Issues with creating Genome Browsers for Whole Genome

Issues with creating Genome Browsers for Whole Genome Assemblies G-On. Ramp Beta Users Workshop Wilson Leung 07/2016

Outline Obtain genome assemblies from NCBI Transfer large genomics datasets to Galaxy Common bioinformatics file formats Datatypes in Galaxy Obtain protein sequences for tblastn searches

Common types of evidence tracks on a Genome Browser Protein alignments RNA-Seq data Repeats Gene predictions Genomic sequence

Obtaining the genome assembly from NCBI Bio. Project http: //www. ncbi. nlm. nih. gov/bioproject/ Entry point to all genomic datasets (e. g. , genome assembly, transcriptome) that pertain to a study

Data from the 1000 Genome Project available through NCBI Bio. Project SRA = Sequence Read Archive Database of high -throughput sequencing data Data available through NCBI, EBI, and DDBJ

Types of genome assemblies Ref. Seq genome categories Reference genome High quality assembly Standard for comparison Example: D. melanogaster Representative genome Best genome assembly available within a clade Example: D. elegans http: //www. ncbi. nlm. nih. gov/assembly/help/

Obtain genome assemblies from the NCBI FTP site Download genome sequence, predicted transcript and protein products Consistent primary sequence IDs (accession. version) for both GFF and FASTA files http: //www. ncbi. nlm. nih. gov/news/08 -26 -2014 -new-genomes-FTP-live/

Naming conventions for Gen. Bank assemblies <accession. version>_<assembly name>_<content type>. <format> Content type Description genomic Genome assembly (Repeats identified by Window. Masker are in lower-case) rm Transposons identified by Repeat. Masker (Eukaryotes only) See the README. txt file within the directory for details

Common data formats used by Gen. Bank assemblies Large data files are compressed by gzip Format Description fna Nucleotide sequence in FASTA format faa Protein sequence in FASTA format Supported by Galaxy gbff Gen. Bank flat file format Built-in support in mac. OS gff General Feature Format Version 3 Use 7 -Zip in MS Windows File suffix =. gz http: //www. 7 -zip. org/ Genome assembly in FASTA format: _genome. fna. gz Example: GCA_000224195. 2_Dele_2. 0_genomic. fna. gz
DEMO: Access the D. elegans genome assembly from the NCBI FTP site

Benefits of using FTP to transfer large files to Galaxy Problems with standard file upload Most servers have a 2 GB file upload size limit Cannot monitor progress of file upload Cannot resume interrupted file upload FTP file upload is supported at Galaxy Main Support transfer of large gzip, bzip 2, and zip files https: //wiki. galaxyproject. org/FTPUpload Install and configure Pro. FTPD to enable FTP file upload in a local Galaxy instance https: //wiki. galaxyproject. org/Admin/Config/Uploadvia. FTP

Overview of the File Transfer Protocol (FTP) Data transfer protocol between a client and a server May allow anonymous access Insecure connection Partial built-in support in most operating systems mac. OS: Go ➜ Connect to Server MS Windows: File Explorer Other graphical clients Cyberduck, File. Zilla, Fugu, …

Use FTP to upload files to Galaxy Use a FTP client to initiate a FTP connection to Galaxy ftp: //usegalaxy. org Use your Galaxy account credentials to authenticate Transfer files to the Galaxy FTP server Use the Upload File tool to import contents of the FTP directory into Galaxy Files available through the “Choose FTP file” button Cannot use mac. OS Finder as FTP client because of @ character in the username (http: //apple. stackexchange. com/questions/89635/how-toconnect-to-ftp-from-finder-with-in-the-credentials)

Use Cyberduck to transfer files from the NCBI FTP site to Galaxy Open Connection Server = usegalaxy. org Enter the username and password for your Galaxy account File ➜ New Browser Copy the FTP link to the Gen. Bank assembly at NCBI Paste link into the “Quick Connect” textbox and press “Enter” Select and drag files from the NCBI connection window to the Galaxy connection window https: //trac. cyberduck. io/wiki/help/en
DEMO: Use FTP to upload the D. elegans genome assembly to Galaxy

Transfer high-throughput sequencing data from the SRA to Galaxy Second and third generation sequencing data available through the Sequence Read Archive (SRA) NCBI SRA stores sequencing data in sra format Use the SRA Toolkit to convert files to fastq (fastq-dump) Paired-end reads might split at the wrong position: https: //www. biostars. org/p/12569/ European Nucleotide Archive (ENA) at EBI SRA sequencing data in fastq format Import data into Galaxy using “Get Data” ➜ “EBI SRA”

