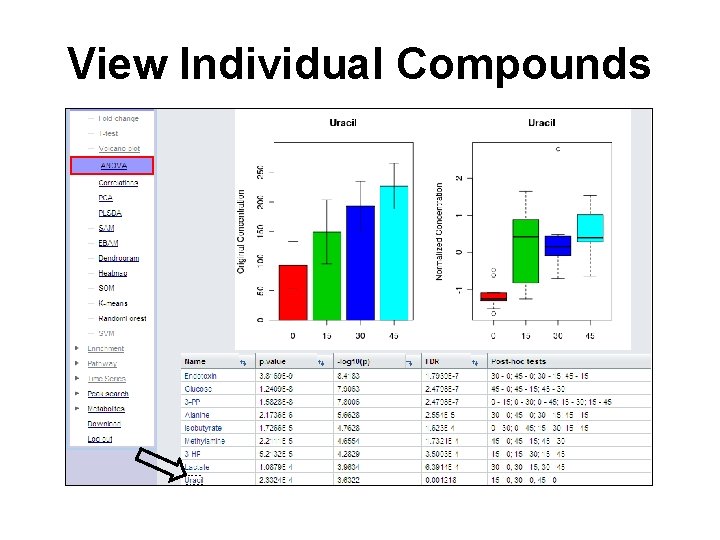
View Individual Compounds

Questions • Q: Which compounds show significant difference among all the neighboring groups (0 -15, 15 -30, and 30 -45)? • Q: For Uracil, are groups 15, 30, 45 significantly different from each other?
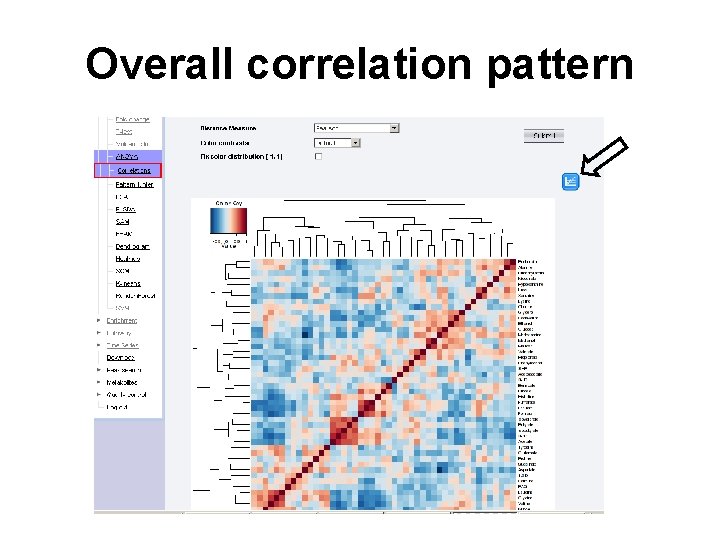
Overall correlation pattern

High resolution image Specify format Specify resolution Specify size

Question • Q: In untargeted metabolomics using NMR, researchers often look for region(s) on the spectra showing biggest change in their correlation patterns under different conditions. Can you do that in Metabo. Analyst? • Hint: check the available parameters of Correlation analysis

Template Matching • Looking for compounds showing interesting patterns of change • Essentially a method to look for linear trends or periodic trends in the data • Best for data that has 3 or more groups
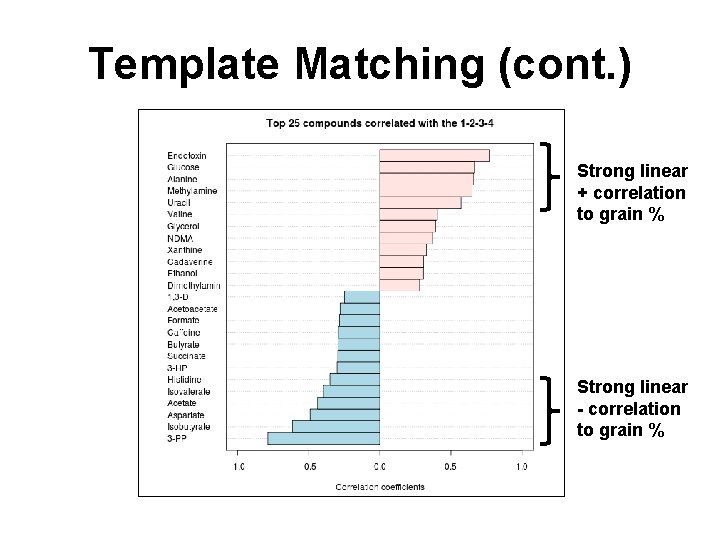
Template Matching (cont. ) Strong linear + correlation to grain % Strong linear - correlation to grain %

