Bioinformatics at Promega Corporation Intro to Bioinformatics Biotec

Bioinformatics at Promega Corporation Intro to Bioinformatics Biotec May 4, 2006 Ethan Strauss Sr. Scientist R&D Bioinformatics, Promega, Ethan. strauss@promega. com http: //q 7. com/~ethan/molbio

My Background • Bachelor’s degree in biology • Ph. D and work experience in Molecular Biology • Eight years in Promega Technical Services • Almost a year in Bioinformatics (officially) No formal computer training No formal bioinformatics training

Bioinformatics at Promega Corporation • Bioinformatics did not exists as a separate function until 2001 • One person 2001 - 2005 • Two people 2005 - ? • Bioinformatics supports primarily R&D (~100 scientists) • Mentor and train R&D scientists • Provide expertise for projects (~120 requests per year) • Propose and evaluate new acquisitions • Liaison to IT department • Manage bioinformatics infrastructure (~15 tools) • Develop new tools and adapt existing tools in house

Bioinformatics Projects Programming • Tools for internal and external Promega customers • Plexor™ Primer Design System • Biomath • si. RNA Designer • Sequence analysis for Excel and Microsoft Word • Analysis of BLAST results • Automated data retrieval (Web services) • Database for tracking vector construction • Database for keeping track of plasmid features • Laboratory Information Management System (LIMS) • Chemical Database

Bioinformatics Projects Biocomputing (use of computers in biological research) • Database searches • data mining • discovery research • Analysis & in silico design of nucleic acid and protein sequence • Molecular visualization • Modeling • Simulation (proteins, ligands)

Programming • Tools for Promega customers – Biomath (http: //www. promega. com/biomath/) • • Basic calculations (Most can be done easily by hand) Simple code (Javascript) Established theory. Universal (not Promega specific) – si. RNA Designer(http: //www. promega. com/si. RNADesigner/ ) • • Complex calculations More complex code (VBScript) Rapidly evolving theory Partially Promega specific

Programming • Tools for Promega customers – Plexor Primer Design (https: //www. promega. com/techserv/tools/plexor) • Complex calculations • Complex code (C#. Net) – – Separate user interface and main calculations Multiple interacting modules Database integration Integration with Genbank (through a web service) • Proprietary improvements on established theory • Very Promega specific

Programming • Tools for internal use – BLAST analysis of Plexor Primers • Primer specificity is important • BLAST can determine specificity, but output is very complex. • Simplify – – Combine all hits from the same “Gene” Only show hits which could mis-prime Groups hits by species Allow sorting by species

Programming • Tools for internal use – BLAST analysis of Plexor Primers Initial BLAST results (1 page out of ~30) Analyzed BLAST results (complete!)

Programming • Tools for internal use – Vector/Insert Database • Promega’s Flexi vector system has a very structured cloning procedure. • R&D has been making many different Flexi vector backbones with many inserts. • Keeping track has been a problem. • A database is in development

Programming • Tools for internal use
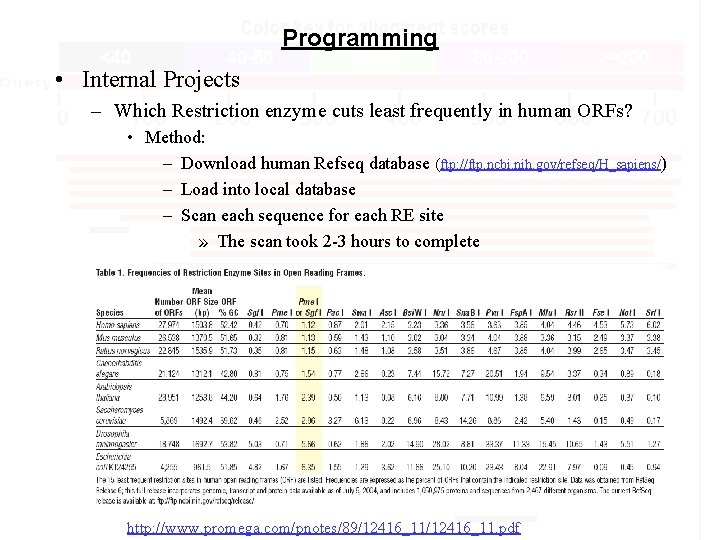
Programming • Internal Projects – Which Restriction enzyme cuts least frequently in human ORFs? • Method: – Download human Refseq database (ftp: //ftp. ncbi. nih. gov/refseq/H_sapiens/) – Load into local database – Scan each sequence for each RE site » The scan took 2 -3 hours to complete http: //www. promega. com/pnotes/89/12416_11. pdf