Proliferation of different bioinformatics file formats Standards (https: //xkcd. com/927/)

Common bioinformatics file formats used by Genome Browsers Format Description BED • Browser Extensible Data • Main data format used by the UCSC Genome Browser big. Bed • Binary indexed version of BED • Main data format used by the UCSC Assembly Hub big. Gene. Pred • Extended version of big. Bed for displaying gene predictions in the UCSC Assembly Hub big. Wig • Used for displaying high density, continuous datasets • (e. g. , RNA-Seq read coverage) GFF 3 • General Feature Format Version 3 • 9 -column, tab-delimited file used to describe genomic features and alignments https: //genome. ucsc. edu/FAQformat. html

BED format specification Three required columns and nine optional columns Tab-delimited text file May include additional fields Coordinates are 0 -based, half open: [start, end) First base in the chromosome = 0 End coordinate is not part of the feature First 100 bases of a sequence: start=0, end=100 Common BED formats: BED 4 = chrom, chrom. Start, chrom. End, name BED 6 = BED 4 + score, strand BED 9 = BED 6 + thick. Start, thick. End, item. Rgb BED 12 = BED 9 + block. Counts, block. Sizes, block. Starts

Common variants of BED format Format BED variant Description bed. Graph BED 3 + data. Value Used by mod. ENCODE and ENCODE to display high density, continuous data. (Scores must be between 0 and 1000) narrow. Peak BED 6 + signal. Value, p. Value, q. Value, peak Used by ENCODE to show narrow signal enrichment and potential binding sites (e. g. , transcription factors ) broad. Peak BED 6 + signal. Value, p. Value, q. Value Used by ENCODE to show broad signal enrichment (e. g. , histone modifications) gapped. Peak BED 12 + signal. Value, p. Value, q. Value Used by ENCODE to show confidence intervals of peak calls TRF BED 4 + 12 Include additional details on the repeats identified by Tandem Repeats Finder https: //genome. ucsc. edu/FAQformat. html
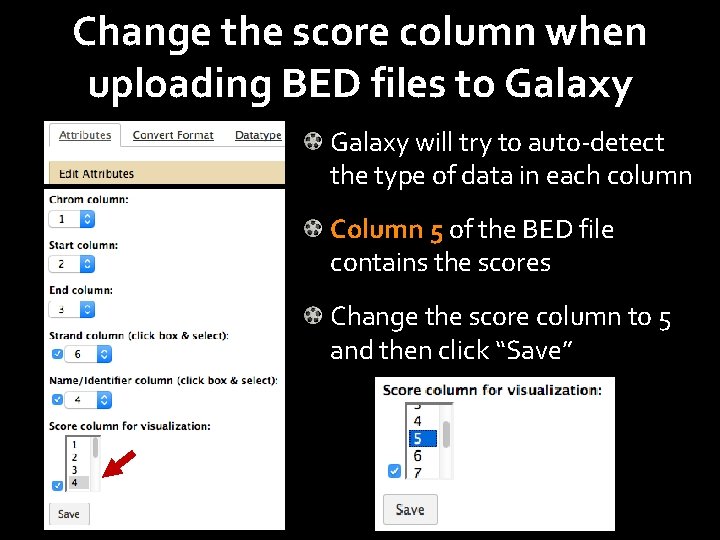
Change the score column when uploading BED files to Galaxy will try to auto-detect the type of data in each column Column 5 of the BED file contains the scores Change the score column to 5 and then click “Save”

General Feature Format Version 3 (GFF 3) Nine-column, tab-delimited text file used to store multiple types of genomic features Coordinates are 1 -based, fully closed: [1, 100] Used by GMOD and many model organisms databases Galaxy tools for manipulating GFF files available under the “Filter and Sort” category Additional tools available at https: //galaxy. cbio. mskcc. org/ Many gene finders report gene models in GFF-like format Most GFF files do not conform to the GFF 3 specification https: //github. com/The-Sequence-Ontology/Specifications/blob/master/gff 3. md

Use the Parent attribute to capture relationships among features Last column of a GFF 3 feature contains its attributes Multiple tag=value attributes are separated by semicolons Attributes are case-sensitive http: //gmod. org/wiki/GFF 3