Question • Q: Identify compounds that decrease in the first three groups but increase in the last group?
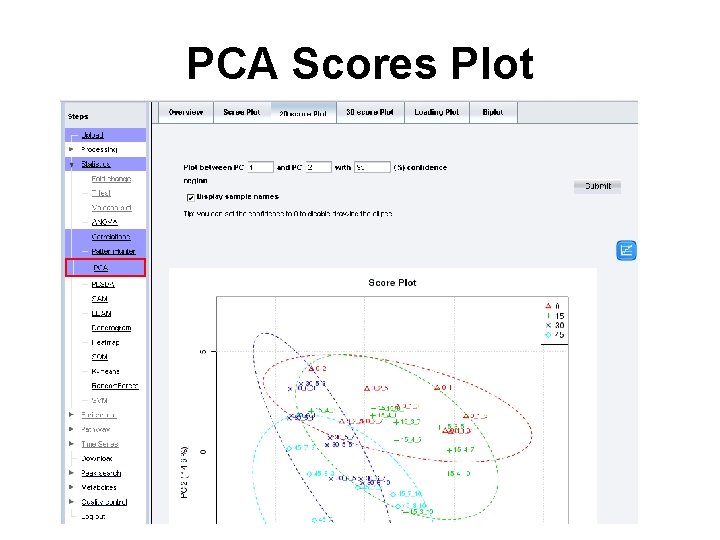
PCA Scores Plot
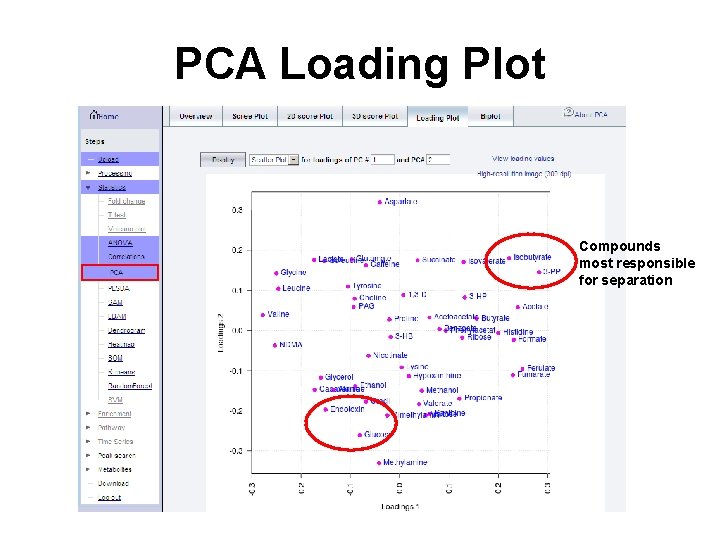
PCA Loading Plot Compounds most responsible for separation

3 D-PCA 48

Question Q: Identify compounds that contribute most to the separation between group 15 and 45
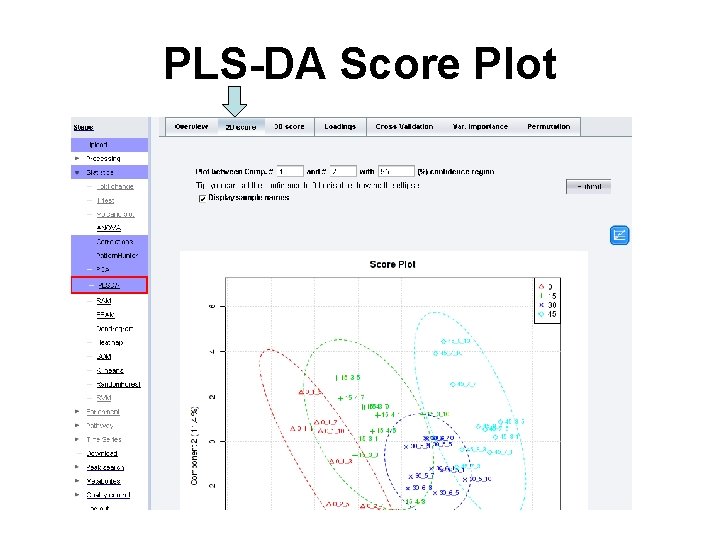
PLS-DA Score Plot
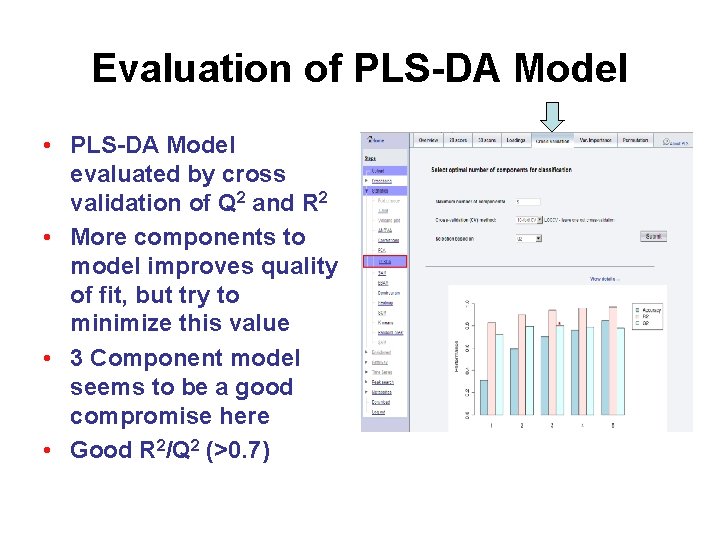
Evaluation of PLS-DA Model • PLS-DA Model evaluated by cross validation of Q 2 and R 2 • More components to model improves quality of fit, but try to minimize this value • 3 Component model seems to be a good compromise here • Good R 2/Q 2 (>0. 7)
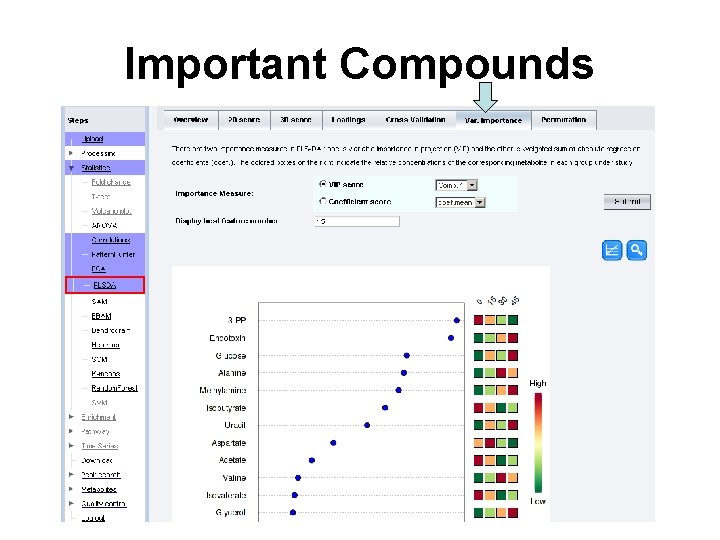
Important Compounds
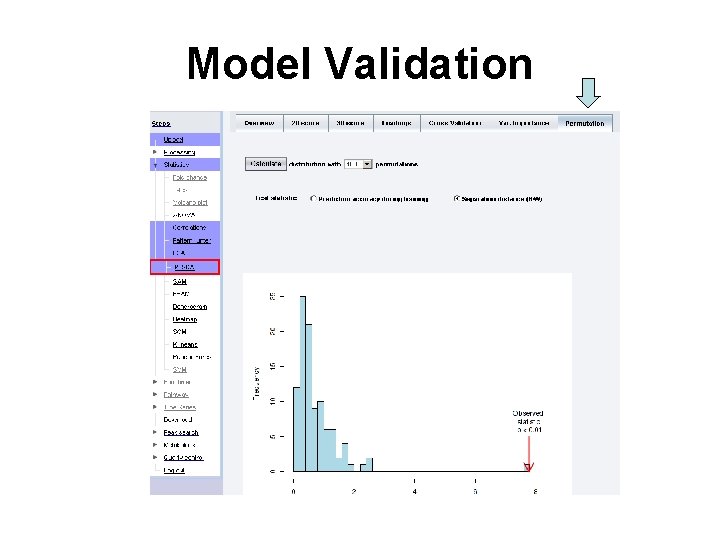
Model Validation