Programming • Internal Projects – Which human genes in Genbank are the most “popular”? • Method – Download “Gene” database (ftp: //ftp. ncbi. nlm. nih. gov/gene/) – Download Gene Ontology information (http: //www. geneontology. org/) – Use web services to get pathway information from KEGG (http: //www. genome. jp/kegg/) – Use web services to get citation information from Pubmed (http: //www. ncbi. nlm. nih. gov/entrez/query. fcgi? db=Pub. Med) – Load all into local database – Rank genes by desired criteria » » » Size Function Localization Pathways Publications
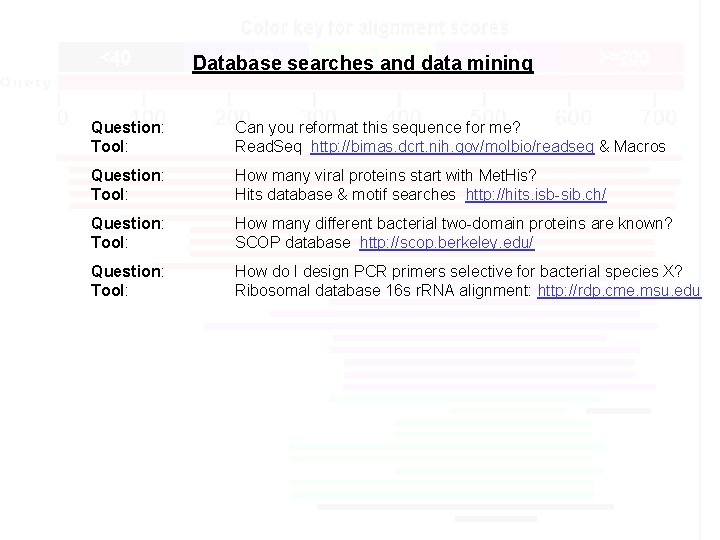
Database searches and data mining Question: Tool: Can you reformat this sequence for me? Read. Seq http: //bimas. dcrt. nih. gov/molbio/readseq & Macros Question: Tool: How many viral proteins start with Met. His? Hits database & motif searches http: //hits. isb-sib. ch/ Question: Tool: How many different bacterial two-domain proteins are known? SCOP database http: //scop. berkeley. edu/ Question: Tool: How do I design PCR primers selective for bacterial species X? Ribosomal database 16 s r. RNA alignment: http: //rdp. cme. msu. edu
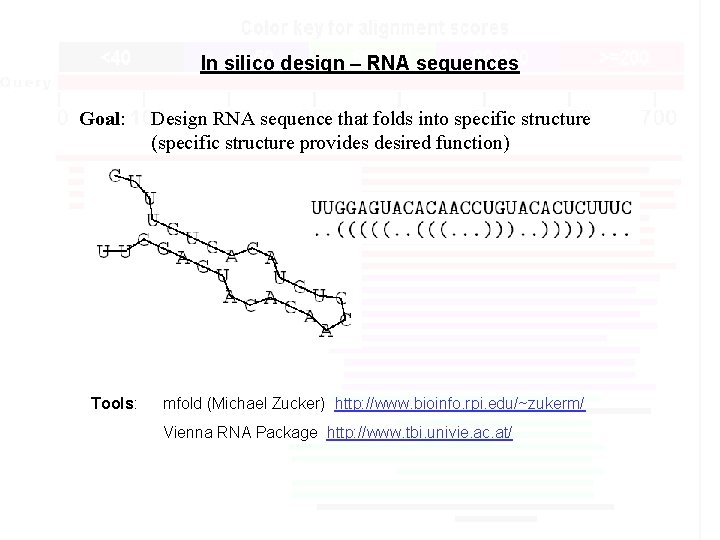
In silico design – RNA sequences Goal: Tools: Design RNA sequence that folds into specific structure (specific structure provides desired function) mfold (Michael Zucker) http: //www. bioinfo. rpi. edu/~zukerm/ Vienna RNA Package http: //www. tbi. univie. ac. at/