Common variants of GFF 2 (Gene Finding Format: http: //gmod. org/wiki/GFF 2) Supports single level of nested features GTF (Gene Transfer Format: http: //mblab. wustl. edu/GTF 22. html) Two mandatory attributes: gene_id and transcript_id UCSC Genome Browser does not support GFF 3

Common data formats for high-throughput sequencing Format Description fastq • Contains nucleotide sequences and quality scores of sequencing reads SAM • Sequence Alignment/Map Format • Contains alignments against a genome • (https: //samtools. github. io/hts-specs/SAMv 1. pdf) BAM • BGZF compressed version of SAM • Usually sorted and indexed by position to enable random access of reads that aligned to specific regions CRAM • Store only differences compared to a reference sequence • More compact than BAM files • (https: //samtools. github. io/hts-specs/CRAMv 3. pdf)

Overview of fastq format Sequence ID Sequence Quality fastq sequences might contain Illumina adapters List of common Illumina adapter sequences: http: //support. illumina. com/downloads/illumina-customer-sequence-letter. html Use Galaxy tools under the “NGS: QC and manipulation” section to remove adapter sequences Trim Galore! Trimmomatic

Different fastq quality encodings Quality encoding depends on version of CASAVA used Sanger quality encoding used in version 1. 8+ https: //en. wikipedia. org/wiki/FASTQ_format

Working with fastq files in Galaxy Default datatype for fastq files = fastq Most Galaxy tools (e. g. , Top. Hat, HISAT) expects fastq files with Sanger quality encoding Datatype = fastqsanger Use Fast. QC to determine quality encoding Use FASTQ Groomer to convert encoding to fastqsanger Examine the “Overrepresented sequences” and “Adapter Content” sections of the Fast. QC report Clip adapter sequences from fastq files as needed
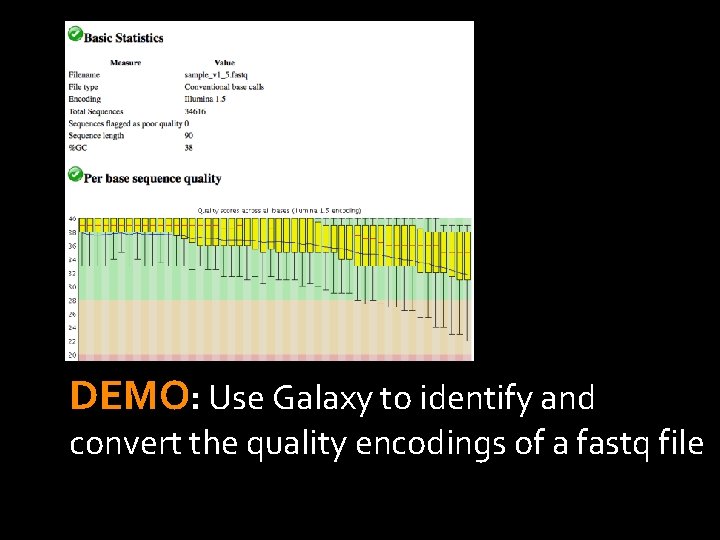
DEMO: Use Galaxy to identify and convert the quality encodings of a fastq file

Obtain protein sequences for tblastn searches Species-specific databases Fly. Base: dmel-all-translation-r 6. 11. fasta. gz http: //flybase. org/static_pages/downloads/bulkdata 7. html Swiss-Prot High quality, manually annotated section of Uni. Prot. KB http: //www. uniprot. org/downloads NCBI Ref. Seq Use only curated Ref. Seq records (accession prefix = NP_) Protein sequences from Ref. Seq reference genomes http: //www. ncbi. nlm. nih. gov/books/NBK 50679/
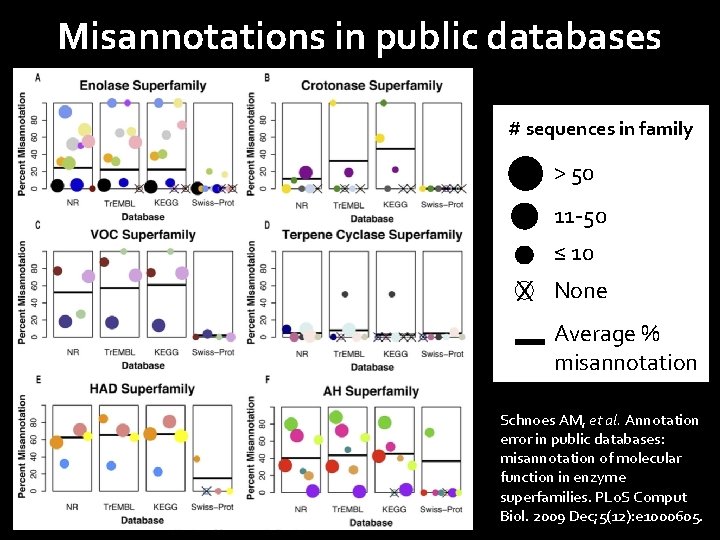
Misannotations in public databases # sequences in family > 50 11 -50 ≤ 10 X None Average % misannotation Schnoes AM, et al. Annotation error in public databases: misannotation of molecular function in enzyme superfamilies. PLo. S Comput Biol. 2009 Dec; 5(12): e 1000605.