Questions • Q: What does p < 0. 01 mean? • Q: How many permutations need to be performed if you want to claim p value < 0. 0001?

Heatmap Visualization Note that the Heatmap is not being clustered on Rows (i. e. the % grain in diet)
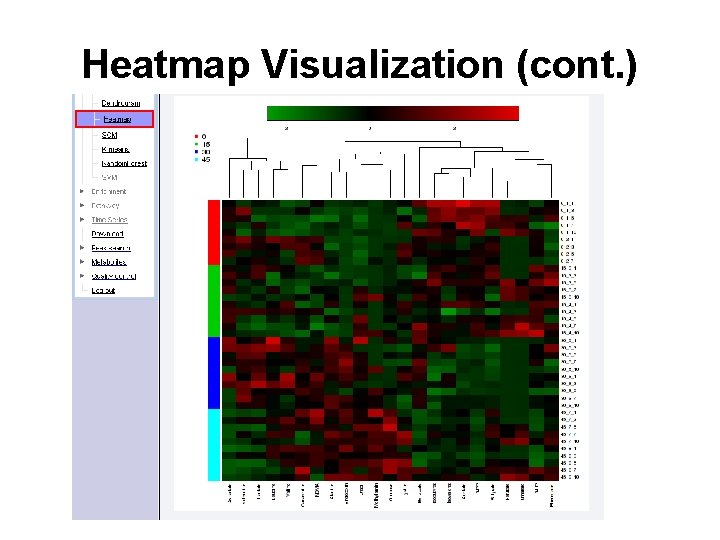
Heatmap Visualization (cont. )

Question Q: Identify compounds with a low concentration in group 0, 15 but increase in the group 35 and 45 Q: Which compound is the only one significantly increased in group 45?
Download Results

Analysis Report

Metabolite Set Enrichment Analysis

Metabolite Set Enrichment Analysis (MSEA) http: //www. msea. ca • Web tool designed to handle lists of metabolites (with or without concentration data) • Modeled after Gene Set Enrichment Analysis (GSEA) • Supports over representation analysis (ORA), single sample profiling (SSP) and quantitative enrichment analysis (QEA) • Contains a library of 6300 pre-defined metabolite sets including 85 pathway sets & 850 disease sets

Enrichment Analysis • Purpose: To test if there are some biologically meaningful groups of metabolites that are significantly enriched in your data • Biological meaningful – Pathways – Disease – Localization • Currently, only supports human metabolomic data

MSEA • Accepts 3 kinds of input files • 1) list of metabolite names only (ORA) • 2) list of metabolite names + concentration data from a single sample (SSP) • 3) a concentration table with a list of metabolite names + concentrations for multiple samples/patients (QEA)
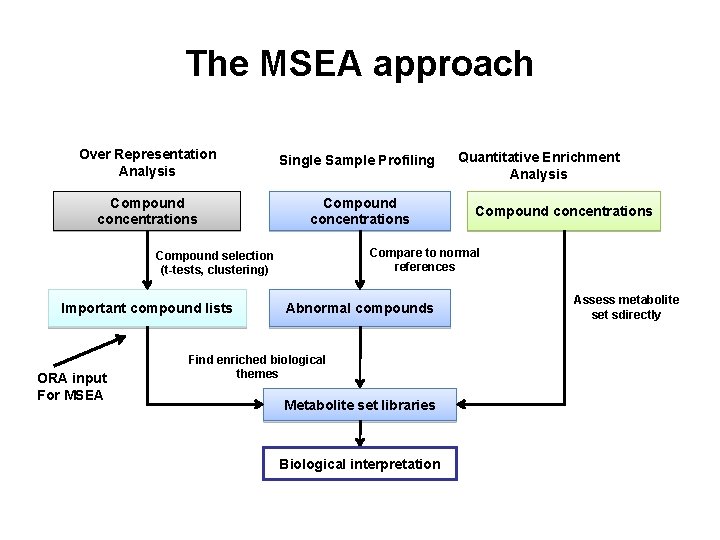
The MSEA approach Over Representation Analysis Single Sample Profiling Compound concentrations ORA input For MSEA Compound concentrations Compare to normal references Compound selection (t-tests, clustering) Important compound lists Quantitative Enrichment Analysis Abnormal compounds Assess metabolite set sdirectly Find enriched biological themes Metabolite set libraries Biological interpretation 64