In silico design – DNA sequences Goal: Express protein of interest in E. coli cells – fastest way Steps: Obtain protein or DNA sequence from database Optimize codon usage for expression in E. coli Match restriction enzyme sites to expression vector Send DNA sequence for synthesis (cost ~$1/base) Tools: NCBI database http: //www. ncbi. nlm. nih. gov Codon usage database http: //www. kazusa. or. jp/codon/ Restriction enzyme database http: //rebase. neb. com/rebase. html Sequence analysis software
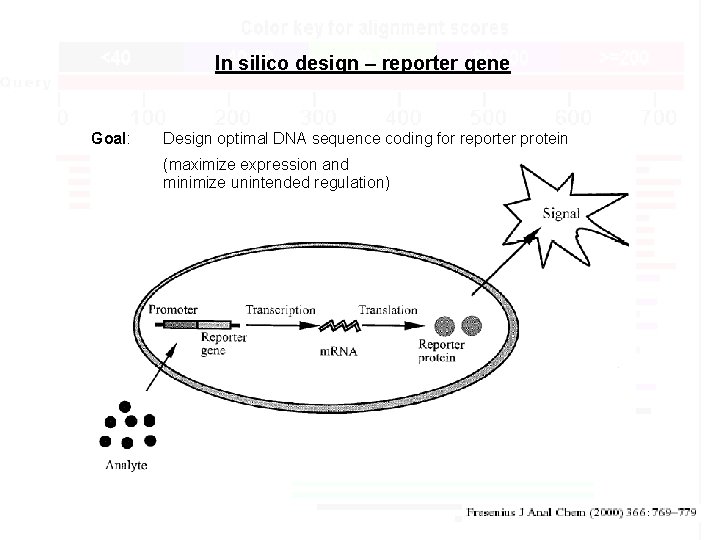
In silico design – reporter gene Goal: Design optimal DNA sequence coding for reporter protein (maximize expression and minimize unintended regulation)
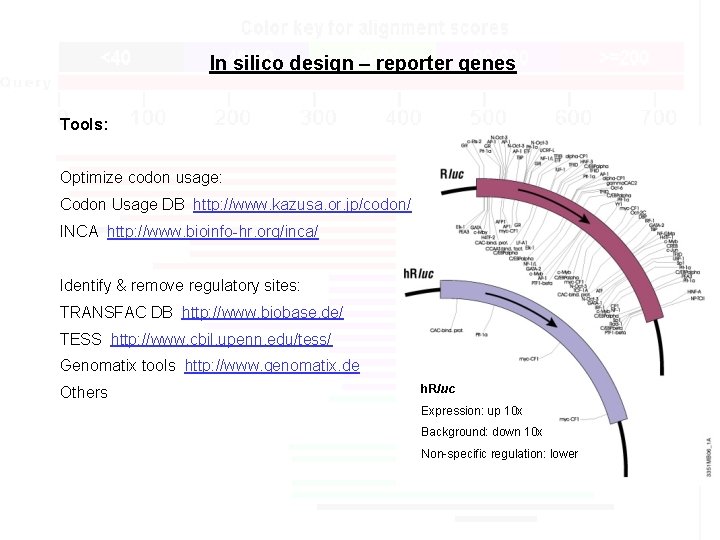
In silico design – reporter genes Tools: Optimize codon usage: Codon Usage DB http: //www. kazusa. or. jp/codon/ INCA http: //www. bioinfo-hr. org/inca/ Identify & remove regulatory sites: TRANSFAC DB http: //www. biobase. de/ TESS http: //www. cbil. upenn. edu/tess/ Genomatix tools http: //www. genomatix. de Others h. Rluc Expression: up 10 x Background: down 10 x Non-specific regulation: lower

Visualization – molecular system of interest Goal: Visualize molecule of interest (blue) and interaction partners Tools: World Index of Molecular Visualization Resources http: //molvis. sdsc. edu/visres/index. html

Modeling – protein fold Goal: Tools: 3 D structure model of enzyme => location of N/C termini => find active site => other NCBI BLink http: //www. ncbi. nlm. nih. gov/ Protein Data Bank http: //www. rcsb. org/pdb Swiss. Model http: //swissmodel. expasy. org/ WHAT IF http: //swift. cmbi. ru. nl/whatif/ Insight. II Modeler http: //www. accelrys. com/insight unknown 3 D structure: Renilla luciferase homologue with known 3 D structure: Hydrolase sequence identity: 36%

Modeling – protein engineering Goal: Alter catalytic activity of enzyme => predict structural effects of different point mutations mutation disrupts structure Tools: mutation does not disrupt structure Insight. II Modeler http: //www. accelrys. com/insight/

Modeling – protein engineering Goal: Improve substrate binding rate of enzyme => identify specific amino acids to mutate constricted binding tunnel Tools: open binding tunnel (mutant) Insight. II Modeler http: //www. accelrys. com/insight/

Modeling – substrate engineering Goal: Find better substrate for enzyme => analyze geometric constraints of substrate binding pocket Tools: Hetero-compound Info Center http: //alpha 2. bmc. uu. se/hicup/ Insight. II Modeler http: //www. accelrys. com/insight/
Database for chemical compounds

LIMS – Laboratory Information Management System Goal: Manage in-house DNA sequences and associated data Eval: UW-Madison Center for Eukaryotic Structural Genomics Sesame http: //www. sesame. wisc. edu/ “…Sesame is designed to organize and record data relevant to complex scientific projects, to launch computer-controlled processes, and to help decide about subsequent steps on the basis of information available. The Sesame system is based on the multi-tier paradigm, and it consists of a framework and application modules that carry out specific tasks. Users interact with Sesame through a series of web-based Java appletapplications designed to organize data. It allows collaborators on a given project to enter, process, view, and extract relevant data, regardless of location, so long as web access is available. Data reside in an Oracle relational database. Sesame serves as a digital laboratory notebook and allows users to attach numerous files and images…”

Bioinformatics Advice • Be aware of bias in databases! – Search Genbank (nucleotide) for Human[Organism] apoptosis. How many hits? – Now try Orcinus[Organism] apoptosis How many hits? – Can you conclude that Orcinus does not have apoptosis?

Bioinformatics Advice • Bioinformatics is changing and advancing very rapidly. – Don’t forget to notice what is new. • NCBI now has ~20 different databases. They had two only 3 -5 years ago – If you want to do something that you know can’t be done, check again in two weeks! • My standard computer can process the entire human genome for Restriction sites, ORFs etc in a few hours. Not long ago, the best computers couldn’t even hold that much data! – If old tools work, don’t feel you need to use the newest tools. • I still do much of my analysis with Microsoft Word…
- Slides: 27