Obtain Swiss-Prot protein sequences Uni. Prot download page (http: //www. uniprot. org/downloads) Complete Swiss-Prot database Swiss-Prot sequences separated by taxonomic divisions Human, invertebrates, mammals, plants, rodents, vertebrates, … Download files with the uniprot_sprot prefix Use the seqret EMBOSS tool in Galaxy to create FASTA file Search for “reviewed: yes” entries in Uni. Prot. KB http: //www. uniprot. org/uniprot/? query=reviewed%3 Ayes Filter protein sequences by taxonomy, keywords, gene ontology, enzyme class or pathways
DEMO: Download Swiss-Prot protein sequences from Uni. Prot

NCBI Reference Sequence database More comprehensive than Swiss-Prot Two major types of Ref. Seq records: Known Ref. Seq: NP_ Model Ref. Seq: XP_ Model Ref. Seq records are based on results from computational pipelines More likely to propagate annotation errors http: //www. ncbi. nlm. nih. gov/refseq/about/
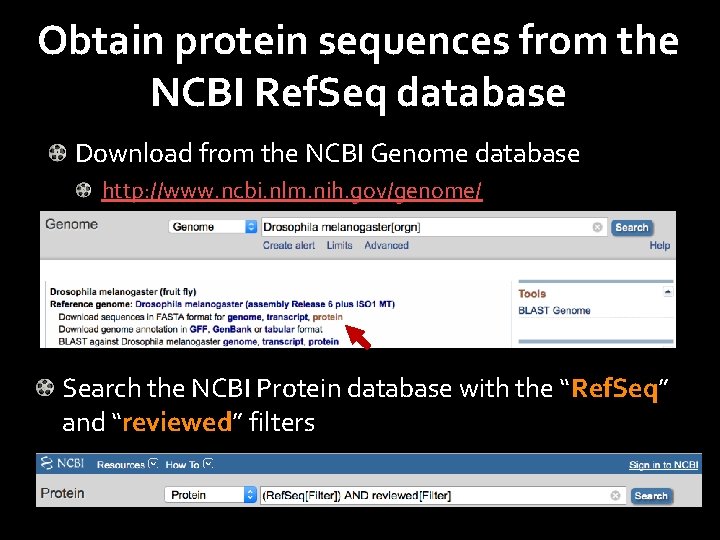
Obtain protein sequences from the NCBI Ref. Seq database Download from the NCBI Genome database http: //www. ncbi. nlm. nih. gov/genome/ Search the NCBI Protein database with the “Ref. Seq” and “reviewed” filters

Summary Obtain genome assemblies from NCBI Use FTP to transfer large genome assemblies to Galaxy Use EBI SRA to transfer fastq files to Galaxy Common variants of BED and GFF file formats Different fastq quality encodings Obtain protein sequences from Swiss-Prot and Ref. Seq databases for tblastn searches

Questions? https: //flic. kr/p/bhy. T 8 B


Improve performance of protein sequence similarity searches Perform initial search to identify high-scoring matches Use large word size (W) and neighborhood threshold (T) Create new protein databased on matches from initial search Use more sensitive parameters and aligner to generate more accurate alignments in target region Compart + Splign tblastn + Exonerate tblastn + SPALN Korf I. Serial BLAST searching. Bioinformatics. 2003 Aug 12; 19(12): 1492 -6.
- Slides: 39