Data Set Selected • Here we are using a collection of metabolites identified by NMR (compound list + concentrations) from the urine from 77 lung and colon cancer patients, some of whom were suffering from cachexia (muscle wasting)
Start with a Compound List
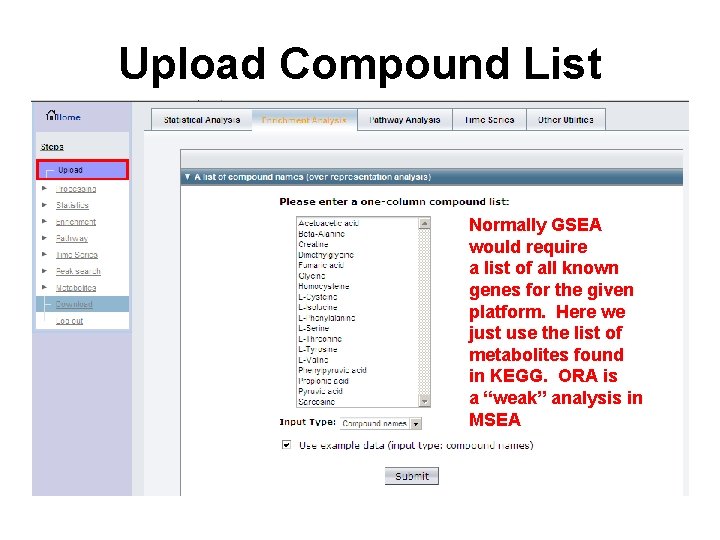
Upload Compound List Normally GSEA would require a list of all known genes for the given platform. Here we just use the list of metabolites found in KEGG. ORA is a “weak” analysis in MSEA
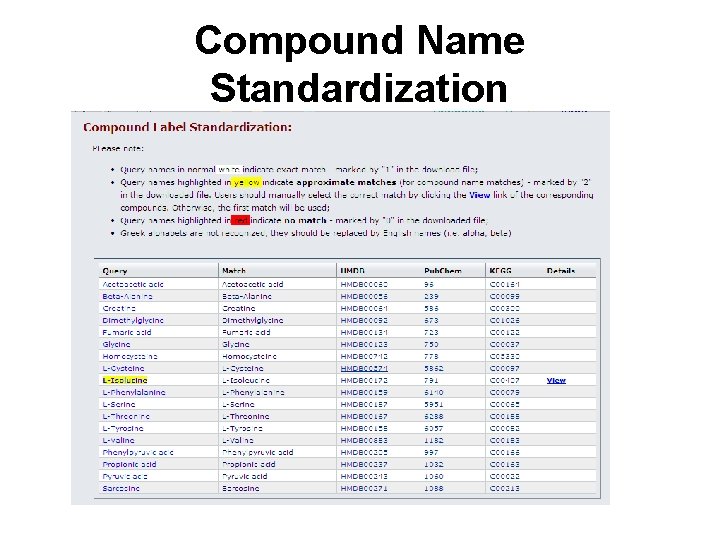
Compound Name Standardization
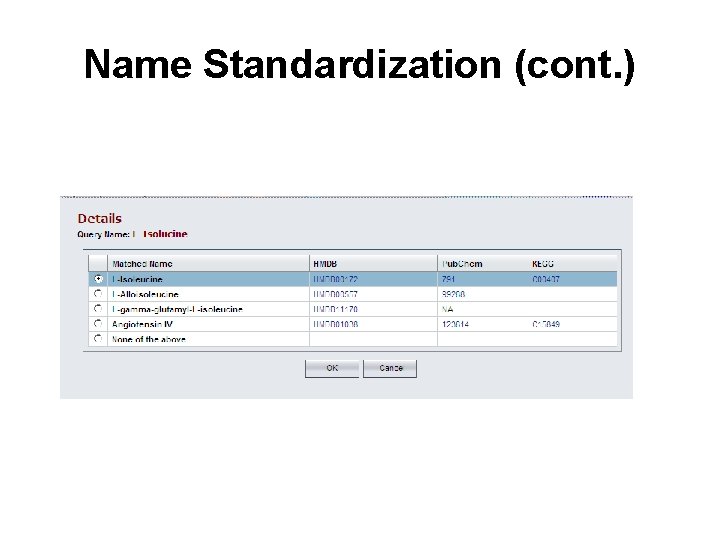
Name Standardization (cont. )
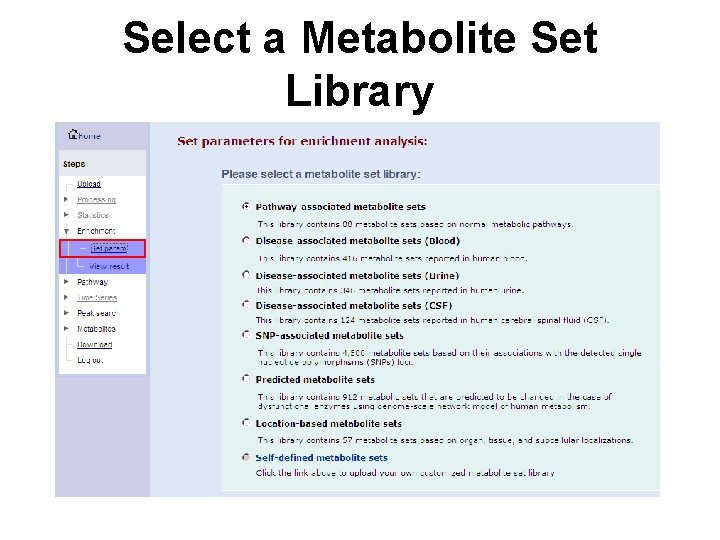
Select a Metabolite Set Library
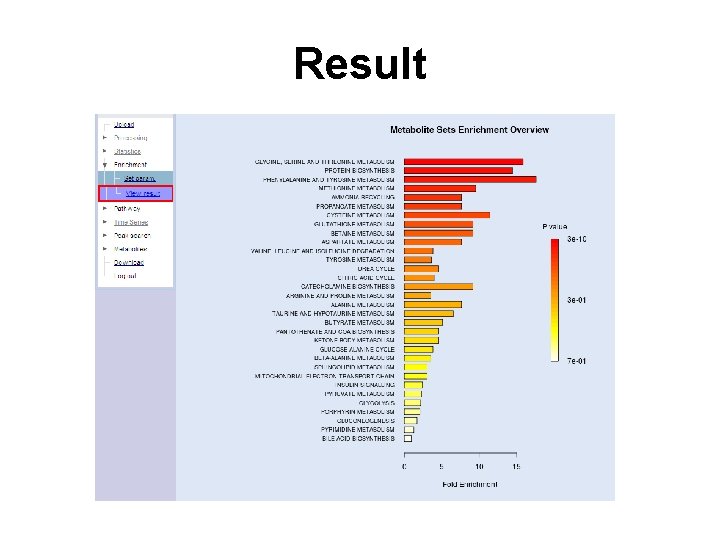
Result
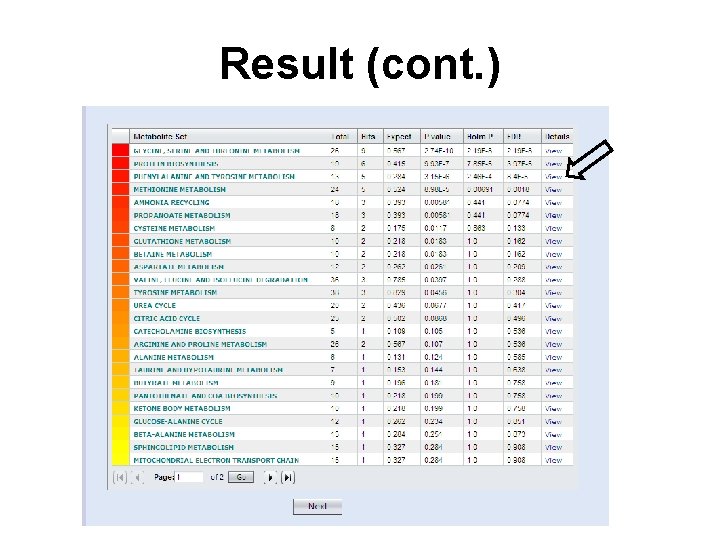
Result (cont. )

The Matched Metabolite Set
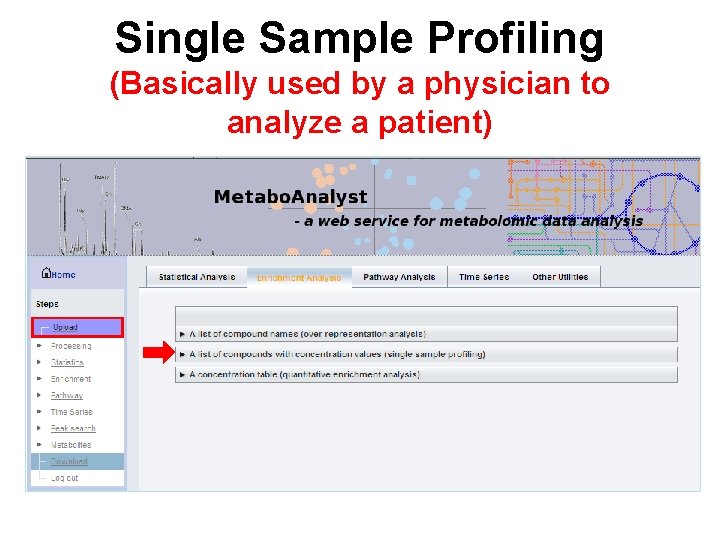
Single Sample Profiling (Basically used by a physician to analyze a patient)
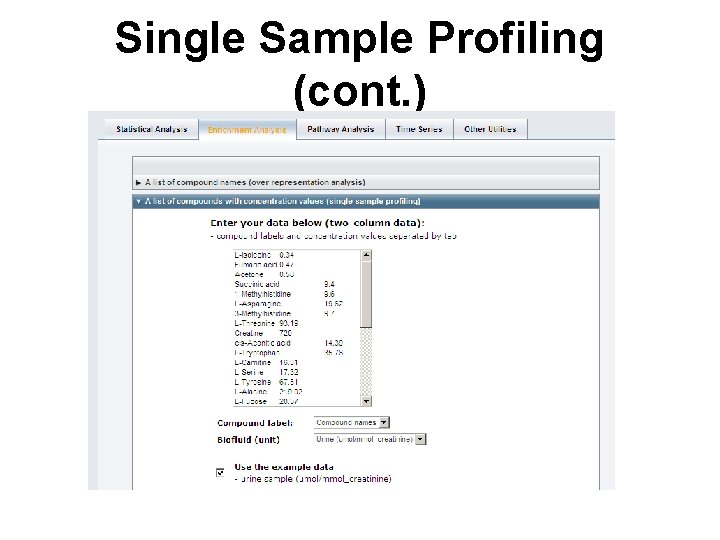
Single Sample Profiling (cont. )
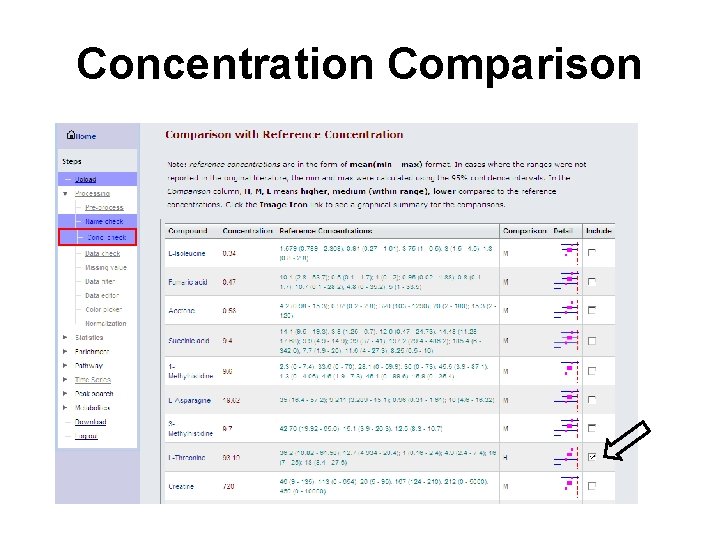
Concentration Comparison
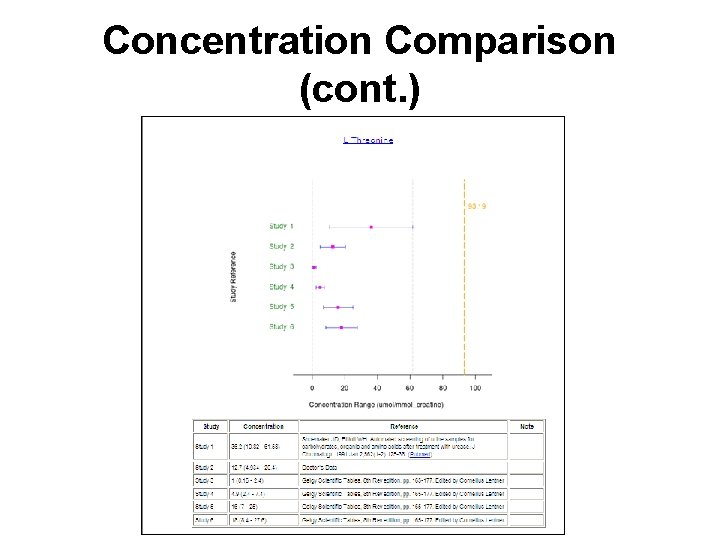
Concentration Comparison (cont. )
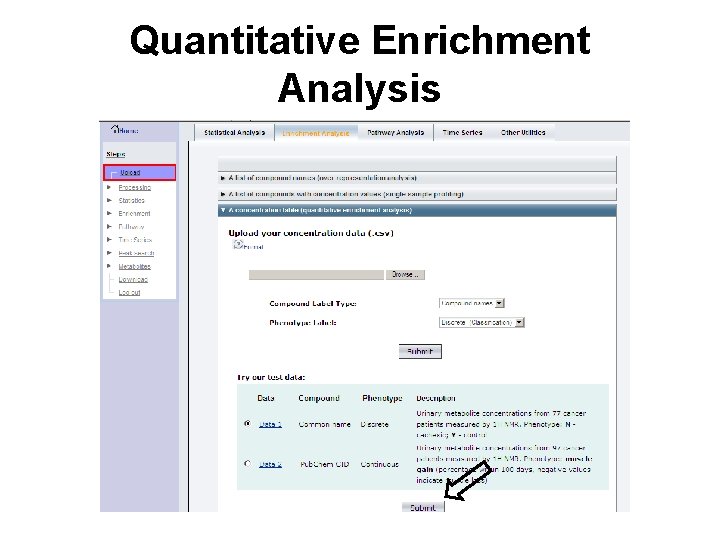
Quantitative Enrichment Analysis
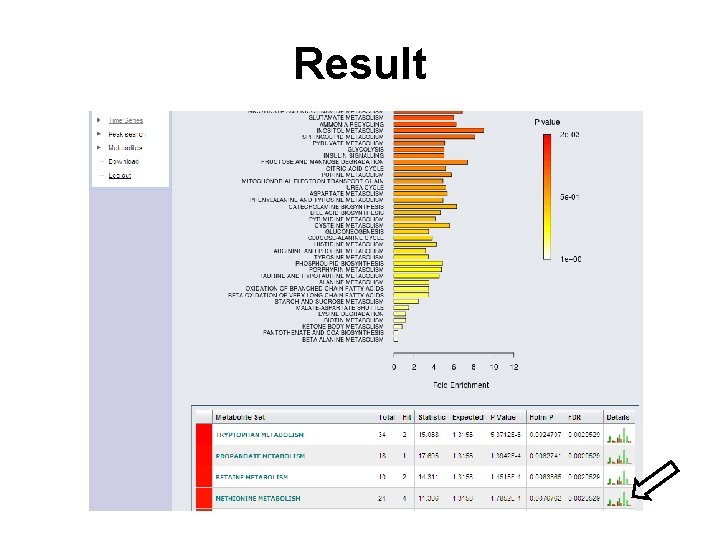
Result
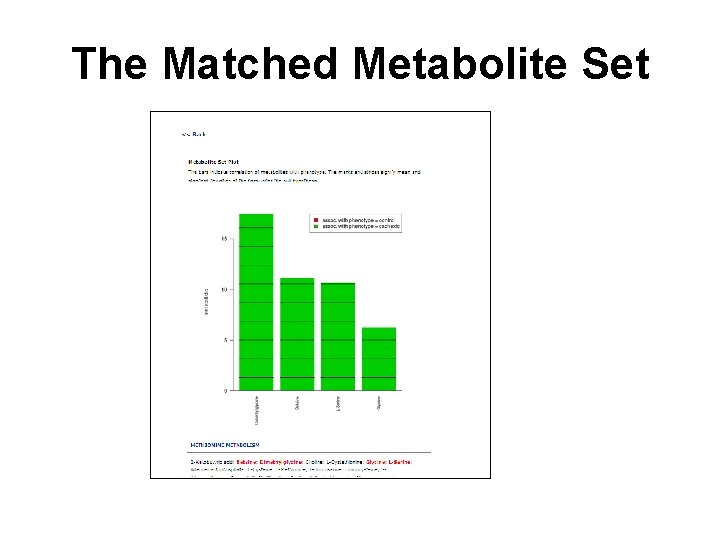
The Matched Metabolite Set

Question • Q: Are these metabolites increased or decreased in the cachexia group?

Metabolic Pathway Analysis with Met. PA

Pathway Analysis • Purpose: to extend and enhance metabolite set enrichment analysis for pathways by – Considering the pathway structures – Supporting pathway visualization • Currently supports 15 organisms
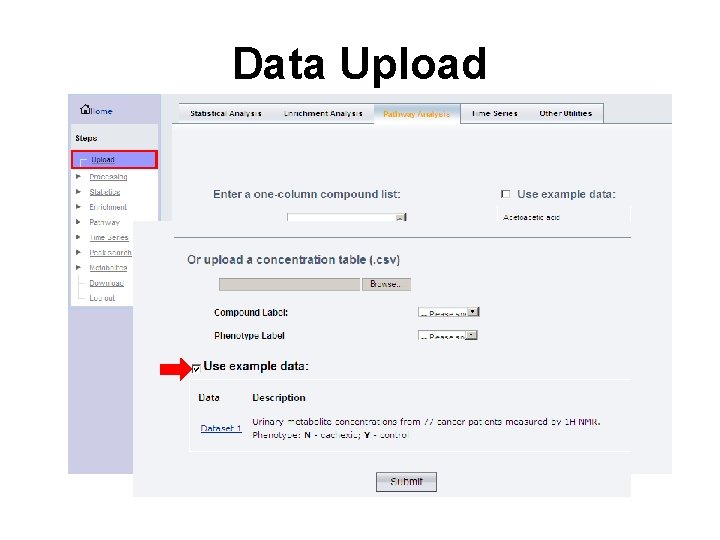
Data Upload

Data Set Selected • Here we are using a collection of metabolites identified by NMR (compound list + concentrations) from the urine from 77 lung and colon cancer patients, some of whom were suffering from cachexia (muscle wasting)
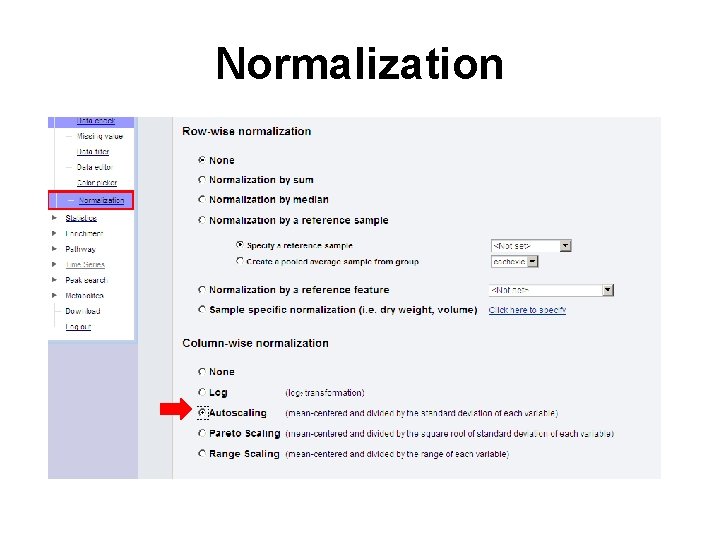
Normalization

Pathway Libraries

Network Topology Analysis
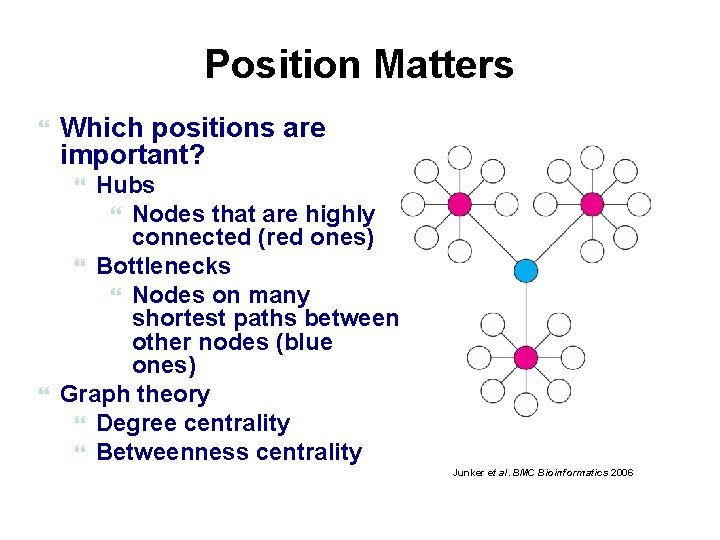
Position Matters Which positions are important? Hubs Nodes that are highly connected (red ones) Bottlenecks Nodes on many shortest paths between other nodes (blue ones) Graph theory Degree centrality Betweenness centrality Junker et al. BMC Bioinformatics 2006 89
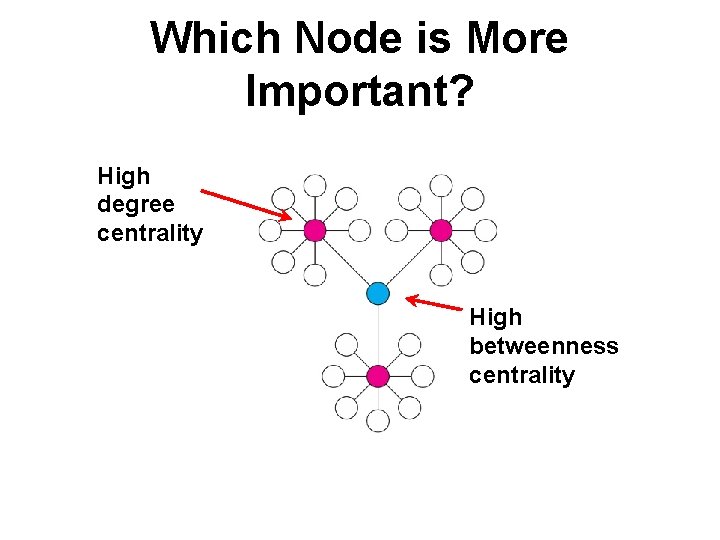
Which Node is More Important? High degree centrality High betweenness centrality
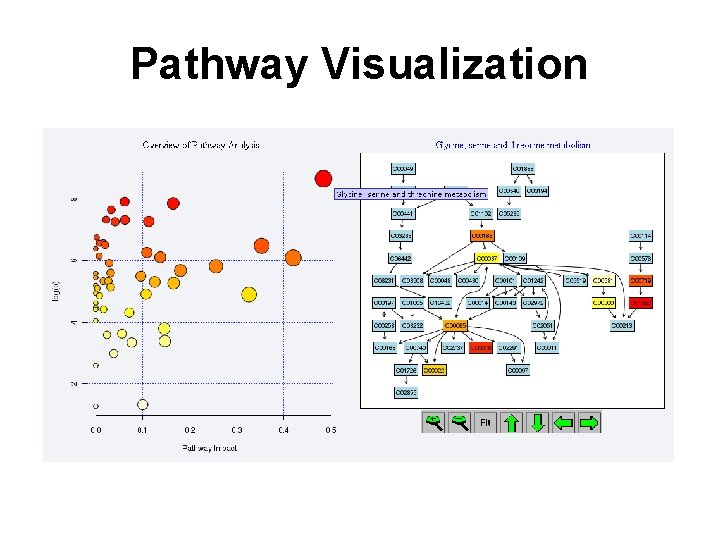
Pathway Visualization
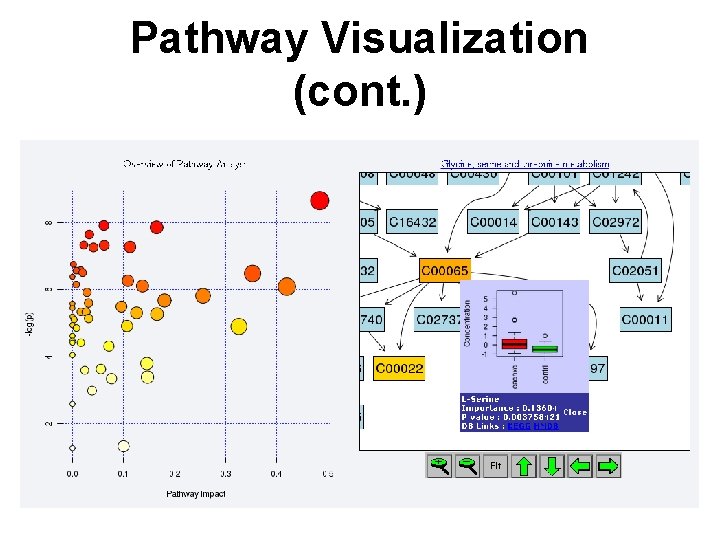
Pathway Visualization (cont. )

Question • Q: Which pathway do you think is likely to be affected the most? Why?
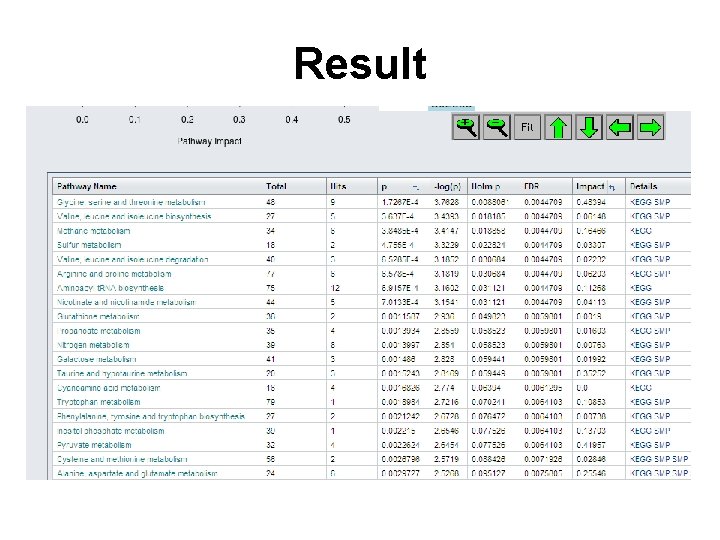
Result

Not Everything Was Covered • • Clustering (K-means, SOM) Classification (SVM, random. Forests) Time-series data analysis Two factor data analysis Data quality checks Peak searching ….
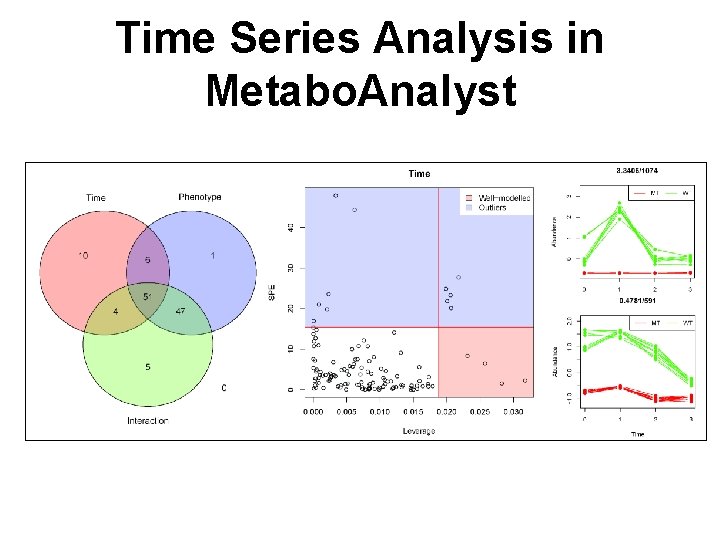
Time Series Analysis in Metabo. Analyst 96
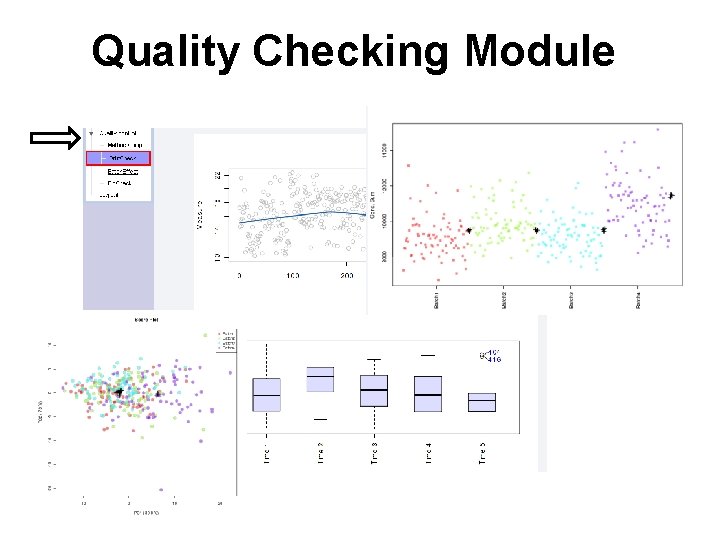
Quality Checking Module
- Slides: 97